Back to Journals » Journal of Pain Research » Volume 11
The Pain Registry for Epidemiological, Clinical, and Interventional Studies and Innovation (PRECISION): registry overview and protocol for a propensity score-matched study of opioid prescribing in patients with low back pain
Authors Licciardone JC, Gatchel RJ, Phillips N, Aryal S
Received 30 March 2018
Accepted for publication 31 May 2018
Published 6 September 2018 Volume 2018:11 Pages 1751—1760
DOI https://doi.org/10.2147/JPR.S169275
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Michael Schatman
John C Licciardone,1 Robert J Gatchel,2 Nicole Phillips,3 Subhash Aryal4
1Department of Family Medicine, University of North Texas Health Science Center, Fort Worth, TX, USA; 2Department of Psychology, College of Science, University of Texas at Arlington, Arlington, TX, USA; 3Department of Microbiology, Immunology and Genetics, University of North Texas Health Science Center, Fort Worth, TX, USA; 4Department of Biostatistics and Epidemiology, Institute for Patient Safety, University of North Texas Health Science Center, Fort Worth, TX, USA
Background: Low back pain is the leading cause of disability worldwide. Nonsteroidal anti-inflammatory drugs (NSAIDs) are recommended as the first-line pharmacologic therapy for subacute or chronic low back pain, with opioids reserved for patients who fail on NSAIDs. CYP2D6, CYP2C9, and CYP2C19 genes have variants that place patients using analgesics at risk for adverse events. However, precision medicine based on pharmacogenetically informed prescribing is becoming more feasible as genotyping costs decline. This study aims to compare opioids vs NSAIDs in treating adults with subacute or chronic low back pain under the alternative models of usual care and precision medicine.
Methods: An observational cohort study within the Pain Registry for Epidemiological, Clinical, and Interventional Studies and Innovation (PRECISION) will be used to simulate a randomized controlled trial. Patients using opioids and NSAIDs will be optimally matched at baseline using propensity scores. A saliva sample will also be collected to determine patient genotypes for drug metabolism based on CYP2D6 (single-gene model) and CYP2D6, CYP2C9, and CYP2C19 (multigene model). Prescribing that is concordant with pharmacogenetically informed care under these models will be considered “low risk”, whereas discordant prescribing will be considered “high risk”. Primary outcomes will be assessed over 6 months using a Numerical Rating Scale for pain, the Roland-Morris Disability Questionnaire, and the Drug Adverse Events Index. Secondary outcomes will be assessed using quality-of-life measures. An estimated 600 patients will be enrolled to acquire at least 400 patients after attrition and allowing for unmatched patients. This will achieve a statistical power of at least 80% in detecting the effect sizes ranging from 0.35 (small–medium effect) to 0.69 (medium–large effect).
Discussion: This PRECISION Pain Research Registry study builds on the concepts espoused in the Precision Medicine Initiative and addresses long-term goals established by the National Institutes of Health by assessing how precision medicine may prevent and treat chronic pain.
Keywords: PRECISION Pain Research Registry, low back pain, physical functioning, quality of life, opioids, codeine, nonsteroidal anti-inflammatory drugs, pharmacogenetics, precision medicine, biopsychosocial model
Background
The Institute of Medicine, in its report on “Relieving Pain in America”, concluded that relieving pain should be a national priority.1 The “Federal Pain Research Strategy” was recently disseminated by the Interagency Pain Research Coordinating Committee and the Office of Pain Policy of the National Institutes of Health to guide long-term strategic planning in the support of pain research.2 The report cited, as a top priority, the development of approaches that incorporate the principles of precision medicine to prevent and effectively treat chronic pain. It also recommended establishing new pain research registries that target the general population to track clinical and patient-reported outcomes.
The Global Burden of Disease Study estimated that 632 million persons worldwide suffer from low back pain, making it the leading cause of disability.3 The societal costs of low back pain are >$100 billion annually in the United States, including health care expenditures and lost productivity.4 Nonsteroidal anti-inflammatory drugs (NSAIDs) are now recommended as the first-line pharmacologic therapy for subacute or chronic low back pain.5 Nevertheless, NSAIDs are associated with potentially serious side effects involving the gastrointestinal, cardiovascular, and renal systems. In particular, nausea, vomiting, heartburn, stomach pain, gastrointestinal ulcer, or abnormal bleeding with short-term NSAID use may herald serious gastrointestinal toxicity.6
Although not recommended as the first-line therapy, many patients who fail on nonpharmacologic treatments and NSAIDs will eventually receive opioids, including tramadol. Such patients are also potentially at risk of drug adverse events. Data from randomized controlled trials have identified six side effects that are significantly associated with opioids: nausea, vomiting, constipation, drowsiness, dizziness, and itching.7 Dry mouth and headache have also been frequently reported with opioid use.8 The Centers for Disease Control and Prevention recently established opioid prescribing guidelines for physicians to help reduce the occurrence of such side effects and stem the national epidemic of opioid abuse.9
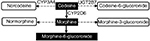
Codeine serves as a prototype for opioid metabolism and analgesia. The conversion of codeine to morphine and subsequently to morphine-6-glucuronide, as shown in Figure 1, represents its primary pathway for analgesia.10 The associations of CYP2D6 metabolizer phenotypes with the formation of morphine via this pathway are well known. Extensive metabolizers experience normal analgesia at recommended opioid doses. However, ultra-rapid metabolizers are at high risk of opioid toxicity due to an increased conversion of codeine to morphine. Poor metabolizers may lack opioid response because of decreased conversion to morphine. Intermediate metabolizers may not achieve adequate analgesia at recommended opioid doses and must be closely monitored to balance the potential benefits and risks of therapy. Pharmacokinetic and pharmacodynamic studies of drugs such as oxycodone and tramadol show that they also depend on CYP2D6 for conversion to active metabolites responsible for analgesia.11
The Clinical Pharmacogenetics Implementation Consortium identified the CYP2D6 gene as having high-risk variants that may inform drug prescribing and strongly recommends that such genetic information be used if available.12 Codeine, oxycodone, and tramadol are drugs for low back pain that fall under this recommendation, and they account for >70 million prescriptions annually in the United States.12 Pharmacogenetically informed prescribing has not been commonly used to identify patients at risk for adverse events or poor response to opioid therapy. However, precision medicine approaches to drug prescribing were encouraged through the Precision Medicine Initiative13 and are becoming more feasible as the cost of genotyping declines. Thus, precision medicine holds the promise of becoming a cost-effective approach to inform drug prescribing for low back pain and therefore yield better effectiveness and safety outcomes than usual care, which is not informed by pharmacogenetics.
Combinatory genotyping of CYP2D6, CYP2C9, and CYP2C19 has been promoted as an advance in the pharmacogenetics of pain management because most analgesics are metabolized by multiple pathways involving enzymes in the CYP450 family.11 Drug metabolism indices for pharmacogenetic functional status based on this multigene model have been developed14 and tested in clinical settings such as those involving pain,11 psychiatric disorders,14 and dyslipidemia.15 The drug metabolism reserve index (DMRI) uses allele scoring for CYP2D6, CYP2C9, and CYP2C19 to define a series of discrete CYP450 metabolic phenotypes that reflect innate drug metabolism capacity, ranging from sub- to supra-functional.11 These phenotypes parallel those defined by the single-gene model based only on CYP2D6.
The primary purpose of this study is to compare the effectiveness and safety outcomes of medical care that is concordant vs discordant with these single- and multigene models for pharmacogenetically informed prescribing of opioids and NSAIDs for patients with low back pain. These precision medicine outcomes will be qualitatively compared with those observed under the prevailing usual care model of drug prescribing without pharmacogenetic input.
Methods
Registry overview
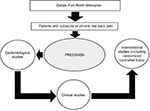
The Pain Registry for Epidemiological, Clinical, and Interventional Studies and Innovation (PRECISION) was established in 2016 to conduct research relating to low back pain. The creation and operations of the registry have been guided by principles established by the Agency for Healthcare Research and Quality.16 The registry relies on patient-reported data to acquire evidence on real-world outcomes in a timely fashion to help impact health care delivery.17,18 At present, the registry focuses on collecting data and biospecimens from patients in the Dallas–Fort Worth Metroplex to conduct observational studies. Both the registry and the study protocol described herein were approved by the Institutional Review Board at the University of North Texas Health Science Center prior to launch (2015-169 and 2017-102, respectively). All patients provide written informed consent prior to enrolling in the registry. The vision for the registry is to eventually increase patient enrollment through geographic expansion and to extend into clinical research and randomized registry trials,19 as shown in Figure 2.
Study design
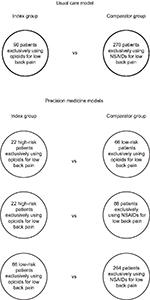
The registry will be used to conduct an observational cohort study with propensity score matching of patients to simulate a randomized controlled trial.20 Baseline data will be used to optimally match patients exclusively using opioids or NSAIDs for low back pain on variables such as age, gender, history of medical conditions, pain sensitivity, pain catastrophizing, pain self-efficacy, low back pain intensity, back-specific functioning, and quality of life. Index patients will be matched, either 1:3 or 1:4, with comparator patients in each model depending on the number of potentially available registry patients, as shown in Figure 3.
Inclusion and exclusion criteria
The registry will aim to enroll patients who are representative of adults with subacute or chronic low back pain in the general population. The inclusion criteria include being 21 years of age or older; self-reporting low back pain for a duration of at least 2 months (subacute) or 6 months (chronic), with a pain frequency of at least half of the days during the relevant period; having the ability to respond to data collection in English; and having a physician who provides medical care for low back pain (primary care or specialist physician). Exclusion criteria include being pregnant (based on self-report) or being institutionalized or incarcerated.
Data elements and survey instruments
Registry patients will complete comprehensive baseline data collection and quarterly follow-ups. Data on potential drug adverse events will be collected at four weekly encounters immediately following the baseline visit or after the next quarterly follow-up encounter for patients enrolled prior to October 2017. Baseline data will be collected during an in-person visit to the registry facilities, which also involves the acquisition of blood and saliva samples to be banked for biomarker and genetic analyses. Subsequent registry encounters may be conducted in person, by telephone, or via online communication. The Qualtrics software (Provo, UT, USA) will be used to collect patient-reported data at all encounters. These data will be exported to the IBM SPSS Statistics software (version 23) (IBM Corporation, Armonk, NY, USA) and other statistical packages as needed for analysis. An overview of the 3-year timetable for the collection of data elements and survey instrument responses is presented in Table 1. Each table entry is briefly described below.
National Institutes of Health minimum dataset for low back pain
This minimum dataset is recommended by the National Institutes of Health Task Force on Research Standards for Chronic Low Back Pain to describe patients participating in research studies.21 It includes 40 items relating to demographic characteristics, medical history, symptoms, and functioning. The items will be adapted to be relevant to patients with either subacute or chronic low back pain.
History of medical conditions
This inventory consists of 10 back-related or general medical conditions that have ever been diagnosed, including herniated disk, sciatica, osteoporosis, osteoarthritis, heart disease, hypertension, diabetes mellitus, asthma, bronchitis, and depression.
Comprehensive pharmacologic treatments
This item will be used to report all medications (both prescription and nonprescription drugs) currently used for low back pain or other conditions. The reporting format includes medication name, dose, and frequency of administration. Such self-reported medication use has generally shown good agreement with pharmacy records.22 It will also be used to estimate the morphine milligram equivalents of opioids used by patients.7
Low back pain-specific opioid use
This item will be used at all encounters to assess the current use and the previous use of any opioid drug for low back pain.
Low back pain-specific NSAID use
This item will be used at all encounters to assess the current use and the previous use of any NSAID for low back pain.
Drug Adverse Events Index
This instrument is adapted from a previous study8 to report 10 drug side effects commonly associated with opioids or NSAIDs, including nausea, vomiting, constipation, drowsiness, dizziness, itching, dry mouth, headache, heartburn, and stomach pain. The index score will be computed for each side effect using the product of the reported presence (1) or absence (0) of the side effect during the previous week and its typical intensity, if present. The latter ranges from 1 (hardly noticeable) to 10 (as bad as possible). The mean index score over 4 weeks will also be computed for each side effect.
History of nonpharmacologic treatments for low back pain
This inventory consists of six common nonpharmacologic treatments that have ever been used for low back pain, including a formal exercise therapy program, yoga, massage therapy, spinal manipulative therapy, acupuncture, and cognitive behavioral therapy.
Numerical Rating Scale (NRS) for low back pain
An 11-point NRS will measure the average low back pain intensity over the past 7 days, ranging from 0 (no pain) to 10 (worst possible pain).
Roland–Morris Disability Questionnaire (RMDQ)
The RMDQ will measure how much low back pain adversely affects patient functioning and activities.23 It consists of 24 items that are scored as either 1 (agree that low back pain has an adverse impact) or 0 (disagree that low back pain has an adverse impact). The RMDQ is scored as the sum of responses to each item, thereby potentially ranging from 0 to 24.
Patient-Reported Outcomes Measurement Information System (PROMIS)
The PROMIS measures the quality of life. A pain behavior item bank was developed as a part of PROMIS to provide static and dynamic measures of pain behavior in clinical studies.24 The seven scales that comprise the PROMIS (29-item version) include physical function, anxiety, depression, fatigue, sleep disturbance, the ability to participate in social roles and activities, and pain interference (with activities). Each scale consists of four ordinal-scale items (the 29th item is an NRS for pain intensity). The crude responses are transformed and standardized using scoring algorithms so that 50 and 10 represent the mean and standard deviation, respectively, for each scale. Higher scores more strongly reflect the scale descriptor (eg, higher scores represent better physical function but more anxiety). Scoring on the scales for physical function and the ability to participate in social roles and activities will be reversed to align with interpretation of the other five scales, wherein higher scores represent worse outcomes.
Pain Sensitivity Questionnaire
The Pain Sensitivity Questionnaire will serve as a self-reported alternative to experimental pain testing.25 It has been validated in chronic pain patients26 and consists of an NRS ranging from 0 (no pain) to 10 (worst possible pain) for 14 items that represent either mildly painful situations or moderately painful situations. The overall score is reported as the mean across all items.
Pain Catastrophizing Scale
The Pain Catastrophizing Scale will measure the phenomenon defined as an exaggerated negative mental set brought to bear during actual or anticipated painful experience.27 It consists of 13 ordinal scale items comprising the three dimensions of rumination, magnification, and helplessness.28 The overall score ranges from 0 (no catastrophizing) to 52 (greatest level of catastrophizing).
Pain Self-Efficacy Questionnaire
The Pain Self-Efficacy Questionnaire will measure the degree to which patients mitigate pain and the negative emotions associated with it to maintain everyday life activities including work.29 It consists of 10 ordinal scale items that assess confidence in performing various activities.30 The total score ranges from 0 (no self-efficacy) to 60 (greatest level of self-efficacy).
Communication Behavior Questionnaire
The Communication Behavior Questionnaire will measure patient preferences with respect to physician communication relating to low back pain.31 It consists of 23 Likert-scale items comprising the four scales of patient participation and orientation, effective and open communication, emotionally supportive communication, and communication about personal circumstances. Scores range from 0 to 100 on each scale, with higher scores reflecting desirable physician communication behavior.
Consultation and Relational Empathy Measure
The Consultation and Relational Empathy Measure will assess physician empathy during encounters for medical care.32 It consists of 10 ordinal scale items. Scores range from 10 to 50, with higher scores representing stronger patient perception of physician empathy.
Patient Satisfaction Questionnaire
The Patient Satisfaction Questionnaire (18-item version) will measure satisfaction with general medical care.33 It consists of seven Likert-scale items, covering communication, general satisfaction, technical quality, interpersonal manner, financial aspects, time spent with provider, and access and convenience. Scale scores range from 1 to 5, with higher scores representing greater satisfaction with medical care.
Scale of Patient Overall Satisfaction with Primary Care Physicians
The Scale of Patient Overall Satisfaction with Primary Care Physicians will measure a patient’s overall satisfaction with a primary care physician or other provider of low back pain care.34 It consists of 10 Likert-scale items. Scores range from 10 to 70, with higher scores representing greater satisfaction with a physician.
Assessment of opioid prescribing risk
Opioid prescribing that is concordant with pharmacogenetically informed care will be considered “low risk”, whereas discordant prescribing will be considered “high risk”. For example, in the single-gene model, opioid prescribing for extensive metabolizers will be considered low risk , whereas opioid prescribing for ultra-rapid, intermediate, or poor metabolizers will be considered high risk. In the multigene model, opioid prescribing for patients with functional metabolic capacity based on the DMRI will be considered low risk, whereas opioid prescribing for patients with sub- or supra-functional metabolic capacity will be considered high risk.
Estimates for high-risk CYP2D6 metabolizer genotypes in the general population range from 15% to 29%;11,12 however, the percentage may be higher in selected patient populations such as those attending pain management programs.11 An estimated 25% of patients in the registry using opioids will be assumed to carry a high-risk metabolizer genotype in the single-gene model. Although less is known about the prevalence of high-risk genotypes in the multigene model, it is reasonable to assume based on the available DMRI data from a community population14 that the prevalence of such high-risk genotypes is comparable to that observed in the single-gene model. Although some patients will undoubtedly be designated as high risk within both models, the disparate patients variously identified as high risk and low risk by these models may serve to determine which model is superior in predicting the effectiveness and safety outcomes.
Outcome measures
The effectiveness outcomes of low back pain treatment will be longitudinally assessed over the 6-month interval following enrollment in the registry. These outcomes will be based primarily on the NRS for pain intensity (0–10) and the RMDQ for deficits in back-specific functioning (0–24). These are the two most common patient-reported outcome measures for low back pain, and research standards for their use and interpretation have been established, including for clinically important changes over time.21,35–37 Each of the seven PROMIS-29 scales will serve as a secondary outcome measure to assess the quality of life in relation to low back pain. The Drug Adverse Events Index score (0–10) will serve to measure the co-primary safety outcomes pertaining to each of the 10 symptoms that may potentially represent a drug side effect over a 4-week period.
Laboratory methods
Patient blood samples will be drawn by certified phlebotomists at the baseline visit and transported to the registry’s affiliated laboratory, where they will be handled in accordance with institutional biosafety policies and procedures. Blood samples will be placed on ice within 20 minutes of collection; fractionation into plasma, serum, and buffy coat will be conducted at 4°C; and all processing will be completed within 1 hour of collection. Blood samples, including whole blood, plasma aliquots, serum aliquots, and buffy coat, will be stored at -80°C until they are processed for future studies. Saliva samples will also be collected at baseline using the Oragene DNA Collection Kit (DNA Genotek, Ottawa, ON, Canada).
DNA genotyping will be conducted using the iScan Array Scanner (Illumina, San Diego, CA, USA) and the Global Screening Array (Illumina). The array content is expert-designed to simultaneously genotype the single-nucleotide polymorphism (SNP) loci that have an established, validated role in absorption, distribution, metabolism, and excretion of drugs. The chip also includes genome-wide SNP coverage, including markers that are ancestrally informative and others that confer known, quantifiable risk for particular human diseases. This global screening aimed at personalized medicine includes the CYP2D6, CYP2C9, and CYP2C19 SNP loci, and the genotypes for all SNPs within these three genes will be specifically mined from the microarray data for the purposes of risk characterization and cohort grouping. CYP2D6 duplications will be assessed as well to identify ultra-rapid metabolizers, either by copy number variant analysis of microarray data or by real-time polymerase chain reaction. The Human Cytochrome P450 Allele Nomenclature Database will be used as the primary resource for allele calling and diplotype designation.38 Although there are >80 known alleles for CYP2D6, the most common alleles have been characterized for enzyme activity and the resulting metabolic functional class can be used to assess the outcomes of pharmacologic treatment and dosing. Risk characterization will be conducted based on the combination of alleles for the SNPs in CYP2D6 in the single-gene model or on the combination of variants in all three cytochrome P450 genes (CYP2D6, CYP2C9, and CYP2C19) in the multigene model.
Statistical methods
Comparison of opioids vs NSAIDs under the usual care model
In the usual care model (ie, wherein drug prescribing is not informed by patient information relating to pharmacogenetics), patients exclusively using opioids for low back pain will be matched with patients exclusively using NSAIDs and compared with respect to effectiveness and safety outcomes. Based on preliminary registry data, an estimated 15% of patients will be treated exclusively with opioids, 50% exclusively with NSAIDs (the recommended first-line pharmacologic therapy), and the remainder with other, combined, or no drug therapy. Using propensity scores modeled from baseline data, 90 patients using opioids will be optimally matched with 270 patients using NSAIDs (Figure 3) through the caliper method without replacement.20 These sample sizes are estimated to provide 80% statistical power to detect an effect size (Cohen’s d) as low as 0.35.
Comparisons involving opioids and NSAIDs under the precision medicine models
In the precision medicine models (ie, wherein opioid prescribing is assessed to determine if, in theory, it is concordant or discordant with pharmacogenetically informed care based on either the single-gene model or the multigene model), patients exclusively using opioids will be similarly matched with comparator patients as illustrated in Figure 3. The sample sizes and statistical power estimates for comparisons under the precision medicine models were based on the assumptions described earlier for opioid prescribing risk and the use of opioids, NSAIDs, and other pharmacologic treatments for low back pain. A summary of sample sizes, effect size detection limits, and statistical power estimates for each model is provided in Table 2. An estimated 600 registry enrollees will be recruited to conduct these analyses because of losses owing to patient attrition and nonsuitable matching of comparator to index patients.
Statistical analyses
Statistical analyses will be performed using parametric methods (or alternate nonparametric methods if indicated) for the effectiveness and safety outcomes described earlier. The primary and secondary outcomes involve data that may be analyzed using methods for continuous variables. Specifically, this will involve the use of hierarchical models for longitudinal data to compare the outcomes of opioid vs NSAID use under the usual care model and high- and low-risk opioid use and NSAID use under the precision medicine models. Such models may explore potential moderators, mediators, or confounders that are not controlled by propensity score matching. If indicated, continuous data may be transformed to categorical or binary data to explore additional outcomes or to conduct responder analysis. Hypotheses will be assessed using two-tailed tests at the 0.05 level of statistical significance, with the exception of analyses involving the 10 co-primary outcome measures for safety. The latter will be assessed using a Bonferroni-adjusted level of statistical significance (0.005). Statistical comparisons will be performed “within models” for usual care and precision medicine but are not feasible “between models” because all patients analyzed under the precision medicine models will be a subset of either the overall groups of 90 patients using opioids or 270 patients using NSAIDs for low back pain under the usual care model (Figure 3). Only a qualitative comparison of precision medicine and usual care may be made under these circumstances.
Discussion
Many patient registries are disease-oriented and focus on cancer or rare conditions that require a critical mass of patients to enable meaningful research. At present, there are few pain research registries in existence. Perhaps this is because conditions such as low back pain and other musculoskeletal disorders are not generally perceived to be life-threatening despite their widespread prevalence and potential for causing disability. Nevertheless, precisely because such painful conditions are so common, pain research registries may facilitate pragmatic studies that truly inform clinical decision-making in real-world settings.
The goals and objectives of the PRECISION Pain Research Registry are highly congruent with the long-term strategic plans for pain research as reported by the National Institutes of Health in their Federal Pain Research Strategy.2 The study protocol described herein addresses the important issues in the pharmacologic treatment of low back pain that have been recently highlighted by the Centers for Disease Control and Prevention in its “Guideline for Prescribing Opioids for Chronic Pain”9 and by the American College of Physicians clinical practice guideline on noninvasive treatments for acute, subacute, and chronic low back pain.5
Conclusion
The linkage of the PRECISION Pain Research Registry, a biobank of patient samples for biomarker and genetic analyses, and a database that includes the biopsychosocial aspects of pain builds on the concepts originally espoused in the Precision Medicine Initiative. By conducting such research as described herein, the registry may contribute to the promise of precision medicine by helping to determine the “right drug for the right patient at the right dose”.39
Acknowledgments
The authors wish to thank the registry research staff, faculty, and community advisory board for their contributions to study implementation. The registry and study protocol described herein have been partially funded by the Osteopathic Heritage Foundation, the American Osteopathic Association (# 131611703 and 751711713), and the Institute for Patient Safety. None of the funding bodies had a role in the design of the registry or study protocol; the plan for collection, analysis, or interpretation of data; or writing the article. JCL is a professor of Family Medicine and Richards-Cohen Distinguished Chair in Clinical Research at the University of North Texas Health Science Center. RJG is a distinguished professor of Psychology and Nancy P. & John G. Penson Endowed Professor of Clinical Health Psychology at the University of Texas at Arlington. NP is an assistant professor of Genetics at the University of North Texas Health Science Center. SA is an associate professor of Biostatistics at the University of North Texas Health Science Center.
Author contributions
JCL conceived the concepts for the registry and study protocol, acquired funding for them, and will be responsible for overall administration of the registry and project. RJG made substantial contributions to the development of the registry and study protocol. NP will be responsible for genetic analyses. SA will provide the oversight of statistical aspects of the study. JCL drafted the original article. RJG, NP, and SA assisted in reviewing and revising the manuscript. All authors contributed toward data analysis, drafting and revising the paper and agree to be accountable for all aspects of the work. All authors approved the final article.
Disclosure
The authors report no conflicts of interest in this work.
References
Institute of Medicine. Relieving Pain in America. A Blueprint for Transforming Prevention, Care, Education, and Research. Washington, DC: The National Academies Press; 2011. | ||
Interagency Pain Research Coordinating Committee. Federal Pain Research Strategy. 2017. Available from: https://iprcc.nih.gov/sites/default/files/FPRS_Research_Recommendations_Final_508C.pdf. Accessed December 17, 2017. | ||
Vos T, Flaxman AD, Naghavi M, et al. Years lived with disability (YLDs) for 1160 sequelae of 289 diseases and injuries 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet. 2013;380(9859):2163–2196. | ||
Katz JN. Lumbar disc disorders and low-back pain: socioeconomic factors and consequences. J Bone Joint Surg Am. 2006;88(Suppl 2):21–24. | ||
Qaseem A, Wilt TJ, McLean RM, Forciea MA; Clinical Guidelines Committee of the American College of Physicians. Noninvasive treatments for acute, subacute, and chronic low back pain: a clinical practice guideline from the American College of Physicians. Ann Intern Med. 2017;166(7):514–530. | ||
Ambegaonkar A, Livengood K, Craig T, Day D. Predicting the risk for gastrointestinal toxicity in patients taking NSAIDs: the Gastrointestinal Toxicity Survey. Adv Ther. 2004;21(5):288–300. | ||
National Opioid Use Guideline Group. Canadian Guideline for Safe and Effective Use of Opioids for Chronic Non-Cancer Pain. Hamilton, ON: McMaster University; 2010. | ||
Jamison RN, Raymond SA, Slawsby EA, Nedeljkovic SS, Katz NP. Opioid therapy for chronic noncancer back pain. A randomized prospective study. Spine. 1998;23(23):2591–2600. | ||
Dowell D, Haegerich TM, Chou R. CDC guideline for prescribing opioids for chronic pain – United States, 2016. MMWR Recomm Rep. 2016;65(1):1–49. | ||
Crews KR, Gaedigk A, Dunnenberger HM, et al. Clinical Pharmacogenetics Implementation Consortium guidelines for cytochrome P450 2D6 genotype and codeine therapy: 2014 update. Clin Pharmacol Ther. 2014;95(4):376–382. | ||
Manworren RC, Jeffries L, Pantaleao A, Seip R, Zempsky WT, Ruano G. Pharmacogenetic testing for analgesic adverse effects: pediatric case series. Clin J Pain. 2016;32(2):109–115. | ||
Dunnenberger HM, Crews KR, Hoffman JM, et al. Preemptive clinical pharmacogenetics implementation: current programs in five US medical centers. Annu Rev Pharmacol Toxicol. 2015;55:89–106. | ||
Ashley EA. The precision medicine initiative: a new national effort. JAMA. 2015;313(21):2119–2120. | ||
Villagra D, Goethe J, Schwartz HI, et al. Novel drug metabolism indices for pharmacogenetic functional status based on combinatory genotyping of CYP2C9, CYP2C19 and CYP2D6 genes. Biomark Med. 2011;5(4):427–438. | ||
Ruano G, Villagra D, Szarek B, et al. Physiogenomic analysis of CYP450 drug metabolism correlates dyslipidemia with pharmacogenetic functional status in psychiatric patients. Biomark Med. 2011;5(4):439–449. | ||
Gliklich RE, Dreyer NA. Registries for Evaluating Patient Outcomes: A User’s Guide. 2nd ed. Rockville, MD: Agency for Healthcare Research and Quality; 2010. | ||
Rotenstein LS, Huckman RS, Wagle NW. Making patients and doctors happier – the potential of patient-reported outcomes. N Engl J Med. 2017;377(14):1309–1312. | ||
Frieden TR. Evidence for health decision making – beyond randomized, controlled trials. N Engl J Med. 2017;377(5):465–475. | ||
Lauer MS, D’Agostino RB Sr. The randomized registry trial – the next disruptive technology in clinical research? N Engl J Med. 2013;369(17):1579–1581. | ||
Austin PC. An introduction to propensity score methods for reducing the effects of confounding in observational studies. Multivariate Behav Res. 2011;46(3):399–424. | ||
Deyo RA, Dworkin SF, Amtmann D, et al. Report of the NIH Task Force on research standards for chronic low back pain. J Pain. 2014;15(6):569–585. | ||
Drieling RL, LaCroix AZ, Beresford SA, Boudreau DM, Kooperberg C, Heckbert SR. Validity of self-reported medication use compared with pharmacy records in a cohort of older women: findings from the Women’s Health Initiative. Am J Epidemiol. 2016;184(3):233–238. | ||
Roland M, Morris R. A study of the natural history of back pain. Part I: development of a reliable and sensitive measure of disability in low-back pain. Spine. 1983;8(2):141–144. | ||
Revicki DA, Chen WH, Harnam N, et al. Development and psychometric analysis of the PROMIS pain behavior item bank. Pain. 2009;146(1–2):158–169. | ||
Ruscheweyh R, Marziniak M, Stumpenhorst F, Reinholz J, Knecht S. Pain sensitivity can be assessed by self-rating: development and validation of the Pain Sensitivity Questionnaire. Pain. 2009;146(1–2):65–74. | ||
Ruscheweyh R, Verneuer B, Dany K, et al. Validation of the Pain Sensitivity Questionnaire in chronic pain patients. Pain. 2012;153(6):1210–1218. | ||
Sullivan MJ, Thorn B, Haythornthwaite JA, et al. Theoretical perspectives on the relation between catastrophizing and pain. Clin J Pain. 2001;17(1):52–64. | ||
Sullivan MJ. The Pain Catastrophizing Scale: User Manual. Montreal, QC: McGill University; 2009. | ||
Miles CL, Pincus T, Carnes D, Taylor SJ, Underwood M. Measuring pain self-efficacy. Clin J Pain. 2011;27(5):461–470. | ||
Nicholas MK. The pain self-efficacy questionnaire: taking pain into account. Eur J Pain. 2007;11(2):153–163. | ||
Farin E, Gramm L, Schmidt E. Taking into account patients’ communication preferences: instrument development and results in chronic back pain patients. Patient Educ Couns. 2012;86(1):41–48. | ||
Mercer SW, Maxwell M, Heaney D, Watt GC. The consultation and relational empathy (CARE) measure: development and preliminary validation and reliability of an empathy-based consultation process measure. Fam Pract. 2004;21(6):699–705. | ||
Marshall GN, Hays RD. The Patient Satisfaction Questionnaire Short-Form (PSQ-18). Santa Monica, CA: RAND Corp; 1994. | ||
Hojat M, Louis DZ, Maxwell K, Markham FW, Wender RC, Gonnella JS. A brief instrument to measure patients’ overall satisfaction with primary care physicians. Fam Med. 2011;43(6):412–417. | ||
Assendelft WJ, Morton SC, Yu EI, Suttorp MJ, Shekelle PG. Spinal manipulative therapy for low back pain. A meta-analysis of effectiveness relative to other therapies. Ann Intern Med. 2003;138(11):871–881. | ||
Ostelo RW, Deyo RA, Stratford P, et al. Interpreting change scores for pain and functional status in low back pain: towards international consensus regarding minimal important change. Spine. 2008;33(1):90–94. | ||
Dworkin RH, Turk DC, Wyrwich KW, et al. Interpreting the clinical importance of treatment outcomes in chronic pain clinical trials: IMMPACT recommendations. J Pain. 2008;9(2):105–121. | ||
PharmGKB [homepage on the Internet]. The Human Cytochrome P450 Allele Nomenclature Database. 2017. Available from: http://www.cypalleles.ki.se/. Accessed December 20, 2017. | ||
National Institutes of Health. The Precision Medicine Initiative. 2018. Available from: https://syndication.nih.gov/multimedia/pmi/infographics/pmi-infographic.pdf. Accessed January 5, 2018. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.