Back to Journals » Infection and Drug Resistance » Volume 13
The First Egyptian Report Showing the Co-Existence of blaNDM-25, blaOXA-23, blaOXA-181, and blaGES-1 Among Carbapenem-Resistant K. pneumoniae Clinical Isolates Genotyped by BOX-PCR
Authors El-Badawy MF, El-Far SW, Althobaiti SS, Abou- Elazm FI, Shohayeb MM
Received 29 December 2019
Accepted for publication 4 April 2020
Published 29 April 2020 Volume 2020:13 Pages 1237—1250
DOI https://doi.org/10.2147/IDR.S244064
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Mohamed F El-Badawy,1,2 Shaymaa W El-Far,1 Shaker S Althobaiti,3 Fatma I Abou- Elazm,2 Mohamed M Shohayeb4
1Division of Pharmaceutical Microbiology, Department of Pharmaceutics and Industrial Pharmacy, College of Pharmacy, Taif University, Taif, Kingdom of Saudi Arabia; 2Department of Microbiology and Immunology, Faculty of Pharmacy, Misr University for Science and Technology, 6th of October City, Egypt; 3King Fahad Armed Forces Hospital, Jeddah, Kingdom of Saudi Arabia; 4Department of Microbiology and Biotechnology, Faculty of Pharmacy, Delta University for Science and Technology, Gamasa, Egypt
Correspondence: Mohamed F El-Badawy
Division of Pharmaceutical Microbiology, Department of Pharmaceutics and Industrial Pharmacy, College of Pharmacy, Taif University, Kingdom of Saudi Arabia
Tel +20-103-205-9964
Email [email protected]
Background and Objective: The emergence of carbapenem-resistant K. pneumoniae (CRKP) continues to escalate and is alarming because of the emergence of pan drug-resistant strains. The objective of this study was to investigate the existence of 12 carbapenemase genes among CRKP clinical isolates.
Methods: Ninety-six Klebsiella spp. clinical isolates were collected. The isolates were identified phenotypically and genotypically. These isolates were screened for susceptibility to 24 different antibiotics. The modified Hodge test (MHT) and the Carba Nordmann/Poirel (NP) test were used to phenotypically screen carbapenem-resistant strains for carbapenemase production. Phenotypic characterization of carbapenemases was performed using the combined disk synergy test (CDST). Additionally, the presence of 12 carbapenemase genes in CRKP isolates was investigated. The DNA sequence of blaNDM and blaGES genes was determined. The BOX-PCR technique was used to determine the clonal relationship between CRKP isolates.
Results: All carbapenem-resistant isolates were related to K. pneumoniae. Susceptibility testing showed that 19.79% (19/96) of the collected isolates were carbapenem-resistant. Of the CRKP isolates, 68.42% (13/19) tested positive for the MHT and Carba NP test. CDST showed that 42.11% (8/19), 63.16% (12/19), 47.37% (9/19), and 73.68% (14/19) of the CRKP isolates tested positive for the inhibitory effect of clavulanic acid, sulbactam, phenylboronic acid, and tazobactam, respectively, while 84.21% (16/19) and 68.42% (13/16) tested positive for the inhibitory effect of EDTA and mercaptopropionic acid, respectively. It was found that 10.53% (2/19) of the isolates tested positive for the inhibitory effect of sodium chloride. Molecular investigation of carbapenemases showed that 26.32% (5/19), 73.68% (14/19), 21.05% (4/19), 10.53% (2/19), and 5.26% (1/19) of the isolates tested positive for blaNDM, blaOXA-48, blaOXA-181, blaOXA-51, and blaOXA-23, respectively. None of the isolates tested positive for blaOXA-40 and blaOXA-58. Two allelic variants of blaNDM (blaNDM-1 and blaNDM-25) were detected. BOX-PCR revealed high clonal relatedness between CRKP isolates.
Conclusion: MHT was more sensitive than Carba NP test for evaluating carbapenemase production and class D carbapenemase genes were the most prevalent of the 12 carbapenemase genes that were evaluated.
Keywords: carbapenemases, CRKP, BOX-PCR, blaNDM-25, MHT, CDST
Introduction
Carbapenems are one of the most important groups of antibiotics used to treat bacterial infections caused by multiple drug-resistant (MDR) Gram-negative enteric isolates.1 Irrational and widespread use of carbapenems in intensive care units (ICUs) and other hospital settings has led to the emergence and spread of carbapenem-resistant strains that can be treated only by antimicrobial agents such as tigecycline and colistin1,2 that are considered to be last resort for treating carbapenem-resistant Enterobacteriaceae (CRE).3
Resistance to carbapenems in Gram-negative bacteria is mediated by three mechanisms, including (i) production of hydrolytic enzymes known as carbapenemases, (ii) extrusion of carbapenems from the interior of bacterial cells by efflux pumps, and (iii) downregulation of the outer membrane proteins (OMPs) that are involved in transmembrane passage of carbapenems to reach their targets inside bacterial cells.1,4
According to the definition proposed by the Center for Disease Control and Prevention (CDC), K. pneumoniae is considered to be MDR when it exhibits resistance to at least one agent in three or more classes of antimicrobial agents.5 Extensive drug resistance (XDR) corresponds to MDR isolates that develop resistance to all antibiotic groups except for two or fewer classes.6 Pan drug resistance (PDR) corresponds to XDR isolates that develop resistance to all antibiotics, including polymyxin and tigecycline.4,6
Production of carbapenemases represents the most important carbapenem resistance mechanism among Gram-negative Enterobacteriaceae in which the genes encoding for these enzymes are easily transferred between bacterial flora in hospitals, which can lead to serious outbreaks within different hospital settings.7
According to their active sites, carbapenemases are classified into two main classes: serine carbapenemases and metallo-carbapenemases.8,9 Serine carbapenemases are further classified into two molecular groups according to the Ambler classification of β-lactamases: class A serine carbapenemases (CASCs) and class D serine carbapenemases, which are also known as OXA-type carbapenemases (OTCs) in which these carbapenemases are actually oxacillinases.10
In clinical settings, CASCs can be inhibited by β-lactamase inhibitors such as clavulanic acid, sulbactam, and tazobactam. In contrast, OTCs are poorly inhibited by the aforementioned β-lactamase inhibitors.11
Class B carbapenemases are metallo-β-lactamases in which the active site requires zinc ions as a cofactor. However, metallo-carbapenemases cannot be clinically inhibited but can be inhibited in vitro by chelating agents such as ethylene diamine tetraacetic acid (EDTA) and mercaptopropionic acid (MPA).12,13
BOX-PCR is a DNA fingerprinting technique that detects repetitive and highly conserved DNA elements located on the bacterial chromosome.14 These repeated conserved DNA elements were termed BOX dispersed-repeat motifs that were first identified on the chromosome of Streptococcus pneumoniae. It is now known that the BOX-motif is commonly found in many bacterial species.14,15
BOX-PCR detects a specific form of repetitive extragenic palindromic (rep) sequences that consist of three different regions called boxA, boxB, and boxC that consist of 59, 45, and 50 base pairs, respectively. BOX-PCR is used to discriminate between clinical isolates of the same genus in order to track genetic differences between them.14
To the best of our knowledge, there are no published studies from Egypt regarding investigation of blaOXA-23, blaOXA-40, blaOXA-51, blaOXA-58, and blaOXA-181 in CRKP clinical isolates, and thus the current study was performed to investigate the existence of these carbapenemase encoding genes in CRKP clinical isolates recovered from Egyptian patients and to identify the clonal relatedness between these isolates.
Patients and Methods
Bacterial Strains
Nineteen CRKP isolates were selected from a total of 96 Klebsiella spp. clinical isolates that had been recovered as a part of routine microbiological laboratory procedures at the hospital involved in the present study. The CRKP clinical isolates were collected from 17 cases (9 males and 8 females) who had been admitted to different hospital departments. The 19 CRKP clinical isolates were recovered from sputum (n = 3), urinary catheter (n = 1), urine (n = 6), wound (n = 7), and central line catheter (n = 2).
Isolation and Identification of K. pneumoniae Isolates
All recovered Klebsiella spp. clinical isolates were primarily isolated on MacConkey’s agar (Oxoid, UK) then on eosin methylene blue (EMB) agar (Scharlau, Spain). The isolated strains were identified phenotypically using API 20E (Biomerieux, France) and were confirmed genotypically based on the amplification of the K. pneumoniae phoE gene as previously described16 using the primers and cycling conditions listed in Table 1.
 |
Table 1 Primers and Cyclic Conditions Used in PCR Targeting Three Classes of Carbapenemases and BoxA Region Among CRKP Clinical Isolates |
Antimicrobial Susceptibility Testing (AST)
All K. pneumoniae clinical isolates were tested for susceptibility to 13 β-lactams, three different β-lactam/β-lactamase inhibitor combinations, and seven other agents representing three different antibiotic classes. Susceptibility testing was performed using the modified Kirby–Bauer disc diffusion method17 and the broth microdilution method.18 Susceptibility results were interpreted based on guidelines from the Clinical Laboratory Standards Institute (CLSI).19 Escherichia coli ATCC 25922 and K. pneumoniae ATCC 700603 were used as standard control strains.
Phenotypic Detection of Carbapenemase Production
Modified Hodge Test (MHT)
All selected CRKP clinical isolates were subjected to the MHT to detect carbapenemase production. MHT was performed as previously described20 using a meropenem disk (10 μg) as a substrate. Test results were considered to be positive when the tested organism gave the characteristic clover leaf-like indentation around the meropenem disk as shown in Figure 1. A culture collection of Acinetobacter baumannii laboratory strains supplied by the microbiology laboratory at Taif University was used as a positive control.
Carba Nordmann/Poirel (CNP) Test
All CRKP isolates were investigated for carbapenemase production using the Carba NP test as previously described21,22 in which both CNP test solutions (A) and (B) were prepared. Test results were considered to be positive when CNP solution (B) was converted from red to yellow color, while CNP solution (A) remained red as shown in Figure 1. A culture collection of Acinetobacter baumannii laboratory strains provided by the microbiology laboratory at Taif University was used as a positive control. E. coli ATCC 22925 was used as a negative control.
Phenotypic Characterization of Carbapenemases
Combined Disk Synergy Test (CDST)
Class A, B, and D carbapenemases expressed by the selected CRKP isolates were subjected to phenotypic characterization using CDST as previously described.23 Phenyl boronic acid (PBA),24 sulbactam, clavulanic acid (CA), and tazobactam were used as class A carbapenemase inhibitors.25 EDTA26 and MPA27 were used as class B carbapenemase inhibitors. Sodium chloride was used as a class D carbapenemase inhibitor.11,26
Genotypic Detection of Carbapenemases
All CRKP clinical isolates were screened for 12 different genes encoding for class A, B, and D carbapenemases. Three class A carbapenemase genes (blaGES, blaSME, and blaKPC), three class B carbapenemase genes (blaIPM, blaVIM, and blaNDM), and six Class D genes (blaOXA-23, blaOXA-40, blaOXA-48, blaOXA-51, blaOXA-58, blaOXA-181) were screened using the primers and cycling conditions listed in Table 1.
Extraction of Total Genomic DNA
To investigate carbapenemase encoding genes, total genomic DNA (chromosomal and plasmid DNA) was extracted by a boiling method. Total genomic DNA was extracted as previously described28 in which 3–5 colonies were picked (according to colony size) off from an overnight tryptic soy agar (TSA) plate using a sterile yellow tip; the selected colonies were suspended in 500 µL of sterile distilled water, boiled for 10 min, then centrifuged at 13200 rpm for 3 min. The supernatant containing total genomic DNA was transferred to a new 0.5 mL sterile DNase-free tube and stored at −20°C until use.
DNA Fingerprinting
DNA fingerprinting was based on the BOX-PCR technique using the primers and cycling conditions listed in Table 1. BOX-PCR was performed only on chromosomal DNA template. Chromosomal DNA was extracted using QIAamp DNA Mini Kit (Qiagen, USA) according to the manufacturer’s instructions.
Polymerase Chain Reaction (PCR)
The following reaction components were transferred into sterile DNase free 0.2 mL PCR tubes (Eppendorf, Germany): 4 µL of DNA template, 4 µL of 5x master mix (Solis BioDyne, Tartu, Estonia), 0.6 µL of forward primer (10 pmol/μL), and 0.6 µL of reverse primer (10 pmol/μL). The volume of the reaction mixture was brought to 20 µL by adding 10.8 µL of sterile DNase-free distilled water. Twelve strains from the laboratory culture collection provided by the microbiology laboratory at Taif University were used as positive controls. E. coli ATCC 25922 was used as a negative control strain.
Gene Amplification and Gel Electrophoresis
Gene amplification was performed using a Mastercycler gradient (Eppendorf, Hamburg, Germany), primers from Macrogen (Korea, Geumcheon-gu, Seoul), and the cycling conditions listed in Table 1.
Amplified PCR products corresponding to carbapenemase gene fragments were run on 1% agarose gels containing 0.5 ug/mL ethidium bromide (EtBr), while amplified fragments of the repetitive palindromic sequences of boxA regions were run on 2.5% agarose gels. Agarose gels were run at 100 V for 1.5 hrs using a continuous electrical current power supply (Labnet International Inc, Taiwan).
Extraction and Purification of PCR Products for Gene Sequencing
Amplified blaNDM and blaGES PCR products were cut out of a 1% agarose gel on a UV transilluminator (Bio-Rad, USA) using sterile DNase-free scalpels. Amplified gene fragments entrapped in the gel slices were extracted using the GenElute extraction kit (Sigma-Aldrich, USA). Extracted amplified gene fragments were purified using the ExoSAP-IT One Step PCR Clean-up® Kit (GE Healthcare Limited, USA).
DNA Sequencing
The purified blaNDM and blaGES genes were sequenced by Macrogen using a 3730xl DNA Analyzer (Thermo Fischer Scientific, USA). The primers used for amplification were also used for sequencing.
Sequence Correction and Identification of Query Sequence
Sequencing data were corrected and edited using molecular evolutionary genetic analysis (MEGA) X software (Biodesign Institute, Tempe, USA), as previously described.29 The corrected blaNDM and blaGES gene sequences were uploaded online to the National Center for Biotechnology and Information (NCBI) and nucleotide basic local alignment search tool (BLASTN) was used to determine the similarity of the query sequences with cited reference sequences.
Phylogenetic Analysis of Corrected of blaNDM Sequence
All corrected blaNDM sequences were phylogenetically analyzed using JalViwe 2.11.0 software30 in which the genetic relatedness between the identified sequences and the selected reference sequence was determined.
DNA Fingerprint Analysis
The generated DNA fingerprint patterns were analyzed using BioNumeric7.5 software (AppliedMaths, Kortrijk, Belgium), as previously described,29 in which the phylogenetic relatedness between the investigated isolates was determined by dendrogram construction using Dice (similarity) coefficient with band matching at 1% tolerance and 1.5% tolerance change. Cluster analysis was performed based on the unweighted pair-group method with arithmetic average (UPGMA).
Results
Isolation of CRKP Clinical Isolates
The majority of CRKP clinical isolates were recovered from the Surgical Intensive Care Unit (SICU) with an incidence rate of 26.32% (5/19). On the other hand, 15.79% (3/19) of CRKP isolates were recovered from the urology department and the Respiratory Intensive Care Unit (RICU). One isolate (8.33%) was recovered from each of the following departments: surgery, cosmetic surgery, hematological diseases, geriatric care, pediatrics, internal medicine, endemic diseases, and ear, nose, and throat (ENT).
Phenotypic Identification and Genotypic Confirmation of CRKP Clinical Isolates
Regarding the phenotypic identification of CRKP clinical isolates, all CRKP isolates were identified as K. pneumoniae subspecies pneumoniae according to API 20E in which eight different biotypes were identified. Most identified biotypes were related to 5215773, 5205773, and 5005573 biotypes, with frequencies of 42.11% (8/19), 21.05% (4/19), and 10.53% (2/19), respectively.
On the other hand, the lowest identified biotypes were 1205773, 5044552, 5205573, 5215573, and 5214773 where each of which were detected at an incidence of 5.26% (1/19). Regarding genotypic confirmation of biotype clinical isolates, 100% of phenotypically identified isolates tested positive for the phoE gene of K. pneumoniae.
Susceptibility Testing
AST showed that 19.79% (19/96) of isolates exhibited XDR pattern (data not shown), while none of the isolates exhibited PDR pattern (data not shown). Of the CRKP clinical isolates, 100% were resistant to ampicillin, amoxicillin, cefazolin, cephradine, cefotaxime and cefoperazone, as shown in Table 2. Regarding the other β-lactams that were tested, 5.26% (1/19) of CRKP isolates were susceptible to ceftriaxone, ceftazidime, and aztreonam. Regarding the carbapenems that were tested, 10.53% (2/19) and 5.26% (1/19) of ertapenem non-susceptible isolates exhibited intermediate resistance to imipenem and meropenem, respectively, with regard to non-β-lactam antibiotics, tigecycline and amikacin were the most effective agents, where 94.47% (18/19) and 31.58% (6/19) of CRKP isolates, respectively, were susceptible. Regarding the quinolones that were tested, levofloxacin was more effective than ciprofloxacin; all CRKP isolates were ciprofloxacin-resistant, while 10.53% (2/19) were levofloxacin susceptible.
 |
Table 2 Antimicrobial Susceptibility Pattern of CRKP Clinical Isolates |
With regards to β-lactam/β-lactamase combination, the current study showed that clavulanic acid failed to reverse the resistance of amoxicillin and sulbactam also failed to reverse the resistance of both ampicillin and cefoperazone, as shown in Table 2.
With regards to the tested β-lactams, the MIC50 values ranged from 4 to >1024 µg/mL, while the MIC90 ranged from 16 to >1024 µg/mL, as shown in Table 2. On the other hand, the lowest detected MIC values corresponded to tigecycline, in which MIC50 and MIC90 were ≤0.5 µg/mL. With regards to the tested aminoglycosides, the MIC50 ranged from 128 to >1024 µg/mL, while the MIC90 was >1024 µg/mL. The current study showed that the MIC50 values of the tested quinolones ranged from 32 to 64 µg/mL, while the MIC90 was 128 µg/mL.
Phenotypic Detection and Characterization of Carbapenemases
The current study revealed that 68.42% (13/19) of CRKP clinical isolates tested positive for MHT and the Carba NP test, as shown in Table 3. On the other hand, phenotypic characterization of class A carbapenemases by CDST revealed that 42.11% (8/19), 63.16% (12/19), 47.37% (9/19), and 73.68% (14/19) of CRKP isolates responded to the inhibitory effect of clavulanic acid, sulbactam, PBA, and tazobactam, respectively, as shown in Table 4 and Figure 1. On the other hand, the present study revealed that 84.21% (16/19) and 68.42% (13/16) of CRKP isolates tested positive for the inhibitory effect of EDTA and MPA, respectively, as class B carbapenemase inhibitors. Regarding class D carbapenemases, 10.53% (2/19) of CRKP isolates responded to the inhibitory effect of sodium chloride by at least 5 mm. Overall, tazobactam and EDTA were the most effective inhibitors of class A and B carbapenemases, respectively.
 |
Table 3 Phenotypic and Genotypic Screening Profiles of CRKP Clinical Isolates |
 |
Table 4 Carbapenemase Inhibitor Profile of CRKP Clinical Isolates |
Genotypic Detection of Carbapenemases
Regarding class A carbapenemases, the current study showed that blaGES was the only detected gene in which 5.26% (1/19) of CRKP isolates tested positive. Both blaKPC and blaSME were not detected. Regarding class B carbapenemases, 26.32% (5/19) of CRKP clinical isolates tested positive for blaNDM, while blaIPM was not detected. Class D carbapenemases were the most prevalent among CRKP clinical isolates; 73.68% (14/19), 10.53% (2/19), and 21.05% (4/19) of CRKP clinical isolates tested positive for blaOXA-48, blaOXA-51, and blaOXA-181, respectively, as shown in Table 5. On the other hand, 5.26% (1/19) of CRKP isolates tested positive for blaOXA-23. None of the isolates tested positive for blaOXA-40 and blaOXA-58.
 |
Table 5 Detailed Resistance and Genetic Profiles of CRKP Clinical Isolates |
DNA Sequencing
Two allelic variants of blaNDM were detected as shown in Figure 2; 40% (2/5) are related to blaNDM-1 and 60% (3/5) are related to blaNDM-25. On the other hand, blaGES-1 was the detected variant of blaGES, which has ESBL activity but not carbapenemase activity.
DNA Fingerprint and Cluster Analysis
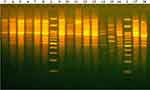
Phylogenetic analysis of the banding pattern generated by Box-PCR (Figure 3) revealed 18 different profiles in which two isolates (K12 and K13) exhibited a single identical profile, whereas the other 17 isolates exhibited 17 different profiles. The generated UPGMA dendrogram classified CRKP isolates into two phylogenetic groups (A&B), as shown in Figure 4. Phylogenetic group A included only one isolate (KP19) that was recovered from the RICU. The genetic profiles of the remaining isolates belonged to phylogenetic group B. Phylogenetic group B was further classified into two main clusters (B1 and B2). Cluster B1 included only one isolate (KP2) that was recovered from the pediatrics department. All the remaining isolates were grouped into the B2 cluster that was further classified into three clades (B2.A, B2.b1, and B2b2). Overall, 73.68% (14/19) of the CRKP isolates were classified into the B2.b2 clade.
 |
Figure 4 Unweighted pair-group method with arithmetic average (UPGMA) dendrogram based on Dice similarity for banding profile generated by BOX-PCR. |
Discussion
K. pneumoniae is one of the most problematic bacterial pathogens and commonly exhibits the MDR pattern. K. pneumoniae is associated with increased mortality and morbidity of hospitalized patients, especially in acute care settings like intensive care units (ICUs) and cardiac care units (CCUs).31
Carbapenems are one of the most effective agents for treating infections caused by MDR K. pneumoniae.32 Unfortunately, resistance to carbapenem among K. pneumoniae continues to increase. Data released by the CDC indicated that 8% of Klebsiella spp. are carbapenem-resistant.33
To the best of our knowledge of the cited literature, the issue of carbapenem resistance in Egypt continues to worsen, which is alarming and represents a serious problem that may be encountered in the future. The current study was designed to investigate the rate and genetic background of carbapenem resistance in Klebsiella spp. clinical isolates recovered from a large tertiary care hospital in Cairo, Egypt. The results of the current study showed that the rate of carbapenem resistance was 19.79%, in which 19 out of 96 Klebsiella spp. were resistant to either imipenem, ertapenem, and/or meropenem.
Our findings are consistent with the findings of Wiener-Well et al34 who reported that the rate of carbapenem resistance in Klebsiella spp. ranged from 8% to 25%.34 Both the findings of the present study and that of Wiener-Well et al34 are far from the findings of the recent Egyptian35 and Iranian36 studies which reported that carbapenem resistance rates among K. pneumoniae clinical isolates were 44.4% and 63.8%, respectively.
Fortunately, the present study did not find any isolates exhibiting the PDR pattern which is consistent with the finding of Wenzi Bi et al37 who reported that no K. pneumoniae isolates exhibited the PDR pattern where all XDR K. pneumoniae isolates were susceptible to tigecycline.
The dissemination of carbapenemase-producing Gram-negative isolates inside hospitals is a major health concern. Early and rapid detection of these isolates is crucial for initiating effective therapeutic options as early as possible in order to achieve desirable outcomes, especially for serious infections and to prevent the spread of these XDR isolates,21,22 based on that, the current study used three phenotypic methods for detection of carbapenemase production among CRKP clinical isolates: (i) MHT (ii), Carb NP test, and (iii) CDST.
Although Carba NP is a cost-effective and rapid test for detecting carbapenemase production, this test in the current study seems to not be highly sensitive, based on the test’s failure to detect carbapenemase production by six isolates that were genotypically confirmed to harbor at least one carbapenemase gene, including OTCs and MBLs, this conclusive finding is identical to the previously reported limitations of the Carba NP test declared by the CLSI.38 Similar reports of low sensitivity for the Carba NP test have been reported.21,22,39
In contrast to the Carba NP test that failed to detect six isolates that were genotypically confirmed to carry at least one carbapenemase gene, the present study revealed that MHT was more sensitive than the Carba NP test, based on failure of MHT to detect only five isolates that were genotypically confirmed to harbor at least one carbapenemase gene.
The current study revealed the superiority of MHT over Carba NP test for detection of carbapenemases, this finding was also reported by a recent Egyptian study35 that showed a high sensitivity for MHT in which 100% (14/14) of CRKP isolates, that were molecularly confirmed to be carbapenemase producers, were tested positive for MHT. Similar findings were reported by AlTamimi et al40 who found that the sensitivity of MHT among CRKP isolates was 92% (23/25). In the current study, we supposed that the negative MHT result for the five isolates that genotypically tested positive for carbapenemases may be due to false negatives rather than true negatives. Similar conclusive false-negative results for MHT were reported by the CLSI.38 The present study suggests that the positive MHT result of the one isolate that tested negative for all investigated carbapenemase genes may be due to (i) false-positive results due to ESBL production or AmpC production coupled with porin loss38 or (ii) the presence of other non-investigated carbapenemase genes.
In contrast to our conclusive findings regarding the higher sensitivity of MHT over Carba NP test, Bayramoğlu et al41 reported that the Carba NP test is more sensitive than MHT.
With regards to the negative results of CDST that was used to characterize carbapenemases in genotypically confirmed isolates that harbor carbapenemases genes, the present study suggests that the negative result may be attributed to one or more of the following four possibilities: (i) low level of expression of the carbapenemase gene(s), (ii) the presence of non-expressed carbapenemase gene(s), (iii) loss of a plasmid that carries the detected carbapenemase gene(s) during the testing period, and (iv) the presence of other resistance mechanisms in addition to the detected carbapenemases.
Regarding the phenotypic characterization of carbapenemases using PBA acid as class A carbapenemase and KPC inhibitor, the current study showed that 47.37% (9/19) of CRKP isolates tested positive, although none of the isolates tested positive for blaKPC, indicating that these results may be false positives due to AmpC hyperproduction42 or the presence of other class A carbapenemase genes that were not investigated in this study.
Although carbapenemase production can be detected by phenotypic tests, molecular detection of these enzymes remains the gold standard.22 The current study revealed that OTCs were the most prevalent carbapenemases that detected in the investigated isolates, in which 78.95% (15/19) of CRKP isolates were found to harbor at least one of the OCTs.
Of the detected OTCs, OXA-48 was the most prevalent one in which 73.68% (14/19) of isolates were found to harbor blaOXA-48. Similar findings were reported by the Turkish,43 Egyptian,31 and Saudi40,44 studies that reported that blaOXA-48 was more prevalent in CRKP clinical isolates.
To the best of our knowledge of the cited literature, few Egyptian studies have investigated the prevalence of blaNDM in CRKP isolates; thus, the current study investigated this issue. The present study detected only five isolates that harbor blaNDM, corresponding to a rate of 26.32%, this finding is somewhat closely related to a recent Egyptian study31 that reported only two CRKP isolates were found to harbor blaNDM at a rate of 4.3%.
Although several reports have been published by some Asian countries like India45 and neighboring countries including Pakistan,46 Bangladesh,47 and Sri Lanka48 indicating the existence of blaOXA-181 in CRKP, there are no Egyptian reports indicating the detection of blaOXA-181 or the coexistence of blaOXA-181 and blaNDM in CRKP clinical isolates.
Based on the above, the current study investigated the existence of blaOXA-181 in CRKP and addressed the first report for the coexistence of blaOXA-181 and blaNDM in CRKP clinical isolates recovered from Egyptian patients. Similar reports have been published by other African countries like Nigeria and Angola.49
With regards to blaOXA-23, many Egyptian reports50–52, have documented the existence of such gene in carbapenem-resistant Acinetobacter baumannii clinical isolates rather than in CRKP clinical isolates.
Based on our knowledge of the cited literature, the current study documents the first African report for the detection of blaOXA-23 in CRKP clinical isolates recovered from Egyptian patients, this finding was confirmed by reviewing the African studies that have investigated such topic and it was found that only one study in this regard has been done in Algeria53 in which blaOXA-23 was detected only in two Acinetobacter baumannii clinical isolates and was not detected in K. pneumoniae clinical isolates.
Although blaOXA-23 blaOXA-40, and blaOXA-58 are plasmid encoded genes that were detected mainly in A. baumannii, while blaOXA-51 is mainly a chromosomal gene in A. baumannii,54 the current study has investigated the four previously mentioned genes in CRKP clinical isolates to check the ability of these genes to transfer from A. baumannii to K. pneumoniae clinical isolates.
As mentioned before, blaOXA-23 blaOXA-40, and blaOXA-58 are mainly plasmid-encoded genes so, they have the chance to transfer to other bacterial genera by conjugation.54 On the other hand, gene transposition from bacterial chromosome to plasmid is well documented in A. baumannii so, blaOXA-511 can be transferred from the chromosome to plasmid in A. baumannii, and hence, plasmid can be transferred from A. baumannii to K. pneumoniae.55 It was documented that blaOXA-51 was detected in K. pneumoniae through the case report conducted by Budak S et al.56
Although blaSME is a carbapenemase gene the carried mainly on the chromosome of Serratia marcescens, it was reported that such gene can be disseminated to other enterobacterial pathogens based on the study that conducted by Lee CR et al57 so, the current study investigated the existence of this gene in CRKP clinical isolates and it was found that no isolate harbored blaSME.
With regards to blaGES that was detected in only one isolate (KP03), it was found that this gene had no carbapenemase activity in which DNA sequencing results revealed that the detected variant was blaGES-1 which is well known to have only extended-spectrum β-lactamases (ESBL). Interestingly, blaGES was not detected in K. pneumoniae clinical isolates in Egypt, but instead it was detected in other Egyptian Gram-negative isolates like Acinetobacter baumannii51 and E. coli.16
The BOX-PCR fingerprint profile indicated that, although CRKP clinical isolates were recovered from different patients in different hospital departments, the circulating K. pneumoniae isolates in this hospital are highly genetically related, where 18 isolates belong to a single phylogenetic group (B).
A limitation of the present study was the limited number of carbapenem-resistant isolates that were tested for the 12 carbapenemase genes; thus, additional studies will be done to further document the prevalence of blaOXA-181 and blaOXA-23 in CRKP clinical isolates.
In conclusion, the current study is the first Egyptian report that addressed the coexistence of blaNDM-25, blaOXA-23, blaOXA-181, and blaGES-1 in CRKP clinical isolates that are genetically related based on DNA fingerprint profiles obtained by BOX-PCR. Also, the current study addressed the limitations of MHT and Carba NP test for the detection of carbapenemases in isolates that tested positive for blaNDM and OTCs.
Acknowledgment
The authors would like to thank the College of Pharmacy at Taif university. Also the authors thanks the AppliedMaths Company, Kortrijk, Belgium for their assistance and giving us the chance to use the trial version of BioNumeric 7.5 software through the assistance of King Fahd Armed Hospital (KFAH), Jeddah, KSA.
Disclosure
The authors declare no conflict of interest in this work.
References
1. Codjoe FS, Donkor ES. Carbapenem resistance: review. Med Sci. 2018;6(1). doi:10.3390/medsci6010001
2. Neuner EA, Yeh J-Y, Hall GS, et al. Treatment and outcomes in carbapenem-resistant Klebsiella pneumoniae bloodstream infections. Diagn Microbiol Infect Dis. 2011;69(4):357–362. doi:10.1016/j.diagmicrobio.2010.10.013
3. Huttner B, Jones M, Rubin MA, Neuhauser MM, Gundlapalli A, Samore M. Drugs of last resort? The use of polymyxins and tigecycline at US Veterans Affairs medical centers, 2005–2010. PLoS One. 2012;7(5):e36649. doi:10.1371/journal.pone.0036649
4. El-Badawy MF, Abdelwahab SF, Alghamdi SA, Shohayeb MM. Characterization of phenotypic and genotypic traits of carbapenem-resistant Acinetobacter baumannii clinical isolates recovered from a tertiary care hospital in Taif, Saudi Arabia. Infect Drug Resist. 2019;12:3113–3124. doi:10.2147/IDR.S206691
5. Wolfensberger A, Kuster SP, Marchesi M, Zbinden R, Hombach M. The effect of varying multidrug-resistance (MDR) definitions on rates of MDR Gram-negative rods. Antimicrob Resist Infect Control. 2019;8(1). doi:10.1186/s13756-019-0614-3
6. Pattnaik D, Panda SS, Singh N, Sahoo S, Mohapatra I, Jena J. Multidrug resistant, extensively drug resistant and pan drug resistant Gram-negative bacteria at a tertiary care center in Bhubaneswar. Int J Community Med Public Heal. 2019;6(2):567–572. doi:10.18203/2394-6040.ijcmph20190170
7. Meletis G. Carbapenem resistance: overview of the problem and future perspectives. Ther Adv Infect Dis. 2016;3(1):15–21. doi:10.1177/2049936115621709
8. Naas T, Dortet L,I, Iorga B. Structural and functional aspects of class A carbapenemases. Curr Drug Targets. 2016;17(9):1006–1028. doi:10.2174/1389450117666160310144501
9. Brouwer MS, Tehrani KH, Rapallini M, et al. Novel carbapenemases FLC-1 and IMI-2 encoded by an Enterobacter cloaca complex isolated from food products. Antimicrob Agents Chemother. 2019;63(6):e02338–02318. doi:10.1128/AAC.02338-18
10. Antunes NT, Lamoureaux TL, Toth M, Stewart NK, Frase H, Vakulenko SB. Class D β- lactamases: are they all carbapenemases? Antimicrob Agents Chemother. 2014;58(4):2119–2125. doi:10.1128/AAC.02522-13
11. Antunes NT, Fisher JF. Acquired class D β-lactamases. Antibiotics. 2014;3(3):398–434. doi:10.3390/antibiotics3030398
12. Abbas HA, Kadry AA, Shaker GH, Goda RM. Impact of specific inhibitors on metallo-β- carbapenemases detected in Escherichia coli and Klebsiella pneumoniae isolates. Microb Pathog. 2019;132:266–274. doi:10.1016/j.micpath.2019.05.022
13. Yong D, Lee K, Yum JH, Shin HB, Rossolini GM, Chong Y. Imipenem- EDTA disk method for differentiation of metallo-β-lactamase-producing clinical isolates of Pseudomonas spp. and Acinetobacter spp. J Clin Microbiol. 2002;40(10):3798–3801. doi:10.1128/jcm.40.10.3798-3801.2002
14. Van Belkum A, Hermans PW. BOX PCR fingerprinting for molecular typing of Streptococcus pneumoniae. In: Antibio Resist. Springer; 2001:159–168. doi:10.1385/1-59259-077-2:159
15. Brusetti L, Malkhazova I, Gtari M, et al. Fluorescent-BOX-PCR for resolving bacterial genetic diversity, endemism and biogeography. BMC Microbiol. 2008;8(1):220. doi:10.1186/1471-2180-8-220
16. El-Badawy MF, Tawakol WM, El-Far SW, et al. Molecular identification of aminoglycoside-modifying enzymes and plasmid-mediated quinolone resistance genes among klebsiella pneumoniae clinical isolates recovered from Egyptian patients. Inter J Microbiol. 2017;2017:1–12. doi:10.1155/2017/8050432
17. Vandepitte J, Verhaegen J, Engbaek K, et al. Basic Laboratory Procedures in Clinical Bacteriology. WHO; 2003.
18. Qaiyumi S. Macro-and microdilution methods of antimicrobial susceptibility testing. In: Antimicrobial Susceptibility Testing Protocols. Taylor & Francis; 2007:75–79.
19. Patel J, Cockerill F, Alder J, et al. National Committee for Clinical Laboratory Standards. Performance Standards for Antimicrobial Susceptibility Testing. 2014.
20. Othman HB, Halim RMA, Abdul-Wahab HEE-A, Atta HA, Shaaban O. Pseudomonas aeruginosa-modified Hodge test (PAE-MHT) and ChromID Carba agar for detection of carbapenemase producing Pseudomonas aeruginosa recovered from clinical specimens. Macedo J Med Sci. 2018;6(12):2283–2289. doi:10.3889/oamjms.2018.414
21. Campana EH, Chuster SG, da Silva IR, et al. modified Carba NP test for the detection of carbapenemase production in Gram-negative rods: optimized handling of multiple samples. Braz J Microbiol. 2017;48(2):242–245. doi:10.1016/j.bjm.2016.09.015
22. Rudresh SM, Ravi GS, Sunitha L, Hajira SN, Kalaiarasan E, Harish BN. Simple, rapid, and cost-effective modified Carba NP test for carbapenemase detection among Gram- negative bacteria. J Lab Physician. 2017;9(4):303–307. doi:10.4103/JLP.JLP_138_16
23. Sachdeva R, Sharma B, Sharma R. Evaluation of different phenotypic tests for detection of metallo-β-lactamases in imipenem-resistant Pseudomonas aeruginosa. J Lab Physician. 2017;9(4):249–253. doi:10.4103/JLP.JLP_118_16
24. Van Dijk K, Voets G, Scharringa J, et al. A disc diffusion assay for detection of class A, B and OXA-48 carbapenemases in Enterobacteriaceae using phenyl boronic acid, dipicolinic acid and temocillin. Clin Microbiol Infect. 2014;20(4):345–349. doi:10.1111/1469-0691.12322
25. Yang Y, Rasmussen BA, Shlaes DM. Class A β-lactamases-enzyme-inhibitor interactions and resistance. Pharma& Therap. 1999;83(2):141–151. doi:doi.10.1016/S0163-7258(99)00027-3
26. Stojanoski V, Chow D-C, Fryszczyn B, et al. Structural basis for different substrate profiles of two closely related class D β-lactamases and their inhibition by halogens. Biochemi. 2015;54(21):3370–3380. doi:10.1021/acs.biochem.5b00298
27. Dobrzaniecka K, Młynarczyk A, Szymanek-Majchrzak K, Młynarczyk G. Comparison of phenotypic methods for the detection of beta-lactamases MBL in strains from the Enterobacteriaceae family and non-fermentative bacilli isolated from clinical specimens. Med Dosw Mikrobiol. 2014;66(3–4):177–184.
28. Dashti AA, Jadaon MM, Abdulsamad AM, Dashti HM. Heat treatment of bacteria: a simple method of DNA extraction for molecular techniques. Kuwait Med J. 2009;41(2):117–122.
29. El-Badawy MF, Tawakol WM, Maghrabi IA, Mansy MS, Shohayeb MM, Ashour MS. Iodometric and molecular detection of ESBL production among clinical isolates of E. coli fingerprinted by ERIC-PCR: the first Egyptian report declares the emergence of E. coli O25b-ST131clone harboring blaGES. Microbiol Drug Resist. 2017;23(6):703–717. doi:10.1089/mdr.2016.0181
30. Waterhouse AM, Procter JB, Martin DM, Clamp M, Barton GJ. Jalview Version 2-a multiple sequence alignment editor and analysis workbench. Bioinformatics. 2009;25(9):1189–1191. doi:10.1093/bioinformatics/btp033
31. Moemen D, Masallat DT. Prevalence and characterization of carbapenem-resistant Klebsiella pneumoniae isolated from intensive care units of Mansoura university hospitals. Egypt J Bas Appli Sci. 2017;4(1):37–41. doi:10.1016/j.ejbas.2017.01.001
32. Sheu -C-C, Chang Y-T, Lin S-Y, et al. Infections caused by carbapenem-resistant Enterobacteriaceae: an update on therapeutic options. Front Microbiol. 2019;10(80). doi:10.3389/fmicb.2019.00080
33. Lledo W, Hernandez M, Lopez E, et al. Guidance for control of infections with carbapenem-resistant or carbapenemase-producing Enterobacteriaceae in acute care facilities. MMWR Morb Mortal Wkly Rep CDC. 2009;58(10):256–260.
34. Wiener-Well Y, Rudensky B, Yinnon A, et al. Carriage rate of carbapenem-resistant Klebsiella pneumoniae in hospitalized patients during a national outbreak. J Hosp Infect. 2010;74(4):344–349. doi:10.1016/j.jhin.2009.07.022
35. Metwally L, Gomaa N, Attallah M, Kamel N. High prevalence of Klebsiella pneumoniae carbapenemase-mediated resistance in K. pneumoniae isolates from Egypt. East Mediterr Health J. 2013;19(11):947–952. doi:10.26719/2013.19.11.947
36. Moghadampour M, Rezaei A, Faghri J. The emergence of blaOXA-48 and blaNDM among ESBL-producing Klebsiella pneumoniae in clinical isolates of a tertiary hospital in Iran. Acta Microbiol Immunol Hung. 2018;65(3):335–344. doi:10.1556/030.65.2018.034
37. Bi W, Liu H, Dunstan RA, et al. Extensively drug resistant Klebsiella pneumoniae causing nosocomial bloodstream infections in China: molecular investigation of antibiotic resistance determinants, informing therapy, and clinical outcomes. Front Microbiol. 2017;8:1230. doi:10.3389/fmicb.2017.01230
38. Patel JB, Weinstein MP, Eliopoulos GM, et al. Carba NP Test for Suspected Carbapenemase Production in Enterobacteriaceae, Pseudomonas Aeruginosa, and Acinetobacter Spp. Clinical and Laboratory Standards Institute (CLSI). CLSI Supplement M100.
39. Morey KE, Vega R, Cassidy PM, et al. Evaluation of the Carba NP test in Oregon, 2013. Antimicrob Agents Chemother. 2017;61(1):e03005–03015. doi:10.1128/AAC.03005-15
40. AlTamimi M, AlSalamah A, AlKhulaifi M, AlAjlan H. Comparison of phenotypic and PCR methods for detection of carbapenemases production by Enterobacteriaceae. Saudi J Biol Sci. 2017;24(1):155–161. doi:10.1016/j.sjbs.2016.07.004
41. Bayramoğlu G, Ulucam G, Gençoğlu ÇÖ, Kılıç A, Aydın F. Comparison of the modified Hodge test and the Carba NP test for detection of carbapenemases in Enterobacteriaceae isolates. Mikrobiyol Bu. 2016;50(1):1–10. doi:10.5578/mb.10861
42. Pournaras S, Poulou A, Tsakris A. Inhibitor-based methods for the detection of KPC carbapenemase-producing Enterobacteriaceae in clinical practice by using boronic acid compounds. J Antimicrob Chemother. 2010;65(7):1319–1321. doi:10.1093/jac/dkq124
43. Kahraman EP, Toptan H, Otlu B, Köroğlu M, Altındiş M. Investigation of blaOXA-48-like genes in carbapenemase producing Klebsiella spp. isolates. Mikrobiyol Bul. 2019;53(2):134–143. doi:10.5578/mb.67914
44. Al-Zahrani IA, Alasiri BA. The emergence of carbapenem-resistant Klebsiella pneumoniae isolates producing OXA-48 and NDM in the Southern (Asir) province, Saudi Arabia. Saudi Med J. 2018;39(1):23–30. doi:10.15537/smj.2018.1.21094
45. Shankar C, Mathur P, Venkatesan M, et al. Rapidly disseminating blaOXA-232 carrying Klebsiella pneumoniae belonging to ST231 in India: multiple and varied mobile genetic elements. BMC Microbiol. 2019;19(1):137. doi:10.1186/s12866-019-1513-8
46. Nahid F, Zahra R, Sandegren L. A blaOXA-181harbouring multi-resistant ST147 Klebsiella pneumoniae isolate from Pakistan that represent an intermediate stage towards pan-drug resistance. PLoS One. 2017;12(12):e0189438. doi:10.1371/journal.Pone
47. Satter S, Mahbub H, Shamsuzzaman S. OXA-181-an emerging threat in Bangladesh. J Mole Stud Medici Res. 3(01):128–134. doi:10.18801/jmsmr.030118.15
48. Zhu C, Liyanapathirana V, Li C, et al. Characterizing mobilized virulence factors and multidrug resistance genes in carbapenemase-producing Klebsiella pneumoniae in a Sri Lankan hospital. Front Microbiol. 2018;9:2044. doi:10.3389/fmicb.2018.02044
49. Uwaezuoke NS, Kieffer N, Iregbu KC, Nordmann P. First report of OXA-181 and NDM- 1 from a clinical Klebsiella pneumoniae isolate from Nigeria. Int J Infect Dis. 2017;61:1–2. doi:10.1016/j.ijid.2017.05.004
50. Lopes BS, Al-Agamy MH, Ismail MA, et al. The transferability of blaOXA-23 gene in multidrug-resistant Acinetobacter baumannii isolates from Saudi Arabia and Egypt. Int J Med Microbiol. 2015;305(6):581–588. doi:10.1016/j.ijmm.2015.07.007
51. Al-Agamy MH, Khalaf NG, Tawfick MM, Shibl AM, El Kholy A. Molecular characterization of carbapenem-insensitive Acinetobacter baumannii in Egypt. Int J Infect Dis. 2014;22:49–54. doi:10.1016/j.ijid.2013.12.004
52. El Bannah AMS, Nawar NN, Hassan RMM, Salem STB. Molecular epidemiology of carbapenem-resistant Acinetobacter baumannii in a tertiary care hospital in Egypt: clonal spread of blaOXA-23. Microb Drug Resist. 2018;24(3):269–277. doi:10.1089/mdr.2017.0057
53. Yousfi K, Touati A, Lefebvre B, et al. Characterization of multidrug-resistant Gram-negative bacilli isolated from hospitals effluents: first report of a blaOXA-48-like in Klebsiella oxytoca, Algeria. Braz J Microbiol. 2019;50(1):175–183. doi:10.1007/s42770-018-0010-9
54. Evans BA, Amyes SG. OXA-type β-lactamases. Clin Microbiol Rev. 2014;27(2):63–72. doi:10.1128/CMR.00117-13
55. Chen TL, Lee YT, Kuo SC, et al. Emergence and distribution of plasmids bearing the blaOXA-51-like gene with an upstream ISAba1 in carbapenem-resistant Acinetobacter baumannii isolates in Taiwan. Antimicrob Agents Chemother. 2010;54(11):4575–4581. doi:10.1128/AAC.00764-10
56. Budak S, Aktaş Z, Oncul O, et al. Detection of OXA-51 Carbapenemase Gene in Klebsiella pneumoniae: a Case Report and a New Dimension on Carbapenemase Resistance. J Mol Genet Med. 2013;7:1747–0862.1000. doi:10.4172/1747-0862.1000063
57. Lee CR, Lee JH, Park KS, et al. Global dissemination of carbapenemase-producing Klebsiella pneumoniae: epidemiology, genetic context, treatment options, and detection methods. Front Microbiol. 2016;7:895. doi:10.3389/fmicb.2016.0089
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.