Back to Journals » International Journal of Nanomedicine » Volume 15
The Efficacy of AgNO3 Nanoparticles Alone and Conjugated with Imipenem for Combating Extensively Drug-Resistant Pseudomonas aeruginosa
Authors Shahbandeh M , Taati Moghadam M, Mirnejad R, Mirkalantari S, Mirzaei M
Received 29 April 2020
Accepted for publication 31 August 2020
Published 21 September 2020 Volume 2020:15 Pages 6905—6916
DOI https://doi.org/10.2147/IJN.S260520
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 5
Editor who approved publication: Prof. Dr. Anderson Oliveira Lobo
Mahsa Shahbandeh,1 Majid Taati Moghadam,2,3 Reza Mirnejad,4 Shiva Mirkalantari,5 Mehrnaz Mirzaei6
1Young Researchers and Elite Club, Saveh Branch, Islamic Azad University, Saveh, Iran; 2Department of Microbiology, Iran University of Medical Sciences, Tehran, Iran; 3Student Research Committee, Iran University of Medical Sciences, Tehran, Iran; 4Molecular Biology Research Center, System Biology and Poisoning Institute, Baqiyatallah University of Medical Sciences, Tehran, Iran; 5Microbiology, Faculty of Medicine, Microbiology Department, Iran University of Medical Sciences, Tehran, Iran; 6Department of Microbiology, Tehran Medical Sciences Branch, Islamic Azad University, Tehran, Iran
Correspondence: : Reza Mirnejad
Molecular Biology Research Center, Systems Biology and Poisonings Institute, Baqiyatallah University of Medical Sciences, Tehran, Iran
Email [email protected]
Shiva Mirkalantari
Microbiology, Faculty of Medicine, Microbiology Department, Iran University of Medical Sciences, Tehran, Iran
Email [email protected]
Introduction: The extensive drug-resistant (XDR) Pseudomonas aeruginosa (P. aeruginosa) causes a range of infections with high mortality rate, which inflicts additional costs on treatment. The use of nano-biotechnology-based methods in medicine has opened a new perspective against drug-resistant bacteria. The aim of this study was to evaluate the effectiveness of the AgNO3 nanoparticles alone and conjugated with imipenem (IMI) to combat extensively drug-resistant P. aeruginosa.
Methods: Antibiotic susceptibility was carried out using disc diffusion method. Detection of different resistant genes was performed using standard polymerase chain reaction (PCR). The chemically synthesized AgNO3 particles were characterized using scanning electron microscope (SEM), dynamic light scattering (DLS) and X-ray diffraction (XRD) methods. Fourier transform infrared spectroscopy (FTIR) was accomplished to confirm the binding of AgNO3 with IMI. The microdilution broth method was used to obtain minimum inhibitory concentration (MIC) of AgNO3 and IMI-conjugated AgNO3. MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay was carried out on L929 cell line to study the cytotoxicity of nanoparticles. The data were analyzed by Eta correlation ratio and chi-square (X2) test.
Results: Analysis of the antibiotic resistance pattern showed that 12 (24%) isolates were XDR, and MIC values of IMI were between 64 and 128 μg/mL. Frequency of SHV, TEM, CTX M, IMP, VIM, OPR, SIM, SPM, GIM, NDM, VEB, PER, KPC, OXA, intI, intII, and intIII genes were 29 (58%), 26 (52%), 26 (52%), 32 (64%), 23 (46%), 43 (86%), 3 (6%), 6 (12%), 3 (6%), 4 (8%), 7 (14%), 6 (12%), 18 (36%), 4 (8%), 19 (38%), 16 (32%), and 2 (4%), respectively. The XRD, SEM, DLS, and FTIR analysis confirmed the synthesis of AgNO3 nanoparticles and their conjugation with IMI. The AgNO3 nanoparticles had antimicrobial activity, and their conjugation with IMI showed enhanced effectiveness against XDR isolates. The synthesized AgNO3 showed no cytotoxic effects.
Conclusion: The results suggest that IMI-conjugated AgNO3 has a strong potency as a powerful antibacterial agent against XDR P. aeruginosa.
Keywords: P. aeruginosa, AgNO3 nanoparticle, XDR, imipenem
Introduction
As shown in the earlier studies, P. aeruginosa is a non-fermenting gram-negative bacterium that correlates clinically and epidemiologically with nosocomial infections. Today, the incidence of antibiotic-resistant P. aeruginosa infections has been increased all over the world.1,2 Although various antibiotic-resistant mechanisms have been identified in P. aeruginosa, one of the most notable mechanisms is extended-spectrum beta-lactamase (ESBL) such as TEM, CTXM, and SHV. Carbapenems are commonly used antibiotics against ESBL-producing P. aeruginosa infection, but today, production of carbapenemases and metallobetalactamases including IMP, VIM, KPC, SIM, GIM, PER, VEB, SPM, OXA143, and NMD have led to the resistance to these antibiotics. By carrying these resistance genes and other mechanisms, P. aeruginosa can become resistant to a wide range of antibiotics and produce multidrug-resistant (MDR) and extensively drug-resistant (XDR) isolates.3–5 Therefore, the treatment of such isolates in hospitals has resulted in several problems for specialists. The MDR isolates are bacteria that are resistant to at least one agent in three or more classes of antimicrobial agents, but XDR isolates are sensitive to only one or two groups of antimicrobials.4 Numerous reports have shown that Carbapenem antibiotics are one of the most effective agents in the treatment of infections caused by gram-negative bacilli. Several characteristics including high affinity to Protein Binding Penicillin 2, excellent permeability across bacterial membranes, and stability against most β-lactamases makes carbapenems as potent antibiotics against gram-negative bacteria.6,7 Although carbapenems are one of the most commonly used antibiotics against drug-resistant P. aeruginosa infection, these antibiotics have lost their effectiveness with advent of XDR P. aeruginosa isolates. The XDR isolates of the P. aeruginosa have been found in different countries, and their clinical impact has been reviewed, however, combat with this challenge is a major concern.8 Other important resistance mechanisms are integrons (int1, int2, and int3) and specialized porin or efflux pumps genes (Opr) which cause resistance to a wide range of antibiotics.9,10 Nosocomial infections by antibiotic-resistant bacteria resulted in 25,000 death cases, an increase of 2.5 days in the hospitalization, and an estimated 1.5 billion € extra costs in Europe per year.11,12 Meanwhile, the United States reports showed that P. aeruginosa infections have put a heavy burden on hospitals, including mortality, morbidity, and health-care costs. High-mortality P. aeruginosa infections (18% to 60%) increase the cost of treatment between $ 20,000 and $ 80,000 per infection.13 Therefore, researchers are looking for new methods for treating antibiotic-resistant bacteria in that nanoparticles are one of the most important solutions.1,3,14,15 The use of nano-biotechnology-based methods in medicine has opened a new perspective against pathogenic bacteria. In order to achieve this purpose, first, the antimicrobial activity of the nanoparticles must be measured, and then, if possible, used for pharmaceutical applications. The chemical and physical properties of nanoparticles offer researchers the opportunity to use them in various medical fields.16 In recent years, many studies have been conducted on the use of silver nanoparticles against antibiotic-resistant bacteria that cause nosocomial infections. In a study by Bardania et al, the antimicrobial effect of silver nanoparticles was reported against antibiotic-resistant P. aeruginosa and S. aureus isolated from the clinical sample.17 Also, Shariati et al showed that silver nanoparticles inhibited MDR isolates of P. aeruginosa and reduced the expression of virulence genes in the bacterium.1 The purpose of this study was to use AgNO3 nanoparticles alone and conjugated with imipenem for combating extensively drug-resistant P. aeruginosa.
Materials and Methods
Chemicals and Apparatus
The reagents including Tryptic Soy Broth (TSB), Stuart transport medium, Mueller–Hinton agar (MHA), and Mueller–Hinton broth (MHB), blood, and MacConkey agar and Dimethyl Sulfoxide (DMSO) were purchased from Merck (Germany). Antibacterial disks and antibiotic powders were provided by Mast (Diagnostics, UK) and Sigma-Aldrich (St. Louis, MO, USA), respectively. The synthetic AgNO3 was also purchased from Sigma-Aldrich. All chemicals were of analytical reagent grade and used as received with no further purification.
Bacterial Isolates
The current study was conducted in Microbiology Section of the Central Laboratory of Iran University of Medical Sciences. A total of 50 clinical isolates of P. aeruginosa from blood, urine, trachea, sputum, and burn wounds were collected from a part of the routine laboratory procedure of hospitals in Tehran. It should be noted that the sputum specimens obtained from the patients were microscopically examined for the presence of salivary contamination. Briefly, sputum was incubated with an equal volume of sputolysin for 15 min at room temperature and plated onto chocolate agar. The deep oropharyngeal swab samples were collected and cultured on blood agar and MacConkey agar media. Wound swabs were added to Stuart transport medium, then immediately transported to the Department of Microbiology, Iran University of Medical Sciences, and inoculated onto TSB, blood, and MacConkey agar for 24 hr. All positive cultures were recognized by the related biochemical tests, including gram-negative bacilli, the ability to grow at 42°C, colony characteristics, motility and oxidase test, citrate utilization, OF-glucose, pigment production on Mueller–Hinton agar and non-fermenting in TSI agar.18 The reference strain used as positive control was P. aeruginosa ATCC 27,853. The isolates identified as P. aeruginosa were preserved at –80°C in TSB supplemented with 30% glycerol until further processing.
Antimicrobial Susceptibility Test and β-Lactamase Determination
Antibiotic susceptibility test was carried out using disc diffusion method according to the Clinical and Laboratory Standard Institute guideline (CLSI). Antibiotic disks used in the current study contained ceftazidime, chloramphenicol, cefotaxime, amikacin, co-trimoxazole, ceftriaxone, ciprofloxacin, meropenem, gentamicin, and tetracycline. P. aeruginosa isolates that were found resistant to IMI by disk diffusion test were re-checked in broth microdilution assay. The MIC for IMI was obtained in the range of 0.5 μg/mL to 256 μg/mL. P. aeruginosa ATCC 27,853 was used as control strain. Double disc synergy test was used for phenotypic ESBL-producing P. aeruginosa detection. In this test, antibiotics, including ceftazidime and cefotaxime, were used alone and adjacent to all two antibiotics combination with clavulanic acid.19
Antibiotic-Resistant Genes Genotyping
Detection of different genes involved in antibiotic resistance including VIM, IMP, OPR, SPM, SIM, GIM, NDM, SHV, TEM, VEB, PER, CTXM, OXA, intI, intII, intII and KPC were performed using standard PCR amplification. DNA was extracted using DNA extraction kit (GeNet Bio Company, Daejeon, Korea) according to the manufacturer’s instruction. PCR reaction was performed in a final volume of 25 µL using 3 µL DNA solution, 12.5 µL Qiagen HotStar Taq Polymerase Master Mix, 10 pmol primer and water. PCR cycling conditions were as follows: denaturation at 94°C for 5 min, followed by 30 amplification cycles at 94°C for 1 min and extension at 72°C for 40 s and a final extension at 72°C for 5 min.20 All primers used in the present study are listed in Table 1. Finally, PCR products were detected by electrophoresis on 1%-1.5% agarose gel stained with safe stain.
 |
Table 1 Primers Used for Detection of Resistance Genes in P. aeruginosa Isolates |
Synthesis of AgNO3
The synthetic AgNO3 had an average purity of 99.7% metals base. Then, 0.05 mg AgNO3 was added to 5 mL sterile distilled water containing 5% polyethylene glycol (PEG). The reason for the use of PEG was to disperse the colloidal nanoparticles, which had no inhibitory effect on the concentration used above. Subsequently, AgNO3 nanoparticles were incubated at 37°C for 30 minutes and homogenized by sonication at 100 w, at 40 kHz to disperse the nanoparticles. Then, the dispersed solutions were autoclaved at 121°C for 15 min and stored at 4°C. The morphology of Ag nanoparticles was evaluated by SEM.
Analysis of Synthesized Nanoparticles
The SEM analysis was conducted using Hitachi S-4500 SEM machine. To perform this evaluation, thin films of the sample were placed on a carbon-coated copper grid by dropping a very small amount of the sample on the grid. The extra solution was removed using a blotting paper, and the films were allowed to dry by putting them under a mercury lamp for 5 min. Also, using DLS equipment on a polystyrene cuvette, the size of the nanoparticles was examined in 3 mL of the reaction mixture. The XRD method was used to identify the phase formation of nanoparticles under different conditions. The samples were prepared as powder and recorded on an X-ray diffractometer (model PW1730, PHILIPS company, Netherlands) with CuKα source radiation (λ=1.5418 Å), accelerator voltage kV40, and current mA30. The sample was analyzed in the range of 5 to 70 degrees.
Conjugation of AgNO3 with Imipenem
The FTIR was performed to confirm the binding of AgNO3 with IMI. The amount of 20 mg of nanoparticles were added to 50 mL distilled water at room temperature under magnetic stirring. Furthermore, 25 mg IMI was poured in 5 mL distilled water and dissolved gently by magnetic stirring. After complete dissolution, the prepared IMI aqueous solution was mixed with AgNO3 suspension under gentle stirring at 25°C. It was kept at room temperature for 24 hr. Then, dissolved suspension was centrifuged at 4000 rpm for about 15–20 min, and the supernatant was separated. Eventually, the resulting solid product was collected by centrifugation and dried at room temperature overnight. The dried structure of the sample was analyzed using FTIR spectrometer in the 400–4000 cm−1 wavenumber region.
Antibacterial Activity Test
Microdilution broth method was used according to CLSI to obtain MIC of nanoparticles and IMI-conjugated AgNO3. To prepare nanoparticles and IMI-conjugated AgNO3 stock solution, 1.024 mg of each substance was completely dissolved in 1 mL sterile water. Then, the stock solution was serially diluted in the range of 0.5–1024 μg/mL, and the nanoparticles and IMI-conjugated AgNO3 dilutions were added to the wells of 96-well plates starting at 256 μg/mL. In the next step, each well was inoculated with 10 μL (5×106 CFU/mL) of the XDR isolates of the P. aeruginosa in Mueller-Hinton broth to a final concentration (5×105 CFU/mL). The positive and negative controls were P. aeruginosa PAO1 and media alone, respectively. The lowest concentration of nanoparticles and IMI-conjugated AgNO3 that prevented the growth of XDR isolates of the P. aeruginosa and did not cause opacity in the well after 24 hr incubation at 37°C was determined as MIC.
Cytotoxicity Assay of Nanoparticles
Cytotoxicity assay was carried out on human epithelial cell lines (L929 ATCC CCL-6364TM purchased from Pasteur Institute of Iran) to study the cytotoxicity level of nanoparticles. To perform the cytotoxicity assay, L929 cell lines were cultured in RPMI 1640 medium (Biosera, USA) supplemented with 10% fetal bovine serum (FBS), 1% L-glutamic acid, 1% non-essential amino acid, and 1% penicillin-streptomycin and incubated at 37°C in CO2 humidified incubator. For cell viability studies, the cells were seeded in 96-well plates (50,000 cells/well). Following 24 hr incubation, the cells reached 80% confluency. Subsequently, different concentrations of nanoparticles were added to the microtiter wells to evaluate the mitochondrial functions of the cells by MTT assay. After 24 hr of incubation, the cell morphology was evaluated with an Olympus IX70 Inverted Microscope. Then, 20 μL of 10X MTT purchased from Roche (Mannheim, Germany) was directly added to the wells. After 4 hr incubation in dark and dissolving formazan crystals in DMSO, the absorbance was obtained at 570 nm with a standard microplate reader.21 The viability of the cells exposed to different concentrations of nanoparticles was measured.
Statistical Analysis
All data were analyzed using SPSS 22. The interpretation of the results was based on the frequencies. Eta correlation ratio was exerted to investigate the relationship between the age of patients and different antibiotic resistance rates. The P-value less than 0.05 was considered statistically significant for association between all antibiotics, resistance genes, type of samples, and clinical isolates by the chi-square (X2) test.
Results
Characteristics of Infectious Samples and Patients
A total of 50 isolates from February 2018 to June 2018 were collected from Tehran hospitals. The specimens were collected from different wards of the hospitals including Burn, Infectious, Intensive Care Unit (ICU), Orthopedic, Gynecological and Outpatient Wards. The number of P. aeruginosa isolates from blood, urine, trachea, sputum, burn wounds and bedsore were 8 (16%), 13 (26%), 4 (8%), 9 (18%), 9 (18%) and 7 (14%), respectively. Eight (16%) isolates were obtained from outpatients and 42 (84%) isolates were from hospitalized patients. The age range of the patients infected with P. aeruginosa was 21 to 87 years (58.73±19.96). The prevalence rate of P. aeruginosa differed among men and women; 20 samples were isolated from females and 30 samples were isolated from males.
Prevalence of ESBL-Producing P. aeruginosa and Antimicrobial Susceptibility
The results of antibiogram showed that the most effective antibiotic against P. aeruginosa isolates was ciprofloxacin, and the highest resistance was observed to cefotaxime. The resistance of P. aeruginosa isolates to all antibiotics was found higher than 50%. The details of the antibiogram results can be seen in Table 2. By disk diffusion test, carbapenem antibiotics susceptibility was evaluated for all the isolates, and the results showed among 50 P. aeruginosa, 25 (50%), and 31 (62%) isolates were resistant to meropenem and imipenem, respectively. The results from MIC demonstrated that all the 31 isolates were resistant to IMI with MIC between 32 and 128 μg/mL. The statistical analysis of data indicated that there was a significant relationship between resistance to all antibiotics and age of the patients. On the other hand, some antibiotics such as meropenem, ceftriaxone, co-trimoxazole, chloramphenicol, and tetracycline, had a significant correlation with the type of samples, especially urine, bed ulcer, and sputum. Different wards of the hospitals, especially the ICU ward, also had a direct relationship with the resistance to a number of antibiotics such as ciprofloxacin, gentamicin, amikacin, meropenem, ceftriaxone, chloramphenicol, and tetracycline. Combination disk test results showed that 45 (90%) isolates produced ESBL.
 |
Table 2 Antibiotics Resistance Pattern of P. aeruginosa Clinical Isolates |
Prevalence and Characterization of Resistance Genes
Among 45 isolates, only 8 (16%) isolates had all three genes (SHV, TEM, and CTXM) together, and 29 (58%) had SHV gene, 26 (52%) had TEM gene, and 26 (52%) carried CTXM gene. The frequency of other resistance genes in P. aeruginosa isolates was different. Among 50 P. aeruginosa isolates, 32 (64%), 23 (46%), 43 (86%), 3 (6%), 6 (12%), 3 (6%), 4 (8%), 7 (14%), 6 (12%), 18 (36%), 4 (8%), 19 (38%), 16 (32%), and 2 (4%) isolates contained IMP, VIM, OPR, SIM, SPM, GIM, NDM, VEB, PER, KPC, OXA, intI, intII, and intIII genes, respectively. Figure 1A shows the band size and location of the resistant genes on agarose gel. The data analysis showed a significant relationship between the presence of all resistance genes and the age of the patients (P<0.05). It was also found that resistance genes were more present in the samples isolated from ICU, and a direct relationship was observed between the hospital ward and the presence of these genes (P<0.05). The specimens isolated from wound, urine, and sputum had higher resistance genes, which showed a significant correlation between the presence of these genes and the type of samples (P<0.05).
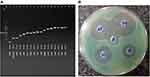 |
Figure 1 (A) The location of resistance genes on electrophoresis gel (B) Antimicrobial activity of AgNO3 nanoparticles against P. aeruginosa. |
Using antibiotics resistance pattern analysis, 12 (24%) isolates were identified as XDR, which were resistant to all antibiotics used in this study. Also, there was a significant relationship between XDR isolates and IMP gene (P<0.05), and the highest number of XDR isolates was isolated from ICU. A significant relationship was observed between these antibiotic-resistant isolates and different parts of the hospital.
Characterization and Antibacterial Effect of AgNO3 on Resistant P. aeruginosa
All XDR isolates were tested to investigate the antimicrobial effects of nanoparticles and IMI-conjugated AgNO3. The SEM analysis confirmed different sizes and shapes of the nanoparticles, as can be seen in Figure 2. According to the SEM image in Figure 2, AgNO3 nanoparticles with a cylindrical shape and diameters of 15–50 nm were fabricated. Also, DLS was employed to obtain the size distribution of AgNO3 nanoparticles. The results demonstrated that the AgNO3 nanoparticles had a very narrow size distribution with an average particle size of about 25 nm. The XRD for the AgNO3 nanoparticles showed that the scattering peaks at angles 37.34, 43.61, and 63.51 were related to the scattering plates (111), (200), and (220), respectively, and were matched with the structure of AgNO3 nanoparticles with the card number JCPDS (04–0783). This confirms the correct synthesis of silver nanoparticles (Figure 3).22,23
 |
Figure 2 AgNO3 Nanoparticles with magnify (A) 40000× and (B) 80000×. |
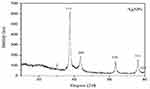 |
Figure 3 XRD pattern of produced silver nanoparticles. Three intense peaks at 37.34°, 43.61°, and 63.51° revealed the presence of produced silver nanoparticles. |
The XDR isolates were resistant to high concentrations of IMI. The antimicrobial activity of the nanoparticles showed that the nanoparticles had antimicrobial activity at low concentrations (Table 3 and Figure 1B). The conjugation of AgNO3 with IMI showed greater antimicrobial effect on XDR isolates compared to AgNO3 alone. These XDR isolates carried different resistance genes; Table 3 shows the details of these genes and the MIC. Data analysis revealed no significant relationship between the presence of resistance genes and XDR isolates.
 |
Table 3 Effect of Nanoparticles Alone and Conjugated with Imipenem Against XDR Isolates of P. aeruginosa |
FTIR spectroscopy was used to investigate the interaction between IMI and AgNO3. Figure 4 shows the presence of O-H stretching bond in the range 3400 cm−1, which was related to water absorption in all samples. The presence of absorption bonds in the range of 3422 cm−1, 2921 cm−1, 2856 cm−1, 1743 cm−1, 1631 cm−1, 1450 cm−1, 1377 cm−1, 1240 cm−1, 1043 cm−1 and 596 cm−1 confirms the presence of AgNO3 nanoparticles (Figure 4B). The bonds in the range of 1368 cm−1 and 1092 cm−1 are characteristic of the absorption bonds of IMI (Figure 4C). These peaks are present in the IMI-conjugated AgNO3 and confirm the interaction between the two substances, which is related to the strong S=O stretching bond (Figure 4A). On the other hand, a minor shift in the peaks in the IMI-conjugated AgNO3 sample indicates chemical bonding or electrostatic adsorption between IMI and AgNO3 nanoparticles. Importantly, the presence of a sulfide group on the surface of IMI has made it predispose to form a bond with AgNO3 nanoparticles. As a result, both components are present in the IMI-conjugated AgNO3, and this observation confirms the correct synthesis of this compound.
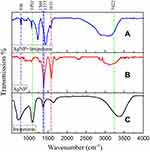 |
Figure 4 FTIR spectra of imipenem and imipenem conjugated AgNO3. As shown in this figure, AgNO3 nanoparticles bond to sulfide groups of imipenem, (A) AgNP+Imipenem, (B) AgNP, (C) Imipenem. |
Cytotoxicity Effect of AgNO3 Nanoparticles
The results of cytotoxic activity of AgNO3 nanoparticles, IMI, IMI-conjugated AgNO3 (AgNO3+IMI), and polystyrene tissue culture (TCPS) in L929 ATCC CCL-6364TM cell line demonstrated that cells exposed to nanoparticles were well proliferated compared to the TCPS. Since IMI alone was more toxic than TCPS, when combined with nanoparticles, its toxic property reduced significantly (Figure 5). The nanoparticles alone did not exhibit any toxicity and changes in cell morphology until 5 days; however, the AgNO3+IMI on day 5 caused minor changes in the cell morphology compared to the TCPS.
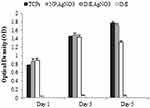 |
Figure 5 MTT assay results confirmed the in vitro cytotoxicity of silver nanoparticles against L929 ATCC CCL-6364TM cell lines after 1, 3, and 5 days exposures. |
Discussion
Antibiotic resistance in the pathogens that cause nosocomial infection is an undeniable challenge that results in mortality and increased hospitalization.24–26 The results of this study showed that the most effective antibiotic against P. aeruginosa isolates was ciprofloxacin, and the highest resistance was observed to cefotaxime. It was also found that the resistance of P. aeruginosa isolates to all antibiotics was above 50%. Similar to our results, a study by Shariati et al confirmed high resistance to most of the antibiotics (70%) in P. aeruginosa isolates.1 Another study found that P. aeruginosa isolates were highly resistant to antibiotics (approximately 50%), and the results showed that the highest resistance was to cefotaxime.27 In this study, the highest number of XDR isolates was isolated from ICU patients, and the antibiotics resistance was higher in the ICU ward than other parts of the hospital. Another study showed a relationship between antibiotics resistant P. aeruginosa, especially MDR, and different wards of the hospital, such as ICU.28 Nseir et al also reported hospitalization in ICU as one of the risk factors for the ones infected with MDR P. aeruginosa, which confirms the results of our study.29 The reason that P. aeruginosa strains isolated from the ICU were more resistant than other wards maybe for the patients admitted to this ward who had been hospitalized for a long time and were on long-term antibiotic therapy. Also, admitting to the ICU room where patients were previously hospitalized with MDR P. aeruginosa infection is a risk factor for the transmission of this resistant bacterium to the new hospitalized patients.29 Among 50 P. aeruginosa isolates, 32 (64%), 23 (46%), 43 (86%), 3 (6%), 6 (12%), 3 (6%), 4 (8%), 7 (14%), 6 (12%), 18 (36%), 4 (8%), 19 (38%), 16 (32%), and 2 (4%) isolates contained IMP, VIM, OPR, SIM, SPM, GIM, NDM, VEB, PER, KPC, OXA, intI, intII, and intIII genes, respectively. Many studies around the world have investigated the genes associated with antibiotic resistance in clinical isolates of P. aeruginosa. In 2014, a study examined common resistance genes in P. aeruginosa including OXA, VEB, VIM, IMP, SIM, GIM, SPM, and NDM and the results showed that the prevalence of VIM, OXA, VEB, NDM, and IMP were 58.1%, 41.7%, 10.4%, 4.2% and 2.1%, respectively. But, contrary to our results, the genes of GIM, SPM and SIM were not found in P. aeruginosa isolates in another study.10 In the other study, researchers also examined the genes affecting the resistance of P. aeruginosa isolates with 3%, 65%, and 52% harbored ESBL, carbapenemase, and AmpC, respectively. The ESBL, carbapenemase and AmpC contained OXA (97%), TEM (61%), VIM (55%), PER (13%), IMP (3%) and AIM (1%). A study investigated the presence of the oprD gene in carbapenem-resistant P. aeruginosa isolates. The rate of this gene was 79.1%, which is very close to the outcome of our study.30 Also, our results demonstrated that 45 (90%) isolates produced ESBL and among these 45 isolates, only 8 (16%) isolates had all three genes (SHV, TEM, and CTXM) together. Chen et al found that 87.5% of P. aeruginosa isolates were ESBL-producers and only 5 isolates (2.7%) carried all of the three TEM, SHV and CTXM genes, simultaneously.31 In 2107, study on integrons in clinical isolates of P. aeruginosa showed that the results of integron 1, integron 2, and integron 3 were 27.5%, 25.5% and 15%, respectively. The results were in agreement with our results that showed class 3 integrons were less abundant than other integrons.27
High or low prevalence of some resistance genes in P. aeruginosa isolates can be due to differences in the geographical areas which indicate that prevalence of resistance genes differs across the world and even in different parts of a country. Other factors that may explain the differences in the prevalence of resistance genes include antibiotic abuse patterns, antimicrobial stewardship program, degree of virulence in strains, and infection control methods. In this study, it was also found that resistance genes were more present in the samples isolated from ICU ward and a direct relationship was observed between hospital ward and the presence of these genes. Other studies have also pointed to the association between the high levels of different antibiotic resistance genes and P. aeruginosa isolated from the ICU ward, which confirms the results of our study.32,33
In the current study, AgNO3 nanoparticles with a cylindrical shape and diameters between 15 and 50 nm were fabricated (average particle size of about 25 nm). Interestingly, AgNO3 nanoparticles had favorable solubility and bioavailability, which could be attributed to the large surface area. Our results also showed that the presence of a sulfide group on the surface of IMI has predisposed it to form a bond with AgNO3 nanoparticles. Previous research has proven that small size and high surface area of the nanoparticles increases the loading of antibiotics, and enhances the concentration of antibiotic at the site of contact with the bacteria and thereby further inhibiting the pathogens.22 Some studies have shown that reducing the size of nanoparticles increases their solubility, which is in agreement with the results of our study.1,34
The antimicrobial activity of the AgNO3 nanoparticles was shown at low concentrations. The conjugation of AgNO3 with IMI showed a greater antimicrobial effect on XDR isolates compared to AgNO3 alone. These results indicated that the AgNO3 nanoparticles increased the antimicrobial effect of IMI. In recent years, the use of nanoparticles as one of the new approaches against lethal antibiotic-resistant bacteria, such as P. aeruginosa has attracted the attention of researchers. Mapara et al investigated the effects of silver nanoparticles against XDR P. aeruginosa and found that the nanoparticles have antimicrobial activity against these resistant isolates.4 It has also been shown that the use of silver nanoparticles with IMI resulted in greater antibiotic efficacy and reduced the effective dose of IMI which is consistent with our study.22 In another study, nanoparticles were used against MDR and XDR isolates of P. aeruginosa and were found to kill these resistant bacteria, which demonstrates the antimicrobial effects of the nanoparticles.35
One of the main concerns of using nanoparticles as a drug is their toxicity. According to our results, human epithelial cell lines exposed to synthesized AgNO3 nanoparticles were well proliferated and no morphological change was observed. Our finding was in accordance with Zendegani et al and Shariati et al studies who demonstrated no toxicity of nanoparticles to the human cells.1,22 In addition, our results showed that IMI alone was more toxic than TCPS. After conjugation with nanoparticles, its toxic property reduced significantly. Various studies confirm that the use of nanoparticles as coating or in combination with organic or inorganic compounds can reduce their toxicity.36,37
Conclusion
In summary, AgNO3 nanoparticles had ideal antibacterial effects on XDR P. aeruginosa, and its conjugation with IMI showed greater antimicrobial effects. Despite widespread antibiotic resistance and the presence of ESBL, metallo-β-lactamase, carbapenemase and other resistance genes in P. aeruginosa, the AgNO3 alone and in combination with IMI indicated strong antibacterial activity. Due to the antimicrobial properties of conjugated AgNO3 nanoparticles with IMI, they have the capacity to be introduced as a suitable candidate for targeting antibiotic-resistant bacteria. However, it is essential to develop new nano-antibiotic formulations that can improve their efficacy and reduce dose and toxicity of the antibiotics. Finally, it is recommended to measure the toxic effects of AgNO3 alone and in combination with IMI in animal models.
Ethics Approval
Ethics Research Committee of Microbial Science Research Center approved this study with ethics code IR.BMSU. REC.1396. 880.
Acknowledgment
The authors kindly thank Baqiyatallah and Iran University of Medical Sciences for their support in providing equipment and facilities.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Shariati A, Asadian E, Fallah F, et al. Evaluation of Nano-curcumin effects on expression levels of virulence genes and biofilm production of multidrug-resistant Pseudomonas aeruginosa isolated from burn wound infection in Tehran, Iran. Infect Drug Resist. 2019;12:2223. doi:10.2147/IDR.S213200
2. Aghamollaei H, Moghaddam MM, Kooshki H, Heiat M, Mirnejad R, Barzi NS. Detection of Pseudomonas aeruginosa by a triplex polymerase chain reaction assay based on lasI/R and gyrB genes. J Infect Public Health. 2015;8(4):314–322. doi:10.1016/j.jiph.2015.03.003
3. Gupta R, Malik A, Rizvi M, Ahmed SM. Incidence of multidrug-resistant Pseudomonas spp. in ICU patients with special reference to ESBL, AMPC, MBL and biofilm production. J Glob Infect Dis. 2016;8(1):25.
4. Mapara N, Sharma M, Shriram V, Bharadwaj R, Mohite K, Kumar V. Antimicrobial potentials of Helicteres isora silver nanoparticles against extensively drug-resistant (XDR) clinical isolates of Pseudomonas aeruginosa. Appl Microbiol Biotechnol. 2015;99(24):10655–10667.
5. Ghazvini H, Taheri K, Edalati E, Miri A, Sedighi M, Mirkalantari S. Virulence factors and antimicrobial resistance in uropathogenic Escherichia Coli strains isolated from cystitis and pyelonephritis. Turk J Med Sci. 2019;49(1):361–367.
6. Kiaei S, Moradi M, Nave HH, Hashemizadeh Z, Taati-Moghadam M, Kalantar-Neyestanaki D. Emergence of co-existence of bla NDM with rmtC and qnrB genes in clinical carbapenem-resistant Klebsiella pneumoniae isolates in burning center from southeast of Iran. Folia Microbiol (Praha). 2019;64(1):55–62.
7. Korbekandi H, Iravani S, Abbasi S. Production of nanoparticles using organisms. Crit Rev Biotechnol. 2009;29(4):279–306.
8. Gill J, Arora S, Khanna S, Kumar KH. Prevalence of multidrug-resistant, extensively drug-resistant, and pandrug-resistant Pseudomonas aeruginosa from a tertiary level intensive care unit. J Glob Infect Dis. 2016;8(4):155.
9. Bokaeian M, Zahedani SS, Bajgiran MS, Moghaddam AA. Frequency of PER, VEB, SHV, TEM and CTX-M genes in resistant strains of Pseudomonas aeruginosa producing extended spectrum β-lactamases. Jundishapur J Microbiol. 2015;8(1).
10. Zafer MM, Al-Agamy MH, El-Mahallawy HA, Amin MA, Ashour MSE-D. Antimicrobial resistance pattern and their beta-lactamase encoding genes among Pseudomonas aeruginosa strains isolated from cancer patients. Biomed Res Int. 2014;2014:1–8. doi:10.1155/2014/101635
11. Kousouli E, Zarkotou O, Polimeri K, Themeli-Digalaki K, Pournaras S. Impact of bloodstream infections caused by carbapenem-resistant Gram-negative pathogens on ICU costs, mortality and length of stay. Infect Prev Pract. 2019;1(2):100020. doi:10.1016/j.infpip.2019.100020
12. Moghadam MT, Amirmozafari N, Shariati A, et al. How phages overcome the challenges of drug resistant bacteria in clinical infections. Infect Drug Resist. 2020;13:45. doi:10.2147/IDR.S234353
13. Bauer KA, West JE, O’Brien JM, Goff DA. Extended-infusion cefepime reduces mortality in patients with Pseudomonas aeruginosa infections. Antimicrob Agents Chemother. 2013;57(7):2907–2912. doi:10.1128/AAC.02365-12
14. Roy K, Sarkar C, Ghosh C. Photocatalytic activity of biogenic silver nanoparticles synthesized using yeast (Saccharomyces cerevisiae) extract. Appl Nanoscie. 2015;5(8):953–959. doi:10.1007/s13204-014-0392-4
15. Ottoni CA, Simões MF, Fernandes S, et al. Screening of filamentous fungi for antimicrobial silver nanoparticles synthesis. AMB Express. 2017;7(1):1–10. doi:10.1186/s13568-017-0332-2
16. Khataminejad MR, Mirnejad R, Sharif M, Hashemi M, Sajadi N, Piranfar V. Antimicrobial effect of imipenem-functionalized Fe2O3 nanoparticles on Pseudomonas aeruginosa producing metallo β-lactamases. Iran J Biotechnol. 2015;13(4):43. doi:10.15171/ijb.1138
17. Bardania H, Mahmoudi R, Bagheri H, et al. Facile preparation of a novel biogenic silver-loaded Nanofilm with intrinsic anti-bacterial and oxidant scavenging activities for wound healing. Sci Rep. 2020;10(1):1–14. doi:10.1038/s41598-020-63032-5
18. Shariati A, Azimi T, Ardebili A, et al. Insertional inactivation of oprD in carbapenem-resistant Pseudomonas aeruginosa strains isolated from burn patients in Tehran, Iran. New Microbes New Infect. 2018;21:75–80. doi:10.1016/j.nmni.2017.10.013
19. Taati Moghadam M, Hossieni Nave H, Mohebi S, Norouzi A. The evaluation of connection between integrons class I and II and ESBL-producing and Non-ESBL klebsiella pneumoniae isolated from clinical samples, Kerman. Iran J Med Microbiol. 2016;10(4):1–9.
20. Moradi M, Norouzi A, Taatimoghadam M. Prevalence of bla-CTX-M, bla-SHV, and bla-TEM genes and comparison of antibiotic resistance pattern in extended-spectrum β-lactamase producing and non-producing groups of Klebsiella pneumoniae isolated from clinical samples in Kerman hospitals. J Fasa Univ Med Sci. 2016;6(1):120–128.
21. Deepa K, Panda T. Biogenic gold nanoparticles from fusarium oxysporum: the impact of fungal morphology and localization studies. J Cluster Sci. 2019;1–13.
22. Zendegani E, Dolatabadi S. The efficacy of imipenem conjugated with synthesized silver nanoparticles against Acinetobacter baumannii clinical isolates, Iran. Biol Trace Elem Res. 2019;1–11.
23. Mollaei S, Habibi B, Amani Ghadim A, Shakouri M. A green approach for the synthesis of silver nanoparticles using Lithospermum officinale root extract and evaluation of their antioxidant activity. J Part Sci Technol. 2017;3(4):187–195.
24. Mohebi S, Hossieni Nave H, Norouzi A, Kandehkar Gharaman M, Taati Moghadam M. Detection of extended spectrum beta lactamases on class I integron in Escherichia coli isolated from clinical samples. J Mazandaran Univ Med Sci. 2016;26(138):66–76.
25. Hadizadeh M, Norouzi A, Taghadosi R, et al. Prevalence of qnr, intI, and intII genes in extended spectrum beta-lactamase (ESBL)-producing Escherichia coli isolated from clinical samples in Iran. Trop J Pharm Res. 2017;16(1):141–147.
26. Rajabi S, Shivaee A, Khosravi MA, Eshaghi M, Shahbazi S, Hosseini F. Evaluation of multidrug efflux pump expression in clinical isolates of Staphylococcus aureus. Gene Rep. 2020;18:100537.
27. Mobaraki S, Aghazadeh M, Barhaghi MHS, et al. Prevalence of integrons 1, 2, 3 associated with antibiotic resistance in Pseudomonas aeruginosa isolates from Northwest of Iran. BioMedicine. 2018;8(1).
28. Walkty A, Lagace-Wiens P, Adam H, et al. Antimicrobial susceptibility of 2906 Pseudomonas aeruginosa clinical isolates obtained from patients in Canadian hospitals over a period of 8 years: results of the Canadian Ward surveillance study (CANWARD), 2008–2015. Diagn Microbiol Infect Dis. 2017;87(1):60–63.
29. Nseir S, Blazejewski C, Lubret R, Wallet F, Courcol R, Durocher A. Risk of acquiring multidrug-resistant Gram-negative bacilli from prior room occupants in the intensive care unit. Clin Microbiol Infect. 2011;17(8):1201–1208.
30. Mathlouthi N, Areig Z, Al Bayssari C, et al. Emergence of carbapenem-resistant Pseudomonas aeruginosa and Acinetobacter baumannii clinical isolates collected from some Libyan hospitals. Microb Drug Resist. 2015;21(3):335–341.
31. Chen Z, Niu H, Chen G, Li M, Li M, Zhou Y. Prevalence of ESBLs-producing Pseudomonas aeruginosa isolates from different wards in a Chinese teaching hospital. Int J Clin Exp Med. 2015;8(10):19400.
32. Goli HR, Nahaei MR, Rezaee MA, et al. Role of MexAB-OprM and MexXY-OprM efflux pumps and class 1 integrons in resistance to antibiotics in burn and Intensive Care Unit isolates of Pseudomonas aeruginosa. J Infect Public Health. 2018;11(3):364–372.
33. Farhan SM, Ibrahim RA, Mahran KM, Hetta HF, El-Baky RMA. Antimicrobial resistance pattern and molecular genetic distribution of metallo-β-lactamases producing Pseudomonas aeruginosa isolated from hospitals in Minia, Egypt. Infect Drug Resist. 2019;12:2125.
34. Kesisoglou F, Panmai S, Wu Y. Nanosizing—oral formulation development and biopharmaceutical evaluation. Adv Drug Deliv Rev. 2007;59(7):631–644.
35. Ashfaq M, Verma N, Khan S. Highly effective Cu/Zn-carbon micro/nanofiber-polymer nanocomposite-based wound dressing biomaterial against the P. aeruginosa multi-and extensively drug-resistant strains. Mater Sci Eng C. 2017;77:630–641.
36. Chen J, Li S, Luo J, Wang R, Ding W. Enhancement of the antibacterial activity of silver nanoparticles against phytopathogenic bacterium Ralstonia solanacearum by stabilization. J Nanomater. 2016;2016.
37. Zhang T, Wang L, Chen Q, Chen C. Cytotoxic potential of silver nanoparticles. Yonsei Med J. 2014;55(2):283–291.
38. Mirkalantari S, Moghadas AJ. Diversity determination of CTX-M1 producing klebsiella pneumoniae using multilocus variable-number tandem repeat analysis, semnan, Iran. Jundishapur J Microbiol. 2018 July; 11(7):e63131. doi:10.5812/jjm.63131
39. Bina M, Pournajaf A, Mirkalantari S, Talebi M, Irajian G. Detection of the Klebsiella pneumoniae carbapenemase (KPC) in K. pneumoniae Isolated from the clinical samples by the phenotypic and genotypic methods. Iran J Pathol. 2015;10(3):199.
40. Jazayeri Moghadas A, Kalantari F, Sarfi M, Shahhoseini S, Mirkalantari S. Evaluation of Virulence Factors and Antibiotic Resistance Patterns in Clinical Urine Isolates of Klebsiella pneumoniae in Semnan, Iran. Jundishapur J Microbiol. 2018;11(7):e63637. doi:10.5812/jjm.63637
41. Reyes A, Bello H, Domínguez M, Mella S, Zemelman R, González G. Prevalence and types of class 1 integrons in aminoglycoside-resistant Enterobacteriaceae from several Chilean hospitals. J Antimicrob Chemother. 2003;51(2):317–321.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
