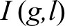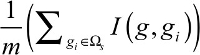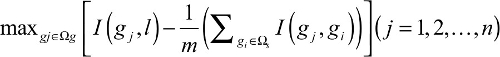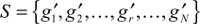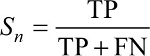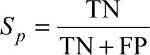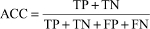Back to Journals » Infection and Drug Resistance » Volume 12
The blood transcriptional signature for active and latent tuberculosis
Authors Deng M, Lv XD, Fang ZX, Xie XS, Chen WY
Received 20 August 2018
Accepted for publication 27 December 2018
Published 30 January 2019 Volume 2019:12 Pages 321—328
DOI https://doi.org/10.2147/IDR.S184640
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eric Nulens
Min Deng,1 Xiao-Dong Lv,2 Zhi-Xian Fang,2 Xin-Sheng Xie,1 Wen-Yu Chen2
1Department of Infectious Diseases, The First Hospital of Jiaxing, The First Affiliated Hospital of Jiaxing University, Jiaxing 314000, China; 2Department of Respiration, The First Hospital of Jiaxing, The First Affiliated Hospital of Jiaxing University, Jiaxing 314000, China
Background: Although the incidence of tuberculosis (TB) has dropped substantially, it still is a serious threat to human health. And in recent years, the emergence of resistant bacilli and inadequate disease control and prevention has led to a significant rise in the global TB epidemic. It is known that the cause of TB is Mycobacterium tuberculosis infection. But it is not clear why some infected patients are active while others are latent.
Methods: We analyzed the blood gene expression profiles of 69 latent TB patients and 54 active pulmonary TB patients from GEO (Transcript Expression Omnibus) database.
Results: By applying minimal redundancy maximal relevance and incremental feature selection, we identified 24 signature genes which can predict the TB activation. The support vector machine predictor based on these 24 genes had a sensitivity of 0.907, specificity of 0.913, and accuracy of 0.911, respectively. Although they need to be validated in a large independent dataset, the biological analysis of these 24 genes showed great promise.
Conclusion: We found that cytokine production was a key process during TB activation and genes like CYBB, TSPO, CD36, and STAT1 worth further investigation.
Keywords: tuberculosis, blood gene expression, support vector machine, minimal redundancy maximal relevance, incremental feature selection
Introduction
Tuberculosis (TB), a pulmonary infectious disease caused by Mycobacterium tuberculosis, is a serious threat to human health. In the early 20th century, TB had spread around the world, killing millions of people.1 Later, with the progress of medical technology and improvement of sanitary conditions, the incidence of TB dropped substantially. However, in recent years the emergence of resistant bacilli and inadequate disease control and prevention has led to a significant rise in the global TB epidemic, which has now become the leading cause of deaths from infectious diseases.2
Currently, about 90%–95% TB patients are with latent tuberculosis infection (LTBI).3 LTBI is defined by persistent immune response caused by M. tuberculosis, but under conditions of a long-term asymptomatic infection.4 During latent infection, body’s immune system can inhibit the growth of bacteria by blocking the replication of the bacteria,5 but not to be completely eliminated. M. tuberculosis can stay dormant for decades, or even during the host’s lifetime.6
However, the risk of developing active TB increases greatly when human immunity declines.7 The main reason bacteria can survive in the body is because it contains high lipid content, which can prevent it from degradation and destruction within macrophages.5 In addition, M. tuberculosis can also affect the normal function of CD8+ T-cells, natural killer cells, and complement membrane attack complex.6
Immune responses to M. tuberculosis is primarily cell-mediated, regulated by the interaction between T-cells and infected macrophages and cytokines secreted by these cells.8 In most cases, infection of M. tuberculosis transits to the dormant state accompanied by the formation of infective granuloma that are well separated from the surrounding tissue.9 Saunders and Cooper indicated that the formation of granulomas is essential for limiting M. tuberculosis growth and tissue damage in TB infections, two major components of active TB.10 Risk factors known for TB reactivation include malnutrition, HIV infection, anti-tumor necrosis factor treatment, insulin-dependent diabetes, alcoholism, and smoking. However, the mechanisms of reactivation of LTBI in most cases are still unknown.11
Early diagnosis and preventive treatment of LTBI are an effective means to eliminate the bacteria and reduce the risk of TB in immunocompromised patients. Currently, the preferred diagnostic tools for LTBI remain tuberculin skin testing (TST) or interferon-γ release assay (IGRA).12 IGRA, based on cellular immunity, was tested to confirm whether people infected with bacteria by detecting the release of IFN-γ to specific antigen of M. tuberculosis. Compared with traditional TST, IGRA has higher specificity and sensitivity, and has been widely used in clinical detection of TB.13 However, they can only detect the infection of TB bacilli, but cannot reflect the risk of developing active TB.
Recently, evidence from researches has demonstrated the importance of genetic factors in the development of active TB,14 but the mechanism of genetic susceptibility for TB remains largely unknown. Since humans are the natural hosts for M. tuberculosis, the precise etiology of infection cannot be studied in any other animal models.15
To understand what actives TB, we analyzed the blood gene expression profiles of 69 latent TB patients and 54 active pulmonary TB (PTB) patients. With advanced feature selection methods, including minimal redundancy maximal relevance (mRMR) and incremental feature selection (IFS), 24 genes were identified as discriminative between latent and active TB patients. What is more, a support vector machine (SVM) classifier for active TB prediction was built based on these 24 genes and its sensitivity, specificity, and accuracy evaluated with leave-one out-cross validation (LOOCV) were 0.907, 0.913, and 0.911, respectively.
Methods
The blood gene expression profiles of latent and active TB patients
To identify the key genes that activate TB, we downloaded the blood gene expression profiles of 69 latent TB patients and 54 active PTB patients from publicly available GEO (Transcript Expression Omnibus) database under accession number of GSE19491.16 The 24 normal samples in the original GSE19491 dataset were excluded, only the 69 latent TB patients and 54 active PTB patients were analyzed to get the key genes that activate TB. These samples were collected from London, UK and Cape Town, South Africa.16 We combined the samples from different cities to get a larger dataset. The age and gender information of these samples are listed in Table S1. Within the 123 samples, there were 64 male and 59 female patients. The average age was 32.5 with an SD of 13.6. The gene expression levels were measured using microarray of Illumina HumanHT-12 V3.0 expression beadchip. There were 48,803 probes corresponding to 25,153 genes. The probes corresponding to the same gene were averaged. The gene expression matrix was quantile normalized.
Identify the genes that are related to the activation of TB
As a basic problem in bioinformatics, many methods have been proposed to identify the phenotype-related genes or proteins. One of these methods, mRMR,17 has been widely used and proved to be effective.18–20
The mRMR method is different from univariate statistical test. To explain its principle, let us use Ω to denote all the 25,153 genes, Ωs to denote the selected m genes, and Ωg to denote the to be selected n genes. First, the relevance of gene g from Ωg with the activeness of TB l was evaluated with mutual information (I) equation:
| (1) |
Then, the average redundancy of gene g with the already selected genes was
| (2) |
At last, to consider both maximum relevance and minimum redundancy, the goal of this algorithm was to find the best gene g from to maximize the function below in which the first part meant relevance and the second represented redundancy
| (3) |
After 25,153 rounds of optimization, all the 25,153 genes can be ranked as
| (4) |
In this gene list, the top ranked ones were discriminative. We considered the top 500 mRMR genes as relevant to the activeness of TB. The top 500 mRMR genes are listed in Table S2.
Find the signature genes for TB activation
The top 500 mRMR genes were related to TB activation, but the number of genes was too large to detect practically. Therefore, we adopted IFS21–27 to get the signature genes for TB activation. Hopefully, a small number of genes were enough to build an actionable predictor for TB activation. The widely used SVM was adopted to construct the classifiers.
During the IFS procedure, the genes were added sequentially based on the mRMR rank, starting with the first mRMR gene, ending with the 500th mRMR gene. In other words, 500 gene sets were selected and 500 SVM classifiers were constructed. Each classifier was evaluated with LOOCV and its sensitivity (Sn), specificity (Sp), and accuracy (ACC) were calculated
| (5) |
| (6) |
| (7) |
TP, TN, FP, and FN were the number of true active TB, true latent TB, false active TB, and latent TB patients.
Based on the IFS results, we can choose the right gene sets as signature and get acceptable prediction accuracy.
Results and discussion
The 500 genes that are related to the activation of TB
Using the mRMR method which considered both the relevance with TB activeness and the redundancy with other genes, we identified 500 genes that were related to the activation of TB. These 500 genes were ranked based on their discriminative ability of latent TB patients and active PTB patients.
The 24 signature genes for TB activation
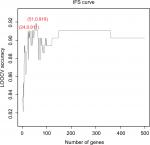
To further narrow down the 500 genes that were related to the activation of TB and obtain a signature for TB activation, we applied the IFS analysis and plotted the IFS curve based on the prediction performances using different number of signature genes (Table S3). As shown in Figure 1, when the top 51 mRMR genes were used, the LOOCV accuracy was the highest, 0.919. But when only 24 genes were used, the accuracy became stable. Therefore, we choose the 24 genes as signature genes for TB activation. The sensitivity, specificity, and accuracy of the 24 signature genes for TB activeness prediction were 0.907, 0.913, and 0.911, respectively. The 24 signature genes are given in Table 1 and the confusion matrix of their prediction performance is shown in Table 2.
  | Table 1 The 24 signature genes for TB activation Abbreviations: mRMR, minimal redundancy maximal relevance; TB, tuberculosis. |
  | Table 2 The confusion matrix of the predicted and actual TB activeness based on the 24 signature genes Abbreviation: TB, tuberculosis. |
To explore the expression levels of these 24 signature genes in latent TB patients and active PTB patients, we plotted their heatmap in Figure 2. It can be seen that most of the latent and active TB patients were clustered into the right groups. TMEM51, TSPO, CD36, CYBB, LOXL3, CYB561, GPR63, LYRM1, and STAT1 were highly expressed in active TB patients while SLC9A3R1, TUSC4, ZNF91, HNRNPD, PRKCQ, QSOX2, BANK1, EPHA4, C15orf15, NELL2, ANAPC1, FNBP4, TRIB2, CA5B, and KIAA1370 were highly expressed in latent TB patients.
The biological analysis of the signature genes for TB activation
We did Gene Ontology (GO) enrichment analysis of these 24 signature genes and the results with false discovery rate <0.10 are given in Table 3. The enriched GO biological processes were GO:0001817 regulation of cytokine production, GO:0001816 cytokine production, GO:0051172 negative regulation of nitrogen compound metabolic process, GO:0042592 homeostatic process, GO:0031324 negative regulation of cellular metabolic process, GO:0010243 response to organonitrogen compound, and GO:0010605 negative regulation of macromolecule metabolic process. There have been many reports of the important roles of cytokine in responding to mycobacteria during the activation of TB.28,29
  | Table 3 The enriched GO biological processes for the 24 signature genes Abbreviations: FDR, false discovery rate; GO, Gene Ontology. |
The following genes from Table 1 showed great promise and were discussed.
Gp91phox, encoded by CYBB ranked second in Table 1, is an essential subunit of NADPH oxidase complex. Alterations in macrophage function, such as defects in NADPH oxidase30 and the vitamin D receptor,31 are known as risk factors for mycobacterial infection. Several studies have shown that mutations in CYBB could result in X-linked chronic granulomatous disease with much higher risk of TB,32–34 which is an immunodeficiency caused by defective activity of NADPH oxidase in phagocytes.33,35 Liu et al have also demonstrated the significant correlation between CYBB polymorphisms and decreased risk of TB, particularly among male smokers.36
TSPO ranked third in Table 1 serves as a trans-mitochondrial membrane channel that transports cholesterol and other endogenous ligands.37 It has been reported that the expression of TSPO is highest in steroidogenic tissues, lung, and immune cells like macrophages.38 Immunofluorescence studies made by Foss et al indicated that TSPO was highly expressed in phagocytic cells and CD68(+) macrophages within TB lesions.39 The increased expression of TSPO will lead to macrophages activation, which is a pivotal component of TB-associated inflammation.39 In PTB, a synthetic ligand for TSPO, radioiodinated DPA-713, is found to be upregulated in activated macrophages.40
CD36 ranked 23rd in Table 1 is a membrane glycoprotein that exist in various cells, including macrophages, monocytes, adipocytes, and platelets.41 It has been implicated in multiple cellular processes and defined as a multiligand scavenger receptor that mediates fatty acid transport, phagocytosis, and inflammation in response to a variety of pathogens, including mycobacteria.42 CD36 facilitates surfactant lipid uptake which can be exploited by M. tuberculosis for growth.43 Lao et al suggested that rs1194182 and rs10499859, two SNPs of CD36, may reduce the risk of PTB, indicating CD36 as an important biomarker for PTB.44 Hawkes et al observed that deficiency of CD36 reduces the susceptibility of mice to mycobacterial infection. In addition, CD36 deficiency of macrophages inhibits the growth of many mycobacterial species in vitro, demonstrating that deficiency of CD36 plays a role in the resistance to mycobacterial infection.45
STAT1 ranked 24th in Table 1, is a member of the STAT protein family and thought to be an important mediator in response to IFN-γ and host defense against M. tuberculosis.46,47 IFN-γ activates macrophages to kill multiple pathogens but cannot activate macrophages to kill M. tuberculosis.48 Much recent evidence shows that M. tuberculosis infection inhibited IFN-γ signaling via blocking several responses to IFN-γ, such as induction of FcγRI,49 and the dysfunction of macrophages in response to IFN-γ depends on an altered regulatory mechanism in STAT1 signaling pathway.50 Sugawara et al discovered that STAT1 knockout mice have higher susceptibility to pulmonary mycobacterial infection in mice, indicating that STAT1 appears to be a key transcription factor in resisting mycobacterial infection.51
The proteomics pattern of the signature genes
Marakalala et al measured the granulomas proteomes of TB patients using mass spectrometry and confocal microscopy.52 They identified 4,406 proteins that were expressed in caseous granuloma caseum, cavitary granuloma cells, cavitary granuloma caseum, and solid granuloma cells. We mapped our signatures onto their proteomics results. Within the 24 signature genes, 6 of them were detected by Marakalala et al. Their log2 label-free quantification intensities in caseous granuloma caseum, cavitary granuloma cells, cavitary granuloma caseum, and solid granuloma cells were shown in Table 4. HNRNPD, CYBB, TSPO, SLC9A3R1, FNBP4, and STAT1 are important on both mRNA and protein levels.
  | Table 4 The log2 label-free quantification intensity of signature genes in caseous granuloma caseum, cavitary granuloma cells, cavitary granuloma caseum, and solid granuloma cells |
Conclusion
As a pulmonary infectious disease caused by M. tuberculosis, TB is a serious threat and can spread through coughing, sneezing, or other ways. But not all infected patients exhibit symptoms. Some patients were active while others were latent. Which factors or genes trigger TB activation is key to understand TB. By analysis, the blood gene expression profiles of 69 latent TB patients and 54 active PTB patients, we found 24 signature genes that can predict TB activeness. In-depth analysis of these 24 genes suggested that cytokine production was a key process during TB activation. Signature genes including CYBB, TSPO, CD36, and STAT1 worth to be further investigated.
Acknowledgments
This study was supported by The Key Discipline of Jiaxing Respiratory Medicine Construction Project, Zhejiang North Regional Anesthesia Special Disease Center, Talent Cultivation in Science and Technology Innovation Project of The First Hospital of Jiaxing (No. 2016-CX-04).
Disclosure
The authors report no conflicts of interest in this work.
References
Mjid M, Cherif J, Ben Salah N, et al. [Epidemiology of tuberculosis]. Rev Pneumol Clin. 2015;71(2–3):67–72. French | ||
Subbian S, Pandey R, Soteropoulos P, Rodriguez GM. Vaccination with an attenuated ferritin mutant protects mice against virulent Mycobacterium tuberculosis. J Immunol Res. 2015;2015:1–12. | ||
Subbian S, Bandyopadhyay N, Tsenova L, et al. Early innate immunity determines outcome of Mycobacterium tuberculosis pulmonary infection in rabbits. Cell Commun Signal. 2013;11(1):60. | ||
Chee CB, Sester M, Zhang W, Lange C. Diagnosis and treatment of latent infection with Mycobacterium tuberculosis. Respirology. 2013;18(2):205–216. | ||
Hernandez-Pando R, Orozco H, Aguilar D. Factors that deregulate the protective immune response in tuberculosis. Arch Immunol Ther Exp (Warsz). 2009;57(5):355–367. | ||
Muñoz L, Stagg HR, Abubakar I. Diagnosis and management of latent tuberculosis infection. Cold Spring Harb Perspect Med. 2015;5(11):a017830. | ||
Salgame P, Geadas C, Collins L, Jones-López E, Ellner JJ. Latent tuberculosis infection—revisiting and revising concepts. Tuberculosis (Edinb). 2015;95(4):373–384. | ||
O’Garra A, Redford PS, McNab FW, Bloom CI, Wilkinson RJ, Berry MP. The immune response in tuberculosis. Annu Rev Immunol. 2013;31(1):475–527. | ||
Ulrichs T, Kaufmann SH. New insights into the function of granulomas in human tuberculosis. J Pathol. 2006;208(2):261–269. | ||
Saunders BM, Cooper AM. Restraining mycobacteria: role of granulomas in mycobacterial infections. Immunol Cell Biol. 2000;78(4):334–341. | ||
Lillebaek T, Dirksen A, Baess I, Strunge B, Thomsen VØ, Andersen AB. Molecular evidence of endogenous reactivation of Mycobacterium tuberculosis after 33 years of latent infection. J Infect Dis. 2002;185(3):401–404. | ||
Esmail H, Barry CE, Wilkinson RJ. Understanding latent tuberculosis: the key to improved diagnostic and novel treatment strategies. Drug Discov Today. 2012;17(9–10):514–521. | ||
Thillai M, Pollock K, Pareek M, Lalvani A. Interferon-gamma release assays for tuberculosis: current and future applications. Expert Rev Respir Med. 2014;8(1):67–78. | ||
Cooke GS, Hill AV. Genetics of susceptibility to human infectious disease. Nat Rev Genet. 2001;2(12):967–977. | ||
Arora G, Misra R, Sajid A. Model systems for pulmonary infectious diseases: paradigms of anthrax and tuberculosis. Curr Top Med Chem. 2017;17(18):2077–2099. | ||
Berry MP, Graham CM, McNab FW, et al. An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis. Nature. 2010;466(7309):973–977. | ||
Peng H, Long F, Ding C. Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Trans Pattern Anal Mach Intell. 2005;27(8):1226–1238. | ||
Li J, Huang T. Predicting and analyzing early wake-up associated gene expressions by integrating GWAS and eQTL studies. Biochim Biophys Acta Mol Basis Dis. 2018;1864(6 Pt B):2241–2246. | ||
Cai L, Huang T, Su J, et al. Implications of newly identified brain eQTL genes and their interactors in schizophrenia. Mol Ther Nucleic Acids. 2018;12:433–442. | ||
Chen L, Pan X, Hu X. Gene expression differences among different MSI statuses in colorectal cancer. Int J Cancer. Epub 2018 Apr 26. | ||
Zhang N, Huang T, Cai YD. Discriminating between deleterious and neutral non-frameshifting indels based on protein interaction networks and hybrid properties. Mol Genet Genomics. 2015;290(1):343–352. | ||
Shu Y, Zhang N, Kong X, Huang T, Cai Y-D. Predicting A-to-I RNA editing by feature selection and random forest. PLoS One. 2014;9(10):e110607. | ||
Li BQ, You J, Huang T, Cai YD. Classification of non-small cell lung cancer based on copy number alterations. PLoS One. 2014;9(2):e88300. | ||
Jiang Y, Huang T, Chen L, Gao Y-F, Cai Y, Chou K-C. Signal propagation in protein interaction network during colorectal cancer progression. BioMed Res Int. 2013;2013(1):1–9. | ||
Zhang PW, Chen L, Huang T, Zhang N, Kong XY, Cai YD. Classifying ten types of major cancers based on reverse phase protein array profiles. PLoS One. 2015;10(3):e0123147. | ||
Huang T, Shu Y, Cai YD. Genetic differences among ethnic groups. BMC Genomics. 2015;16(1):1093. | ||
Chen L, Li J, Zhang YH. Identification of gene expression signatures across different types of neural stem cells with the Monte-Carlo feature selection method. J Cell Biochem. 2018;119(4):3394–3403. | ||
Etna MP, Giacomini E, Severa M, Coccia EM. Pro- and anti-inflammatory cytokines in tuberculosis: a two-edged sword in TB pathogenesis. Semin Immunol. 2014;26(6):543–551. | ||
Mayer-Barber KD, Sher A. Cytokine and lipid mediator networks in tuberculosis. Immunol Rev. 2015;264(1):264–275. | ||
Lau YL, Chan GC, Ha SY, Hui YF, Yuen KY. The role of phagocytic respiratory burst in host defense against Mycobacterium tuberculosis. Clin Infect Dis. 1998;26(1):226–227. | ||
Sun YP, Cai QS. Vitamin D receptor FokI gene polymorphism and tuberculosis susceptibility: a meta-analysis. Genet Mol Res. 2015;14(2):6156–6163. | ||
Khan TA, Kalsoom K, Iqbal A, et al. A novel missense mutation in the NADPH binding domain of CYBB abolishes the NADPH oxidase activity in a male patient with increased susceptibility to infections. Microb Pathog. 2016;100:163–169. | ||
Bustamante J, Arias AA, Vogt G, et al. Germline CYBB mutations that selectively affect macrophages in kindreds with X-linked predisposition to tuberculous mycobacterial disease. Nat Immunol. 2011;12(3):213–221. | ||
Barese C, Copelli S, Zandomeni R, Oleastro M, Zelazko M, Rivas EM. X-linked chronic granulomatous disease: first report of mutations in patients of Argentina. J Pediatr Hematol/Oncol. 2004;26(10):656–660. | ||
Ishibashi F, Mizukami T, Kanegasaki S, et al. Improved superoxide-generating ability by interferon gamma due to splicing pattern change of transcripts in neutrophils from patients with a splice site mutation in CYBB gene. Blood. 2001;98(2):436–441. | ||
Liu Q, Wang J, Sandford AJ, et al. Association of CYBB polymorphisms with tuberculosis susceptibility in the Chinese Han population. Infect Genet Evol. 2015;33:169–175. | ||
Rupprecht R, Papadopoulos V, Rammes G, et al. Translocator protein (18 kDa) (TSPO) as a therapeutic target for neurological and psychiatric disorders. Nat Rev Drug Discov. 2010;9(12):971–988. | ||
Zavala F, Haumont J, Lenfant M. Interaction of benzodiazepines with mouse macrophages. Eur J Pharmacol. 1984;106(3):561–566. | ||
Foss CA, Harper JS, Wang H, Pomper MG, Jain SK. Noninvasive molecular imaging of tuberculosis-associated inflammation with radioiodinated DPA-713. J Infect Dis. 2013;208(12):2067–2074. | ||
Tucker EW, Pokkali S, Zhang Z, et al. Microglia activation in a pediatric rabbit model of tuberculous meningitis. Dis Model Mech. 2016;9(12):1497–1506. | ||
Park YM. CD36, a scavenger receptor implicated in atherosclerosis. Exp Mol Med. 2014;46(6):e99. | ||
Febbraio M, Hajjar DP, Silverstein RL. CD36: a class B scavenger receptor involved in angiogenesis, atherosclerosis, inflammation, and lipid metabolism. J Clin Invest. 2001;108(6):785–791. | ||
Dodd CE, Pyle CJ, Glowinski R, Rajaram MV, Schlesinger LS. CD36-mediated uptake of surfactant lipids by human macrophages promotes intracellular growth of Mycobacterium tuberculosis. J Immunol. 2016;197(12):4727–4735. | ||
Lao W, Kang H, Jin G, et al. Evaluation of the relationship between MARCO and CD36 single-nucleotide polymorphisms and susceptibility to pulmonary tuberculosis in a Chinese Han population. BMC Infect Dis. 2017;17(1):488. | ||
Hawkes M, Li X, Crockett M, et al. CD36 deficiency attenuates experimental mycobacterial infection. BMC Infect Dis. 2010;10(1):299. | ||
Meraz MA, White JM, Sheehan KC, et al. Targeted disruption of the STAT1 gene in mice reveals unexpected physiologic specificity in the JAK-STAT signaling pathway. Cell. 1996;84(3):431–442. | ||
Durbin JE, Hackenmiller R, Simon MC, Levy DE. Targeted disruption of the mouse STAT1 gene results in compromised innate immunity to viral disease. Cell. 1996;84(3):443–450. | ||
Flynn JL, Chan J, Triebold KJ, Dalton DK, Stewart TA, Bloom BR. An essential role for interferon gamma in resistance to Mycobacterium tuberculosis infection. J Exp Med. 1993;178(6):2249–2254. | ||
Ting LM, Kim AC, Cattamanchi AA, Ernst JD. Mycobacterium tuberculosis inhibits IFN-gamma transcriptional responses without inhibiting activation of STAT1. J Immunol. 1999;163(7):3898–3906. | ||
Esquivel-Solís H, Quiñones-Falconi F, Zarain-Herzberg A, Amieva-Fernández RI, López-Vidal Y. Impaired activation of STAT1 and c-Jun as a possible defect in macrophages of patients with active tuberculosis. Clin Exp Immunol. 2009;158(1):45–54. | ||
Sugawara I, Yamada H, Mizuno S. STAT1 knockout mice are highly susceptible to pulmonary mycobacterial infection. Tohoku J Exp Med. 2004;202(1):41–50. | ||
Marakalala MJ, Raju RM, Sharma K, et al. Inflammatory signaling in human tuberculosis granulomas is spatially organized. Nat Med. 2016;22(5):531–538. |
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.