Back to Journals » ImmunoTargets and Therapy » Volume 10
Soluble IL-7Rα/sCD127 in Health, Disease, and Its Potential Role as a Therapeutic Agent
Authors Barros PO , Berthoud TK, Aloufi N, Angel JB
Received 2 December 2020
Accepted for publication 16 February 2021
Published 8 March 2021 Volume 2021:10 Pages 47—62
DOI https://doi.org/10.2147/ITT.S264149
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Michael Shurin
Priscila O Barros,1 Tamara K Berthoud,1 Nawaf Aloufi,2 Jonathan B Angel3
1Chronic Disease Program, Ottawa Hospital Research Institute, Ottawa, Ontario, Canada; 2Department of Biochemistry, Microbiology and Immunology, Faculty of Medicine, University of Ottawa, Ottawa, Ontario, Canada; 3Division of Infectious Diseases, Department of Medicine, University of Ottawa and the Ottawa Hospital, Ottawa, Ontario, Canada
Correspondence: Jonathan B Angel
The Ottawa Hospital, 501 Smyth Road, Room G-8, Ottawa, ON, K1H 8L6, Canada
Tel +1 (613) 737-8442
Fax +1 (613) 739-6866
Email [email protected]
Abstract: Soluble cytokine receptors can influence immune responses by modulating the biological functions of their respective ligands. These effects can be either agonistic or antagonistic and a number of soluble cytokine receptors have been shown to play critical roles in both maintenance of health and disease pathogenesis. Soluble IL-7Ra (sCD127) is one such example. With its impact on the IL-7/CD127 pathway, which is fundamental for the development and homeostasis of T cells, the role of sCD127 in health and disease has been extensively studied in recent years. Within this review, the role of sCD127 in maintaining host immune function is presented. Next, by addressing genetic factors affecting sCD127 expression and the associated levels of sCD127 production, the roles of sCD127 in autoimmune disease, infections and cancer are described. Finally, advances in the field of soluble cytokine therapy and the potential for sCD127 as a biomarker and therapeutic agent are discussed.
Keywords: sCD127, IL7RA, sIL-7Rα, IL-7, soluble cytokine receptor, rs6897932 SNP
Introduction
It was not until 1985 with the discovery of the soluble IL-2 receptor (sIL-2R) that it was recognized that cytokine receptors could exist as soluble entities.1 Over the following years, many other cytokine receptors have been cloned, resulting in the identification of cytokine receptor forms lacking the transmembrane domain, and it became clear that soluble cytokine receptors were a widespread phenomenon. Since then, it has become apparent that soluble cytokine receptors can play a valuable role in regulating the immune response, affecting the balance between their ligands and the membrane-bound receptors. It is therefore not unexpected that alterations in the expression of soluble cytokine receptors may result in human disease.
Soluble cytokine receptors usually arise from the extracellular portion of the membrane-bound protein. These proteins can be generated by proteolytic cleavage of the membrane-bound receptor, a mechanism termed ectodomain shedding, or by alternative mRNA splicing, which gives rise to a form of the receptor lacking the transmembrane domain. The role of alternative mRNA splicing in the generation of soluble receptors has been demonstrated in various studies where the presence of cDNA encoding the respective receptors, lacking the transmembrane domains, was detected. Soluble versions of IL-9Rα, IL-4Rα, IL-5Rα and IL-7Rα were shown to be generated in this manner.2–5 Interestingly, some soluble cytokine receptors, such as the IL-1RII6,7 and soluble IL-6R (sIL-6R)8,9 can be formed by both ectodomain shedding and alternative mRNA splicing. A third mechanism involved with the generation of soluble cytokine receptors is the release of exosomes containing the protein. Since exosomes carry molecules expressed on the membrane of the cell from which they originated, the cytokine receptors expressed on these vesicles are usually present in the full-length form, as observed with TNFR1, IL-6R, and IL-15Rα.10–12
Due to their ability to modulate the effects of their cytokine ligands, either in an agonistic or antagonistic fashion, soluble cytokine receptors can mediate a variety of immunological functions. For example, the soluble IL-1R2 can bind to both IL-1α and IL-1β and act as a decoy receptor to inhibit IL-1 signaling.13,14 Another intriguing example is found in the IL-6/IL-6R pathway, generally associated with pro-inflammatory responses. Indeed, IL-6 signaling through sIL-6R was shown to induce a pro-inflammatory response, however, the membrane-bound counterpart (mIL-6R) has been associated with anti-inflammatory and protective effects. This happens due to different signaling pathways that are triggered by the different forms of the receptor, as the soluble form induces trans-signaling pathway, whereas the membrane-bound form is involved with the classic signaling pathway.15
Ectodomain shedding by proteases has recently been shown to be important in the regulation of the immune system. For instance, the cytokine receptor sheddase, tumor necrosis factor-α converting enzyme (TACE), was originally identified as a proteolytic enzyme capable of cleaving the TNF family receptors.16 More recently, TACE was renamed ADAM1717,18 and is one of the most well studied among these proteases whose cytokine receptor substrates include IL-6R, IL-1Rα, IL-15Rα, TNF-RI and TNF-RII.19 Further, ADAM17 can mediate cleavage of IL-6R and, therefore, be involved in the pathogenic effects mediated by the sIL-6R. Indeed, the production of sIL-6R was shown to be responsible for the involvement of IL-6 in the generation of pathogenic Th17 cells and their deleterious effects in the context of autoimmunity.20,21
The importance of the IL-7/CD127 in T cells function and biology is well established, and the presence of soluble IL-7Rα (sIL-7Rα, also known as sCD127) in the plasma has been described.22–24 A number of studies have demonstrated the impact of this soluble cytokine receptor both in health and in disease and will be reviewed here.
Biology of IL-7 and Its Receptor
IL-7 is a crucial cytokine for T cells homeostasis, as it plays a non-redundant role in the generation, development, survival, and function of T cells in humans.25–28 This cytokine is primarily produced by non-haematopoietic stromal cells in the bone marrow and thymus.29 Additionally, in humans, the production of IL-7 has been shown to occur in numerous non-lymphoid sites, such as intestinal epithelium,30,31 hepatic tissue,32,33 endothelial cells,34 keratinocytes,34 and neuronal progenitor cells in the brain.34 With regard to its production by immune cells, low amounts have been detected from DCs.35–38
IL-7 induces its response via signaling through its receptor, IL-7R, a heterodimer composed of two chains, an alpha chain, IL-7Rα (also known as CD127), and the common cytokine receptor γ chain (γc, also known as CD132). CD132 is shared with other cytokine receptors (ie IL-2, IL-4, IL-9, IL-15, IL-21) and expressed by most hematopoietic cells,39–41 whereas CD127 is also expressed on the surface of hematopoietic cells, but almost exclusively by the lymphoid lineage cells, throughout different stages of differentiation.42 CD127 is also shared with thymic stromal-derived lymphopoietin (TSLP) as part of a complex comprised of CD127 and TSLPR (thymic stromal-derived lymphopoietin receptor).43 IL-7 engagement with CD127 induces its heterodimerization with CD132, enabling the complex to induce phosphorylation of JAK1 and JAK3.44 Activated JAKs can, in turn, phosphorylate Y449, present on the intracellular tail of CD127, inducing anchoring and subsequent phosphorylation of STAT5. Activated STAT5 translocates to the nucleus, where it activates various transcription factors.40,45–47 Further, phosphorylated Y449 also activates PI3K and induces another downstream signaling cascade that results in AKT activation.48 These signaling cascades induce the increased expression of anti-apoptotic members of the Bcl-2 family, Bcl-2 and MCL1, and decreased expression of pro-apoptotic molecules, such as BAX and BIM, among other effects that culminate with the enhancement of cell survival and proliferation.49 The importance of this pathway is illustrated by the observation that T cells (particularly naïve and memory T cells) need IL-7 to be constantly available for their survival.50–53
As discussed above, IL-7 plays a pivotal role in T cells homeostasis, thus, dysregulation of IL-7-induced pathways has been linked to several immune-mediated conditions. Given its role in T cell proliferation, IL-7 is thought to be involved with abnormal immune responses associated with autoimmune diseases, such as multiple sclerosis and diabetes.54,55 One mechanism by which IL-7 is believed to play a role in the pathogenesis of disease is the alteration in the expression of its own receptor CD127.
The expression of membrane-bound CD127 (mCD127) is modulated, at least in part, by the amount of available IL-7, as high concentrations of the cytokine induce a decrease in mCD127 expression, as demonstrated in vitro.22,56 Further, genetic factors may also influence the levels of expression of mCD127. In this context, it is known that CD127, encoded by the IL7RA gene, is polymorphic and different single nucleotide polymorphisms (SNPs) are involved with generation of non-synonymous amino acid substitutions, potentially presenting negative impact on the expression of this cytokine receptor.57 Some IL7RA SNPs have been shown to impact the splicing of IL7RA transcripts, resulting in truncated proteins. Such a mechanism has been proposed as being the main mechanism responsible for the generation of the soluble form of CD127 (sCD127) as we describe further below.
Biological Functions of sCD127
The soluble form of CD127 (sCD127) was first described in 1990 by Goodwin and colleagues.3 The group performed an analysis of cDNA clones that encode mCD127. Interestingly, they found cDNA clones that encoded a truncated form of the protein lacking the transmembrane domain, corresponding to the secreted CD127 form.3 Since then, a number of studies have been performed to understand the mechanisms involved with sCD127 expression. In this regard, it has been shown that IL-7 induces up-regulation of sCD127, possibly a result of alternative splicing.3,58,59 Additionally, shedding of the protein from the cell membrane has been proposed to play a role.22 Given that sCD127 is capable of binding to IL-7 with a similar binding affinity as to the membrane-bound form,3 and is detected in plasma,60 several studies have been performed to shed light on the physiological roles of this soluble cytokine receptor.
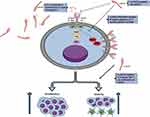
Initial studies from our group demonstrated that both plasma-derived and recombinant sCD127 significantly inhibited IL-7-mediated STAT5 and Akt phosphorylation, cell proliferation, and Bcl-2 expression in primary CD8+ T cells. These results suggest sCD127 has an antagonistic effect on IL-7 activity.24 Conversely, more recent studies demonstrated that sCD127 increases IL-7 bioactivity61 and enhances both IL-7-induced proliferation and viability,62 suggesting sCD127 has agonistic effects on IL-7-induced activity. Differences in the methodologies of these studies might explain the apparently conflicting findings. Specifically, in work from our group, the inclusion or exclusion of serum during critical steps of the experiments influenced the activity of sCD127.24,62 Given that fetal bovine serum is composed of a wide variety of proteins, lipids, and hormones, it is possible that, with the addition of serum, the interaction of any of these molecules with sCD127 might modulate the impact of sCD127 on IL-7-induced activity. More recently, preliminary results from our group demonstrate that sCD127 did not influence IL-7-mediated down regulation of mCD127 but did enhance IL-7-mediated proliferation of both CD4+ and CD8+ T cells when administered to healthy mice.63 Although the mechanism is not completely understood, these data also suggest sCD127 has agonistic effects on IL-7-induced activity. Figure 1 summarizes the mechanisms that have been considered to influence IL-7-induced effects on CD8+ T cells.
As demonstrated above, limited evidence supports a primarily agonistic role of sCD127 on IL-7-induced effects, though further studies are needed in order to clarify the impact of this soluble cytokine receptor on IL-7-mediated effects in health.
sCD127 in Disease
Given the clear importance of this soluble cytokine receptor in modulating IL-7-induced effects on T cells function, sCD127 has also been extensively studied in the context of various immune-mediated and infectious diseases, as discussed below.
Genetic Factors Affecting sCD127 Expression and Their Influence on Disease
Genetic polymorphism of the IL7RA gene, a number of which influence sCD127 expression, has been associated with predisposition to various immune-mediated diseases, including multiple sclerosis (MS), type I diabetes, and graft versus host disease.64–67 A SNP in the CD127 gene, rs6897932, is associated with increased levels of sCD127 in individuals carrying the C allele when compared to the T allele of this gene. This non-synonymous SNP has this functional impact, as it leads to skipping of CD127 exon 6 during gene transcription, and has been associated with various conditions, as discussed here and shown in Table 1.68 Further, beyond its impact on sCD127 expression, two studies reported that rs6897932-C allele was also associated with lower expression of mCD127.68,69 To our knowledge, there are no other published reports of an association between rs6897932 SNP and alteration in mCD127 expression.
 |
Table 1 Immune-Mediated Disorders Associated with rs6897932, the Risk Alleles, Known Functional Effects, and Disease-Specific Impact |
Another SNP that has been implicated in influencing sCD127 expression is the rs2523506 SNP in the DDX39B gene.70 DDX39B is a protein involved in pre-mRNA splicing and nuclear export of mRNAs.71–74 Galarza-Muños and colleagues have proposed that the DDX39B protein promotes the inclusion of exon 6, thus limiting sCD127 expression. The rs2523506 SNP risk allele A is, in fact, associated with the reduced expression of DDX39B, and higher levels of sCD127. Interestingly, this study also demonstrated a robust epistatic interaction between rs2523506 in DDX39B and rs6897932 in IL7RA, as an elevated risk of MS was only observed in carriers of at least one copy of the risk alleles at both loci. These results suggest these alleles can work in concert to increase skipping of IL7R exon 6, thereby inducing higher levels of sCD127.70 To date, there are no published reports of an association between rs2523506 and alterations in mCD127 expression.
Multiple Sclerosis (MS) and Other Demyelinating Autoimmune Disorders of the Central Nervous System (CNS)
MS is a potentially disabling disease of the CNS and the leading cause of non-traumatic neurological disability among young adults.75 This CNS autoimmune disorder is characterized by immune attack to the myelin sheath of neurons by auto-reactive T cells, leading to demyelination.76 By means of large, international genome wide association studies (GWAS), various genetic factors associated with the susceptibility to MS have been identified. Both HLA alleles, and some non-HLA polymorphisms have also been shown to be involved, including those related to CD127 expression.77–80
Polymorphisms in cytokine receptor genes have been associated with MS77 and one of the first identified was that of CD127.64,68,81 As mentioned above, in 2007, Gregory et al identified an SNP in the CD127 gene, rs6897932, where the C allele was associated with a significant risk for MS in a population of European descent and true for all clinical subtypes of MS (relapsing-remitting MS, secondary progressive MS, progressive-relapsing MS, primary progressive MS). Additionally, this study showed that the rs6897932 C allele also has a functional impact on gene expression, resulting in an increase in sCD127 levels. Thus, individuals carrying the C allele of rs6897932 express higher amounts of sCD127, when compared with the individuals with the T allele. This alteration would impact the ratio of sCD127/mCD127 expression, potentially affecting the IL-7/CD127 pathway, and therefore influencing T cells homeostasis.68
Since then, major studies have corroborated the findings that the rs6897932 C allele is associated with a higher susceptibility to MS in cohorts of Caucasian individuals.81–83 However, a number of other studies with individuals from Europe are not completely in line with the original study by Gregory et al No association between this SNP and an elevated risk for MS in individuals of European cohorts was reported by Broux et al, Rubio et al and Stankovic et al.84–86 Moreover, O’ Doherty et al observed this association in only one of two different MS cohorts.87 Similarly, Weber et al demonstrated an association between MS and the rs6897932 C allele in their French cohort, whereas it was not observed in their German cohort, however, a positive trend was observed in this population.88 Further, Kreft and colleagues were not able to detect higher levels of sCD127 in MS patients, observing lower concentrations of this soluble receptor, when compared to healthy controls. When stratifying the results according to rs6897932 allele, however, higher sCD127 levels correlated with rs6897932-C risk allele carriership in both MS and healthy controls.89
With regard to non-Caucasians, two studies performed with Japanese and Korean cohorts also demonstrated an association between rs6897932 SNP and the susceptibility to MS,90,91 whereas results from Iranian and Indian cohorts are not completely consistent with these findings.92–95 Despite having found an association between rs6897932 SNP and MS susceptibility, along with higher expression of sCD127 mRNA in an Iranian MS cohort, Sayad et al observed a higher frequency of the T allele in the MS patients, rather than the C allele, when compared to a control group.92 An association between the rs6897932 SNP and MS risk, also characterized by a higher frequency of the T allele in the MS patients was observed in three additional studies, with Iranian and Indian cohorts.93–95 Interestingly, a number of other studies performed with Iranian, Jordanian, and Chinese Han cohorts did not find any association between the rs6897932 SNP and the MS risk.96–98
With the conflicting observations outlined above, a number of meta-analyses have been performed in an attempt to clarify the impact of the CD127 rs6897932 C allele in the susceptibility to MS. In this sense, in 2014, Kim and colleagues reported that the SNP rs6897932 was associated with a higher risk for the development of MS in both Caucasians and Asians, potentially being a greater risk factor for individuals of Asian descent.90 In 2016, Tavakolpour et al concluded that there was not enough evidence to determine if SNP is indeed linked to a higher risk of MS, while Wu et al believe that this SNP is linked to a higher risk in Europeans, but not in Asians.99,100 Finally, very recently, Sahami-Fard and colleagues published an updated meta-analysis of the role of this SNP in MS susceptibility. Similarly, they concluded that the rs6897932 SNP might be a risk factor for MS in Europe but not in the Middle East.101
A growing body of evidence indicates that ethnicity might impact clinical manifestations of MS, possibly leading to different phenotypes of the disease.102,103 Interestingly, despite not demonstrating an association with susceptibility to develop MS, some groups have observed that rs6897932 SNP is associated with greater disease severity. Kulakova and colleagues observed that, in their Russian cohort, MS patients that carried the T allele showed favorable manifestations of the autoimmune disorder (optic neuritis or sensory disturbances), whereas carriers of the C allele presented with manifestations that are associated with worse prognosis (eg, motor disorders, brain stem disorders, impaired coordination).104 Furthermore, some studies have reported that the rs6897932 SNP was associated with a progressive courses of MS, indicating that this SNP may be a determinant of the disease course.105,106
Taken together, although MS is a complex and multifactorial disease, with various genetic and environmental factors involved, the balance of evidence suggests that the rs6897932 SNP is involved in its pathogenesis, either being associated with a higher susceptibility to MS or a determinant of the disease severity. The exact impact of each allele is, however, yet to be defined, particularly in non-Caucasian populations.
In addition to MS, the impact of the rs6897932 SNP has also been studied in a less common inflammatory demyelinating disease, known as Neuromyelitis Optica (NMO). NMO is a CNS autoimmune disease characterized by recurrent immune attacks to the spinal cord and optic nerve.107 For decades, NMO was considered as a severe subtype of MS, however, due to its different immunopathology, NMO and MS are now considered distinct entities. With regard to the influence of ethnicity, NMO is more frequently observed in non-Caucasian populations and non-Caucasian patients with NMO tend to have a more severe disease course than Caucasian individuals, particularly in the early phase of the disease.108–111
NMO has also been linked to alterations in the IL-7/CD127 pathway, with diminished expression of sCD127 and mCD127,112 however, evidence of a relationship between the rs6897932 SNP and NMO risk is limited and only available in few Asian and one Israeli cohort. Two studies have found an association between this SNP and NMO risk in Korean cohorts.90,96 Interestingly, Zhuang and colleagues observed an association of the rs6897932 SNP with NMO and not MS, whereas Kim et al demonstrated an association between this SNP and both NMO and MS. The association with NMO was observed especially in the group that was seropositive for aquaporin-4 (AQP4)-IgG, an antibody for AQP4, the most common water channel in the CNS, the detection of which is associated with a poorer prognosis.96 Conversely, three studies with Southern Han Chinese, Japanese and Israeli cohorts could not find an association between rs6897932 SNP and NMO risk.91,112,113 In summary, given the limited sample sizes and small number of studies performed in NMO, there is not enough evidence to confirm the role of the rs6897932 SNP in this disorder.
Other Autoimmune and Immune-Mediated Conditions
Beyond its association with CNS autoimmunity, numerous studies have suggested that the rs6897932 SNP might be involved with other autoimmune disorders and immune-mediated conditions. These include asthma,114 type 1 diabetes,115 sarcoidosis,116 rheumatoid arthritis,117 and systemic lupus erythematosus (SLE).118
In this context, Wang and colleagues reported that the rs6897932 C allele might be a susceptibility allele to SLE, as demonstrated in their Chinese cohort.118 With regard to asthma, an association of the rs6897932 C allele and increased risk of disease was demonstrated in an Indian cohort.114 Similarly, in a cohort with Type 1 Diabetes, Todd et al have observed an association between rs6897932 SNP and this autoimmune disorder.65 Subsequently, Santiago and colleagues have confirmed this association. Additionally, when the patients were stratified by age, they showed that rs6897932 T allele might be protective, particularly in patients with early age at disease onset.115 Also in line with these studies and what has been observed in most studies in MS, it was demonstrated that T1D patients homozygous for the C risk allele had higher circulating levels of sCD127 when compared to heterozygotes (CT) and patients without the risk allele (TT).119 One study of patients with sarcoidosis, a chronic granulomatous inflammatory condition of unknown etiology that most frequently affects the lungs, reported an association between the rs10213865 SNP, one other polymorphism in the IL7RA gene, and the disease. Despite rs10213865 SNP not being associated with a functional impact on CD127, a complete linkage disequilibrium between the rs10213865 SNP and the rs6897932 SNP was observed. These data suggest an association between IL7RA and the risk for sarcoidosis, however, a study evaluating the possible link between rs6897932 and sarcoidosis is still lacking.116 The rs6897932 SNP has also been shown to have a weak association with risk for rheumatoid arthritis (RA). Interestingly, the genotype that was associated with RA risk was TT,117 distinct from what is seen with MS. In graft versus host disease (GVHD), a common complication following allogeneic hematopoietic stem cell transplant (HSCT),120 donor carriage of the rs6897932 T allele was associated with lower levels of circulating sCD127 in the donors and a higher risk of acute GVHD in recipients. The patients whose donors carried the T allele also had a higher transplant-related mortality.121 Interestingly, Shamim and colleagues observed an association between rs6897932 T carriership in the donors and increased relapse of leukemia in patients that had gone through HSCT, however, in contrast with the findings previously mentioned, a trend towards reduced risk for GVHD was observed.122 Intriguingly, an association between the carriership of this SNP in the patients themselves and the risk for developing GVHD was not observed in any of these studies.
The general picture emerging from these studies is that the rs6897932 SNP probably plays a role in these various immune-mediated conditions, most likely due to its impact in IL-7/CD127 pathway, as a result of the shift in the relative expression of mCD127 and sCD127. In most cases, the T allele was shown to be protective, however, for some diseases, the T allele was associated with greater risk of disease. This is most likely due to the distinct immunopathological mechanisms involved with the various diseases that have been studied. Further evaluation of the possible association of this SNP with some disorders for which data is scarce (eg, sarcoidosis and asthma) is needed. Additionally, more research needs to be done in cohorts of different ethnic backgrounds, especially in conditions where ethnicity was shown to impact immunopathological aspects of the disease, like MS and NMO.
Cancer
The association between various cancers and the rs6897932 SNP has also been evaluated. Recently, with regard to breast cancer, patients homozygous for the rs6897932-T allele were shown to have an increased susceptibility for a subtype of a particularly aggressive breast tumor.123 Interestingly, despite being associated with the risk for this aggressive subtype, TT patients tended to have lower disease stages. This phenomenon is possibly due to enhanced IL-7R signaling that, through STAT5 activation, might induce a decrease in stem and mesenchymal cell markers in the tumor cells (eg, CD44), lower levels of which have been shown to be associated with better prognosis.123,124 Conversely, one study evaluated the relationship between rs6897932 and lymphoma (Hodgkin lymphoma and B cell lymphoma) and no significant difference was found in the distribution of the individual alleles or genotype frequencies between the controls and patients group.125
Infectious Diseases
Impairments in the IL-7/CD127 pathway have also been described in the context of different infectious diseases, and so, the impact on sCD127 expression and its association with genetic background have also been investigated. The potential relevance of the rs6897932 SNP in the context of infectious diseases and most frequently in the setting of HIV disease has been evaluated. HIV-1 infection, the cause of acquired immunodeficiency syndrome (AIDS), remains a great threat to the global public health. The introduction of HAART (highly active antiretroviral therapy), a cocktail of multiple drugs targeting different steps of the viral life cycle, has transformed HIV infection into a manageable chronic disease, dramatically reducing its morbidity and mortality. Despite the clear beneficial effects of HAART in HIV+ individuals, a degree of immune dysfunction persists, including altered expression of mCD127 and sCD127. Interestingly, a factor that might affect the impact of HAART on the altered expression of both mCD127 and sCD127 is how soon after infection treatment is initiated. In this regard, T cells derived from late presenting HIV+ individuals (CD4 counts <200 cells/ul before HAART) were shown to express lower mCD127 levels and higher sCD127 in the plasma when compared with both non-late presenters and healthy controls. Both mCD127 expression and sCD127 levels normalized after several months of HAART.126 Additionally, late presenting HIV+ individuals with the rs6897932-T allele were shown to have faster CD4+ recovery and higher CD4+ T cell counts at the end of the follow-up.127
As previously mentioned, alterations in the circulating levels of sCD127 in HIV+ individuals have been reported by a number of groups. Indeed, this was observed both in treated and untreated cohorts. Rose and colleagues observed lower levels of sCD127 in chronically HIV-infected individuals not receiving treatment.60 In contrast, studies from our group and Carini et al demonstrated increased sCD127 circulating levels in treated and untreated HIV infection.22,24,128 Moreover, one group reported no significant differences in the sCD127 levels in HIV+ patients.23 The discrepancies observed in these results are likely, at least in part, related to distinct methodologies used for the quantification of sCD127 in each of the studies. Interestingly, lower plasma levels of sCD127 have been associated with the rs6897932 TT genotype in HIV+ individuals.129 Further, the rs6897932 SNP has been also associated with immune recovery in HIV patients. In two Caucasian cohorts, the T allele was associated with a faster rate of CD4+ T cell recovery after initiation of HAART.129,130
In line with what was observed in MS, ethnic background or other factors appear to influence how the rs6897932 SNP impacts cell-mediated immune responses in the context of HIV. In a study performed with two cohorts, the rs6897932-T allele was not associated with faster recovery of CD4+ T cells in the African cohort, whereas this association was observed in the Caucasians group. In spite of these divergent results, an association was found between the rs6897932 T allele and lower sCD127 levels in both cohorts.131 With regard to disease progression, in a cohort from Zimbabwe, the rs6897932-T allele was shown to be associated with higher rates of mortality.132 In contrast, a study performed with two Spanish cohorts did not observe an association between this SNP and HIV disease progression.133
One frequent comorbidity observed in HIV-infected individuals is co-infection with hepatitis C virus (HCV).134 HCV causes chronic infection in about two-thirds of those infected, being responsible for chronic liver disease, liver failure and, hepatocellular carcinoma in a significant proportion of the patients. In HCV mono-infected individuals, the rs6897932-T allele has been shown to be associated with higher risk of liver fibrosis progression.135 With regard to HIV/HCV co-infected patients, however, María Guzmán-Fulgencio et al reported that those who were homozygous for the rs6897932-C allele were at increased risk of developing severe liver disease. They also observed that HIV/HCV coinfected individuals who were homozygous for the C allele had a higher likelihood of pathologically graded severe necroinflammatory activity.136 Given the limited data, it is not possible to conclude how each allele of the rs6897932 SNP impacts HCV progression, however, together, these data suggest this SNP might be a factor involved in the immunopathogenesis of HCV mono- and/or co-infection with HIV.
Role of sCD127 in Disease Pathogenesis
Association Between sCD127 Levels and Disease
sCD127 levels are altered in the plasma of patients suffering from a number of immune-related conditions where impairment in the IL-7 pathway has been suggested to play a role. As such, beyond the impact of genetic factors, the levels of this soluble cytokine receptor might be independently involved with the immune dysfunction observed in the context of the various diseases.
As previously discussed, sCD127 levels are altered in HIV infection and given the importance of CD8+ T cells function in virus-specific cell-mediated immune responses, the role of this cell compartment in the sCD127 expression has been evaluated. In this context, Carini and co-workers observed increased expression of sCD127 from CD8+ T cells isolated from HIV infected individuals.128 As suggested by the authors, this phenomenon might be related to HIV-1 proteins inducing down-regulation of mCD127.128 Indeed, downregulation of mCD127, in both CD4+ and CD8+ T cells derived from HIV+ individuals is well documented.137–139 Accordingly, Tat protein was shown to be involved in mCD127 downregulation from the surface of CD8+ T cells.140
In the setting of septic shock, sCD127 levels appear to be altered. Results, however, seem conflicting as sCD127 levels have been shown to be upregulated141 or downregulated142 when compared to healthy controls. Additionally, Delwarde et al observed lower levels of mRNA encoding total CD127, only mCD127 or only sCD127 in patients with septic shock.143 Interestingly, when stratifying the patients group into survivors and non-survivors, both Demaret et al and Peronnet et al showed sCD127 levels are higher in patients who survived when compared to non-survivors.142,144 It must be noted that the study performed by Peronnet and colleagues included a mixed ICU population and not only individuals with septic shock.142
Impaired sensitivity to IL-7 has been demonstrated in patients with tuberculosis.145 In one study, patients with tuberculosis also presented with lower sCD127 and higher IL-7 levels in the plasma when compared to healthy controls. sCD127 levels increased in response to the treatment, reaching levels that were comparable to the ones observed in the healthy controls.145 Interestingly, in non-human primates, it was shown that animals that were vaccinated with rBCG had increased survival and higher CD127 mRNA levels in the lungs, with most of it encoding for sCD127.146
As for autoimmune disorders, in the context of T1D, detection of autoantibodies against antigens found in secretory granules within pancreatic islet cells and that are involved in the pathogenesis of T1D was shown to be associated with higher circulating levels of CD127.119 Interestingly, the authors also demonstrated that sCD127 is glycated in patients with T1D and that glycated sCD127 is not as effective at inhibiting T cells expansion as is un-glycated sCD127.119 In patients with SLE, plasma levels of sCD127 were correlated with SLEDAI score (a marker of kidney disease activity) and anti-C1q antibody levels, suggesting an association between the soluble cytokine receptor and SLE disease activity.147,148 With regard to patients with RA who were refractory to disease-modifying anti-rheumatic drugs therapy and treated with a TNFα inhibitor, elevated sCD127 serum levels strongly predicted a poor response to the anti-TNF therapy.149 Additionally, regarding primary Sjogren’s syndrome, an autoimmune condition characterized by inflammation of exocrine glans, especially salivary and lacrimal glands, has been shown to be associated with elevated levels of circulating sCD127. Further, the levels of CCL25, a chemokine involved with recruiting CCR9+ Th cells to the salivary glands, were strongly correlated with sCD127 levels.150 As it has been suggested that sCD127 enhances IL-7 activity in vivo,61 this correlation between sCD127 and CCL25 levels may indicate that this soluble cytokine receptor plays a role in the inflammation of the salivary glands.150
Finally, with regard to Sézary Syndrome, a rare and severe variant of cutaneous T cell lymphoma, has been associated with impaired adaptive immune responses, along with IL-7/IL-7R pathway disturbances and has recently been shown to be associated with higher levels of sCD127 in the serum, along with lower levels of IL-7.151
sCD127 as a Potential Biomarker and Therapeutic Agent
As sCD127 levels are altered in several disorders, it is possible that sCD127 levels could be used as a biomarker, to better define the prognosis or even as a tool to evaluate disease progression or the response to the therapy. For instance, it is known that due to the heterogeneous immunopathology associated with septic shock, deciding on the optimal therapeutic approach for each case may not be straightforward.152 In this setting, plasma sCD127 levels were proposed as a possible biomarker for the identification of septic shock patients who were at higher risk of death, as the circulating levels of this soluble cytokine receptor were elevated in non-survivors within the first 4 days after the onset of the shock.144 These results are in line with a study performed with ICU patients, where sCD127 levels were also higher in patients who did not survive when compared to survivors.142 Further studies should be performed to evaluate the possibility of using sCD127 plasma levels as a tool that would help determine which septic shock patients would have a higher risk of death.
As for tuberculosis, Lundtoft et al suggest sCD127 along with IL-7 levels could be used as a biomarker for diagnosis of the disease. By receiver operator curve analysis of a cohort of adult patients with tuberculosis and health controls in Ghana, accurate prediction was achieved for 73% of all donors. Given that the plasma levels of both IL-7 and sCD127 normalize following the treatment, the combined use of IL-7 and sCD127 could, thus, be used as a biomarker of a successful tuberculosis chemotherapy. Future studies in the setting of tuberculosis are clearly needed.145
Concerning autoimmune disorders, in patients with MS, in an attempt to find a less expensive and more practical alternative to MRI to use for predicting clinical outcomes, Bedri and colleagues evaluated a number of soluble cytokine receptors in the plasma of MS patients treated with a variety of drugs. They observed that in patients homozygous for the r6897932-C allele, treatment with fingolimod induced a more significant increase in sCD127 levels, than in the other groups. The group also reported that different treatments have been shown to impact sCD127 levels differently, thus, sCD127 levels might be exploited as a possible biomarker of treatment outcome, as they propose.153
In individuals with SLE, the use of sCD127 serum levels has also been proposed to be used as a biomarker. This is the result of two studies that demonstrated sCD127 levels to be correlated to SLEDAI score, a marker of kidney disease activity in SLE, especially in patients with lupus nephritis.147,148 Interestingly, Chi and collaborators also observed a correlation between sCD127 plasma levels and anti-C1q antibodies in SLE patients.148 These studies suggest that screening of sCD127 plasma levels might be a useful tool to assess disease activity in SLE patients. In the context of RA, given that elevated sCD127 levels predicted poor response to anti-TNF therapy, the quantification of the soluble cytokine receptor could potentially be used as a biomarker to predict response to treatment, potentially helping avoid an ineffective therapy.149
Finally, with regard to GVHD, studies have suggested that quantification of plasma levels of sCD127 in the donor might be useful to predict the risk of development of GVHD. In this sense, POIRET and colleagues have shown that low sCD127 levels were linked to higher risk of developing GVHD in any grade.154 Additionally, an elevated IL-7/sCD127 ratio was shown to be a predictor of moderate to severe aGVHD.121
Given their ability to influence the effects induced by their ligands, soluble cytokine receptors have been studied in the treatment of several immune-mediated disorders characterized by the detrimental effects caused by the cytokine ligand. One interesting example, currently being studied as a possible therapy, is the administration of sgp130 in autoimmune disorders where IL-6-induced pathogenesis was identified. gp130 (CD130) is a co-receptor and part of the membrane-bound IL-6R receptor complex, which mediates signal transduction following IL-6 engagement. Interestingly, its soluble form, sgp130, has an antagonistic effect as it inhibits the trans-signaling pathway by binding to IL-6/sIL-6R complexes. Given that the trans-signaling pathway has been associated with IL-6-induced pathogenesis, whereas the classic pathway not only possesses a great deal of importance in innate immunity but also does not seem to be associated with disease pathogenesis, this strategy is expected to be safer than current therapies that also block mIL-6R signaling (ie, tocilizumab). The use of sgp130 fused to Fc (Olamkicept) is currently in Phase II clinical trials for inflammatory bowel disease.155 Moreover, since IL-6 may play an important role in the cytokine storm associated with infection by SARS-CoV2 (COVID-19), a clinical trial for the use of tocilizumab has started and the possible use of sgp130 has also been considered in this setting.156,157
One other example of a soluble cytokine receptor, currently in use for treatment of rheumatoid arthritis, psoriasis, chronic plaque psoriasis, and others, is the TNFR2-Fc (Etanercept).158,159 Etanercept is a protein composed of two molecules of the soluble cytokine receptor TNFR2 fused to human IgG1. Through binding to TNF-α, Etanercept interferes with its interaction with membrane-bound TNF receptors, thereby acting as an effective TNF-α inhibitor.158,159 Indeed, this medication is the first line biologic disease-modifying antirheumatic drug for treatment of individuals with psoriasis with active dactylitis or enthesitis, despite treatment with methotrexate.160,161
Since sCD127 has been shown to modulate IL-7 activity, this soluble cytokine receptor might also be a candidate for use in treatment of a number of conditions in which IL-7 is implicated in the pathophysiology. In a murine model of lung cancer, administration of IL-7/IL-7Rα-Fc resulted in higher levels of inflammation in the tumor environment, enhancing APC activity and, consequently, T cell activity, when compared to IL-7 alone. Further, this treatment was able to induce an anti-tumor environment, attributed to higher levels of IFN-γ, CXCL-9 and CXCL-10. The treatment also resulted in tumor size reduction, along with enhanced survival.162
Administration of sCD127, in a mouse model of MS (ie, experimental autoimmune encephalomyelitis (EAE)), induces an increase in the bioactivity of IL-7. This was associated with the exacerbation of disease as characterized by greater symptoms of EAE in the animals that were injected with both IL-7 and sCD127 when compared to animals receiving IL-7 alone. The sCD127-induced increase in IL-7 bioactivity in these experiments may be due to limiting the consumption of IL-7, but also because sCD127 impairs the induction of negative regulators of IL-7 signaling (SOCS-1 and CD95), facilitating inducing IL-7-mediated homeostatic expansion.61 Given that exacerbation of EAE was observed in this study, the administration of this soluble cytokine receptor must be considered carefully in future studies in order to avoid the possibility of triggering autoimmunity.61 Lastly, preliminary results by our group demonstrate that, following T-cell depletion in healthy mice, administration of sCD127, along with IL-7, induces a more rapid reconstitution of T cells, particularly in the CD8+ T cell compartment.63 These data suggest administration of sCD127, in combination with IL-7, could potentially be useful in the treatment of lymphopenia in situations, such as following chemo or radiation therapy.
Concluding Statement
Given the abundance of evidence, it is clear that sCD127 plays an important role in modulating IL-7-dependent activities, both in health and in disease. Several studies suggest a role for sCD127 in the immunopathogenesis of a variety of conditions, as is the case with other soluble cytokine receptors. Future studies should better elucidate the impact of rs6897932 SNP in different ethnicities, especially in conditions where ethnic background was shown to impact clinical outcomes. The possible role of sCD127 as a biomarker, as supported by studies in septic shock, SLE and GVHD, could prove to be useful in guiding prognosis and be an important tool used to optimize individual therapeutic approaches. As for the potential use of sCD127 in therapy, this possibility should be better studied, with appropriate caution, given the association between sCD127 and autoimmunity. Collectively, limited but accumulating data suggest sCD127 plays an important role in modulating the IL-7/CD127 pathway, likely in an agonistic fashion. Due to the heterogenicity of the disorders discussed here, this activity of sCD127 likely has different implications. Ultimately sCD127 might prove to be a useful tool for the therapy in settings where there is a need to restore or enhance IL-7 activity or a potential biomarker to better guide clinical decision-making.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Rubin LA, Kurman CC, Fritz ME, et al. Soluble interleukin 2 receptors are released from activated human lymphoid cells in vitro. J Immunol. 1985;135(5):3172–3177.
2. Renauld JC, Druez C, Kermouni A, et al. Expression cloning of the murine and human interleukin 9 receptor cDNAs. Proc Natl Acad Sci U S A. 1992;89(12):5690–5694. doi:10.1073/pnas.89.12.5690
3. Goodwin RG, Friend D, Ziegler SF, et al. Cloning of the human and murine interleukin-7 receptors: demonstration of a soluble form and homology to a new receptor superfamily. Cell. 1990;60(6):941–951. doi:10.1016/0092-8674(90)90342-C
4. Mosley B, Beckmann MP, March CJ, et al. The murine interleukin-4 receptor: molecular cloning and characterization of secreted and membrane bound forms. Cell. 1989;59(2):335–348. doi:10.1016/0092-8674(89)90295-X
5. Tavernier J, Devos R, Cornelis S, et al. A human high affinity interleukin-5 receptor (IL5R) is composed of an IL5-specific α chain and a β chain shared with the receptor for GM-CSF. Cell. 1991;66(6):1175–1184. doi:10.1016/0092-8674(91)90040-6
6. Orlando S, Sironi M, Bianchi G, et al. Role of metalloproteases in the release of the IL-1 type II decoy receptor. J Biol Chem. 1997;272(50):31764–31769. doi:10.1074/jbc.272.50.31764
7. Jensen LE, Muzio M, Mantovani A, Whitehead AS. IL-1 signaling cascade in liver cells and the involvement of a soluble form of the IL-1 receptor accessory protein. J Immunol. 2000;164(10):5277–5286. doi:10.4049/jimmunol.164.10.5277
8. Mülberg J, Schooltink H, Stoyan T, et al. The soluble interleukin‐6 receptor is generated by shedding. Eur J Immunol. 1993;23(2):473–480. doi:10.1002/eji.1830230226
9. Horiuchi S, Koyanagiu Y, Zhouu Y, et al. Soluble interleukin‐6 receptors released from T cell or granulocyte/macrophage cell lines and human peripheral blood mononuclear cells are generated through an alternative splicing mechanism. Eur J Immunol. 1994;24(8):1945–1948. doi:10.1002/eji.1830240837
10. Schumacher N, Meyer D, Mauermann A, et al. Shedding of endogenous interleukin-6 receptor (il-6r) is governed by a disintegrin and metalloproteinase (ADAM) proteases while a full-length IL-6r isoform localizes to circulating microvesicles. J Biol Chem. 2015;290(43):26059–26071. doi:10.1074/jbc.M115.649509
11. Hawari FI, Rouhani FN, Cui X, et al. Release of full-length 55-kDa TNF receptor 1 in exosome-like vesicles: a mechanism for generation of soluble cytokine receptors. Proc Natl Acad Sci U S A. 2004;101(5):1297–1302. doi:10.1073/pnas.0307981100
12. Viaud S, Terme M, Flament C, et al. Dendritic cell-derived exosomes promote natural killer cell activation and proliferation: a role for NKG2D ligands and IL-15Rα. Pockley G ed. PLoS One. 2009;4(3):e4942. doi:10.1371/journal.pone.0004942
13. Arend WP, Malyak M, Smith MF, et al. Binding of IL-1 alpha, IL-1 beta, and IL-1 receptor antagonist by soluble IL-1 receptors and levels of soluble IL-1 receptors in synovial fluids. J Immunol. 1994;153(10):4766–4774.
14. Cui X, Rouhani FN, Hawari F, Levine SJ. Shedding of the type II IL-1 decoy receptor requires a multifunctional aminopeptidase, aminopeptidase regulator of TNF receptor type 1 shedding. J Immunol. 2003;171(12):6814–6819. doi:10.4049/jimmunol.171.12.6814
15. Zegeye MM, Lindkvist M, Fälker K, et al. Activation of the JAK/STAT3 and PI3K/AKT pathways are crucial for IL-6 trans-signaling-mediated pro-inflammatory response in human vascular endothelial cells. Cell Commun Signal. 2018;16(1):55. doi:10.1186/s12964-018-0268-4
16. Mohler KM, Sleath PR, Fitzner JN, et al. Protection against a lethal dose of endotoxin by an inhibitor of tumour necrosis factor processing. Nature. 1994;370(6486):218–220. doi:10.1038/370218a0
17. Black RA, Rauch CT, Kozlosky CJ, et al. A metalloproteinase disintegrin that releases tumour-necrosis factor-α from cells. Nature. 1997;385(6618):729–733. doi:10.1038/385729a0
18. Moss ML, Jin S-LC, Milla ME, et al. Cloning of a disintegrin metalloproteinase that processes precursor tumour-necrosis factor-α. Nature. 1997;385(6618):733–736. doi:10.1038/385733a0
19. Zunke F, Rose-John S. The shedding protease ADAM17: physiology and pathophysiology. Biochim Biophys Acta. 2017;1864(11):2059–2070. doi:10.1016/J.BBAMCR.2017.07.001
20. Wolf J, Waetzig GH, Reinheimer TM, Scheller J, Rose-John S, Garbers C. A soluble form of the interleukin-6 family signal transducer gp130 is dimerized via a C-terminal disulfide bridge resulting from alternative mRNA splicing. Biochem Biophys Res Commun. 2016;470(4):870–876. doi:10.1016/j.bbrc.2016.01.127
21. Heink S, Yogev N, Garbers C, et al. Trans-presentation of IL-6 by dendritic cells is required for the priming of pathogenic TH17 cells. Nat Immunol. 2016;18(1):74–85. doi:10.1038/ni.3632
22. Vranjkovic A, Crawley AM, Gee K, Kumar A, Angel JB. IL-7 decreases IL-7 receptor (CD127) expression and induces the shedding of CD127 by human CD8+ T cells. Int Immunol. 2007;19(12):1329–1339. doi:10.1093/intimm/dxm102
23. Blom-Potar M-C, Bugault F, Lambotte O, Delfraissy J-F, Thèze J. Soluble IL-7Ralpha (sCD127) and measurement of IL-7 in the plasma of HIV patients. J Acquir Immune Defic Syndr. 2009;51(1):104–105. doi:10.1097/QAI.0b013e31819d8a20
24. Crawley AM, Faucher S, Angel JB. Soluble IL-7Rα (sCD127) inhibits IL-7 activity and is increased in HIV infection. J Immunol. 2010;184(9):4679–4687. doi:10.4049/jimmunol.0903758
25. Schluns KS, Lefrançois L. Cytokine control of memory T-cell development and survival. Nat Rev Immunol. 2003;3(4):269–279. doi:10.1038/nri1052
26. Okamoto Y, Douek DC, McFarland RD, Koup RA. IL-7, the thymus, and naiïve T cells. Adv Exp Med Biol. 2002;512:81–90. doi:10.1007/978-1-4615-0757-4_11
27. Marrack P, Kappler J. Control of T cell viability. Annu Rev Immunol. 2004;22(1):765–787. doi:10.1146/annurev.immunol.22.012703.104554
28. Khaled AR, Durum SK. Lymphocide: cytokines and the control of lymphoid homeostasis. Nat Rev Immunol. 2002;2(11):817–830. doi:10.1038/nri931
29. Zaunders JJ, Lévy Y, Seddiki N. Exploiting differential expression of the IL-7 receptor on memory T cells to modulate immune responses. Cytokine Growth Factor Rev. 2014;25(4):391–401. doi:10.1016/J.CYTOGFR.2014.07.012
30. Madrigal-Estebas L, McManus R, Byrne B, et al. Human small intestinal epithelial cells secrete interleukin-7 and differentially express two different interleukin-7 mRNA transcripts: implications for extrathymic T-cell differentiation. Hum Immunol. 1997;58(2):83–90. doi:10.1016/S0198-8859(97)00230-9
31. Watanabe M, Ueno Y, Yajima T, et al. Interleukin 7 is produced by human intestinal epithelial cells and regulates the proliferation of intestinal mucosal lymphocytes. J Clin Invest. 1995;95(6):2945–2953. doi:10.1172/JCI118002
32. Golden-Mason L, Kelly AM, Traynor O, et al. Expression of interleukin 7 (IL-7) mRNA and protein in the normal adult human liver: implications for extrathymic T cell development. Cytokine. 2001;14(3):143–151. doi:10.1006/cyto.2001.0852
33. G-r JC, Olsson C, Palacios R. Interleukin (IL1 to IL7) gene expression in fetal liver and bone marrow stromal clones: cytokine-mediated positive and negative regulation. Exp Hematol. 1992;20(8):986–990.
34. Heufler C, Topar G, Grasseger A, et al. Interleukin 7 is produced by murine and human keratinocytes. J Exp Med. 1993;178(3):1109–1114. doi:10.1084/jem.178.3.1109
35. de Saint-vis B, Fugier-Vivier I, Massacrier C, et al. The cytokine profile expressed by human dendritic cells is dependent on cell subtype and mode of activation. J Immunol. 1998;160(4):1666–1676.
36. Sorg RV, Mclellan AD, Hock BD, Fearnley DB, Hart DNJ. Human dendritic cells express functional interleukin-7. Immunobiology. 1998;198(5):514–526. doi:10.1016/S0171-2985(98)80075-2
37. Kröncke R, Loppnow H, Flad H-D, Gerdes J. Human follicular dendritic cells and vascular cells produce interleukin-7: a potential role for interleukin-7 in the germinal center reaction. Eur J Immunol. 1996;26(10):2541–2544. doi:10.1002/eji.1830261040
38. Guimond M, Veenstra RG, Grindler DJ, et al. Interleukin 7 signaling in dendritic cells regulates the homeostatic proliferation and niche size of CD4+ T cells. Nat Immunol. 2009;10(2):149–157. doi:10.1038/ni.1695
39. Noguchi M, Nakamura Y, Russell SM, et al. Interleukin-2 receptor gamma chain: a functional component of the interleukin-7 receptor. Science. 1993;262(5141):1877–1880. doi:10.1126/science.8266077
40. Jiang Q, Wen QL, Aiello FB, et al. Cell biology of IL-7, a key lymphotrophin. Cytokine Growth Factor Rev. 2005;16(4–5):513–533. doi:10.1016/j.cytogfr.2005.05.004
41. Spolski R, Gromer D, Leonard WJ. The γc family of cytokines: fine-tuning signals from IL-2 and IL-21 in the regulation of the immune response. F1000Research. 2017;6:1872. doi:10.12688/f1000research.12202.1
42. Mazzucchelli R, Durum SK. Interleukin-7 receptor expression: intelligent design. Nat Rev Immunol. 2007;7(2):144–154. doi:10.1038/nri2023
43. Park LS, Martin U, Garka K, et al. Cloning of the murine thymic stromal lymphopoietin (TSLP) receptor: formation of a functional heteromeric complex requires interleukin 7 receptor. J Exp Med. 2000;192(5):659–669. doi:10.1084/jem.192.5.659
44. McElroy CA, Holland PJ, Zhao P, et al. Structural reorganization of the interleukin-7 signaling complex. Proc Natl Acad Sci U S A. 2012;109(7):2503–2508. doi:10.1073/pnas.1116582109
45. Palmer MJ, Mahajan VS, Trajman LC, Irvine DJ, Lauffenburger DA, Chen J. Interleukin-7 receptor signaling network: an integrated systems perspective. Cell Mol Immunol. 2008;5(2):79–89. doi:10.1038/cmi.2008.10
46. Jiang Q, Li WQ, Aiello FB, Klarmann KD, Keller JR, Durum SK. Retroviral transduction of IL-7Rα into IL-7Rα-/- bone marrow progenitors: correction of lymphoid deficiency and induction of neutrophilia. Gene Ther. 2005;12(24):1761–1768. doi:10.1038/sj.gt.3302558
47. Jiang Q, Li WQ, Hofmeister RR, et al. Distinct regions of the interleukin-7 receptor regulate different bcl2 family members. Mol Cell Biol. 2004;24(14):6501–6513. doi:10.1128/mcb.24.14.6501-6513.2004
48. Venkitaraman AR, Cowling RJ. Interleukin‐7 induces the association of phosphatidylinositol 3‐kinase with the α chain of the interleukin‐7 receptor. Eur J Immunol. 1994;24(9):2168–2174. doi:10.1002/eji.1830240935
49. MacKall CL, Fry TJ, Gress RE. Harnessing the biology of IL-7 for therapeutic application. Nat Rev Immunol. 2011;11(5):330–342. doi:10.1038/nri2970
50. Sprent J, Surh CD. Normal T cell homeostasis: the conversion of naive cells into memory-phenotype cells. Nat Immunol. 2011;12(6):478–484. doi:10.1038/ni.2018
51. Fry TJ, Mackall CL. Interleukin-7: from bench to clinic. Blood. 2002;99(11):3892–3904. doi:10.1182/blood.V99.11.3892
52. Opferman JT, Letai A, Beard C, Sorcinelli MD, Ong CC, Korsmeyer SJ. Development and maintenance of B and T lymphocytes requires antiapoptotic MCL-1. Nature. 2003;426(6967):671–676. doi:10.1038/nature02067
53. Rathmell JC, Farkash EA, Gao W, Thompson CB. IL-7 enhances the survival and maintains the size of naive T cells. J Immunol. 2001;167(12):6869–6876. doi:10.4049/jimmunol.167.12.6869
54. Lee LF, Axtell R, Tu GH, et al. IL-7 promotes TH1 development and serum IL-7 predicts clinical response to interferon-β in multiple sclerosis. Sci Transl Med. 2011;3(93):93ra68–93ra68. doi:10.1126/scitranslmed.3002400
55. Lee L-F, Logronio K, Tu GH, et al. Anti-IL-7 receptor-α reverses established type 1 diabetes in nonobese diabetic mice by modulating effector T-cell function. Proc Natl Acad Sci U S A. 2012;109(31):12674–12679. doi:10.1073/pnas.1203795109
56. Park J-H, Yu Q, Erman B, et al. Suppression of IL7Rα transcription by IL-7 and other prosurvival cytokines: a novel mechanism for maximizing IL-7-dependent T cell survival. Immunity. 2004;21(2):289–302. doi:10.1016/j.immuni.2004.07.016
57. Teutsch SM, Booth DR, Bennetts BH, Heard RNSN, Stewart GJ. Identification of 11 novel and common single nucleotide polymorphisms in the interleukin-7 receptor-a gene and their associations with multiple sclerosis. Eur J Hum Genet. 2003;11(7):509–515. doi:10.1038/sj.ejhg.5200994
58. Korte A, Köchling J, Badiali L, et al. Expression analysis and characterization of alternatively spliced transcripts of human IL-7Rα chain encoding two truncated receptor proteins in relapsed childhood all. Cytokine. 2000;12(11):1597–1608. doi:10.1006/cyto.2000.0777
59. Rane L, Vudattu N, Bourcier K, et al. Alternative splicing of interleukin-7 (IL-7) and interleukin-7 receptor alpha (IL-7Rα) in peripheral blood from patients with multiple sclerosis (MS). J Neuroimmunol. 2010;222(1–2):82–86. doi:10.1016/j.jneuroim.2010.02.014
60. Rose T, Lambotte O, Pallier C, Delfraissy J-F, Colle J-H. Identification and biochemical characterization of human plasma soluble IL-7R: lower concentrations in HIV-1-infected patients. J Immunol. 2009;182(12):7389–7397. doi:10.4049/jimmunol.0900190
61. Lundström W, Highfill S, Walsh STR, et al. Soluble IL7Rα potentiates IL-7 bioactivity and promotes autoimmunity. Proc Natl Acad Sci U S A. 2013;110(19):E1761–1770. doi:10.1073/pnas.1222303110
62. Côté S, Matte J, Sad S, Angel JB, Crawley AM. Complexed soluble IL-7 receptor α and IL-7 increase IL-7-mediated proliferation and viability of CD8+ T-cells in vitro. Cell Immunol. 2015;293(2):122–125. doi:10.1016/j.cellimm.2015.01.001
63. Aloufi N The role of sCD127 in mediating IL-7 function on T cell homeostasis in vivo, PhD thesis, University of Ottawa, 2020. Available from: https://ruor.uottawa.ca/bitstream/10393/41089/1/Aloufi_Nawaf_2020_Thesis.pdf. Accessed February 26, 2021.
64. Hafler DA, Compston A, Sawcer S, et al. Risk alleles for multiple sclerosis identified by a genomewide study. N Engl J Med. 2007;357(9):851–862. doi:10.1056/NEJMoa073493
65. Todd JA, Walker NM, Cooper JD, et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat Genet. 2007;39(7):857–864. doi:10.1038/ng2068
66. Shamim Z, Ryder LP, Heilmann C, et al. Genetic polymorphisms in the genes encoding human interleukin-7 receptor-α: prognostic significance in allogeneic stem cell transplantation. Bone Marrow Transplant. 2006;37(5):485–491. doi:10.1038/sj.bmt.1705277
67. Shamim Z, Ryder LP, Christensen IJ, et al. Prognostic significance of interleukin-7 receptor-α gene polymorphisms in allogeneic stem-cell transplantation: a confirmatory study. Transplantation. 2011;91(7):1. doi:10.1097/TP.0b013e31820f08b2
68. Gregory SG, Schmidt S, Seth P, et al. Interleukin 7 receptor α chain (IL7R) shows allelic and functional association with multiple sclerosis. Nat Genet. 2007;39(9):1083–1091. doi:10.1038/ng2103
69. Jäger J, Schulze C, Rö Sner S, Martin R. IL7RA haplotype-associated alterations in cellular immune function and gene expression patterns in multiple sclerosis. Genes Immun. 2013;14(7):453–461. doi:10.1038/gene.2013.40
70. Galarza-Muñoz G, Briggs FBS, Evsyukova I, et al. Human epistatic interaction controls IL7R splicing and increases multiple sclerosis risk. Cell. 2017;169(1):72–84.e13. doi:10.1016/j.cell.2017.03.007
71. Masuda S, Das R, Cheng H, Hurt E, Dorman N, Reed R. Recruitment of the human TREX complex to mRNA during splicing. Genes Dev. 2005;19(13):1512–1517. doi:10.1101/gad.1302205
72. Luo MJ, Zhou Z, Magni K, et al. Pre-mRNA splicing and mRNA export linked by direct interactions between UAP56 and aly. Nature. 2001;413(6856):644–647. doi:10.1038/35098106
73. Shen J, Zhang L, Zhao R. Biochemical characterization of the ATPase and helicase activity of UAP56, an essential pre-mRNA splicing and mRNA export factor. J Biol Chem. 2007;282(31):22544–22550. doi:10.1074/jbc.M702304200
74. Shen H, Zheng X, Shen J, Zhang L, Zhao R, Green MR. Distinct activities of the DExD/H-box splicing factor hUAP56 facilitate stepwise assembly of the spliceosome. Genes Dev. 2008;22(13):1796–1803. doi:10.1101/gad.1657308
75. Koch-Henriksen N, Sørensen PS. The changing demographic pattern of multiple sclerosis epidemiology. Lancet Neurol. 2010;9(5):520–532. doi:10.1016/S1474-4422(10)70064-8
76. Ghasemi N, Razavi S, Nikzad E. Multiple sclerosis: pathogenesis, symptoms, diagnoses and cell-based therapy. Cell J. 2017;19(191):1–10. doi:10.22074/cellj.2016.4867
77. Beecham AH, Patsopoulos NA, Xifara DK, et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat Genet. 2013;45(11):1353–1360. doi:10.1038/ng.2770
78. Sawcer S, Hellenthal G, Pirinen M, et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature. 2011;476(7359):214–219. doi:10.1038/nature10251
79. Steinman L, Bar-Or A, Behne JM, et al. Restoring immune tolerance in neuromyelitis optica: part II. Neurol Neuroimmunol Neuroinflamm. 2016;3(5):e277. doi:10.1212/NXI.0000000000000277
80. Liu H, Huang J, Dou M, et al. Variants in the IL7RA gene confer susceptibility to multiple sclerosis in Caucasians: evidence based on 9734 cases and 10436 controls. Sci Rep. 2017;7(1):1–10. doi:10.1038/s41598-017-01345-8
81. Lundmark F, Duvefelt K, Iacobaeus E, et al. Variation in interleukin 7 receptor alpha chain (IL7R) influences risk of multiple sclerosis. Nat Genet. 2007;39(9):1108–1113. doi:10.1038/ng2106
82. Alcina A, Fedetz M, Ndagire D, et al. The T244I variant of the interleukin-7 receptor-alpha gene and multiple sclerosis. Tissue Antigens. 2008;72(2):158–161. doi:10.1111/j.1399-0039.2008.01075.x
83. Sombekke MH, Van Der Voort LF, Kragt JJ, et al. Relevance of IL7R genotype and mRNA expression in Dutch patients with multiple sclerosis. Mult Scler J. 2011;17(8):922–930. doi:10.1177/1352458511402411
84. Stanković A, Dinčić E, Ristić S, et al. Interleukin 7 receptor alpha polymorphism rs6897932 and susceptibility to multiple sclerosis in the Western Balkans. Mult Scler. 2010;16(5):533–536. doi:10.1177/1352458509360548
85. Broux B, Hellings N, Venken K, et al. Haplotype 4 of the multiple sclerosis-associated interleukin-7 receptor alpha gene influences the frequency of recent thymic emigrants. Genes Immun. 2010;11(4):326–333. doi:10.1038/gene.2009.106
86. Rubio JP, Stankovich J, Field J, et al. Replication of KIAA0350, IL2RA, RPL5 and CD58 as multiple sclerosis susceptibility genes in Australians. Genes Immun. 2008;9(7):624–630. doi:10.1038/gene.2008.59
87. O’Doherty C, Kantarci O, Vandenbroeck K. IL7RA polymorphisms and susceptibility to multiple sclerosis. N Engl J Med. 2008;358(7):753–754. doi:10.1056/NEJMc0707553
88. Weber F, Fontaine B, Cournu-Rebeix I, et al. IL2RA and IL7RA genes confer susceptibility for multiple sclerosis in two independent European populations. Genes Immun. 2008;9(3):259–263. doi:10.1038/gene.2008.14
89. Kreft KL, Verbraak E, Wierenga-Wolf AF, et al. Decreased systemic IL-7 and soluble IL-7Rα in multiple sclerosis patients. Genes Immun. 2012;13(7):587–592. doi:10.1038/gene.2012.34
90. Kim JY, Cheong HS, Kim HJ, Kim LH, Namgoong S, Shin HD. Association analysis of IL7R polymorphisms with inflammatory demyelinating diseases. Mol Med Rep. 2014;9(2):737–743. doi:10.3892/mmr.2013.1863
91. Fang L, Isobe N, Yoshimura S, et al. Interleukin-7 receptor alpha gene polymorphism influences multiple sclerosis risk in Asians. Neurology. 2011;76(24):2125–2127. doi:10.1212/WNL.0b013e31821f466c
92. Sayad A, Omrani MD, Solgi G, et al. Interleukin 7 receptor alpha gene variants are correlated with gene expression in patients with relapsing-remitting multiple sclerosis. Iran J Allergy Asthma Immunol. 2017;16(4):338–346.
93. Taheri M, Sayad A. Investigating the exon 6 sequence changes of interleukin 7 receptor A (IL7RA) gene in patients with relapsing-remitting multiple sclerosis. Hum Antibodies. 2017;26(2):43–48. doi:10.3233/HAB-170320
94. Majdinasab N, Hosseini Behbahani M, Galehdari H, Mohaghegh M. Association of interleukin 7 receptor gene polymorphism rs6897932 with multiple sclerosis patients in Khuzestan. Iran J Neurol. 2014;13(3):168–171.
95. Pandit L, Ban M, Sawcer S, et al. Evaluation of the established non-MHC multiple sclerosis loci in an Indian population. Mult Scler J. 2011;17(2):139–143. doi:10.1177/1352458510384011
96. Zhuang JC, Wu L, Qian MZ, et al. Variants of interleukin-7/interleukin-7 receptor alpha are associated with both neuromyelitis optica and multiple sclerosis among Chinese han population in southeastern China. Chin Med J. 2015;128(22):3062–3068. doi:10.4103/0366-6999.169093
97. Ibayyan L, Zaza R, Dahbour S, et al. The promoter SNP, but not the alternative splicing SNP, is linked to multiple sclerosis among Jordanian patients. J Mol Neurosci. 2014;52(4):467–472. doi:10.1007/s12031-013-0151-0
98. Haj MS, Nikravesh A, Kakhki MP, Rakhshi N. Association study of four polymorphisms in the interleukin-7 receptor alpha gene with multiple sclerosis in Eastern Iran. Iran J Basic Med Sci. 2015;18(6):593–598.
99. Tavakolpour S. Interleukin 7 receptor polymorphisms and the risk of multiple sclerosis: a meta-analysis. Mult Scler Relat Disord. 2016;8:66–73. doi:10.1016/j.msard.2016.05.001
100. Wu S, Liu Q, Zhu JM, Wang MR, Li J, Sun MG. Association between the IL7R T244I polymorphism and multiple sclerosis risk: a meta analysis. Neurol Sci. 2016;37(9):1467–1474. doi:10.1007/s10072-016-2608-8
101. Sahami-Fard MH, Mozhdeh M, Izadpanah F, Kashani HH, Nezhadi A. Interleukin 7 receptor T244I polymorphism and the multiple sclerosis susceptibility: a meta-analysis. J Neuroimmunol. 2020;341:577166. doi:10.1016/j.jneuroim.2020.577166
102. Alvarenga RMP, Araújo ACRDAE, Nascimento ACB. Is Asian type MS an MS phenotype, an NMO spectrum disorder, or a MOG-IgG related disease? Mult Scler Relat Disord. 2020;42:102082. doi:10.1016/j.msard.2020.102082
103. Amezcua L, McCauley JL. Race and ethnicity on MS presentation and disease course. Mult Scler J. 2020;26(5):561–567. doi:10.1177/1352458519887328
104. Kulakova OG, Bashinskaya VV, Tsareva EY, Boyko AN, Favorova OO, Gusev EI. Association analysis of cytokine receptors’ genes polymorphisms with clinical features of multiple sclerosis. Zh Nevrol Psikhiatr Im S S Korsakova. 2016;116(SpecialIssue 10):10–15. doi:10.17116/jnevro201611610210-15
105. Akkad DA, Hoffjan S, Petrasch-Parwez E, Beygo J, Gold R, Epplen JT. Variation in the IL7RA and IL2RA genes in German multiple sclerosis patients. J Autoimmun. 2009;32(2):110–115. doi:10.1016/j.jaut.2009.01.002
106. Traboulsee AL, Bernales CQ, Ross JP, Lee JD, Sadovnick AD, Vilariño-Güell C. Genetic variants in IL2RA and IL7R affect multiple sclerosis disease risk and progression. Neurogenetics. 2014;15(3):165–169. doi:10.1007/s10048-014-0403-3
107. Wingerchuk DM, Banwell B, Bennett JL, et al. International consensus diagnostic criteria for neuromyelitis optica spectrum disorders. Neurology. 2015;85(2):177–189. doi:10.1212/WNL.0000000000001729
108. Kim S-H, Mealy MA, Levy M, et al. Racial differences in neuromyelitis optica spectrum disorder. Neurology. 2018;91(22):e2089–e2099. doi:10.1212/WNL.0000000000006574
109. Sepúlveda M, Armangué T, Sola-Valls N, et al. Neuromyelitis optica spectrum disorders: comparison according to the phenotype and serostatus. Neurol Neuroimmunol NeuroInflamm. 2016;3(3):e225. doi:10.1212/NXI.0000000000000225
110. Kitley J, Leite MI, Nakashima I, et al. Prognostic factors and disease course in aquaporin-4 antibody-positive patients with neuromyelitis optica spectrum disorder from the United Kingdom and Japan. Brain. 2012;135(6):1834–1849. doi:10.1093/brain/aws109
111. Cabrera-Gómez JA, Kurtzke JF, González-Quevedo A, Lara-Rodríguez R. An epidemiological study of neuromyelitis optica in Cuba. J Neurol. 2009;256(1):35–44. doi:10.1007/s00415-009-0009-0
112. Brill L, Lavon I, Vaknin-Dembinsky A. Reduced expression of the IL7Ra signaling pathway in neuromyelitis optica. J Neuroimmunol. 2018;324(May):81–89. doi:10.1016/j.jneuroim.2018.08.011
113. Dai Y, Li J, Zhong X, et al. IL2RA allele increases risk of neuromyelitis optica in Southern Han Chinese. Can J Neurol Sci. 2013;40(6):832–835. doi:10.1017/S0317167100015973
114. Sinha S, Singh J, Jindal SK. Association of interleukin 7 receptor (rs1494555 and rs6897932) gene polymorphisms with asthma in a north Indian population. Allergy Rhinol. 2015;6(3):168–176. doi:10.2500/ar.2015.6.0137
115. Santiago JL, Alizadeh BZ, Martínez A, et al. Study of the association between the CAPSL-IL7R locus and type 1 diabetes. Diabetologia. 2008;51(9):1653–1658. doi:10.1007/s00125-008-1070-4
116. Heron M, Grutters JC, van Moorsel CHM, et al. Variation in IL7R predisposes to sarcoid inflammation. Genes Immun. 2009;10(7):647–653. doi:10.1038/gene.2009.55
117. O’Doherty C, Alloza I, Rooney M, Vandenbroeck K. IL7RA polymorphisms and chronic inflammatory arthropathies. Tissue Antigens. 2009;74(5):429–431. doi:10.1111/j.1399-0039.2009.01342.x
118. Wang XS, Wen PF, Zhang M, et al. Interleukin-7 receptor single nucleotide polymorphism rs6897932 (C/T) and the susceptibility to systemic lupus erythematosus. Inflammation. 2014;37(2):615–620. doi:10.1007/s10753-013-9777-x
119. Monti P, Brigatti C, Krasmann M, Ziegler AG, Bonifacio E. Concentration and activity of the soluble form of the interleukin-7 receptor α in type 1 diabetes identifies an interplay between hyperglycemia and immune function. Diabetes. 2013;62(7):2500–2508. doi:10.2337/db12-1726
120. Socié G, Ritz J. Current issues in chronic graft-versus-host disease. Blood. 2014;124(3):374–384. doi:10.1182/blood-2014-01-514752
121. Kielsen K, Shamim Z, Thiant S, et al. Soluble interleukin-7 receptor levels and risk of acute graft-versus-host disease after allogeneic haematopoietic stem cell transplantation. Clin Immunol. 2018;187:26–32. doi:10.1016/j.clim.2017.08.015
122. Shamim Z, Spellman S, Haagenson M, et al. Polymorphism in the interleukin-7 receptor-alpha and outcome after allogeneic hematopoietic cell transplantation with matched unrelated donor. Scand J Immunol. 2013;78(2):214–220. doi:10.1111/sji.12077
123. Vitiello GAF, Losi Guembarovski R, Amarante MK, Ceribelli JR, Carmelo ECB, Watanabe MAE. Interleukin 7 receptor alpha Thr244Ile genetic polymorphism is associated with susceptibility and prognostic markers in breast cancer subgroups. Cytokine. 2018;103:121–126. doi:10.1016/j.cyto.2017.09.019
124. Collina F, Di Bonito M, Li Bergolis V, et al. Prognostic value of cancer stem cells markers in triple-negative breast cancer. Biomed Res Int. 2015;2015:2015. doi:10.1155/2015/158682
125. Sun D-P, Wang L, Ding C-Y. Investigating factors associated with thymic regeneration after chemotherapy in patients with lymphoma. Front Immunol. 2016;7(DEC):654. doi:10.3389/fimmu.2016.00654
126. Hartling HJ, Jespersen S, Gaardbo JC, et al. Reduced IL-7R T cell expression and increased plasma sCD127 in late presenting HIV-infected individuals. J Acquir Immune Defic Syndr. 2017;74(1):81–90. doi:10.1097/QAI.0000000000001153
127. Resino S, Navarrete-Muñoz MA, Blanco J, et al. IL7RA rs6897932 polymorphism is associated with better CD4+ T-cell recovery in HIV infected patients starting combination antiretroviral therapy. Biomolecules. 2019;9(6):233. doi:10.3390/biom9060233
128. Carini C, McLane MF, Mayer KH, Essex M. Dysregulation of interleukin‐7 receptor may generate loss of cytotoxic T cell response in human immunodeficiency virus type 1 infection. Eur J Immunol. 1994;24(12):2927–2934. doi:10.1002/eji.1830241202
129. Rajasuriar R, Booth D, Solomon A, et al. Biological determinants of immune reconstitution in HIV-infected patients receiving antiretroviral therapy: the role of interleukin 7 and interleukin 7 receptor α and microbial translocation. J Infect Dis. 2010;202(8):1254–1264. doi:10.1086/656369
130. Hartling HJ, Thørner LW, Erikstrup C, et al. Polymorphism in interleukin-7 receptor α gene is associated with faster CD4+ T-cell recovery after initiation of combination antiretroviral therapy. AIDS. 2014;28(12):1739–1748. doi:10.1097/QAD.0000000000000354
131. Rajasuriar R, Booth DR, Gouillou M, et al. The role of SNPs in the α-chain of the IL-7R gene in CD4+ T-cell recovery in HIV-infected African patients receiving suppressive cART. Genes Immun. 2012;13(1):83–93. doi:10.1038/gene.2011.65
132. Hartling HJ, Thørner LW, Erikstrup C, et al. Polymorphisms in the interleukin-7 receptor α gene and mortality in untreated HIV-infected individuals. AIDS. 2013;27(10):1615–1620. doi:10.1097/QAD.0b013e3283606c2d
133. Medrano LM, Jiménez JL, Jiménez-Sousa MA, et al. IL7RA polymorphisms are not associated with AIDS progression. Eur J Clin Invest. 2017;47(10):719–727. doi:10.1111/eci.12797
134. Maier I, Wu GY. Hepatitis C and HIV co-infection: a review. World J Gastroenterol. 2002;8(4):577–579. doi:10.3748/wjg.v8.i4.577
135. Jiménez-Sousa MÁ, Gómez-Moreno AZ, Pineda-Tenor D, et al. The IL7RA rs6897932 polymorphism is associated with progression of liver fibrosis in patients with chronic hepatitis C: repeated measurements design. PLoS One. 2018;13(5):e0197115. doi:10.1371/journal.pone.0197115
136. Guzmán-Fulgencio M, Berenguer J, Jiménez-Sousa MA, et al. Association between IL7R polymorphisms and severe liver disease in HIV/HCV coinfected patients: a cross-sectional study. J Transl Med. 2015;13:206. doi:10.1186/s12967-015-0577-y
137. Rethi B, Fluur C, Atlas A, et al. Loss of IL-7Ralpha is associated with CD4 T-cell depletion, high interleukin-7 levels and CD28 down-regulation in HIV infected patients. AIDS. 2005;19(18):2077–2086. doi:10.1097/01.aids.0000189848.75699.0f
138. MacPherson PA, Fex C, Sanchez-Dardon J, Hawley-Foss N, Angel JB. Interleukin-7 receptor expression on CD8(+) T cells is reduced in HIV infection and partially restored with effective antiretroviral therapy. J Acquir Immune Defic Syndr. 2001;28(5):454–457. doi:10.1097/00042560-200112150-00008
139. Paiardini M, Cervasi B, Albrecht H, et al. Loss of CD127 expression defines an expansion of effector CD8 + T cells in HIV-infected individuals. J Immunol. 2005;174(5):2900–2909. doi:10.4049/jimmunol.174.5.2900
140. Faller E, Kakal J, Kumar R, MacPherson P. IL-7 and the HIV Tat protein act synergistically to down-regulate CD127 expression on CD8 T cells. Int Immunol. 2009;21(3):203–216. doi:10.1093/intimm/dxn140
141. Mouillaux J, Allam C, Rimmelé T, et al. Regulation of soluble CD127 protein release and corresponding transcripts expression in T lymphocytes from septic shock patients. Intensive Care Med Exp. 2019;7(1):3. doi:10.1186/s40635-018-0220-3
142. Peronnet E, Mouillaux J, Monneret G, et al. Elevated soluble IL-7 receptor concentration in non-survivor ICU patients. Intensive Care Med. 2016;42(10):1639–1640. doi:10.1007/s00134-016-4445-x
143. Delwarde B, Peronnet E, Venet F, et al. Low interleukin-7 receptor messenger RNA expression is independently associated with day 28 mortality in septic shock patients. Crit Care Med. 2018;46(11):1739–1746. doi:10.1097/CCM.0000000000003281
144. Demaret J, Villars-Méchin A, Lepape A, et al. Elevated plasmatic level of soluble IL-7 receptor is associated with increased mortality in septic shock patients. Intensive Care Med. 2014;40(8):1089–1096. doi:10.1007/s00134-014-3346-0
145. Lundtoft C, Afum-Adjei Awuah A, Rimpler J, et al. Aberrant plasma IL-7 and soluble IL-7 receptor levels indicate impaired T-cell response to IL-7 in human tuberculosis. PLoS Pathog. 2017;13(6):e1006425. doi:10.1371/journal.ppat.1006425
146. Rane L, Rahman S, Magalhaes I, et al. Increased (6 exon) interleukin-7 production after M. tuberculosis infection and soluble interleukin-7 receptor expression in lung tissue. Genes Immun. 2011;12(7):513–522. doi:10.1038/gene.2011.29
147. Badot V, Luijten RKMAC, van Roon JA, et al. Serum soluble interleukin 7 receptor is strongly associated with lupus nephritis in patients with systemic lupus erythematosus. Ann Rheum Dis. 2013;72(3):453–456. doi:10.1136/annrheumdis-2012-202364
148. Chi S, Xue J, Li F, et al. Correlation of serum soluble interleukin-7 receptor and anti-C1q antibody in patients with systemic lupus erythematosus. Autoimmune Dis. 2016;2016:1–11. doi:10.1155/2016/8252605
149. Badot V, Durez P, Van den Eynde BJ, Nzeusseu-Toukap A, Houssiau FA, Lauwerys BR. Rheumatoid arthritis synovial fibroblasts produce a soluble form of the interleukin-7 receptor in response to pro-inflammatory cytokines. J Cell Mol Med. 2011;15(11):2335–2342. doi:10.1111/j.1582-4934.2010.01228.x
150. Blokland SLM, Hillen MR, Kruize AA, et al. Increased CCL25 and T helper cells expressing CCR9 in the salivary glands of patients with primary sjögren’s syndrome: potential new axis in lymphoid neogenesis. Arthritis Rheumatol. 2017;69(10):2038–2051. doi:10.1002/art.40182
151. Torrealba MP, Manfrere KC, Miyashiro DR, et al. Chronic activation profile of circulating CD8+ T cells in Sézary syndrome. Oncotarget. 2018;9(3):3497–3506. doi:10.18632/oncotarget.23334
152. Leligdowicz A, Matthay MA. Heterogeneity in sepsis: new biological evidence with clinical applications. Crit Care. 2019;23(1):80. doi:10.1186/s13054-019-2372-2
153. Bedri SK, Fink K, Manouchehrinia A, et al. Multiple sclerosis treatment effects on plasma cytokine receptor levels. Clin Immunol. 2018;187:15–25. doi:10.1016/j.clim.2017.08.023
154. Poiret T, Rane L, Remberger M, et al. Reduced plasma levels of soluble interleukin-7 receptor during graft-versus-host disease (GVHD) in children and adults. BMC Immunol. 2014;15(1):25. doi:10.1186/1471-2172-15-25
155. Rose-John S. The soluble interleukin 6 receptor: advanced therapeutic options in inflammation. Clin Pharmacol Ther. 2017;102(4):591–598. doi:10.1002/cpt.782
156. Zhang C, Wu Z, Li J, Zhao H, Wang G. Cytokine release syndrome in severe COVID-19: interleukin-6 receptor antagonist. Int J Antimicrob Agents. 2020;55(2020):105954–105960. doi:10.1016/j.ijantimicag.2020.105954
157. Magro G. SARS-CoV-2 and COVID-19: is interleukin-6 (IL-6) the ‘culprit lesion’ of ARDS onset? What is there besides tocilizumab? SGP130Fc. Cytokine X. 2020;2(2):100029. doi:10.1016/j.cytox.2020.100029
158. Murdaca G, Colombo BM, Puppo F. Anti-TNF-α inhibitors: a new therapeutic approach for inflammatory immune-mediated diseases: an update upon efficacy and adverse events. Int J Immunopathol Pharmacol. 2009;22(3):557–565. doi:10.1177/039463200902200301
159. Murdaca G, Colombo BM, Cagnati P, Gulli R, Span F, Puppo F. Update upon efficacy and safety of TNF-α inhibitors. Expert Opin Drug Saf. 2012;11(1):1–5. doi:10.1517/14740338.2012.630388
160. Merola JF, Lockshin B, Mody EA. Switching biologics in the treatment of psoriatic arthritis. Semin Arthritis Rheum. 2017;47(1):29–37. doi:10.1016/j.semarthrit.2017.02.001
161. Carrascosa JM, Belinchón I, De-la-cueva P, Izu R, Luelmo J, Ruiz-Villaverde R. Expert recommendations on treating psoriasis in special circumstances. Actas Dermosifiliogr. 2015;106(4):292–309. doi:10.1016/j.adengl.2015.03.019
162. Andersson Å, Srivastava MK, Harris-White M, et al. Role of CXCR3 ligands in IL-7/IL-7Rα-Fc-mediated antitumor activity in lung cancer. Clin Cancer Res. 2011;17(11):3660–3672. doi:10.1158/1078-0432.CCR-10-3346
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.