Back to Journals » International Journal of General Medicine » Volume 14
SET-NUP214 Fusion Gene Involved Early T-Cell Precursor Acute Lymphoblastic Leukemia in Adult with B Marker Expression
Authors Xu X , Zhai Q, Jin H, Yu Y, Han D, Zhang H, Fu K , Meng B
Received 4 December 2020
Accepted for publication 28 January 2021
Published 25 February 2021 Volume 2021:14 Pages 659—664
DOI https://doi.org/10.2147/IJGM.S294715
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Xiaoying Xu,1 Qiongli Zhai,1 Hao Jin,2 Yong Yu,3 Dongmei Han,2 Huilai Zhang,4 Kai Fu,5 Bin Meng1
1Department of Pathology, Tianjin Medical University Cancer Institute and Hospital, National Clinical Research Center for Cancer, Key Laboratory of Cancer Prevention and Therapy, Tianjin’s Clinical Research Center for Cancer, Tianjin, People’s Republic of China; 2International Personalized Cancer Center, Tianjin Cancer Hospital Airport Hospital, Tianjin, People’s Republic of China; 3Department of Hematology, Tianjin Medical University Cancer Institute and Hospital, Tianjin, People’s Republic of China; 4Department of Lymphoma, Tianjin Medical University Cancer Institute and Hospital, Tianjin, People’s Republic of China; 5Department of Pathology, Roswell Park Comprehensive Cancer Center, Buffalo, NY, USA
Correspondence: Bin Meng
Department of Pathology, Tianjin Medical University Cancer Institute and Hospital, National Clinical Research Center of Cancer, Key Laboratory of Cancer Prevention and Therapy, Tianjin’s Clinical Research Center for Cancer, Tianjin, 300060, People’s Republic of China
Tel +86-18622221609
Fax +86-22-23340123-6202
Email [email protected]
Kai Fu
Department of Pathology, Roswell Park Comprehensive Cancer Center, Buffalo, NY, 14263, USA
Tel +1-716-8451300-4219
Email [email protected]
Abstract: Early T-cell precursor acute lymphoblastic leukemia (ETP-ALL) is rare and associated with poor clinical outcome especially in adults. ETP tumor cells that express cross-lineage antigens or lack pan T markers usually pose big challenges to diagnosis, and their prognostic implications are therefore more uncertain. This study reports the unique case of a 44-year-old woman with breast mass as the initial presentation of acute leukemia possessing both T- and B-cell features (cytoplasmic CD3+CD7+CD19+CD79a+). Despite the presence of gene rearrangements of IGH and IGK probably in a small amount of B cells, the patient was diagnosed with T-ALL mainly according to WHO criteria, and further ETP-ALL rather than mixed phenotype ALL based on additional positive expression of stem/myeloid lineage antigens (CD34+CD13+CD33+HLA-DR+). Moreover, in spite of normal karyotype, SET-NUP214 gene fusion is identified, which has not been described in ETP-ALL with bi-phenotype. After intensive chemotherapy, the patient achieved short-term morphologic complete remission but relapsed within one month. This report may expand immunophenotype and clinical behavior of ETP-ALL in adults. Comprehensive evaluations are emphasized in making a differential diagnosis and distinguishing subtypes of acute leukemia.
Keywords: adult acute lymphoblastic leukemia, early T-cell precursor, mixed phenotype acute leukemia, breast infiltration, SET-NUP214 fusion gene
Introduction
Early T-cell precursor acute lymphoblastic leukemia (ETP-ALL) accounts for about 17% of T-ALL and is highly aggressive.1 Besides the concurrence of myeloid/stem cell signatures, this hematopoietic malignancy with B marker expression is exceptionally rare, and usually poses big challenges to diagnosis. Additionally, patients with infiltration of the breast were only sporadically recorded over the last decades either at initial presentation or during relapse of acute leukemia, and their clinical outcomes varied.2,3 Due to a general absence of ALL manifestations, differential diagnosis on ultrasonography and immunophenotyping is therefore essential in case patients are mistaken for benign breast mass or primary mammary carcinoma. Furthermore, the SET-NUP214 fusion gene is an uncommon genetic event. It has been observed in mixed phenotype acute leukemia (MPAL), like acute undifferentiated leukemia and B/M MPAL,4 and exists in only <6% of T-ALL. Data regarding its effects on prognosis remained controversial as some cases with SET-NUP214 demonstrated no significant differences on survival, but better prognosis in a study containing a series of adult T-ALL was shown by Chen et al.5,6 However, it was recently proposed as a risk predictor.7,8 To date, clinical manifestations of ETP-ALL with this fusion gene are quite limited. Herein, we present a unique SET-NUP214 fusion involved ETP-ALL with B marker expression. The patient experienced mild clinical course with breast masses as first presentation, but progressed fast. After ALL strategy treatment, she just got temporary complete remission (CR) and soon relapsed. This case may help expand the expression profile and clinical behavior of ETP-ALL, and might provide more clues in tracking the cell of origin of ETP-ALL in adults.
Case Presentation
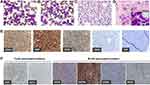
A 44-year-old woman was admitted with a chief complaint of painless masses in the breast found by chance, and had no history of malignancy. Results of physical examination including body temperature, blood pressure and superficial lymph node were within normal limits. Serum laboratory tests showed normal hemoglobin (140 g/L) and platelets (111×109/L), but high levels of lactic acid dehydrogenase (507 U/L), β2-microglobulin (1.47 mg/L) and white blood cells (21.1×109/L). Subsequently, the hemoglobin decreased to 97 g/L along with severe thrombocytopenia (33×109/L), and white blood cells soared to 78.95×109/L in a half-month. Morphologically, hypercellular marrow was observed, and aberrantly increased white blood cells were detected in peripheral blood (PB). The majority of cells in both bone marrow (BM) and PB were medium- to large-sized leukemic blasts (>90%), which were irregularly shaped with high nucleus/cytoplasm ratio, fine chromatin and inconspicuous nucleoli (Figure 1A–C). Less residual mammary ducts remained and were shown by tumor biopsy (Figure 1D).
Immunohistochemistry (IHC) performed on core-needle biopsy at the mammary gland exhibited strong CD43 expression on tumor cells illustrating lymphoid/hematopoietic lineage commitment. Ki67 was high (>90%). The expression of CD38 was positive, while CD34 was merely punctate positive and terminal deoxynucleotidyl transferase (TdT) was negative (Figure 1E). Besides scattered positive expression of CD3, most lineage markers were negative (CD4–CD8–CD1a–CD10–CD20–PAX5–MPO–CD117–) except for CD19 and CD79a (partial results are shown by Figure 1F, Supplementary Figure S1). Consistently, flow cytometry (FCM) analyses using BM aspirate indicated a large single population of CD34+CD38+HLA-DR+nTdT– tumor cells (75.31% of karyocytes) (Figure 2A). Myeloid marker MPO and additional B markers including cytoplasmic CD22 (cCD22), CD24 and cIgM were undetectable. As regards clonality assessment, restricted expression of B-cell immunoglobulin light chain was not found (Figure 2A and B). Noteworthy, these cells expressed cCD3, generally the most specific marker in T lineage assignment, and CD7, whereas other T antigens including surface CD3, CD2, CD4, CD8, CD5, CD1a and CD99 were negative (partial results are shown by Figure 2B).
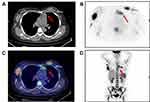
Gene rearrangements of BCR immunoglobulin genes (VH–JH framework region of IGH and IGK) and TCR-beta (TCRB) and -gamma (TCRG) were determined by the BIOMED-2 multiplex RT-PCR system (effective region scope is shown by Supplementary Table 1).9 Both IGH and IGK rearrangements rather than TCR genes were revealed (Figure 2C). Moreover, a gene fusion test was carried out using a 43-gene panel that was frequently involved in ALL by quantitative RT-PCR (listed in Supplementary Table 2), and the SET-NUP214 fusion gene was identified. Karyotype was normal (46, XX) (data not shown). Positron emission tomography and computed tomography (PET-CT) scan showed the largest mass at the right internal mammary region (3.0×3.6 cm2). Additional suspicious malignant tumors observed at the anterior mediastinum, bilateral axillar, uterus corpus and cervix suggested multiple organ involvement (Figure 3A–D). Upon comprehensive evaluations, diagnosis of T-ALL was made mainly according to the WHO principle, and ETP-ALL was concluded finally. The patient did not respond to primary VDCLP (vincristine, daunorubicin, cyclophosphamide,
Discussion
Among ALL cases presenting as a breast mass, MPAL is extremely rare,2,10 and to our knowledge, there has been no similar case report of ETP-ALL infiltration of the breast. Upon combination therapy or hematopoietic stem cell transplantation (HSCT), lasting clinical remission could be achievable at initial presentation, but not for relapsed settings. Besides, bilateral breast infiltration may imply a diffuse leukemic process involving sanctuary organs. Unlike those previous cases of predominantly young girls (median age is about 20),2 the patient described here is older. She remained asymptomatic except for breast masses initially, but the immunophenotype was fairly complex, making it challenging to distinguish the diagnosis of ETP-ALL from MPAL. Despite limited lineage clues upon primary IHC results, from the view of the European Group for Immunological Characterization scoring system, MPAL (B/T bi-phenotype) was possible for the patient (>2 points for either lineage due to co-expression of CD19, cCD79a and cCD3).11 However, the WHO definition cannot be satisfied because of dim or negative expression of CD10, CD22 and cCD79a besides dim CD19, and instead, a T-ALL diagnosis was made.12 Actually, an arbitrary conclusion often occurs, partially because the latter further takes expression intensity into consideration although sometimes underpowered.13 Additionally, the mediastinal mass, frequently presented in T-cell lymphoblastic lymphoma or T-ALL, might also prompt the T lineage malignancy assignment. Finally, based on concurrence of myeloid/stem cell-associated markers (CD34+CD13+CD33+HLA-DR+CD4–CD8–CD1a–CD5–), the patient was diagnosed with ETP-ALL. In addition to the absence of TdT expression as extra evidence, this diagnosis could be further confirmed by other immunophenotypic score-based algorithms.14–16
Co-expression of B antigens has been already reported in both T-ALL and mature T-cell lymphoma with controversial prognosis.17,18 However, to our knowledge, there were only three ETP-ALL cases with B marker expression once discussed. They were immunophenotypically and genetically distinct.19,20 The seemingly confusing result here was conjunction of gene rearrangements of both IGH and IGK, which were highly sensitive in detecting tumorous B cells,21 although they could occasionally occur in either T-cell malignancy or in non-cancerous condition.22 Considering that no detectable clonality of light chain protein expression was observed among either B cells or ETP tumor cells by FCM analyses, there is probably reactive proliferation existing in a small amount of mature monoclonal B cells, while less likely is a B lineage commitment potential of tumor cells. In contrast, the absence of TCR gene rearrangement could be explained by the most immature stage of T-cell development before CD1a starts to express.
ETP-ALL is believed to originate from thymus seeding progenitors (TSP). Since the immunophenotype of migrants from BM is highly heterogeneous as lineage commitment progresses, the cell of origin especially in adult ETP-ALL remains elusive. Here, the CD10–CD7+CD24– tumor cells could be postulated to originate from lineage-negative CD34+CD10+CD24– progenitors capable of circulating in blood throughout life and generating a thymic TSP1 population.23,24 Meanwhile, they also phenotypically overlap with TSP2 (CD10–CD7+), which was primed by NOTCH signals but transcriptionally distinct from TSP1 (Supplementary Figure S2).24 Further gene expression profiling is needed to exploit the potential path involved.
Furthermore, NUP214 gene fusion, critical in nuclear export of proteins and mRNA, is frequently involved in glucocorticoid chemotherapy resistance. Mechanically, it is capable of inhibiting histone 3 acetylation, which could eventually repress transcription activity of corticosteroid receptor. Thus far, 65% of patients involved are T-ALL (including ETP-ALL) with a male predominance. As far as we know, this is the first report showing SET-NUP214 fusion in ETP-All with B antigen expression, suggesting its potential role in hematopoietic differentiation. In terms of clinical practice, ALL-based treatment is likely more effective than an acute myeloid leukemia-based strategy for ETP-ALL. However, the treatment outcome of this case is worse in comparison to that of three previous cases who relapsed within 120 to 505 days. Considering that SET-NUP214 fusion was proven to be a good factor for clinical outcome when an intensive Hyper-CVAD-based regimen was used,6 the poor clinical course is probably mainly ascribed to the ETP immunophenotype and concurrence of cross-lineage markers. In addition, although HSCT is performed more frequently in ETP-ALL than that in non-ETP-ALL,25 it is not clear whether such cases with rapid progression could be benefited or not once the first morphologic CR is achieved. More recently, a “triple mutation” model was provided by Booth et al to highlight mutations of RUNX1, EZH2 and FLT3-ITD genes in transformation of ETP, upon which bromodomain inhibitor involved treatment could be hopefully effective.26 Meanwhile, certain novel strategies like monoclonal antibody against CD33, CD38 or CD123, and combination treatment are potentially promising.27
Collectively, we report the first SET-NUP214 fusion involved ETP-ALL case with B-cell features, and emphasize the importance of comprehensive evaluations for diagnosis of ETP-ALL with extramedullary infiltration of the breast and ambiguous immunophenotype. Also, this case might add more evidence to blocked extrathymic differentiation of T precursors in adults. Further follow-up and more similar cases will facilitate the exploration of consensus on ETP-ALL treatment.
Ethics Statement
This study was approved by the Ethics Committee of Tianjin Medical University Cancer Institute and Hospital. Written informed consent was obtained from the patient’s family for the publication of all data and accompanying images.
Author Contributions
All authors made substantial contributions to conception and design of this manuscript, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Disclosure
The authors declare no conflicts of interest in this work.
References
1. Coustan-Smith E, Mullighan CG, Onciu M, et al. Early T-cell precursor leukaemia: a subtype of very high-risk acute lymphoblastic leukaemia. Lancet Oncol. 2009;10(2):147–156. doi:10.1016/S1470-2045(08)70314-0
2. Karbasian-Esfahani M, Wiernik PH, Yeddu M, Abebe L. Leukemic infiltration of the breast in acute lymphocytic leukemia (ALL). Hematology. 2008;13(2):101–106. doi:10.1179/102453308X315933
3. McCrea ES, Johnston C, Haney PJ. Metastases to the breast. AJR Am J Roentgenol. 1983;141(4):685–690. doi:10.2214/ajr.141.4.685
4. Alexander TB, Gu Z, Iacobucci I, et al. The genetic basis and cell of origin of mixed phenotype acute leukaemia. Nature. 2018;562(7727):373–379. doi:10.1038/s41586-018-0436-0
5. Ben Abdelali R, Roggy A, Leguay T, et al. SET-NUP214 is a recurrent γδ lineage-specific fusion transcript associated with corticosteroid/chemotherapy resistance in adult T-ALL. Blood. 2014;123(12):1860–1863. doi:10.1182/blood-2013-08-521518
6. Chen B, Jiang L, Zhong ML, et al. Identification of fusion genes and characterization of transcriptome features in T-cell acute lymphoblastic leukemia. Proc Natl Acad Sci U S A. 2018;115(2):373–378. doi:10.1073/pnas.1717125115
7. Yang Q, Qian H, Jin Z, et al. SET-CAN fusion gene as poor prognosis predictor in adult T-cell acute lymphoblastic leukemia. Leuk Lymphoma. 2020;61(1):217–220. doi:10.1080/10428194.2019.1660966
8. Gorello P, La Starza R, Varasano E, et al. Combined interphase fluorescence in situ hybridization elucidates the genetic heterogeneity of T-cell acute lymphoblastic leukemia in adults. Haematologica. 2010;95(1):79–86. doi:10.3324/haematol.2009.010413
9. Van Dongen JJ, Langerak AW, Bruggemann M, et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 concerted action BMH4-CT98-3936. Leukemia. 2003;17(12):2257–2317. doi:10.1038/sj.leu.2403202
10. Shin HC. Relapse of biphenotypic acute leukemia as a breast mass. J Breast Cancer. 2016;19(4):455–458. doi:10.4048/jbc.2016.19.4.455
11. Bene MC, Castoldi G, Knapp W, et al. Proposals for the immunological classification of acute leukemias. European Group for the Immunological Characterization of Leukemias (EGIL). Leukemia. 1995;9(10):1783–1786.
12. Arber DA, Orazi A, Hasserjian R, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127(20):2391–2405. doi:10.1182/blood-2016-03-643544
13. Weinberg OK, Seetharam M, Ren L, Alizadeh A, Arber DA. Mixed phenotype acute leukemia: a study of 61 cases using World Health Organization and European Group for the immunological classification of leukaemias criteria. Am J Clin Pathol. 2014;142(6):803–808. doi:10.1309/AJCPPVUPOTUVOIB5
14. Zhou Y, Fan X, Routbort M, et al. Absence of terminal deoxynucleotidyl transferase expression identifies a subset of high-risk adult T-lymphoblastic leukemia/lymphoma. Mod Pathol. 2013;26(10):1338–1345. doi:10.1038/modpathol.2013.78
15. Khogeer H, Rahman H, Jain N, et al. Early T precursor acute lymphoblastic leukaemia/lymphoma shows differential immunophenotypic characteristics including frequent CD33 expression and in vitro response to targeted CD33 therapy. Br J Haematol. 2019;186(4):538–548. doi:10.1111/bjh.15960
16. Zuurbier L, Gutierrez A, Mullighan CG, et al. Immature MEF2C-dysregulated T-cell leukemia patients have an early T-cell precursor acute lymphoblastic leukemia gene signature and typically have non-rearranged T-cell receptors. Haematologica. 2014;99(1):94–102. doi:10.3324/haematol.2013.090233
17. Rizzo K, Stetler-Stevenson M, Wilson W, Yuan CM. Novel CD19 expression in a peripheral T cell lymphoma: a flow cytometry case report with morphologic correlation. Cytometry B Clin Cytom. 2009;76(2):142–149. doi:10.1002/cyto.b.20442
18. Fujisawa S, Tanioka F, Matsuoka T, Ozawa T, Naito K, Kobayashi M. CD7/CD19 double-positive T-cell acute lymphoblastic leukemia. Int J Hematol. 2006;83(4):324–327. doi:10.1532/IJH97.05130
19. Garg S, Gupta SK, Bakhshi S, Mallick S, Kumar L. ETP-ALL with aberrant B marker expression: case series and a brief review of literature. Int J Lab Hematol. 2019;41(2):e32–e37. doi:10.1111/ijlh.12942
20. Khurana S, Melody ME, Ketterling RP, et al. Molecular and phenotypic characterization of an early T-cell precursor acute lymphoblastic lymphoma harboring PICALM-MLLT10 fusion with aberrant expression of B-cell antigens. Cancer Genet. 2020;240:40–44. doi:10.1016/j.cancergen.2019.11.002
21. Kokovic I, Jezersek Novakovic B, Novakovic S. Diagnostic value of immunoglobulin kappa light chain gene rearrangement analysis in B-cell lymphomas. Int J Oncol. 2015;46(3):953–962. doi:10.3892/ijo.2014.2790
22. Langerak AW, Groenen PJ, Jm van Krieken JH, Van Dongen JJ. Immunoglobulin/T-cell receptor clonality diagnostics. Expert Opin Med Diagn. 2007;1(4):451–461. doi:10.1517/17530059.1.4.451
23. Six EM, Bonhomme D, Monteiro M, et al. A human postnatal lymphoid progenitor capable of circulating and seeding the thymus. J Exp Med. 2007;204(13):3085–3093. doi:10.1084/jem.20071003
24. Lavaert M, Liang KL, Vandamme N, et al. Integrated scRNA-seq identifies human postnatal thymus seeding progenitors and regulatory dynamics of differentiating immature thymocytes. Immunity. 2020;52(6):1088–1104.e1086. doi:10.1016/j.immuni.2020.03.019
25. Brammer JE, Saliba RM, Jorgensen JL, et al. Multi-center analysis of the effect of T-cell acute lymphoblastic leukemia subtype and minimal residual disease on allogeneic stem cell transplantation outcomes. Bone Marrow Transplant. 2017;52(1):20–27. doi:10.1038/bmt.2016.194
26. Booth CAG, Barkas N, Neo WH, et al. Ezh2 and Runx1 mutations collaborate to initiate lympho-myeloid leukemia in early thymic progenitors. Cancer Cell. 2018;33(2):274–291.e8. doi:10.1016/j.ccell.2018.01.006
27. Castaneda Puglianini O, Papadantonakis N. Early precursor T-cell acute lymphoblastic leukemia: current paradigms and evolving concepts. Ther Adv Hematol. 2020;11:2040620720929475. doi:10.1177/2040620720929475
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.