Back to Journals » OncoTargets and Therapy » Volume 10
Role of downregulated miR-133a-3p expression in bladder cancer: a bioinformatics study
Authors Gao L, Li SH, Tian YX, Zhu QQ, Chen G , Pang YY, Hu XH
Received 18 March 2017
Accepted for publication 9 May 2017
Published 20 July 2017 Volume 2017:10 Pages 3667—3683
DOI https://doi.org/10.2147/OTT.S137433
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr William C. Cho
Li Gao,1,* Sheng-Hua Li,2,* Yi-Xin Tian,1 Qing-Qing Zhu,1 Gang Chen,3 Yu-Yan Pang,3 Xiao-Hua Hu1
1Department of Medical Oncology, 2Department of Urology Surgery, 3Department of Pathology, First Affiliated Hospital of Guangxi Medical University, Nanning, People’s Republic of China
*These authors contributed equally to this work
Abstract: It has been discovered that miR-133a-3p acts as a tumor suppressor in bladder cancer (BC). Nevertheless, the function of miR-133a-3p in BC remains unclarified. Thus, we carried out this study to validate the expression of miR-133a-3p in BC and provide insights into the molecular mechanism underlying it. To assess the expression of miR-133a-3p in BC, we searched eligible studies from literature and Gene expression Omnibus (GEO) to perform a meta-analysis. We also plotted the summary receiver operating characteristic (SROC) curve to evaluate the diagnostic ability of miR-133a-3p in BC. Additionally, the potential target genes of miR-133a-3p were acquired from 14 online software programs and GEO database. Protein-protein interaction (PPI) network was created to identify the hub genes. Then, Gene Ontology (GO) functional annotation analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis were carried out to investigate the regulatory network of the target genes. From the meta-analysis, miR-133a-3p was remarkably downregulated in BC tissues compared with that in non-cancer tissues (standard mean difference =-3.84, 95% confidence interval =-6.99–0.29). Moreover, results from SROC suggested that miR-133a-3p exhibited the ability to diagnose BC (area under curve =0.8418). As for the bioinformatics study, 488 genes were chosen as the potential targets of miR-133a-3p in BC, among which 10 genes were defined as hub genes (all degrees >5). Further GO and KEGG pathway analysis indicated that the target genes of miR-133a-3p aggregated in specific biological process and pathways. In conclusion, miR-133a-3p possessed great diagnostic potential with its downregulation in BC, and miR-133a-3p might serve as a novel biomarker for BC.
Keywords: bladder cancer, target genes, miR-133a-3p, meta-analysis, bioinformatics study
Introduction
Bladder cancer (BC) is the most common malignant tumor in the urinary tract neoplasms with a high prevalence in the world, posing a threat to patients.1–10 According to the statistical data from Siegel et al, there were estimated 79,030 new cases of BC and 16,870 BC deaths in 2017.11 BC can be divided into two types: non-muscle-invasive BC and muscle-invasive BC. Treatment for non-muscle-invasive tumor consisted of transurethral resection of bladder tumor (TURBT) and post-TURBT adjuvant therapy based on risk level, while radical cystectomy and cisplatin-based chemotherapy constitute major treatments for muscle-invasive BC.12–14 However, muscle-invasive BC had high recurrence and distant metastasis in patients undergoing the aforementioned treatment.15 Therefore, there is a urgent need to seek a novel biomarker for BC.
MicroRNA (miRNA) is an endogenous non-coding small single-stranded RNA in eukaryotes, about 19–24 nucleotides in length. This small, non-coding molecule inhibited the expression of downstream mRNAs either by degrading mRNAs or suppressing the translation process by binding to the 3′untranslated site of target mRNAs.16–21 Previous researches revealed that miRNAs are involved in fundamental biological process such as proliferation, differentiation, apoptosis, metabolism and immunity, playing roles of oncogenes or tumor suppressor genes in a wide type of human cancers.22–24
MiR-133a-3p, a member of the miRNA family, plays a pivotal role in the oncogenesis of various cancers, such as ovarian cancer, colorectal cancer, breast cancer and prostate cancer.25–28 Although several studies reported that miR-133a-3p was downregulated in BC and acted as a tumor suppressor by interacting with related molecules including IGF1R, MMP-9 and EGFR,29–33 the exact mechanism of miR-133a-3p in the pathogenesis of BC was far from elucidated. Therefore, in the present study, we assessed the expression and diagnostic significance of miR-133a-3p in BC by conducting meta-analysis of literature and GSE datasets. Furthermore, bioinformatics study was performed to probe into the underlying molecular mechanism of miR-133a-3p in BC.
Materials and methods
The extraction of literature studies
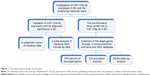
To validate the expression of miR-133a-3p in BC, we conducted a systematic search in the following database: PubMed, Chinese VIP, CNKI, WanFang database, SinoMed, Embase, Web of science, Science Direct and Wiley Online Library. The search was conducted with the following keywords: (miR-133a OR miRNA-133a OR microRNA-133a OR miR-133a OR miRNA-133a OR microRNA-133a OR “miR-133a” OR “miRNA-133a” OR “microRNA-133a”) AND (malignan* OR cancer OR tumor OR tumour OR neoplas* OR carcinoma) AND (bladder OR urinary OR BC). Studies that met the following inclusion criteria were included in our studies: 1) studies provided the expression data of miR-133a-3p between BC and non-cancer tissues, 2) the subjects of the experiment were human beings and 3) the studies should be published in English or Chinese. In addition, studies were excluded according to the following exclusion criteria: 1) the expression of miR-133a-3p was detected in cell lines, 2) there were no non-cancer tissues as the control for the BC tissues and 3) the articles belonged to the types of review, meta-analysis, letters, commentaries and conference abstracts. After the selection of qualified literature studies, the basic information and data were extracted from each of the included studies: first author, ID, year of publication, country, ethnicity of the samples, sample sizes of the experimental and control group, male and female population in experimental and control group, types of specimen, types of experiment, platform and the mean (M) and standard deviation (SD) of miR-133a-3p expression in experimental and control group. The experimental design of the paper is presented in Figure 1.
Meta-analysis
Chip selection and data extraction
Apart from literature database, Gene expression Omnibus (GEO) database proved to be a powerful tool to mine the expression of miRNAs in human cancers. Thus, we also searched GEO database to investigate miR-133a-3p expression in BC. Eligible chip data for the meta-analysis were searched with the following strategies: (miR-133a OR miRNA-133a OR microRNA-133a OR miR-133a OR miRNA-133a OR microRNA-133a OR “miR-133a” OR “miRNA-133a” OR “microRNA-133a”) AND (malignan* OR cancer OR tumor OR tumour OR neoplas* OR carcinoma). The inclusion and exclusion criteria of chip data were the same as those of literature studies. The selected chip data were also extracted for the following basic information and data: first author, ID, year of publication, country, ethnicity of the samples, sample sizes of the experimental and control group, male and female population in experimental and control group, types of specimen, types of experiment, platform and the mean (M) and standard deviation (SD) of miR-133a-3p expression in experimental and control groups. The expression patterns of miR-133a-3p in BC and non-cancer tissues from literature studies and GEO microarray chips were visualized by GraphPad Prism 5.
Statistical analysis
Pooled standard mean difference (SMD) with 95% confidential interval (CI) of miR-133a-3p expression in BC was summarized for literature studies and GEO microarray data. Cochran’s Q test and Higgins I2 statistic were employed to check the heterogeneity of chip data. The pooled effect was calculated by random-effects model when great heterogeneity existed between chip data (P<0.05 or I2>50%); otherwise, fixed-effect model was applied (P>0.05 or I2<50%).34 To detect the source of heterogeneity, we performed subgroup analysis and sensitivity analysis to assess the influence of a single chip data on the overall SMD of the whole study cohorts. Additionally, publication bias was identified by the funnel plot from Begg’s and Egger’s test. STATA v.12.0 was used for all the statistical analysis for meta-analysis.
The diagnostic significance of miR-133a-3p
We employed MedCalc to extract the sensitivity and positivity of studies included in our meta-analysis by creating receiver operating characteristic curve for studies that provided expression value of miR-133a-3p in each specimen of BC and normal tissues. Then, we calculated true positivity (TP), false positivity (FP), false negativity (FN) and true negativity (TN) based on sensitivity and specificity from each study. Summary receiver operating characteristic (SROC) curve was generated to evaluate the diagnostic capacity of miR-133a-3p in BC by putting TP, FP, FN and TN of each study into MetaDiSc v.1.4.
Potential target genes of miR-133a-3p
To obtain the potential targets of miR-133a-3p, we used online prediction software and GEO dataset in combination. A total of 14 online prediction software, miRWalk, Microt4, miRanda, mirbridge, miRDB, miRMap, miRNAMap, Pictar2, PITA, RNA22, RNAhybrid, Targetscan, mirTarbase and PolymiRTS, were employed to collect the target genes of miR-133a-3p. Predicted genes that appear in more than five of all the online software were screened out. The differentially expressed genes (DEGs) from GSE datasets with the transfection of miR-133a-3p were another source of the targets of miR-133a-3p. Interpretation, normalization and log2 transformation were applied to process the primitive GSE data through GCBI (https://www.gcbi.com.cn). DEGs between experiment and control samples of the selected datasets were also identified via GCBI according to the threshold of fold-change >1.5 and P-value <0.05. Finally, the intersection of predicted target genes and DEGs were considered as the potential target genes of miR-133a-3p.
The construction of PPI network
The potential target genes of miR-133a-3p were put into the Search Tool for the Retrieval of Interacting Genes (STRING) to analyze the interactions between target genes. The interaction pairs with a combined score of more than 4 were illustrated by the protein-protein interaction (PPI) network composed of nodes and edges. Nodes and edges in the network represented the target genes and the interactions between them, respectively. Furthermore, hub genes were selected by calculating the degrees of the nodes.
Expression of hub genes from Gene Expression Profiling Interactive Analysis
Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancer-pku.cn/) is a web-based tool that provides comprehensive expression analyses of genes on the basis of TCGA and GTEx data, which facilitated the data mining from cancer genomics.35 In this study, box plots of hub genes from PPI network in BC and normal tissues were downloaded from GEPIA for the purpose of indirectly verifying whether these genes were targeted by miR-133a-3p.
Functional and pathway enrichment analysis for target genes
Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis were conducted to explore the functions and metabolic pathways that the potential target genes of miR-133a-3p participated in. The functions of the target genes were aggregated according to GO terms of biological process (BP), cellular component (CC) and molecular functions (MF). Both GO and KEGG pathway analyses were performed in Database for Annotation, Visualization and Integrated Discovery. GO terms and pathways with P<0.05 were of significance. Furthermore, interactions between the function of target genes were visualized by three GO maps in Cytoscape v.5.3.0.
Consequence
The extraction of literature studies
The selection process for eligible studies from literature was illustrated by the flowchart in Figure 2. Finally, six studies consisting of 322 BC tissues and 95 non-cancer tissues were included to analyze the expression of miR-133a-3p in BC and non-cancer tissues.32,33,36–39 Due to the lack of sufficient data for pooling estimates, meta-analysis was not conducted for literature studies. Basic information of the selected literature studies is listed in Table 1. All the samples of the included studies were tissues from BC or non-cancer bladder. In the study by Pignot et al, samples with small and large sizes were included to evaluate the expression of miR-133a-3p through qRT-PCR.36 Yoshino et al also detected miR-133a-3p expression in samples with small and large sizes by miRNA expression signature and qRT-PCR, respectively.37 Expression data of miR-133a-3p were only available from the experiment of miRNA expression signature in Yoshino et al’s study. Similarly, Song et al assessed miR-133a-3p expression via microarray and qRT-PCR in the same set of sample.32 We only recorded expression data of miR-133a-3p from the microarray experiment considering the accessibility of the data. MiR sequencing and digital gene expression sequencing were applied to examine miR-133a-3p expression in Han et al’s study and Zhu et al’s study, respectively.38,39 As for Ichimi et al’s study, miRNA expression signatures and qRT-PCR were performed to examine miR-133a-3p expression in the same samples.33 MiR-133a-3p expression data from the group of miRNA expression signature was excluded because cell lines were included into the samples to detect miR-133a-3p expression. Data extracted from the four studies indicated that miR-133a-3p expression was obviously lower in BC tissues than in non-cancer tissues.
  | Figure 2 The flow chart. |
  | Table 1 The basic information and data of the included studies |
Meta-analysis
Having searched eligible chip data, a total of 4 chips, GSE36121, GSE39093, GSE20414 and GSE2564, that contained miR-133a-3p expression data between BC and non-cancer tissues were included into the meta-analysis (Table 1). As shown in Figure 3, miR-133a-3p expression was significantly lower in BC tissues than in normal tissues in the study by Zhu et al (Figure 3A) and GSE36121 (Figure 3B).39 All the expression data of miR-133a-3p from two literature studies and all the GEO microarray data yielded an overall SMD of −3.64 (−6.99–0.29) (Figure 4),38,39 which indicated that miR-133a-3p expression was downregulated in BC tissues compared with that in non-cancer tissues. Due to the significant heterogeneity between chips, random-effects model was used to pool the estimates. According to the characteristics of the chip data, subgroup analysis and sensitivity analysis were chosen for tracking the source of heterogeneity. Since only six studies were included to calculate the overall SMD, meta-regression analysis was inappropriate for identifying the source of heterogeneity.40,41 By excluding one study each time to compare pooling effects of the remaining studies with the initial one, we found that no study exerted significant influence to the whole study cohorts (Figure 5). Then, we expected to find out the source of heterogeneity by conducting subgroup analysis. We considered ethnicity and experiment type as the possible sources of heterogeneity. Unfortunately, both the subgroups of ethnicity and experiment type failed to explain heterogeneity between studies. As shown in Table 2, 95% CI of SMD for subgroups of Asian, non-Asian, other experiment and non-coding RNA profiling by array was 0, which indicated no statistical significance. With regard to the publication bias, the symmetrical funnel plot created by Begg’s test revealed that no publication bias was detected (P>0.05) (Figure 6). Now that, only one literature study and all the four GEO microarray data provided detailed data of miR-133a-3p expression in each specimen, SROC was plotted based on the five qualified studies.36 The area under curve (AUC) value (AUC =0.8418) (Figure 7) indicated that miR-133a-3p showed significant ability to differentiate BC tissues from non-cancer tissues.
  | Figure 3 The pattern of miR-133a-3p expression in BC tissues and normal tissues. |
  | Figure 5 The sensitivity analysis. |
  | Figure 6 The publication bias. |
Potential target genes of miR-133a-3p and PPI network
From the result of 14 online software, a total of 18,103 genes were predicted to be the target genes of miR-133a-3p and 2,722 genes that appeared in more than five of the online software were screened out. Two GSM microarray chips, GSM492572 and GSM492574, were included to obtain DEGs between control cell lines and cell lines transfected with miR-133a-3p. After removing the duplicate, a total of 1976 DEGs were defined as the target genes of miR-133a-3p. Eventually, a total of 488 genes overlapping in the online software and GSM microarray chips were selected as the potential targets of miR-133a-3p. Then, we generated a PPI network comprising 110 edges and 75 nodes by putting the 488 potential target genes into STRING (Figure 8). Furthermore, we identified 10 hub genes (all degrees >5), PHLPP1, ESR1, ABL1, RPS6KB1, PAK2, STK38, IGF1R, UBE2I, CDK16 and CSK, from the PPI network by calculating the degree of each node.
Expression of hub genes from GEPIA
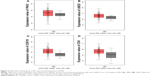
Among all the 10 hub genes from PPI network, four genes, PAK2, UBE2I, CDK16 and CSK, were observed to present a higher expression in 404 BC tissues than in 28 normal tissues (Figure 9).
Functional and pathway enrichment analysis for target genes
According to the results from GO analysis, there were 20, 16 and 5 GO terms significantly enriched by target genes in BP, CC and MF (Table 3). The 3 most significant GO terms of BP assembled by miR-133a-3p were membrane protein ectodomain proteolysis, collagen biosynthetic process and response to oxidative stress. In CC, target genes most largely gathered in the following GO terms: extracellular exosome, focal adhesion and cytosol. As for MF, ATP binding, RNA polymerase II transcription factor activity, sequence-specific DNA binding and protein serine/threonine kinase activity were the three most significant GO terms. Moreover, three GO maps in Figures S1–S3 illustrated the interactions between the functions of target genes. Each node and link in the map represented specific GO term and interactions between them. A darker color of the node reflected a greater significance of the GO term, and sizes of the nodes indicated the number of genes assembled in each GO term.
  | Table 3 GO enrichment analysis of the target genes of miR-133a-3p |
We also analyzed metabolic pathways of the target genes via KEGG pathway analysis. As shown in Table 4, 13 significant pathways were recorded, among which endocytosis, AMPK signaling pathway and proteoglycans in cancer were the three most significant ones.
  | Table 4 KEGG pathway analysis of the target genes of miR-133a-3p |
Discussion
Although miR-133a-3p has been reported to inhibit the deterioration of BC and exhibit downregulated expression in BC,37,42,43 the molecular mechanism of miR-133a-3p in BC was far from elucidated. Therefore, we validated the expression and diagnostic significance of miR-133a-3p in BC and further explored the underlying molecular mechanism through bioinformatics study.
Firstly, we searched literature studies and employed high throughput database, GEO, to mine the expression of miR-133a-3p in BC and non-cancer tissues, which is one of the highlights of our study. Meta-analysis of literature studies and GEO microarray data validated that miR-133a-3p presented significantly lower expression in BC tissues than in non-cancer tissues. Nevertheless, there was distinct heterogeneity between the included studies. Even sensitivity analysis and subgroup analysis failed to detect the source of heterogeneity. We assumed that the significant heterogeneity might originate from the different numbers of samples in BC and non-cancer bladder groups between studies or the different platforms used for the evaluation of miR-133a-3p expression. The number of samples in GEO database was obviously less than that from literature studies. Particularly, only two BC tissues and two non-cancer bladder tissues were recruited in GSE20414. Additionally, the platform for examining miR-133a-3p expression differed from one another.
Though the expression of miR-133a-3p in BC has been researched by several studies, no study up to date has evaluated the diagnostic significance of miR-133a-3p in BC. Therefore, we also explored the diagnostic ability of miR-133a-3p in BC by drawing SROC curve. The AUC value of SROC implied that miR-133a-3p might serve as a future diagnostic target for BC. Nevertheless, we collected limited number of samples; future studies with larger sample size are necessary to further assess the diagnostic ability of miR-133a-3p in BC.
Now that we have found out the diagnostic significance of miR-133a-3p expression in BC, we endeavor to clarify the underlying molecular mechanism of miR-133a-3p in BC by conducting bioinformatics study. Although previous studies have elaborated on the interaction between miR-133a-3p and several downstream targets in the pathogenesis of BC,37,44,45 the miR-133a-3p-centered regulation network in BC remained elusive on account that a single miRNA could target multiple downstream genes involved in diversified biological process and signaling pathways. Thus, we firstly identified potential targets of miR-133a-3p. Since predicted genes of miR-133a-3p from online software might express in wide types of human tissues, potential targets of miR-133a-3p in BC were obtained from the intersection of DEGs in GEO microarray chips transfected with miR-133a-3p and the selected predicted genes from online software, which enhanced the reliability of our result. As a result, a total of 488 genes were identified as the target genes of miR-133a-3p, and we focused on the hub genes to facilitate our understanding of the functions of target genes in regulating the development of BC. According to the results from GEPIA, four of the 10 hub genes, PAK2, UBE2I, CDK16 and CSK, had a higher expression in BC tissues than in normal tissues. Since we have confirmed downregulation of miR-133a-3p in BC tissues, overexpression of the four hub genes in BC tissues proved indirectly that these genes might be targeted by miR-133a-3p. Among all the 10 hub genes, some of them such as PHLPP1, IGF1R and CSK played essential roles in the regulation of various cellular activities such as cell proliferation, survival, migration and apoptosis.46–48 Furthermore, IGF1R has been validated to be the target of miR-133a-3p in various cancers such as osteosarcoma, hepatocellular carcinoma and ovarian cancer.25,29,49 In BC, IGF1R was reported to be targeted by miR-145,50 which strengthened the possibility of IGF1R being targeted by miR-133a-3p. We conceived that PHLPP1, IGF1R and CSK might influence the progression of BC in several malignant aspects. Additionally, the hub genes RPS6KB1, STK38, and CDK16 engaged in specific biological processes. RPS6KB1 is a serine/threonine kinase with important roles in integrating signals related to cell growth and proliferation as well as in mediating protein synthesis.51–53 STK38 is a member of the AGC kinase family and its phosphorylation is pivotal for the regulation of apoptosis and cell cycle-related processes.54,55 CDK16 belongs to the PCTAIRE family that plays an indispensable role in cancer cell proliferation and anti-apoptosis.56–59 The growth, proliferation and apoptosis of cancer cells in BC might be affected by these genes as potential targets of miR-133a-3p. Although the genes ESR1 and ABL1 functioned mainly in breast cancer and chronic myeloid leukemia, they were also listed as the hub genes.60,61 Moreover, ESR1 expression in NMIBC was significantly different between stage and grade.62 ABL1 was considered as a candidate tumor suppressor in the study by Amira et al with its significantly reduced expression in BC tissues compared with normal bladder tissues.73 These findings suggested that ESR1 and ABL1 might also participate in the development of BC through some unknown interactions with miR-133a-3p. Further studies are needed to validate the interactions between miR-133a-3p and these hub genes via experiment.
To achieve a comprehensive understanding of the functions of the potential target genes, we further carried out GO annotation analysis and KEGG pathway analysis. It can be deduced from GO analysis that the target genes of miR-133a-3p exerted an influence on the progression of BC significantly by regulating biological processes such as membrane protein ectodomain proteolysis, collagen biosynthetic process and response to oxidative stress as well as MF such as ATP binding, RNA polymerase II transcription factor activity, sequence-specific DNA binding and protein serine/threonine kinase activity. Results from KEGG pathway analysis also indicated how target genes of miR-133a-3p functioned in BC. Significant pathways such as endocytosis, AMPK signaling pathway, proteoglycans in cancer and bacterial invasion of epithelial cells enlightened us that the target genes of miR-133a-3p might participate in these pathways to impact the initiation and development of BC. Among the 14 significant signaling pathways, several of them such as proteoglycans in cancer, AMPK signaling pathway, adherens junction and tight junction closely correlated with human cancers. AMPK signaling pathway plays crucial roles in cellular metabolism, preserving cellular energy homeostasis and cell apoptosis.63,64 There is abundant evidence supporting that AMPK is critical for the survival and growth of tumor cells.65,66 In BC, AMPK signaling pathway serves as a therapeutic target for certain drugs such as metformin and gefitinib, which reveal the importance of AMPK signaling pathway in BC. Adherens junction and tight junction are two pathways related to epithelial-mesenchymal transition (EMT), a process that confers invasive phenotype to various cancers.67 EMT is also reported to accelerate the deterioration of BC.68,69 We hypothesized that the target genes of miR-133a-3p might modulate the process of EMT to influence the progression of BC.
Although this study investigated the clinical pathological significance of miR-133a-3p and explored the potential underlying molecular mechanism, it should be noted that we only hypothesized the MF of the target genes, and greater efforts are needed to verify the relationship between miR-133a-3p and the hypothesized hub genes and signaling pathways.
In summary, the present study confirmed that miR-133a-3p acts as a tumor suppressor in BC with its downregulated expression in BC tissues. MiR-133a-3p might perform its inhibitory function in BC by regulating network of target genes through specific signaling pathways.
Acknowledgments
The study was supported by the Fund of Promoting Project of Basic Capacity for University Young and Middle-aged Teachers in Guangxi (KY2016LX034) and Innovation Project of Guangxi Graduate Education (201610598107).
Disclosure
The authors report no conflicts of interest in this work.
References
Tadin T, Krpina K, Štifter S, Babarović E, Jonjić N. Significance of uroplakin III expression in recurrence of solitary muscle non-invasive bladder cancer. Pathol Res Pract. 2014;210(5):279–284. | ||
Fu D, Li P, Cheng W, et al. Impact of vascular endothelial growth factor gene-gene and gene-smoking interaction and haplotype combination on bladder cancer risk in Chinese population. Oncotarget. 2017;8(14):22927–22935. | ||
Qi D, Li J, Que B, et al. Long non-coding RNA DBCCR1-003 regulate the expression of DBCCR1 via DNMT1 in bladder cancer. Cancer Cell Int. 2016;16:81. | ||
Xiao J, Lin HY, Zhu YY, Zhu YP, Chen LW. MiR-126 regulates proliferation and invasion in the bladder cancer BLS cell line by targeting the PIK3R2-mediated PI3K/Akt signaling pathway. Onco Targets Ther. 2016;9:5181–5193. | ||
Gil D, Ciołczyk-Wierzbicka D, Dulińska-Litewka J, Laidler P. Integrin-linked kinase regulates cadherin switch in bladder cancer. Tumour Biol. 2016;37(11):15185–15191. | ||
Wu Z, Wang C, Zhang Z, et al. High expression of Derlin-1 is associated with the malignancy of bladder cancer in a Chinese Han population. PLoS One. 2016;11(12):e0168351. | ||
Cui X, Shen D, Kong C, et al. NF-kB suppresses apoptosis and promotes bladder cancer cell proliferation by upregulating survivin expression in vitro and in vivo. Sci Rep. 2017;7:40723. | ||
Heidegger I, Tulchiner G, Schäfer G, Horninger W, Pichler R. Long term disease free survival with multimodal therapy in small cell bladder cancer. Eur J Med Res. 2016;21(1):40. | ||
Yu Y, Li X, Liang C, et al. The relationship between GSTA1, GSTM1, GSTP1, and GSTT1 genetic polymorphisms and bladder cancer susceptibility: a meta-analysis. Medicine (Baltimore). 2016;95(37):e4900. | ||
Du J, Wang SH, Yang Q, Chen QQ, Yao X. p53 status correlates with the risk of progression in stage T1 bladder cancer: a meta-analysis. World J Surg Oncol. 2016;14:137. | ||
Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. | ||
Vashistha V, Quinn DI, Dorff TB, Daneshmand S. Current and recent clinical trials for perioperative systemic therapy for muscle invasive bladder cancer: a systematic review. BMC Cancer. 2014;14:966. | ||
Malmström PU, Wijkström H, Lundholm C, Wester K, Busch C, Norlén BJ. 5-year followup of a randomized prospective study comparing mitomycin C and bacillus Calmette-Guerin in patients with superficial bladder carcinoma. Swedish-Norwegian Bladder Cancer Study Group. J Urol. 1999;161(4):1124–1127. | ||
Tsuji S, Chen X, Hancock B, et al. Preclinical evaluation of VAX-IP, a novel bacterial minicell-based biopharmaceutical for nonmuscle invasive bladder cancer. Mol Ther Oncolytics. 2016;3:16004. | ||
Ye F, Wang L, Castillo-Martin M, et al. Biomarkers for bladder cancer management: present and future. Am J Clin Exp Urol. 2014;2(1):1–14. | ||
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–297. | ||
Bartel DP, Chen CZ. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet. 2004;5(5):396–400. | ||
Huang Y, Li X, Tao G, Zhu T, Lin J. Comparing serum microRNA levels of acute herpes zoster patients with those of postherpetic neuralgia patients. Medicine (Baltimore). 2017;96(8):e5997. | ||
Wang C, Liao H, Cao Z. Role of osterix and microRNAs in bone formation and tooth development. Med Sci Monit. 2016;22:2934–2942. | ||
Zhou YL, Xu YJ, Qiao CW. MiR-34c-3p suppresses the proliferation and invasion of non-small cell lung cancer (NSCLC) by inhibiting PAC1/MAPK pathway. Int J Clin Exp Pathol. 2015;8(6):6312–6322. | ||
Yang G, Zhang X, Shi J. MiR-98 inhibits cell proliferation and invasion of non-small cell carcinoma lung cancer by targeting PAK1. Int J Clin Exp Med. 2015;8(11):20135–20145. | ||
Kim VN. MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol. 2005;6(5):376–385. | ||
Shi M, Liu D, Duan H, Shen B, Guo N. Metastasis-related miRNAs, active players in breast cancer invasion, and metastasis. Cancer Metastasis Rev. 2010;29(4):785–799. | ||
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834–838. | ||
Guo J, Xia B, Meng F, Lou G. miR-133a suppresses ovarian cancer cell proliferation by directly targeting insulin-like growth factor 1 receptor. Tumour Biol. 2014;35(2):1557–1564. | ||
Dong Y, Zhao J, Wu CW, et al. Tumor suppressor functions of miR-133a in colorectal cancer. Mol Cancer Res. 2013;11(9):1051–1060. | ||
Cui W, Zhang S, Shan C, Zhou L, Zhou Z. microRNA-133a regulates the cell cycle and proliferation of breast cancer cells by targeting epidermal growth factor receptor through the EGFR/Akt signaling pathway. FEBS J. 2013;280(16):3962–3974. | ||
Kojima S, Chiyomaru T, Kawakami K, et al. Tumour suppressors miR-1 and miR-133a target the oncogenic function of purine nucleoside phosphorylase (PNP) in prostate cancer. Br J Cancer. 2012;106(2):405–413. | ||
Zhang W, Liu K, Liu S, Ji B, Wang Y, Liu Y. MicroRNA-133a functions as a tumor suppressor by targeting IGF-1R in hepatocellular carcinoma. Tumour Biol. 2015;36(12):9779–9788. | ||
Chen X, Bo L, Zhao X, Chen Q. MicroRNA-133a inhibits cell proliferation, colony formation ability, migration and invasion by targeting matrix metallopeptidase 9 in hepatocellular carcinoma. Mol Med Rep. 2015;11(5):3900–3907. | ||
Zhou Y, Wu D, Tao J, Qu P, Zhou Z, Hou J. MicroRNA-133 inhibits cell proliferation, migration and invasion by targeting epidermal growth factor receptor and its downstream effector proteins in bladder cancer. Scand J Urol. 2013;47(5):423–432. | ||
Song T, Xia W, Shao N, et al. Differential miRNA expression profiles in bladder urothelial carcinomas. Asian Pac J Cancer Prev. 2010;11(4):905–911. | ||
Ichimi T, Enokida H, Okuno Y, et al. Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int J Cancer. 2009;125(2):345–352. | ||
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327(7414):557–560. | ||
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. Epub 2017 Apr 12. | ||
Pignot G, Cizeron-Clairac G, Vacher S, et al. microRNA expression profile in a large series of bladder tumors: identification of a 3-miRNA signature associated with aggressiveness of muscle-invasive bladder cancer. Int J Cancer. 2013;132(11):2479–2491. | ||
Yoshino H, Chiyomaru T, Enokida H, et al. The tumour-suppressive function of miR-1 and miR-133a targeting TAGLN2 in bladder cancer. Br J Cancer. 2011;104(5):808–818. | ||
Han Y, Chen J, Zhao X, et al. MicroRNA expression signatures of bladder cancer revealed by deep sequencing. PLoS One. 2011;6(3):e18286. | ||
Zhu J, Jiang Z, Gao F, et al. A systematic analysis on DNA methylation and the expression of both mRNA and microRNA in bladder cancer. PLoS One. 2011;6(11):e28223. | ||
Higgins J, Thompson S, Deeks J, Altman D. Statistical heterogeneity in systematic reviews of clinical trials: a critical appraisal of guidelines and practice. J Health Serv Res Policy. 2002;7(1):51–61. | ||
Schmid CH, Stark PC, Berlin JA, Landais P, Lau J. Meta-regression detected associations between heterogeneous treatment effects and study-level, but not patient-level, factors. J Clin Epidemiol. 2004;57(7):683–697. | ||
Uchida Y, Chiyomaru T, Enokida H, et al. MiR-133a induces apoptosis through direct regulation of GSTP1 in bladder cancer cell lines. Urol Oncol. 2013;31(1):115–123. | ||
Wei Y, He R, Wu Y, et al. Comprehensive investigation of aberrant microRNA profiling in bladder cancer tissues. Tumour Biol. 2016;37(9):12555–12569. | ||
Chiyomaru T, Enokida H, Tatarano S, et al. miR-145 and miR-133a function as tumour suppressors and directly regulate FSCN1 expression in bladder cancer. Br J Cancer. 2010;102(5):883–891. | ||
Yamasaki T, Yoshino H, Enokida H, et al. Novel molecular targets regulated by tumor suppressors microRNA-1 and microRNA-133a in bladder cancer. Int J Oncol. 2012;40(6):1821–1830. | ||
Hou Y, Deng J, Zhang L, et al. Lower expression of PH domain leucine-rich repeat protein phosphatase 1 (PHLPP1) association with poor prognosis of gastric cancer. Int J Clin Exp Med. 2015;8(11):20481–20489. | ||
Ter Braak B, Siezen CL, Lee JS, et al. Insulin-like growth factor 1 receptor activation promotes mammary gland tumor development by increasing glycolysis and promoting biomass production. Breast Cancer Res. 2017;19(1):14. | ||
Qayyum T, Fyffe G, Duncan M, et al. The interrelationships between Src, Cav-1 and RhoGD12 in transitional cell carcinoma of the bladder. Br J Cancer. 2012;106(6):1187–1195. | ||
Chen G, Fang T, Huang Z, et al. MicroRNA-133a inhibits osteosarcoma cells proliferation and invasion via targeting IGF-1R. Cell Physiol Biochem. 2016;38(2):598–608. | ||
Zhu Z, Xu T, Wang L, et al. MicroRNA-145 directly targets the insulin-like growth factor receptor I in human bladder cancer cells. FEBS Lett. 2014;588(17):3180–3185. | ||
Khotskaya YB, Goverdhan A, Shen J, et al. S6K1 promotes invasiveness of breast cancer cells in a model of metastasis of triple-negative breast cancer. Am J Transl Res. 2014;6(4):361–376. | ||
Fleckenstein DS, Dirks WG, Drexler HG, Quentmeier H. Tumor necrosis factor receptor-associated factor (TRAF) 4 is a new binding partner for the p70S6 serine/threonine kinase. Leuk Res. 2003;27(8):687–694. | ||
de Groot RP, Ballou LM, Sassone-Corsi P. Positive regulation of the cAMP-responsive activator CREM by the p70 S6 kinase: an alternative route to mitogen-induced gene expression. Cell. 1994;79(1):81–91. | ||
Hergovich A, Stegert MR, Schmitz D, Hemmings BA. NDR kinases regulate essential cell processes from yeast to humans. Nat Rev Mol Cell Biol. 2006;7(4):253–264. | ||
Hergovich A. Regulation and functions of mammalian LATS/NDR kinases: looking beyond canonical Hippo signalling. Cell Biosci. 2013;3(1):32. | ||
Cole AR. PCTK proteins: the forgotten brain kinases? Neurosignals. 2009;17(4):288–297. | ||
Yanagi T, Krajewska M, Matsuzawa S, Reed JC. PCTAIRE1 phosphorylates p27 and regulates mitosis in cancer cells. Cancer Res. 2014;74(20):5795–5807. | ||
Yanagi T, Reed JC, Matsuzawa S. PCTAIRE1 regulates p27 stability, apoptosis and tumor growth in malignant melanoma. Oncoscience. 2014;1(10):624–633. | ||
Yanagi T, Matsuzawa S. PCTAIRE1/PCTK1/CDK16: a new oncotarget? Cell Cycle. 2015;14(4):463–464. | ||
Li T, Zhao J, Yang J, et al. A meta-analysis of the association between ESR1 genetic variants and the risk of breast cancer. PLoS One. 2016;11(4):e0153314. | ||
Lyu X, Yang J, Wang X, et al. A novel BCR-ABL1 fusion gene identified by next-generation sequencing in chronic myeloid leukemia. Mol Cytogenet. 2016;9:47. | ||
Breyer J, Wirtz RM, Laible M, et al. ESR1, ERBB2, and Ki67 mRNA expression predicts stage and grade of non-muscle-invasive bladder carcinoma (NMIBC). Virchows Arch. 2016;469(5):547–552. | ||
Hardie DG, Alessi DR. LKB1 and AMPK and the cancer-metabolism link – ten years after. BMC Biol. 2013;11:36. | ||
Liang J, Mills GB. AMPK: a contextual oncogene or tumor suppressor? Cancer Res. 2013;73(10):2929–2935. | ||
Steinberg GR, Kemp BE. AMPK in health and disease. Physiol Rev. 2009;89(3):1025–1078. | ||
Viollet B, Horman S, Leclerc J, et al. AMPK inhibition in health and disease. Crit Rev Biochem Mol Biol. 2010;45(4):276–295. | ||
Wiklund ED, Bramsen JB, Hulf T, et al. Coordinated epigenetic repression of the miR-200 family and miR-205 in invasive bladder cancer. Int J Cancer. 2011;128(6):1327–1334. | ||
Zhu B, Qi L, Liu S, et al. CLASP2 is involved in the EMT and early progression after transurethral resection of the bladder tumor. BMC Cancer. 2017;17(1):105. | ||
Wang X, Liang Z, Xu X, et al. miR-148a-3p represses proliferation and EMT by establishing regulatory circuits between ERBB3/AKT2/c-myc and DNMT1 in bladder cancer. Cell Death Dis. 2016;7(12):e2503. | ||
Ratert N, Meyer HA, Jung M, et al. Reference miRNAs for miRNAome analysis of urothelial carcinomas. PLoS One. 2012;7(6):e39309. | ||
Xu C, Zeng Q, Xu W, et al. miRNA-100 inhibits human bladder urothelial carcinogenesis by directly targeting mTOR. Mol Cancer Ther. 2013;12(2):207–219. | ||
Meiri E, Levy A, Benjamin H, et al. Discovery of microRNAs and other small RNAs in solid tumors. Nucleic Acids Research. 2010;38(18):6234–6246. | ||
Amira N, Cancel-Tassin G, Bernardini S, et al. Expression in bladder transitional cell carcinoma by real-time quantitative reverse transcription polymerase chain reaction array of 65 genes at the tumor suppressor locus 9q34.1-2: identification of 5 candidates tumor suppressor genes. Int J Cancer. 2004;111(4):539–542. |
Supplementary materials
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.