Back to Archived Journals » Gastrointestinal Cancer: Targets and Therapy » Volume 5
Prospective prediction of resistance to neoadjuvant therapy in patients with locoregional esophageal adenocarcinoma
Authors Rosen D, Shan W, Lassen N, Johnson C, Oelschlager K, Bierman-Harrar Y, Kesler K, Maetzold D, Badve S, Cook R , Saxena R
Received 16 October 2014
Accepted for publication 12 December 2014
Published 19 February 2015 Volume 2015:5 Pages 53—59
DOI https://doi.org/10.2147/GICTT.S75904
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Eileen O'Reilly
Daniel G Rosen,1 Weiwei Shan,2 Natalie Lassen,2 Clare Johnson,2 Kristen Oelschlager,2 Yaeli Bierman-Harrar,1 Kenneth A Kesler,3 Derek Maetzold,2 Sunil Badve,3 Robert W Cook,2 Romil Saxena3
1Baylor College of Medicine, Houston TX, USA; 2Castle Biosciences, Incorporated, Friendswood, TX, USA; 3Indiana University, Indianapolis, IN, USA
Background: To clinically validate a multianalyte algorithmic immunohistochemistry (IHC) assay that has been previously shown to accurately identify patients with locoregional esophageal adenocarcinoma (EC) who will exhibit extreme resistance to neoadjuvant chemoradiotherapy.
Methods: Archived biopsy specimens of EC were subject to IHC examination of compartmentalized immunoreactivity of nuclear factor kappa B (NF-κB), Sonic Hedgehog (SHH), and GLI family zinc finger 1 (Gli-1), and a labeling index score was assigned to each biomarker. Test prediction was generated by logistic regression predictive modeling, using the labeling index scores for all three analytes from each sample, referring to a validated training set of 167 EC patients. Accuracy of the test was determined by comparing the predicted outcomes with pathologically determined College of American Pathologists tumor response grade. Analytical validity of the test was measured by comparing validation set prediction results obtained in two independent Clinical Laboratory Improvement Amendment-certified laboratories, and by measuring concordance between two trained labeling index readers.
Results: Specimens from 64 patients that met specific criteria were collected. No technical failure was encountered during the IHC labeling procedures. The logistic regression algorithm generated an area under the curve of 0.96 and 0.85 for the 64 sample cohort in two independent clinical laboratories, respectively, comparing predictive results with the established training set. Positive predictive values of 88% and 82% were also achieved in each laboratory, respectively. A negative predictive value of 83% was reported by both laboratories. Interobserver concordance was 97%.
Discussion: We report the second validation of a multianalyte algorithmic IHC-based predictive test that accurately identifies EC patient response to fluorouracil-based neoadjuvant chemoradiotherapy regimens under College of American Pathologists-accredited Clinical Laboratory Improvement Amendment-certified laboratory protocols. The validated assay provides the opportunity to identify patients with EC who have extreme resistance to neoadjuvant chemoradiotherapy who are resistant to fluorouracil-based neoadjuvant chemoradiotherapy, allowing for more effective treatment planning by clinicians and less toxicity for patients.
Keywords: neoadjuvant chemoradiation, immunohistochemistry, predictive test, validation
Introduction
Neoadjuvant chemoradiotherapy (neoCTRT), followed by surgery or trimodality therapy, has generally been considered beneficial to the survival of locoregional esophageal adenocarcinoma (EC) patients and is the current standard of care recommended by the National Comprehensive Cancer Network.1 The literature shows that an estimated 25% of EC patients have optimal response to treatments (pathologic complete response), and another 25% have minimal, if any, response (extreme resistance to neoCTRT; exCTRT), irrespective of the particular regimens received.2–6 If prospectively identified before treatment begins, an EC patient who is most likely to be resistant to neoadjuvant treatment could be spared significant morbidity, time, and cost by avoiding standard fluorouracil (5FU)-based regimens, and might instead proceed directly to surgery, enter a clinical trial with alternative therapy, or receive targeted therapy regimens that may be effective. Existing pathologic tools, however, are unable to identify exCTRT patients.
Recent evidence suggests that increased activity of the nuclear factor kappa B (NF-κB) and Hedgehog (HH) pathway members confer resistance to neoCTRT and are promising candidates as biomarkers for the prediction of patient response to neoCTRT.7 A multianalyte immunohistochemistry (IHC)-based proprietary assay was previously developed, using a cohort of 60 EC patients.8 The assay interrogates the activation and localization of the NF-κB p65 subunit, Sonic HH (SHH), and its downstream effector Gli-1, in pretreatment tumor biopsies, and classifies patients into responders (non-exCTRT; College of American Pathologists [CAP] tumor response grade [TRG] =0–2) and nonresponders (exCTRT; TRG =3). A first validation study was subsequently completed with an independent cohort of 167 patients that proved the accuracy of the test for predicting patients’ response.8,9 The present study had two objectives: to migrate the test from a research laboratory to a centralized clinical environment and to further validate the test’s ability to predict treatment response in patients with EC.
Materials and methods
Patient characteristics of the training set
Previous validation of the IHC test was conducted with 167 histologically confirmed EC tumor specimens accrued from the University of Texas MD Anderson Cancer Center.8,9 This cohort was used as a training set for the current validation study. All 167 patients received concurrent neoCTRT, and 67 patients (40%) underwent induction chemotherapy before maintenance chemoradiotherapy. All cases were given 5FU as part of neoCTRT regimens and subsequently underwent surgery. In addition to 5FU, 96 patients received a taxane-based therapy, 86 received platinum-based therapy, and 21 received both therapies. Radiation therapy was administered at 50.4 Gy for all patients, given in 28 fractions. Of the 167 patients, 27 (16.2%) patients had a clinical outcome of exCTRT, and the remaining 140 (84.8%) were identified as non-exCTRT (partial and complete responders).8,9
Patient characteristics of the validation set
A total of 64 patients from two independent institutions were enrolled in the current validation study. Studies were conducted under approved protocols by the corresponding institutional review boards. Patients with American Joint Committee on Cancer TNM Stages II and III EC were eligible. Each enrolled patient was confirmed to have undergone neoadjuvant chemotherapy, and 91% of patients also received concurrent radiotherapy. Formalin-fixed and paraffin-embedded pretreatment diagnostic biopsy and posttreatment surgically resected tumor specimens were collected from each patient. Clinical characteristics for the EC cases making up the current validation cohort are shown in Table 1. Sixty (94%) patients had T1, T2, and T3 disease, and one (2%) patient had more advanced T4 stage cancer. Of these 64 patients, 43 (67%) received a chemotherapy regimen containing 5FU, three (5%) had non-5FU-based chemotherapy, and 18 (28%) lacked regimen information. Tumor response to neoCTRT, as assessed by a clinically trained pathologist who determined TRG from surgically resected tissue, was exCTRT in 22 (34%) cases and non-exCTRT in 42 (66%) cases.
  | Table 1 Patient demographics |
Determination of pathologic response
Assessment of tumor regression was performed on hematoxylin and eosin-stained sections from posttreatment, surgically resected tumors by a board-certified surgical pathologist (RS) with specialty training in gastrointestinal pathology, who was blinded to the results of the IHC assay performed on the corresponding pretreatment biopsies. Tumor response to treatment was assessed by two methods. The first method employed the TRG system initially proposed by Mandard et al,10 later modified by Becker et al,11 and accepted by the CAP.12 Specifically, pathologic complete response is defined as no residual tumor (TRG =0), marked response shows minimally residual tumor (<10%; TRG =1), moderate response exhibits between 10% and 50% residual tumor (TRG =2), and exCTRT is characterized by more than 50% residual disease (TRG =3) after presurgical neoCTRT.11 The second method employed the three-class grading scale described by Rohatgi et al as complete (TRG =0), partial/intermediate (TRG =1–2), and no response (TRG =3)13 and was also used in clinical validation studies of the test. The Rohatgi grading system13 is, in principle, 100% concordant with the CAP-approved TRG.
Immunohistochemistry
IHC staining for the 64 patient specimens was independently performed in two CAP-accredited/Clinical Laboratory Improvement Amendment (CLIA) certified laboratories. Briefly, IHC detection of the predictive biomarkers was performed on 5 μm formalin-fixed and paraffin-embedded sections prepared from 64 diagnostic biopsy specimens. Slides were deparaffinized in Xylene Substitute (Thermo Fisher Scientific, Waltham, MA, USA), rehydrated in serially diluted ethanol solutions (Thermo Fisher Scientific), and incubated in primary antibody against NF-κB p65 subunit (Abcam 7970; 1:1500) or Gli-1 (Abcam 92611; 1:150) at 4°C overnight, or against SHH (Abcam 53281; 1:1,000) for 1 hour at room temperature. Stained slides were mounted with Ecomount solution (Biocare Medical, Concord, CA, USA) before pathologic review for determination of tumor cell positivity. Positive and negative controls included normal lung, normal testes, or normal gastric mucosa (Flagship Biosciences, Aurora, CO, USA) and were processed concurrently with EC tissue.
Microscopic examination and generation of labeling index scores
Stained slides from 64 specimens were independently reviewed by a clinical pathologist and a trained PhD-level clinical scientist, both blinded to patient’s actual pathologic outcome and each other’s review. Each region of the tumor was marked and analyzed separately, and peripheral staining resulting from “edge effect” was excluded from the analysis. A positive EC tumor cell was defined as a cell displaying nuclear localization of NF-κB or Gli-1, or cytoplasmic localization of SHH. The labeling index score for each biomarker was calculated as the ratio of positively stained cells to the total number of tumor cells in the fields reviewed.8
Statistical methods
Labeling index scores from the three biomarkers for each of the 64 samples were compared with labeling index scores for the 167 sample training set, using a multivariate logistic regression model (SAS JMP Genomics, Cary, NC, USA) to predict treatment response outcomes. Accuracy was determined by comparing algorithm-based predictions with pathologically assigned TRG. The parameters of area under the curve (AUC), accuracy, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were reported for both the training and current validation sets.
Concordance of predicted outcomes based on labeling index scores independently determined by a trained pathologist and a clinical scientist, each of whom reviewed IHC slides in the current validation study, was separately analyzed. Kappa analysis was performed to determine the degree of agreement between the two readers, using an online Kappa calculator supported by GraphPad software (http://graphpad.com/quickcalcs/kappa1.cfm).
Results
Immunohistochemical detection of NF-κB, Gli-1, and SHH
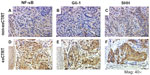
IHC staining of NF-κB, Gli-1, and SHH in pretreatment biopsied formalin-fixed and paraffin-embedded tumor tissue was performed for all 167 patients in a study previously conducted at MD Anderson Cancer Center8 and for the 64 multicenter EC patients in the current validation study. The technical success rate for IHC staining was 100% at both CLIA laboratories. Representative staining patterns for each analyte in an exCTRT and a non-exCTRT case are shown in Figure 1. Localization of NF-κB and Gli-1 proteins, characterized by dark nuclear staining, is markedly elevated in the exCTRT cases, with less localization of the two biomarkers present in the nuclear compartment of non-exCTRT cases (P<0.0001 for both NF-κB and Gli-1). Similarly, enhanced cytoplasmic localization of SHH is found in exCTRT specimens compared with that observed in non-exCTRT cases (P<0.0001). These characteristics of the staining process are reflected by the labeling index scores determined by pathologic review.
Cross-validation and establishment of the training set
Fivefold cross validation of the 167 sample cohort with 20 iterations, using multivariate logistic regression modeling, achieved a markedly high average AUC of 0.957 (95% confidence interval, 0.919–0.971) and accuracy of 90% (95% confidence interval, 88%–92%) for exCTRT and non-exCTRT classifications. These results accurately reproduced findings from the initial validation conducted at the MD Anderson Cancer Center8 and suggested that the 167 patient validation cohort could be used as an independent training set for the current validation study.
Accuracy of the IHC predictive test
Using the 167 sample cohort as a training set, prediction of treatment response for each of the 64 cases in the second validation cohort was carried out in two independent CLIA-certified clinical laboratories. Logistic regression modeling provided a validation set AUC of 0.96 and 0.85, respectively, in each laboratory, reflecting a highly accurate and clinically significant assay (Table 2). Validation set PPVs were 88% and 82%, and NPV was 83% and 83%, in the two clinical laboratories, respectively (Table 2). Specificity reached 95% and 93%, respectively. In other words, when compared with patients’ pathologic tumor response, determined by TRG, 40 (95%) of 42 and 39 (93%) of 42 non-exCTRT cases were accurately classified. In comparison with results attained for the validation cohort, AUC, specificity, sensitivity, PPV, and NPV values for the training set were 0.96, 90%, 93%, 64%, and 98%, respectively (Table 2), which is highly consistent with results reported by the initial validation study from the MD Anderson Cancer Center.8,14
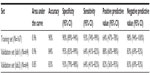  | Table 2 Accuracy of the IHC predictive test |
Concordance between the two clinical reviewers
Interobserver concordance between the two independent, blinded slide reviewers was assessed by kappa analysis. A concordance of 97% (62 of 64) was achieved. Specifically, 16 patients were predicted to be exCTRT, and 46 patients to be non-exCTRT, by both readers (Table 3). Next, kappa concordance analysis (GraphPad Software; http://graphpad.com/quickcalcs/kappa1.cfm) was performed to determine the degree of agreement between the two readers. As shown in Table 3, a kappa of 0.92 (95% confidence interval, 0.81–1.00) was reached. These results suggest that the strength of agreement, or concordance, between the judgments of the two observers was very good.
  | Table 3 Concordance between the two pathologic reviewers |
Discussion
The present study was designed to achieve two goals. The first goal was migration of a research laboratory protocol for an IHC-based assay for prediction of treatment response to a standardized clinical laboratory platform. Standard operating protocols for the IHC test were modified to meet all the qualifications for CAP-accredited/CLIA-certified laboratory protocols. To prove the analytic validity of the procedures, the test was performed in two independent CAP-approved and CLIA-certified laboratories, and interobserver concordance was measured by comparing treatment response predictions determined by two independent, blinded reviewers who determined labeling index scores. Excellent concordance (97%) was achieved. These data support the conclusion that the multianalyte algorithm IHC test is analytically valid under these CLIA procedures.
The second goal of the study was to independently validate the accuracy of the assay for predicting treatment response outcomes. This goal was also accomplished using a cohort of 64 patients accrued from multiple centers. As discussed earlier, the initial validation study with 167 patients conducted at the MD Anderson Cancer Center reached an AUC, PPV, and NPV of 0.96, 64%, and 98%, respectively, for the IHC predictive test.8 That study reported the significant stratification of overall survival outcomes for patients predicted to be exCTRT or non-exCTRT.8 The current validation established the 167 patient cohort as a training set and compared labeling index scores for the 64 samples with those of the training set to predict treatment responses. The test performed with an AUC of 0.96, accuracy of 84%, and PPV of 88% for exCTRT and non-exCTRT classifications in the CAP-accredited CLIA laboratory that will be performing this test for clinical use (Table 2). When performed in a second CLIA laboratory, the test performed similarly, with an AUC of 0.85 and a PPV of 82%, confirming the validity of the predictive modeling (Table 2). The consistency between the two sets of results obtained under independent clinical laboratory conditions with the same patient sample set supports the analytical validity of the assay protocols. In addition, interobserver concordance for the two clinical scientists that each independently assigned labeling scores for the 64 cases was 97%, further strengthening the analytical robustness of this test.
The clinical value of the reported test lies in the opportunity to personalize management in patients identified as unlikely to respond to neoadjuvant treatment, rather than subjecting all patients to delayed surgery and the known extensive toxicity associated with the neoCTRT regimen. Indeed, recent results from a group of investigators suggest that Stage II treatment-insensitive EC patients who received neoCTRT followed by surgery had shorter overall and disease-free survival when compared with patients who underwent surgery only,15 suggesting that treating nonresponders with CTRT could be counterproductive.
The molecular mechanisms behind the functions of the biomarkers of this test, NF-κB, SHH, and Gli-1, are well understood. Specifically, NF-κB16 and HH17 target genes have been suggested to mediate CTRT resistance by accelerating cell proliferation, repressing apoptosis, promoting angiogenesis, and enhancing epithelial-mesenchymal transition. Recently, Izzo et al compared the gene expression profiles of EC patients with good or poor clinical outcomes and showed that NF-κB and HH activators and/or targets were constitutively active in tumors of patients with a poor outcome.18 In addition, the same group demonstrated that pretreatment nuclear NF-κB immune-positivity in locoregional EC was significantly associated with shorter overall survival and worse disease-free survival, as well as the insensitivity to CTRT (defined by the lack of pathologic complete response) when compared with those with immune-negative NF-κB.18 Similarly, SHH and the oncoprotein Gli-1 were detected in a higher proportion of EC patients with resistance to CTRT compared with those with good response and caused proliferation of esophageal cancer cells when ectopically expressed.7 Further, initial trials with NF-κB, SHH, and Gli-1 showed that the panel of the three biomarkers was adequate and proficient at discriminating neoCTRT responders and nonresponders.8
There are limitations to the reported study. First, the majority of the patients enrolled in the validation process had chemotherapy regimens containing 5FU and cisplatin (92% and at least 67% in the 167 and 64 sets, respectively). In the last few years, clinicians have begun to replace 5FU and cisplatin-based regimens with chemoreagents of markedly lower toxicity, such as carboplatin and paclitaxel (CROSS regimen5,19). However, stratification of EC patients by treatment response appears to be independent of the specific regimens used, as many trials using cisplatin and 5FU,2,20–22 oxaliplatin and 5FU,3 or the CROSS regimen5,19 all found 25%–30% patients to have exhibited exCTRT. These observations suggest that exCTRT in EC is mediated by the inherent genetic composition of the tumor. A third independent clinical validation study is underway to specifically accrue specimens of non-5FU-treated EC patients, and we expect to be able to assess the use of the test in response prediction to these additional regimens in the near future.
Second, the reported test, similar to all tests performed on IHC platforms, has inherent susceptibility to human subjectivity. However, one of the advantages offered by this assay is that instead of relying on an intensity-based scoring system (eg, “weak,” “intermediate,” “strong,” or “1, 2, 3”), labeling scores are determined by quantitative identification of cellular localization (presence or absence in the desired compartment), thus minimizing human errors. In addition, the multianalyte and algorithm-based requirements of the test minimize analytic variability.
In conclusion, the current study confirms the analytic validity of the reported assay through interlaboratory and interobserver comparisons under CAP-accredited CLIA-certified laboratory protocols. The studies also represent the second independent, multicenter clinical validation of the assay. The clinical validation study confirmed the high AUC, accuracy, and predictive value, ensuring a low error rate when identifying patients at high risk for extreme resistance to standard neoadjuvant chemoradiation therapy. Future studies will focus on testing the ability of the IHC assay for prediction of complete pathologic response and combining the test results with other pathoclinical factors to optimize treatment response prediction. Clinical use of the test should enable improved treatment decision making as patients identified as exCTRT can consider alternative treatment plans, including surgery, targeted therapy, or clinical trials.
Disclosure
The authors report no conflicts of interest in this work.
References
Ajani JA. NCCN Clinical Practice Guidelines in Oncology: Esophageal and Esophagogastric Junction Cancers Version 2.2013. 2013. | |
Bollschweiler E, Metzger R, Drebber U, et al. Histological type of esophageal cancer might affect response to neo-adjuvant radiochemotherapy and subsequent prognosis. Ann Oncol. 2009;20(2):231–238. | |
Leichman LP, Goldman BH, Bohanes PO, et al. S0356: a phase II clinical and prospective molecular trial with oxaliplatin, fluorouracil, and external-beam radiation therapy before surgery for patients with esophageal adenocarcinoma. J Clin Oncol. 2011;29(34):4555–4560. | |
Slotta-Huspenina J, Wolff C, Drecoll E, et al. A specific expression profile of heat-shock proteins and glucose-regulated proteins is associated with response to neoadjuvant chemotherapy in oesophageal adenocarcinomas. Br J Cancer. 2013;109(2):370–378. | |
van Meerten E, Muller K, Tilanus HW, et al. Neoadjuvant concurrent chemoradiation with weekly paclitaxel and carboplatin for patients with oesophageal cancer: a phase II study. Br J Cancer. 2006;94(10):1389–1394. | |
Mealy K, Feely J, Reid I, McSweeney J, Walsh T, Hennessy TP. Tumour marker detection in oesophageal carcinoma. Eur J Surg Oncol. 1996;22(5):505–507. | |
Sims-Mourtada J, Izzo JG, Apisarnthanarax S, et al. Hedgehog: an attribute to tumor regrowth after chemoradiotherapy and a target to improve radiation response. Clin Cancer Res. 2006;12(21):6565–6572. | |
Ajani JA, Wang X, Hayashi Y, et al. Validated biomarker signatures that predict pathologic response to preoperative chemoradiation therapy (CTRT) with high specificity and desirable sensitivity levels in patients with esophageal cancer (EC). Presented at: 2011 ASCO Annual Meeting; June 3–7; 2011; Chicago, IL. Abstract 4027. | |
Ajani JA, Wang X, Song S, et al. ALDH-1 expression levels predict response or resistance to preoperative chemoradiation in resectable esophageal cancer patients. Mol Oncol. 2014;8(1):142–149. | |
Mandard AM, Dalibard F, Mandard JC, et al. Pathologic assessment of tumor regression after preoperative chemoradiotherapy of esophageal carcinoma. Clinicopathologic correlations. Cancer. 1994;73(11):2680–2686. | |
Becker K, Mueller JD, Schulmacher C, et al. Histomorphology and grading of regression in gastric carcinoma treated with neoadjuvant chemotherapy. Cancer. 2003;98(7):1521–1530. | |
Kay Washington JB. Philip Branton, Lawrence J Burgart, David K Carter, Patrick Fitzgibbons, Wendy L Frankel, John Jessup, Sanjay Kakar, Bruce Minsky, Raouf Nakhleh, Carolyn C Compton. Protocol for the Examination of Specimens from Patients with Carcinoma of the Esophagus. Baylor College of Medicine; 2009. | |
Rohatgi P, Swisher SG, Correa AM, et al. Characterization of pathologic complete response after preoperative chemoradiotherapy in carcinoma of the esophagus and outcome after pathologic complete response. Cancer. 2005;104(11):2365–2372. | |
Ajani JA, Barthel JS, Bentrem DJ, et al; National Comprehensive Cancer Network. Esophageal and esophagogastric junction cancers. J Natl Compr Canc Netw. 2011;9(8):830–887. | |
Dittrick GW, Weber JM, Shridhar R, et al. Pathologic nonresponders after neoadjuvant chemoradiation for esophageal cancer demonstrate no survival benefit compared with patients treated with primary esophagectomy. Ann Surg Oncol. 2012;19(5):1678–1684. | |
Chaturvedi MM, Sung B, Yadav VR, Kannappan R, Aggarwal BB. NF-κB addiction and its role in cancer: ‘one size does not fit all’. Oncogene. 2011;30(14):1615–1630. | |
Amakye D, Jagani Z, Dorsch M. Unraveling the therapeutic potential of the Hedgehog pathway in cancer. Nat Med. 2013;19(11):1410–1422. | |
Izzo JG, Correa AM, Wu TT, et al. Pretherapy nuclear factor-kappaB status, chemoradiation resistance, and metastatic progression in esophageal carcinoma. Mol Cancer Ther. 2006;5(11):2844–2850. | |
van Hagen P, Hulshof MC, van Lanschot JJ, et al; CROSS Group. Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med. 2012;366(22):2074–2084. | |
Burmeister BH, Walpole ET, Burmeister EA, et al. Feasibility of chemoradiation therapy with protracted infusion of 5-fluorouracil for esophageal cancer patients not suitable for cisplatin. Int J Clin Oncol. 2005;10(4):256–261. | |
Smithers BM, Devitt P, Jamieson GG, et al. A combined modality approach to the management of oesophageal cancer. Eur J Surg Oncol. 1997;23(3):219–223. | |
Walsh TN, Noonan N, Hollywood D, Kelly A, Keeling N, Hennessy TP. A comparison of multimodal therapy and surgery for esophageal adenocarcinoma. N Engl J Med. 1996;335(7):462–467. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.