Back to Journals » Infection and Drug Resistance » Volume 13
Pre-S Deletions are Predominant Quasispecies in HIV/HBV Infection: Quasispecies Perspective
Authors Nie Y , Deng XZ, Lan Y, Li F, Hu FY
Received 30 March 2020
Accepted for publication 16 May 2020
Published 8 June 2020 Volume 2020:13 Pages 1643—1649
DOI https://doi.org/10.2147/IDR.S255473
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Yuan Nie,1,2,* Xi-Zi Deng,1,* Yun Lan,1 Feng Li,1 Feng-Yu Hu1
1Research Institute, Guangzhou Eighth People’s Hospital Affiliated to Guangzhou Medical University, Guangzhou, Guangdong, People’s Republic of China; 2Department of Gastroenterology, The First Affiliated Hospital of Nanchang University, Nanchang, Jiangxi, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Feng-Yu Hu
Research Institute,Guangzhou Eighth People’s Hospital Affiliated to Guangzhou Medical University, No. 627, Dongfeng Road, Yuexiu District, Guangzhou 510030, Guangdong, People’s Republic of China
Email [email protected]
Background: Combined HIV infection can accelerate HBV-induced liver disease. It is known that HBV Pre-S deletion is closely related to HBV-associated terminal liver disease in HBV mono-infection. Currently, data on HBV Pre-S quasispecies feature deletion in HIV/HBV co-infected patients are lacking.
Methods: The characteristics and blood samples of patients with chronic HBV infection were collected and classified into an HIV/HBV co-infection group and an HBV mono-infection group according to HIV antibody results before treatment. HBV DNA in serum was extracted. The HBV Pre-S region was amplified by nested-PCR and was further T-A cloned. Using the standard sequence of the matched genotype HBV as a reference, BioEdit 7.0 software was employed for sequence alignment.
Results: HBV Pre-S regions were successfully amplified from 147 patients, including 71 cases in the HIV/HBV co-infected group and 76 cases in the HBV mono-infected group. The proportion of the HIV/HBV co-infected group with Pre-S quasispecies deletion was lower than that of the HBV mono-infected group. By analyzing the frequency of Pre-S quasispecies in the two groups, the frequency of Pre-S quasispecies in HIV/HBV co-infected patients with Pre-S quasispecies was higher than HBV mono-infected patients. The frequency of Pre-S quasispecies deletion of the S protein promoter region in the HIV/HBV co-infected group was significantly higher than that in the HBV mono-infected group.
Conclusion: High-frequency Pre-S quasispecies deletions are predominant in HIV/HBV co-infected patients; however, low-frequency Pre-S deletions are predominant in HBV mono-infected patients, providing a reference for the pathogenesis of the accelerated progression of liver disease in HIV/HBV co-infection.
Keywords: human immunodeficiency virus, hepatitis B virus, quasispecies, Pre-S region, deletion
Introduction
Co-infection with hepatitis B virus (HBV) is common among human immunodeficiency virus (HIV)-infected patients because of similar transmission routes. It is estimated that 10% of HIV-infected patients have chronic hepatitis B worldwide, and the prevalence of HBV co-infection is as high as 20% in high HBV endemic areas.1,2 HIV and HBV co-infection accelerate the progression of liver disease during anti-HIV treatment. HIV/HBV co-infection has been a global public health problem, and end-stage liver disease is now the leading cause of death in AIDS patients. However, the pathogenesis mechanism of the accelerated progression of liver disease in HIV/HBV co-infection needs to be further studied.2,3
The HBV genome consists of four open reading frames (ORFs): Pre-core/core, polymerase, X ORF and Pre-S/S ORF. The peptide chain encoded by the HBV Pre-S gene is located on the surface of virus particles and plays an important role in the HBV life cycle.4 As documented in many studies, HBV Pre-S gene deletion can affect virus packaging, secretion, infection, immune recognition and other functions, is one of the most important genetic variations in the development of end-stage liver disease and is closely related to the severity of liver disease.5,6 In HBV quasispecies, the quasispecies that evade the host immune response gradually develop into the dominant quasispecies, which is an important cause of persistent and chronic HBV and is closely related to the development of drug resistance, antiviral treatment effects, and liver disease processes.7–9 Previous studies on combined HIV infection with HBV Pre-S deletion are based on direct sequencing, and there are few studies from the quasispecies perspective. The aim of this study was to investigate the quasispecies feature of HBV Pre-S deletion in HIV/HBV co-infected patients to help us understand the pathogenesis mechanisms of the accelerated progression of liver disease in HIV/HBV co-infection.
Materials and Methods
Study Subjects
The study subjects were taken from a chronic HBV infection cohort that was set up from January 2009 to December 2011 in Guangzhou Eighth People’s Hospital Infectious Disease Center. All patients in this cohort were positive for hepatitis B surface antigen for more than 6 months, and all HIV/HBV co-infected patients were confirmed to be HIV-positive by ELISA and protein imprinting. Exclusion criteria: (1) treatment with antiviral therapy; (2) liver cirrhosis, liver cancer, and liver failure; (3) hepatitis A virus (HAV) infection, hepatitis C virus infection (HCV), hepatitis D virus infection (HDV) and other apparent opportunistic infections; (4) <18 years old, pregnant or lactating women; and (5) cardiovascular disease or renal failure. According to the results of laboratory examinations, the individuals in this study were divided into an HIV/HBV co-infected group and an HBV mono-infected group. The study protocol was met with the declaration of Helsinki and was approved by the Institutional Ethics Committee of Guangzhou Eighth People’s Hospital. Written informed consent was obtained from all the study participants.
Serological Examination
HBsAg and HBeAg/anti-HBe were determined by ELISA (Zhong Shan Biological Technology Company, Limited, Guangzhou, China). Serum HBV DNA levels were monitored using the COBAS TaqMan HBV Test (Roche Diagnostics, Branchburg NJ. USA). Quantification of HBV DNA was performed by real-time PCR. Alanine aminotransferase (ALT) and aspartate aminotransferase (AST) levels were determined using an AU-2700 automatic biochemical detector (OLYMPUS, Japan). To determine absolute CD4 and CD8 cell counts, whole blood was processed using CD3/CD8/CD45/CD4 antibody multitest reagents (BD Biosciences, San Jose, CA, USA) on a BD FACS flow cytometer (CantoⅡ).
Sequence Cloning and Analysis
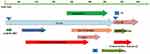
Total DNA was extracted from 200 µL serum samples from each individual with QIAamp DNA mini kits (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The HBV Pre-S region was amplified by nested PCR. The primers for the first round of PCR were 5ʹ-GCCTCATTTTGYGGGTCACCATATTC-3ʹ (F-out), and 5ʹ-AATTCGTTGACANACTTTCCAATCAAT-3ʹ (R-out). The primers for the second round of PCR were 5ʹ-GGGTCACCATATTCTTGGGAACAAGA-3ʹ (F-in) and 5ʹ-AATTCGTTGACANACTTTCCAATCAAT-3ʹ (R-in). Amplification was performed with 4 µL of DNA template for the first round of PCR (extracted DNA from serum) in a 25-µL reaction system under the following conditions: 20 cycles of 98ºC denaturation for 10 s, annealing at 56ºC for 30 s, and extension at 72ºC for 30 s. The second round of PCR was performed with 25 µL of DNA template (first-round PCR product) in a 100-µL reaction system with 35 cycles of 98ºC denaturation for 10 s, annealing at 56ºC for 30 s, and extension at 72ºC for 30s. A schematic diagram of the amplified region was shown in Figure 1. The second PCR amplification range was 1339 nucleotides, including the HBV Pre-S1 and HBV Pre-S2 regions. Final PCR products were purified with the TaKaRa Agarose Gel DNA extraction Kit (Japan) and then recombined with the PMD-19T/A cloning vector and transformed into JM109 Escherichia coli competent cells. Almost 75 recombinant clones of each sample were picked randomly and then sequenced. Multiple alignments were carried out using BioEdit 7.0. HBV Pre-S deletion was defined by nucleotide deletion of the quasispecies sequence that is present in many individuals. The deletion of functional regions and deletion frequencies of quasispecies were defined by comparing with the functional regions of standard sequences. The reference sequence accession numbers were as follows: B1 D00329; B2 AF121249; B3 M54923; B4 AY033072; B5 AB219429; C1 AY057947; C2 AY217378; C3 X75665; C4 AB048705; and C5 AB241111.
 |
Figure 1 Schematic diagram of the amplified region. |
Statistical Analysis
Statistical analyses were performed using SPSS software version 22.0 (SPSS Inc, Chicago, IL). Continuous variables were compared using Student’s t-test or the Mann–Whitney U-test, and categorical variables were compared using the chi-squared test. All statistical tests were two-sided, and P value<0.05 was considered statistically significant.
Results
Demographic Data
PCR products and clone sequences were successfully obtained from 71 HIV/HBV co-infected patients and 76 HBV mono-infected patients. The characteristics of the two groups are summarized in Table 1. Among the HBV mono-infected group, the median age was 35 years (interquartile range [IQR], 27–39 years), and 59 (77.6%) were male. Among the HIV/HBV co-infected group, the median age was 38 (31–47) years, and 56 (78.9%) were male. All HIV/HBV co-infected patients were HIV-1 patients; 26 cases of them were HBV genotype C, and 31 cases of them were HBeAg-negative. In the HBV mono-infected group, there were 29 cases of HBV genotype C and 29 cases of HBeAg-negative. Before ART start, the median baseline CD4 cell count and CD4/CD8 ratio of HIV/HBV co-infected patients were 63 cells/mL (IQR: 16–92) and 0.106 (0.035–0.181), respectively.
 |
Table 1 Demographic and Clinical Features of Patients of Patients |
Proportion Comparisons of Patients with Pre-S Quasispecies Deletion in the Two Groups
In the HIV/HBV co-infected group, there were 42 (59.2%) patients with Pre-S quasispecies deletion, 37 (52.1%) patients with Pre-S1 quasispecies deletion, and 22 (30.9%) patients with Pre-S2 quasispecies deletion. In the HBV mono-infected group, there were 55 (72.3%) patients with Pre-S quasispecies deletion, 44 (57.9%) patients with Pre-S1 quasispecies deletion, and 24 (31.5%) patients with Pre-S2 quasispecies deletion. The proportions of patients with Pre-S, Pre-S1 and Pre-S2 quasispecies deletion in the HBV mono-infected group were higher than those in the HIV/HBV co-infected group. The proportions of the two groups with Pre-S quasispecies deletion are summarized in Table 2. The basic information of patients with Pre-S quasispecies deletion in the HIV/HBV co-infected and HBV mono-infected groups is shown in Supplementary Tables 1 and 2, respectively.
 |
Table 2 Proportion Comparison of Patients with Pre-S Quasispecies Deletion Patients in the Two Groups |
Quasispecies Frequency Comparisons of Patients with Pre-S, Pre-S1, Pre-S2 Quasispecies Deletions Between the Two Groups
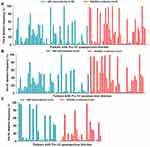
The median (IQR) of quasispecies deletion frequencies of patients with Pre-S deletion were 30.3% (4.4–66.7%) in the HIV/HBV co-infected group and 18.9% (5.1–49.0%) in the HBV mono-infected group. The frequency of Pre-S quasispecies deletion in HIV/HBV co-infected patients was higher than in HBV mono-infected patients, and the comparison was statistically significant (P=0.046). The frequency of Pre-S1 quasispecies deletion in the HIV/HBV co-infected group was significantly higher than that in the HBV mono-infected group (34.0% (5.7–70.9%); 16.6% (3.5–48.7%); P=0.027). The median of Pre-S2 quasispecies deletion frequency in the HIV/HBV co-infected group was higher than that in the HBV mono-infected group (30.3% (4.4% −66.7%); 14.7% (2.2–30.3%); P=0.461). Quasispecies frequencies of patients with Pre-S deletion is shown in Figure 2.
Comparisons of Deletion Quasispecies Frequencies in Functional Domains Between the Two Groups
The functional domain distributions of the HBV Pre-S region are shown9 in Figure 3. In the HIV/HBV co-infected group, the frequencies of quasispecies deletion were higher than in the HBV mono-infection group in the transactivator domain; the hepatocyte binding site; the S-promoter; the heat shock protein 70 binding site; the cytosolic anchorage determinant; the CCAAT binding site; the nucleocapsid binding site; the pHSA binding site; and the viral secretion site. The comparisons of the deletion quasispecies frequencies in the S-promoter and the heat shock protein 70 binding site were statistically significant (P=0.020; P=0.022). The comparisons of deletion quasispecies frequencies in the different functional domains are shown in Table 3.
 |
Table 3 Frequency Comparison of Functional Deletion in HBV Pre-S Region Between the Two Groups |
Discussion
After HBV infection, the proportion of HIV/HBV co-infections developing chronic HBV infection is 9 times higher than in HBV mono-infection, and the mortality rate due to liver-related diseases is 19 times higher than in HBV mono-infection;10 however, the mechanism is unknown. Currently, mutations in HBV genes of HIV/HBV co-infected patients are known to involve Pre-core/core region-1G, rt181T/sw172*A1762T/G1764A, and Pre-S deletion.11–14 Li K’s results showed that variation in Pre-C A1762T/G1764A and Pre-S deletion were more common in HIV/HBV co-infected patients in China through direct sequencing analysis.12 However, from the quasispecies perspective, there are limited studies on minor variants that may be undetected, leaving out useful information. Audsley J et al from the United States found that the frequency of Pre-S deletion was higher in the HIV/HBV co-infected group than in the HBV mono-infected group by HBV full-length direct sequencing.11 However, these studies are limited to direct sequencing analysis and do not involve quasispecies. There has been some study on the quasispecies comparison of HBV mono-infected patients and HIV/HBV co-infected patients.15,16 Their study suggested that there are low evolutionary in the HIV/HBV co-infected group and did not analyze dominant quasispecies from the quasispecies perspective.
The highlight of our study is that the basic characteristics of the HIV/HBV co-infected group and the HIV mono-infected group before antiviral treatment were roughly matched (P>0.05), especially the CD4 cell count. First of all, the proportion of Pre-S quasispecies deletion in the HIV/HBV co-infected group is lower than that in the HBV mono-infected group. Further analysis of the frequency of Pre-S quasispecies in the two groups showed that the frequency of Pre-S quasispecies in HIV/HBV patients with Pre-S quasispecies was higher. This interesting phenomenon reveals that high-frequency Pre-S deletions are predominant in HIV/HBV co-infected patients, while low-frequency Pre-S deletions are predominant in HBV mono-infected patients. Analysis of Pre-S1 and Pre-S2 indicated that the quasispecies frequency of Pre-S1 in HIV/HBV co-infected patients was higher than in HBV mono-infection, and the comparison of Pre-S1 was statistically significant. The comparison of the quasispecies frequency of Pre-S deletion in the functional region showed that the frequency of Pre-S quasispecies deletion in the HIV/HBV co-infected group was higher than that in the HBV mono-infected group, and the comparisons of the S protein promoter region and the heat shock protein 70 binding site were statistically significant (P<0.05). The S protein promoter region (nt 3045–3180) overlaps with the heat shock protein 70 binding site (nt 3067–3201).9
The S protein promoter region transcribes a 2.1-kb mRNA, which encodes median and medium proteins (MHBsAg) and small proteins (HBsAg) and plays an important role in maintaining the ratios of LHBsAg, MHBsAg, and HBsAg.17 Excessive LHBsAg can induce transcription of the S protein promoter by inducing endoplasmic reticulum stress, and positive feedback promotes increased protein production.18 A study from Taiwan reported that the Pre-S deletion produced a large number of truncated LHBsAg proteins, which were retained to form ground-glass-like hepatocytes (GGH) and caused endoplasmic reticulum stress to increase DNA oxidative damage, thereby stimulating the abnormal expression of tumor-related genes.19 Pre-S polypeptide has the function of maintaining the spatial distance between the nucleocapsid binding site and the first transmembrane region (TM-1/Sig-1) of the S protein and maintaining the conformation of the viral particle protein; it can also affect the assembly and transmembrane organization of the virus.20 The deletion protein encoded by HBV Pre-S can act as trans activator, upregulate the expression of cyclin A, inhibit the expression of cyclin D1, and finally induce liver nodule growth in transgenic mice.21 The deletion protein can upregulate the expression of Cox-2 through the NF-kB and MAPK signaling pathways, thus causing abnormal proliferation of hepatocytes in mice.22 The polypeptide encoded by the Pre-S/S region contains multiple important HBV epitopes. Deletion of the Pre-S region causes the immunogenicity of the epitope to change, allowing the virus to escape the host’s immune surveillance and to cause a sustained immune response.23
This study also has some limitations. First, it was a cross-sectional study, and we were not able to evaluate the quasispecies complexity and diversity of the dynamic changes, especially after treatment. Second, the main route of HBV is mother-to-child transmission, and the genotype of HBV is B or C, which is different from the populations of European and American countries. Finally, the study population lacks liver biopsy and liver stiffness results before HAART; thus, we can only use ARPI and platelet indicators instead.
In conclusion, this study investigated the quasispecies feature of HBV Pre-S deletion between HBV mono-infection and HIV/HBV co-infection. Our results showed that high-frequency Pre-S quasispecies deletions are predominant in HIV/HBV co-infected patients, while low-frequency Pre-S deletions are predominant in HBV mono-infected patients, providing a reference for the pathogenesis of the accelerated progression of liver disease in HIV/HBV co-infection.
Ethical Approval
This study was performed in accordance with institutional committee protocols of Guangzhou Eighth People’s Hospital (No.20180307) and written informed consent was obtained from all patient.
Disclosure
The authors declare that there are no conflicts of interest.
References
1. Kourtis AP, Bulterys M, Hu DJ, et al. HIV-HBV coinfection–a global challenge. N Engl J Med. 2012;366(19):1749–1752. doi:10.1056/NEJMp1201796
2. Zhang F, Zhu H, Wu Y, et al. HIV, hepatitis B virus, and hepatitis C virus co-infection in patients in the China national free antiretroviral treatment program, 2010–12: a retrospective observational cohort study. Lancet Infect. Dis. 2014;14(11):1065–1072. doi:10.1016/S1473-3099(14)70946-6
3. Tanday S. HBV and HCV co-infection increases cancer risk in HIV patients. Lancet Oncol. 2016;17(11):e484. doi:10.1016/S1470-2045(16)30512-5
4. Lian M, Zhou X, Chen B, et al. Identification of the critical regions in hepatitis B virus preS required for its stability. J Pept Sci. 2008;14(3):307–312. doi:10.1002/psc.929
5. Yeung P, Wong DK, Lai CL, et al. Association of hepatitis B virus pre-S deletions with the development of hepatocellular carcinoma in chronic hepatitis B. J Infect Dis. 2011;203(5):646–654. doi:10.1093/infdis/jiq096
6. Yeung P, Wong DK, Lai CL, et al. Profile of pre-S deletions in the natural history of chronic hepatitis B infection. J Med Virol. 2010;82(11):1843–1849. doi:10.1002/jmv.21901
7. Liu F, Chen L, Yu DM, et al. Evolutionary patterns of hepatitis B virus quasispecies under different selective pressures: correlation with antiviral efficacy. Gut. 2011;60(9):1269–1277. doi:10.1136/gut.2010.226225
8. Shen T. Hepatitis B virus genetic mutations and evolution in liver diseases. World J Gastroenterol. 2014;20(18):5435–5441. doi:10.3748/wjg.v20.i18.5435
9. Chen BF, Liu CJ, Jow GM, et al. High prevalence and mapping of pre-S deletion in hepatitis B virus carriers with progressive liver diseases. Gastroenterology. 2006;130(4):1153–1168. doi:10.1053/j.gastro.2006.01.011
10. Thio CL, Seaberg EC, Skolasky RJ, et al. HIV-1, hepatitis B virus, and risk of liver-related mortality in the Multicenter Cohort Study (MACS). Lancet. 2002;360(9349):1921–1926. doi:10.1016/S0140-6736(02)11913-1
11. Audsley J, Littlejohn M, Yuen L, et al. HBV mutations in untreated HIV-HBV co-infection using genomic length sequencing. Virology. 2010;405(2):539–547. doi:10.1016/j.virol.2010.06.038
12. Li KW, Kramvis A, Liang S, et al. Higher prevalence of cancer related mutations 1762T/1764A and PreS deletions in hepatitis B virus (HBV) isolated from HBV/HIV co-infected compared to HBV-mono-infected Chinese adults. Virus Res. 2017;227:88–95. doi:10.1016/j.virusres.2016.10.002
13. Revill PA, Littlejohn M, Ayres A, et al. Identification of a novel hepatitis B virus precore/core deletion mutant in HIV/hepatitis B virus co-infected individuals. Aids. 2007;21(13):1701–1710. doi:10.1097/QAD.0b013e32826fb305
14. Warner N, Locarnini S. The antiviral drug selected hepatitis B virus rtA181T/sW172* mutant has a dominant negative secretion defect and alters the typical profile of viral rebound. Hepatology. 2008;48(1):88–98. doi:10.1002/hep.22295
15. Mondal RK, Khatun M, Ghosh S, et al. Immune-driven adaptation of hepatitis B virus genotype D involves preferential alteration in B-cell epitopes and replicative attenuation–an insight from human immunodeficiency virus/hepatitis B virus coinfection. Clin Microbiol Infect. 2015;21(7):710–711. doi:10.1016/j.cmi.2015.03.004
16. Cassino L, Laufer N, Salomon H, et al. Hepatitis B precore/core promoter mutations in isolates from HBV-monoinfected and HBV-HIV coinfected patients: a 3-yr prospective study. J Clin Virol. 2009;46(4):354–359. doi:10.1016/j.jcv.2009.09.015
17. Wu CC, Chen YS, Cao L, et al. Hepatitis B virus infection: defective surface antigen expression and pathogenesis. World J Gastroenterol. 2018;24(31):3488–3499. doi:10.3748/wjg.v24.i31.3488
18. Hsieh YH, Su IJ, Wang HC, et al. Pre-S mutant surface antigens in chronic hepatitis B virus infection induce oxidative stress and DNA damage. Carcinogenesis. 2004;25(10):2023–2032. doi:10.1093/carcin/bgh207
19. Teng YC, Neo JC, Wu JC, et al. Expression of a hepatitis B virus pre-S2 deletion mutant in the liver results in hepatomegaly and hepatocellular carcinoma in mice. J Pathol. 2017;241(4):463–474. doi:10.1002/path.4850
20. Ni Y, Sonnabend J, Seitz S, et al. The pre-s2 domain of the hepatitis B virus is dispensable for infectivity but serves a spacer function for L-protein-connected virus assembly. J Virol. 2010;84(8):3879–3888. doi:10.1128/JVI.02528-09
21. Mun HS, Lee SA, Kim H, et al. Novel F141L pre-S2 mutation in hepatitis B virus increases the risk of hepatocellular carcinoma in patients with chronic genotype C infections. J Virol. 2011;85(1):123–132. doi:10.1128/JVI.01524-10
22. Wang HC, Huang W, Lai MD, et al. Hepatitis B virus pre-S mutants, endoplasmic reticulum stress and hepatocarcinogenesis. Cancer Sci. 2006;97(8):683–688. doi:10.1111/j.1349-7006.2006.00235.x
23. Desmond CP, Bartholomeusz A, Gaudieri S, et al. A systematic review of T-cell epitopes in hepatitis B virus: identification, genotypic variation and relevance to antiviral therapeutics. Antivir Ther. 2008;13(2):161–175.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.