Back to Journals » International Journal of Nanomedicine » Volume 13
Plasma membrane is the target of rapid antibacterial action of silver nanoparticles in Escherichia coli and Pseudomonas aeruginosa
Authors Bondarenko OM , Sihtmäe M, Kuzmičiova J, Ragelienė L, Kahru A , Daugelavičius R
Received 13 June 2018
Accepted for publication 6 August 2018
Published 26 October 2018 Volume 2018:13 Pages 6779—6790
DOI https://doi.org/10.2147/IJN.S177163
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Thomas Webster
Olesja M Bondarenko,1 Mariliis Sihtmäe,1 Julia Kuzmičiova,2 Lina Ragelienė,2 Anne Kahru,1,3 Rimantas Daugelavičius2
1Laboratory of Environmental Toxicology, National Institute of Chemical Physics and Biophysics, Tallinn 12618, Estonia; 2Department of Biochemistry, Vytautas Magnus University, Kaunas LT-44404, Lithuania; 3Estonian Academy of Sciences, Tallinn 10130, Estonia
Introduction: Silver nanoparticles (AgNP) are widely used in consumer products and in medicine, mostly due to their excellent antimicrobial properties. One of the generally accepted antibacterial mechanisms of AgNP is their efficient contact with cells and dissolution in the close vicinity of bacterial cell envelope. Yet, the primary mechanism of cell wall damage and the events essential for bactericidal action of AgNP are not elucidated.
Materials and methods: In this study we used a combination of various assays to differentiate the adverse effects of AgNP on bacterial cell envelope: outer membrane (OM) and plasma membrane (PM).
Results: We showed that PM was the main target of AgNP in gram-negative bacteria Escherichia coli and Pseudomonas aeruginosa: AgNP depolarized PM, induced the leakage of the intracellular K+, and inhibited cellular respiration. The results of bacterial bioluminescence inhibition assay in combination with AgNP dissolution and oxidation assays demonstrated that the adverse effects of AgNP occurred at concentrations 7–160 µM. These toxic effects occurred already within the first few seconds of contact of bacteria and AgNP and were driven by dissolved Ag+ ions targeting bacterial PM. However, the irreversible inhibition of bacterial growth detected after 1-hour exposure occurred at 40 µM AgNP for P. aeruginosa and at 320 µM AgNP for E. coli. In contrast to effects on PM, AgNP and Ag+ ions had no significant effect on the permeability and integrity of bacterial OM, implying that AgNP indeed targeted mainly PM via dissolved Ag+ ions.
Conclusion: AgNP exhibited antibacterial properties via rapid release of Ag+ ions targeting the PM and not the OM of gram-negative bacteria.
Keywords: antimicrobials, mechanism of toxicity, gram-negative, inner membrane, outer membrane, nanomaterials, collargol, bioluminescence
Plain language summary
The resistance of bacteria to conventional antibiotics is steadily increasing, necessitating development of novel antibacterials. One of the alternatives to conventional antibiotics is silver nanoparticles (AgNP) that, due to their excellent antimicrobial properties, are already widely used in consumer products and in medicine. It is commonly accepted that AgNP interact with bacterial cell envelope, but the primary cellular targets of AgNP are not known. The significance of our research is in identifying that AgNP act on inner membrane (ie, plasma membrane; PM) of gram-negative bacteria Escherichia coli and Pseudomonas aeruginosa and target cellular processes associated with PM. More precisely, the rapid antibacterial action of AgNP on PM was mediated via shed Ag+ ions. Therefore, we proposed that the design of AgNP-based antimicrobials should focus on the burst release of Ag+ ions to ensure rapid killing of bacteria, thereby avoiding the development of resistant bacterial strains. Moreover, a suite of methods applied in this study is useful for evaluation of various rapidly acting antibacterials.
Introduction
Silver nanoparticles (AgNP) (1–100 nm) are the most widely used nanoparticles (NP) in consumer products, mostly due to their excellent antimicrobial properties.1
However, the mechanisms of antibacterial action of AgNP are still disputable. A recent comprehensive review by Djurišić et al2 summarized four main tentative mechanisms of antibacterial action of metal oxide NP: 1) generation of ROS, 2) partial dissolution accompanied by release of toxic metal ions, 3) NP accumulation on membrane surface, and 4) cellular uptake of NP. Although all of these four mechanisms are also relevant for antimicrobial action of AgNP, the prevailing mechanism of dissolving AgNP is the oxidative dissolution on bacterial surface and subsequent release of Ag ions.3 Indeed, in our previous study we showed that the interaction of NP with bacterial cell wall accompanied by the release of toxic Ag+ ions significantly contributed to the antibacterial action of various AgNP on different gram-negative and gram-positive bacteria.4 In addition, we observed that medically important pathogenic bacteria Pseudomonas aeruginosa were inherently more susceptible to AgNP (especially to the colloidal form of AgNP, Collargol) than Escherichia coli cells. The current study is a continuation of our previous work and aims to provide detailed insight into the interactions of AgNP with gram-negative bacteria E. coli and P. aeruginosa with the focus on the effects on bacterial cell membranes. We selected the colloidal form of AgNP, Collargol, as the model AgNP since it has a long history of usage and is prescribed as a medication in several countries.1 In addition, Collargol is a commercial product with defined composition and, different from many other NP, has exceptional stability in suspension.
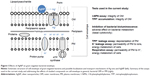
Human opportunistic pathogens E. coli and P. aeruginosa were selected as bacterial models. Both are gram-negative bacteria, ie, have two membranes in their cell envelope – the outer membrane (OM) and the plasma membrane (PM) that possess different compositions and therefore, distinct functions (Figure 1). OM is covered by the dense layer of highly negatively charged lipopolysaccharides (LPS) and serves as a selective permeability barrier. OM prevents the entrance of hydrophobic substances and macromolecules,5 but allows the entrance of small charged molecules into the periplasm through the porins – narrow channels passable for ions and nutrients. In contrast, PM is relatively permeable for hydrophobic compounds but not for the inorganic ions and hydrophilic compounds.6,7 PM is of utmost importance for bacteria since all the vital membrane-related functions of prokaryotic cells are performed in the PM (eg, respiratory chain functions, lipid biosynthesis, protein secretion, and transport events).5,8
To reach the PM of gram-negative bacteria, Ag ion/AgNP should first pass OM. Most likely, Ag+ ions pass the OM through porins9–11 inducing several toxic events hampering the vital functions of PM. For instance, Dibrov et al12 showed that Ag+ ions induced dissipation of pH gradient on membrane of inside-out membrane vesicles of Vibrio cholerae. Authors suggested that H+ leakage is the main mechanism behind the bactericidal action of Ag+ ions in V. cholerae, and assumed that it occurs due to the unspecific, Ag+-induced modification of proteins or phospholipid bilayer.
In contrast to Ag+ ions, the mechanism by which AgNP interact with the cells and possess an antibacterial effect has not yet been convincingly revealed. It is hypothesized that AgNP enter the cells by creating irregular shaped pits on the bacterial surface, inducing the LPS release and damaging the bacterial OM.10,13–16 These events allow the penetration of AgNP into periplasm where AgNP act similarly to Ag+ ions, ie, inhibit SH-groups containing enzymes due to the very high affinity of Ag ions toward sulfur-containing groups.15
The main aim of this study was to differentiate the adverse effects of Ag ions (AgNO3) and AgNP (Collargol, protein-coated AgNP) on bacterial OM and PM. To achieve this, different approaches were used: 1) bacterial bioluminescence inhibition assay to estimate rapid changes in bacterial general metabolism; 2) bacterial viability testing to determine minimal bactericidal concentration (MBC) of Ag compounds; 3) 1-N-phenylnaphthylamine (1-NPN) assay to estimate the integrity of OM; 4) tetraphenylphosphonium (TPP+) assay to discriminate the effect of Ag compounds on PM and OM; and 5) K+ content and cellular respiration to evaluate the effects of Ag compounds on PM. All the assays addressed the rapid effects of studied compounds on bacterial metabolism in general, bacterial OM or PM, as depicted in Figure 1.
Materials and methods
Chemicals
All the chemicals were at least of analytical grade. Polymyxin B (PMB) and 3,5-dichlorophenol (3,5-DCP) Pestanal were purchased from Sigma-Aldrich Co. (St Louis, MO, USA), AgNO3 from J.T. Baker (Phillipsburg, NJ, USA), protein (casein)-coated colloidal AgNP from Laboratorios Argenol S. L. (Zaragoza, Spain) (ordered as a suspension, batch no 297), propidium iodide from Fluka (Steinheim, Germany), and TPP+ chloride from Sigma-Aldrich Co. The stock solutions of AgNO3 and AgNP were prepared at 2 mM (215.7 mg Ag/L, 20 mL) in autoclaved deionized water and further stored in the dark at +4°C. The stock solutions were vortexed before each use. If not stated otherwise, all subsequent dilutions and experiments were performed in Ag+ ions noncomplexing buffer consisting of 50 mM 3-(N-morpholino)propanesulfonic acid (MOPS) adjusted to pH 8 with (hydroxymethyl)aminomethane (Tris) and further referred to as MOPS-Tris buffer.
Characterization of AgNP
The shape, primary size, and coating (casein) content of AgNP were determined in our previous publication (Table 1). Hydrodynamic size (measured by dynamic light scattering), polydispersity index (PDI), and ζ-potential in MOPS-Tris buffer were measured at a concentration of 200 mg/L using Malvern Zetasizer (Nano-ZS; Malvern Instruments, Malvern, UK). Dissolution of AgNP was quantified using atomic absorption spectroscopy (AAS). For that, AgNP were incubated in MOPS-Tris at a concentration of 10 mg/L for 10 minutes, 1 hour, or 24 hours, ultra-centrifuged (390,000 g, 30–60 minutes), and the supernatant was analyzed by AAS in an accredited17 laboratory of the Institute of Chemistry of Tallinn University of Technology, Estonia. The ratio of determined dissolved Ag to total nominal metal in AgNP was designated as dissolution (%).
  | Table 1 Characterization of silver nanoparticles (AgNP) used in the current study |
Bacterial strains and cultivation of bacteria
E. coli MC1061 (obtained from Prof Matti Karp, University of Tampere, Finland) and P. aeruginosa DS10-129 (isolated from soil)18 were used in all experiments. All cultivations were performed on a shaker at 200 rpm and at 30°C instead of conventional 37°C since some of the tests involved bioluminescent bacteria. The production of bioluminescence in bioluminescent bacteria consumes a considerable amount of energy, and we observed that the lower temperature was preferred for the enhanced stability of bioluminescence-encoding elements. Bacteria were pre-grown overnight in 3 mL of NaCl-free Luria-Bertani medium (LB, 10 g tryptone and 5 g yeast extract per liter, pH =7). In case of genetically modified bioluminescent E. coli and P. aeruginosa (more information in the following paragraph), medium was supplemented with appropriate antibiotics (100 mg ampicillin per liter for E. coli and 10 mg tetracycline for P. aeruginosa). Ten to fifty mL of fresh medium was inoculated with 1/20 diluted overnight culture and bacteria were grown until mid-exponential phase (OD600 of ~1), centrifuged at 6,000 g for 5 minutes, washed twice with equal amount of 50 mM MOPS-Tris, and adjusted to OD600=1 corresponding to approximately 109 colony forming units (cfu)/mL.
ζ-potential of bacteria was measured in MOPS-Tris buffer using Malvern Zetasizer.
Bacterial bioluminescence inhibition assay 10-minute EC50
Constitutively luminescent recombinant bacteria E. coli MC1061 (pSLlux)19 and P. aeruginosa DS10-129 (pDNcadlux)18 bearing bioluminescence encoding plasmid were used. Bacteria were grown and prepared as described previously (ie, approximately 109 cfu/mL were used for the test). Bacterial acute bioluminescence inhibition assay (exposure time 10 minutes) was conducted at room temperature on white sterile 96-well polypropylene microplates (Greiner Bio-One GmbH, Kremsmünster, Austria) following the Flash-assay protocol,20 essentially as described by Kurvet et al.21 To obtain the concentration-effect curves, 5–7 sequential exponential dilutions of each compound were analyzed for inhibition of bacterial luminescence. The following concentrations of test chemicals were used: AgNP and AgNO3 from 2.5 to 320 μM (0.27–35 mg Ag/L); PMB from 1.2 to 160 μM (1.66–222 mg/L); 3,5-DCP from 47.9 to 6,135 μM (7.8–1,000 mg/L). Testing was performed in MOPS-Tris. Three to six independent experiments were performed in duplicate. The 10-minute EC50 the concentration of a compound reducing the bacterial bioluminescence by 50% after being in contact for 10 minutes) values were calculated from the concentration vs 10-minute inhibition curves based on nominal exposure concentrations as described by Kurvet et al21 using the log-normal model of MS Excel macro Regtox.22
Evaluation of the MBC
Since bacterial bioluminescence inhibition test does not fully reflect the viability of cells, bacterial viability assay as described by Suppi et al,23 was performed in addition. Briefly, at the end of the bioluminescence inhibition assay (previous paragraph) microplates were incubated for 1 hour at room temperature in the dark, and 3 μL of bacterial suspension from each test well was pipetted onto toxicant-free LB agar plate to assess the viability (1-hour MBC) of the cells after 1-hour exposure to studied compounds. The inoculated agar plates were incubated at 30°C for 24 hours and the 1-hour MBC was defined as the lowest tested concentration of the toxicant yielding no visible bacterial growth (ie, irreversible growth inhibition) on nutrient agar. Three independent experiments were performed in duplicate.
Evaluation of bacterial OM integrity by 1-NPN assay
The cell wall permeabilization of E. coli and P. aeruginosa by AgNO3, AgNP, and PMB was assayed by the cellular uptake of a hydrophobic probe 1-NPN essentially as described by Helander and Mattila-Sandholm.24 Differently from hydrophilic environment, the fluorescence of 1-NPN is significantly enhanced in a hydrophobic environment (eg, membrane lipid bilayer), rendering it an appropriate dye to probe OM integrity of gram-negative bacteria.24 Briefly, 50 μL of 40 μM 1-NPN and 50 μL of tested compound in 50 mM MOPS-Tris buffer were pipetted into wells of black microplates. An amount of 100 μL of bacterial suspension in 50 mM MOPS-Tris was dispensed into each well and the fluorescence was immediately measured (Fluoroskan Ascent FL plate luminometer [Thermo Fisher Scientific Oy, Vantaa, Finland]; excitation/emission filters 350/460 nm). The 1-NPN cell uptake factor was calculated as a ratio between intensity of fluorescence values of the bacterial suspension incubated with and without test compounds. Three independent experiments were performed in duplicate.
Electrochemical measurements performed on bacterial cells
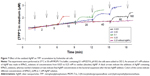
Bacterial cultures were grown overnight in LB broth containing 0.5% NaCl (Sigma-Aldrich Co.), then diluted 1:50 in fresh medium and further incubated until OD600 of 1.0. The cells were harvested by centrifugation at 4°C for 10 minutes at 3,000 g (Heraeus™ Megafuge™ 16R, Thermo Fisher Scientific, Waltham, MA, USA). The pelleted cells were re-suspended in LB medium without NaCl (to avoid the formation of insoluble AgCl while exposing bacteria to Ag compounds), pH 8.0, to obtain ~2×1011 cfu/mL. The concentrated cell suspensions were kept on ice until used, but not longer than 4 hours. TPP+ and K+ concentrations in the incubation media were potentiometrically monitored using selective electrodes as described previously.25,26 In the experiment with TPP+, EDTA was used to increase OM permeability of bacteria to TPP+ and to equilibrate TPP+ across the cell envelope. Since P. aeruginosa cells are sensitive to incubation in EDTA-containing MOPS-Tris buffer,26 100 mM NaPi was used instead of MOPS-Tris in the experiments with these cells. The thermostated magnetically stirred glass vessels were filled with 5 mL of 50 mM MOPS-Tris, pH 8.0. After calibration of the electrodes, the concentrated cell suspension was added to obtain an OD600 of 1 or 2. To register potential of the electrodes we used the electrode potential-amplifying system with an ultralow-input bias current operational amplifier AD549JH (Analog Devices, Norwood, MA, USA). The data acquisition system PowerLab 8/35 (ADInstruments, Oxford, UK) was used to connect the amplifying system to a computer. Agar salt bridges were used for indirect connection of the Ag/AgCl reference electrodes (Orion model 9001; Thermo Fisher Scientific) with cell suspensions in the vessels. The measurements were performed simultaneously in 2–4 reaction vessels. The representative sets of curves from three independent series of measurements are presented in Figures 4–7.
Dissolved oxygen level in the medium was monitored by dissolved oxygen probe (Orion model 9708; Thermo Fisher Scientific), as described previously.25 At the end of every experiment, solid Na2S2O5 (Sigma-Aldrich Co.) was added to vessels to obtain a final concentration of approximately 20 mM and deplete the dissolved oxygen (0% baseline). Concentration of the dissolved oxygen in the medium before addition of cells was set as 100%.
In the oxidation experiment, AgNP were step-wise oxidized (dissolved) using KMnO4. Solutions of 2 mM AgNP and 0.05–0.5 mM KMnO4 were mixed 1:1 16–18 hours before the experiment. Oxidized AgNP were added to bacterial suspensions and TPP+ concentration was potentiometrically monitored as described previously.
Statistical analysis
All tests were performed in duplicate and in at least three independent experiments. The data were expressed as mean ± SD or shown as representative figure. To define statistically significant differences, the data were analyzed with unpaired two-tailed Student’s t-test assuming equal variances at P<0.01.
Results
Physico-chemical characterization of AgNP
Physico-chemical characteristics of studied AgNP are shown in Table 1. AgNP have been described for primary size (12.5±4 nm) and percentage of coating (30% casein of the dry weight of the particles, analyzed by thermogravimetry) earlier by Blinova et al.27 The AgNP formed stable dispersion in the MOPS-Tris buffer (PDI of 0.31) with the hydrodynamic diameter of 50 nm (Table 1). The dissolution of AgNP in the test conditions was time-dependent and relatively high: from 26.9% after 10 minutes of incubation to 66.3% after 24 hours of incubation.
To rule out the ROS-associated toxicity, the potential of AgNP to generate ROS was measured. No ROS were induced by 0.3–100 μM AgNP in abiotic conditions after 1-hour incubation in 50 mM MOPS-Tris according to 2′,7′-dichlorofluorescein assay (data not shown).
Effect of studied compounds on bacterial bioluminescence
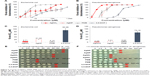
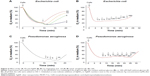
The bacterial bioluminescence assay was used to assess the effects of the studied compounds on bacterial metabolic activity. The bioluminescence of bacteria is an outcome of a complex chain of biochemical reactions, where reduced flavin mononucleotide (FMNH2), a long-chain fatty acid aldehyde, luciferase, and a cofactor NADH are the key players. Thus, any decrease of bacterial metabolic activity can be registered as a decrease in light output (inhibition of bioluminescence).29 All tested compounds had rapid (acting on second-scale) effects on bacterial bioluminescence (Figure 2A–D) and thus, interfered with the systems connected to the energy production. 3,5-DCP, a chemical included as the control due to its wide use as a standard in various toxicological tests,30 caused comparable bactericidal effect on both bacteria (Figure 2E and F). Differently from 3,5-DCP, Ag+ ions and AgNP were more toxic to P. aeruginosa than to E. coli, while PMB was more toxic to E. coli (Figure 2A–D).
Compared with the bioluminescence inhibition test (Figure 2A–D), in the viability assay exposure to higher concentrations of chemicals was needed to irreversibly inhibit bacterial growth (Figure 2E–F). This difference was less remarkable for PMB (24-hour MBC values were nearly comparable to the respective 10-minute EC50 values) and more pronounced for Ag compounds, implying that inhibition of bacterial metabolism by Ag compounds did not instantly lead to cell death. Ag compounds completely abolished bacterial growth at concentrations from 10 μM (AgNO3) to 40 μM (AgNP) for P. aeruginosa and from 80 μM (AgNO3) to 320 μM (AgNP) for E. coli (Figure 2C and D). On the basis of 4× difference between the bactericidal effect of AgNO3 and AgNP evident for both bacteria, we hypothesized that dissolution of AgNP (26.9% in the test medium, Table 1) could have accounted for that result.
Evaluation of bacterial OM integrity by 1-NPN assay
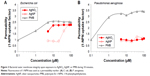
The impact of Ag+ ion and AgNP on the barrier properties of the cell envelope was studied using nonpolar dye 1-NPN that has strong fluorescence in the lipid environment but only weak fluorescence in the hydrophilic medium.24 The access of 1-NPN to lipid environment depends on the OM permeability (on structural order of LPS in the OM) and/or activity of the efflux pumps. When the membrane-active compounds disturb the integrity of OM, the lipophilic phase of cell envelope becomes accessible to hydrophobic agents, such as 1-NPN. Thus, the increase of 1-NPN fluorescence could be interpreted as the loss of OM integrity.24 Alternatively, the fluorescence can increase in case of the general de-energization of cells leading to the inhibition of efflux of lipophilic compounds, including 1-NPN.31
A cationic peptide, PMB, was used as a positive control due to its ability to disrupt the OM barrier and diminish the membrane potential.7 PMB increased the 1-NPN fluorescence approximately 3-fold in both E. coli and P. aeruginosa suspensions (Figure 3A and B). AgNO3 affected the 1-NPN fluorescence in case of E. coli cells starting from 5 μM and AgNP starting from 40 μM, although their effect was weaker than that in case of PMB. As the dissolution of AgNP was 26.9% at these conditions (Table 1), the effects of AgNP on 1-NPN fluorescence were most probably caused by Ag+ ions released from AgNP.
Differently from E. coli, no effect of Ag compounds (neither AgNO3, nor AgNP) was observed in the case of P. aeruginosa suspensions (Figure 3B). This was in contrast to higher toxicity of AgNO3 and to AgNP toward P. aeruginosa than to E. coli cells (Figure 2) indicating that the OM was not a primary target of Ag compounds. Thus, we analyzed AgNO3 and AgNP effects on bacteria in more detail, monitoring their interaction with PM and OM using TPP+ assay.
Evaluation of the permeability of bacterial membranes by TPP+ assay
To assay effects of AgNP and Ag+ ions on the permeability of bacterial OM and PM, TPP+ accumulation inside the cells (in the cytosol) was analyzed. Accumulation of TPP+ in the cytosol depends on the difference of electrical potential across the PM (membrane potential) and the OM barrier. The OM forms a permeability barrier to lipophilic ions like TPP+, whereas PM is permeable to TPP+. Thus, accumulation of TPP+ by the cells would indicate permeabilization of the OM by the Ag compounds, whereas leakage of the accumulated TPP+ would indicate the depolarization of PM.7 E. coli and P. aeruginosa cells with the intact OM accumulated low amounts of TPP+ and additions of increasing concentrations of AgNO3 to bacterial suspensions did not induce any additional binding of TPP+ to the cells, indicating that Ag compounds did not act on OM (data not shown).
To induce the accumulation of TPP+ and to measure the effect of Ag compounds on PM, EDTA was added to the cells. EDTA removes divalent cations from the LPS layer, increasing the permeability of the OM to lipophilic compounds.7 Both bacteria accumulated considerable amounts of TPP+ when 0.1 mM EDTA was added to the cell suspensions (Figure 4). Addition of 1–2.5 μM AgNO3 did not induce any changes in the amount of cell-accumulated TPP+, but 5 μM and higher concentrations of Ag+ ions induced release of TPP+ from the cells. Addition of 20 μM Ag+ ions induced release of the maximal amount of TPP+. In experiments with AgNP, 160 μM concentration was needed to depolarize the bacterial PM and induce leakage of accumulated TPP+ (Figure 4C and D).
Evaluation of the permeability of the bacterial PM by K+ leakage assay
Concentration of K+ ions in incubation medium was monitored to evaluate the effects of AgNP and Ag+ ions on barrier functions of the PM. In contrast to TPP+, K+ ions easily cross the OM through porins but the PM forms a barrier to this ion. Therefore, K+ gradient specifically characterizes the PM.
Addition of 10 μM AgNO3 induced leakage of accumulated K+ from the cells of both bacteria (Figure 5). Notably, accumulation of K+ in cells is an energy-dependent process that is sensitive to depolarization of the PM. Taking into account that depolarization of the PM observed in TPP+ assay (Figure 4) preceded the leakage of accumulated K+ ions (Figure 5), we suggest that depolarization of the PM was the reason of dissipation of K+ gradient. Unfortunately, it was technically not possible to register AgNP-induced K+ release since high concentrations of AgNP affected the K+ electrode.
Evaluation of the permeability of bacterial PM using cell respiration assay
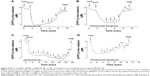
To understand the mechanism of Ag+ ion and AgNP action on bacterial PM, we analyzed the respiration activity of the cells. Starting from 1 μM concentration, the addition of AgNO3 stimulated the respiration of E. coli cells, but in 5 minutes the activation period was followed by a considerable, Ag+ concentration-dependent inhibition (Figure 6A). With the increase of Ag+ concentration, the period of enhanced respiration activity became shorter and the effect of inhibition stronger. The period of stimulated respiration of E. coli cells was considerably less expressed in the medium without glucose (data not shown). AgNP did not have any respiration stimulating activity in these conditions (Figure 6B). In the case of P. aeruginosa cells, the respiration enhancing activity of AgNO3 was the highest at 5 μM, and only the inhibitory activity of Ag+ was observed at 10 μM (Figure 6C). However, AgNP started to stimulate respiration at 40 μM and the inhibition started at 160 μM (Figure 6D). Such activation of respiration at low concentrations and inhibition at higher ones is typical for uncouplers of oxidative phosphorylation, ie, compounds permeabilizing the PM to H+ ions.32
Evaluation of the permeability of bacterial membrane by TPP+ assay in response to oxidized NP
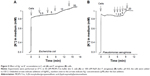
To get direct evidence that Ag+ ions are responsible for the PM-damaging effects of AgNP, AgNP were step-wise oxidized (dissolved) using KMnO4. Suspensions of AgNP (final concentration 1 mM Ag) were made in KMnO4 solutions of concentrations from 0.025 to 0.25 mM (Figure 7). When the concentration of KMnO4 in the solution was 0.025 mM, 80 μM of AgNP was needed to induce the depolarization. However, when the concentration of KMnO4 was increased to 0.25 mM, only 10 μM of AgNP was needed to induce the depolarization of the PM. Thus, the efficiency of AgNP depended on the concentration of KMnO4 in the solution, ie, on the oxidized fraction of Ag in the AgNP. Remarkably, the efficiency of maximally oxidized AgNP (Figure 7) was very close to the efficiency of AgNO3 solution (5 μM, Figure 4). Thus, the experiments with suspensions of AgNP in KMnO4 solutions clearly showed, that Ag+ ions are responsible for the rapid PM depolarizing effect of AgNP.
Discussion
Discussions on the mechanisms of AgNP interaction with bacterial cells are continuing despite a remarkable amount of experimental data on their toxicity and possible mode of action.33–36 One among the well-established mechanisms of action of AgNP on bacteria is their adhesion to the surface of bacterial envelope, disruption of the integrity of cellular membranes, and subsequent damage of (intra)cellular biomolecules.37 To the best of our knowledge, the exact mechanism of damage of the envelope barrier and the toxicity initiating event of AgNP have not yet been elucidated.
In this study we showed that the PM is the main target of AgNP in E. coli and P. aeruginosa. In the bacterial whole-cell bioluminescence inhibition assay (Figure 2) we observed that P. aeruginosa was inherently more susceptible to Ag compounds than E. coli, probably because the energy transformation of the former bacteria mainly depends on PM-linked ATP-ase.38 Using TPP+, K+, and respiration assays, we confirmed that the antibacterial effects of Ag and AgNP occur via challenging the PM. Indeed, exposure of bacteria to AgNP induced a set of second-scale acting toxic events targeting PM: inhibition of the respiration, depolarization of the PM, and leakage of the intracellular K+ (Figures 4–6). In addition, as shown in many previous studies, we confirmed that AgNP toxicity is mediated via Ag+ ions (Figure 7). This was observed in bacterial bioluminescence inhibition assay (Figure 2) in combination with AgNP dissolution studies (Table 1), and confirmed by AgNP oxidation assay which showed that Ag+ ions dissolving from AgNP were driving the quick toxic effects of AgNP on the bacterial PM (Figure 7). Most likely, the action of Ag+ ions on PM is unspecific and occurs through binding of Ag+ ions to SH-groups of vitally important enzymes localized on the PM.12,15
Surprisingly, we did not observe any specific effects of Ag compounds on bacterial OM (Figures 3 and 4). Several studies involving transmission electron microscopy have previously shown structural changes in the surface of bacteria after exposure to AgNP, and proposed that physical interaction with AgNP and/or ROS was the primary cause of cell death.10,13,16 It is highly possible that different AgNP have distinct antibacterial mechanisms that seem to depend on AgNP dissolution. For example, the study by Ramalingam et al16 revealed that biosynthesized AgNP with low dissolution rates caused physical and ROS-mediated damage to bacterial cells. Authors showed that in the absence of AgNP, ζ-potential values of E. coli and P. aeruginosa cells in 50 mM phosphate buffer were −28.5 mV and −20.6 mV, respectively, whereas they became less negative after incubation with AgNP, suggesting cell-NP interaction and neutralization of the cell surface charge by AgNP. Our surface charge measurements revealed results similar to Ramalingam et al:16 −39.1 mV and −22.4 mV for E. coli and P. aeruginosa cells in MOPS-Tris buffer, respectively, that, however, did not change after incubation with AgNP. Also, in contrast to the study by Ramalingam et al,16 AgNP used in our study did not induce ROS. Thus, it can be suggested that AgNP with high dissolution rates kill bacteria via Ag+ ion-mediated action before the potential induction of ROS and the effect of physical interaction can be observed. On the contrary, AgNP with lower dissolution rates are usually less cytotoxic4,39 and enable observation of the contribution of ROS and physical interaction to net cytotoxicity.
Since Ag+ ions dissolved from AgNP reached bacterial PM without causing significant OM damage, these data support the idea that Ag+ transport occurs via cation selective OM porins.40,41 Previous studies showed that E. coli mutant strains deficient in OmpF or OmpC porins were more resistant to Ag+ ions9 and to AgNP11 as compared to the wild-type strain. In addition, ompF gene was downregulated in response to Ag+ ions and AgNP,10 suggesting that porins are crucial in antibacterial effects of Ag+ ions and AgNP. On the basis of our study and the previous observations, we suggest that Ag+ ions dissolved from AgNP enter the bacterial periplasm through porin channels and interact directly with the components of PM.
Finally, the assays used in our study showed the rapid (seconds to minutes scale) dissolution-dependent antimicrobial mechanisms of Ag compounds on both bacteria. We observed that the rapid toxicity of AgNP to E. coli and P. aeruginosa differed only a few times (Figure 2) in this study, while the long-term toxicity (exposure time of 4 hours) of AgNP was almost two orders of magnitude different for these two bacteria, as shown in our previous article.4 Thus, the rapid dissolution-dependent effects of AgNP might be universal but the time-dependent effects seem to be specific to different bacteria. For example, the long-term effects may be related to the capacity of bacteria to induce oxidative dissolution of adsorbed AgNP to Ag+ ions and/or alter the efficiency of Ag+ efflux. For instance, it has been shown that E. coli with the more active efflux transporters of Ag+ ions (eg, SilCFBA) is more resistant to Ag.42
Conclusion
We demonstrated that Ag+ ions and dissolving AgNP induced a cascade of rapid toxic effects targeting PM of E. coli and P. aeruginosa, including inhibition of respiration, depolarization of the PM, leakage of the intracellular K+, and inhibition of metabolic activity. These fast (seconds to minutes scale) adverse effects of AgNP and AgNO3 depended on released Ag+ ions targeting the PM, whereas the effect on bacterial OM in this time scale was negligible. Therefore, we propose that the design of AgNP-based antimicrobial materials should focus on the burst release of Ag+ ions to ensure the rapid killing of bacteria, thereby avoiding the development of resistant bacterial strains. The set of methods applied in this study is suitable for evaluation of rapidly acting antibacterials.
Acknowledgments
This work was supported by Estonian Research Council grants IUT23-5 and PUT1015 and by Research Council of Lithuania, funding grant no MIP-040/2015. Mr Ilja Makarenkov is acknowledged for the scheme in Figure 1. The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Disclosure
The authors report no conflicts of interest in this work.
References
Nowack B, Krug HF, Height M. 120 years of nanosilver history: implications for policy makers. Environ Sci Technol. 2011;45(4):1177–1183. | ||
Djurišić AB, Leung YH, Ng AM, et al. Toxicity of metal oxide nanoparticles: mechanisms, characterization, and avoiding experimental artefacts. Small. 2015;11(1):26–44. | ||
Le Ouay B, Stellacci F. Antibacterial activity of silver nanoparticles: A surface science insight. Nano Today. 2015;10(3):339–354. | ||
Bondarenko O, Ivask A, Käkinen A, Kurvet I, Kahru A. Particle-cell contact enhances antibacterial activity of silver nanoparticles. PLoS One. 2013;8(5):e64060. | ||
Silhavy TJ, Kahne D, Walker S. The bacterial cell envelope. Cold Spring Harb Perspect Biol. 2010;2(5):a000414. | ||
Amro NA, Kotra LP, Wadu-Mesthrige K, Bulychev A, Mobashery S, Liu G-Y. High-Resolution Atomic Force Microscopy Studies of the Escherichia coli Outer Membrane: Structural Basis for Permeability. Langmuir. 2000;16(6):2789–2796. | ||
Daugelavičius R, Bakienė E, Bamford DH. Stages of polymyxin B interaction with the Escherichia coli cell envelope. Antimicrob Agents Chemother. 2000;44(11):2969–2978. | ||
Holt KB, Bard AJ. Interaction of silver(I) ions with the respiratory chain of Escherichia coli: an electrochemical and scanning electrochemical microscopy study of the antimicrobial mechanism of micromolar Ag+. Biochemistry. 2005;44(39):13214–13223. | ||
Li XZ, Nikaido H, Williams KE. Silver-resistant mutants of Escherichia coli display active efflux of Ag+ and are deficient in porins. J Bacteriol. 1997;179(19):6127–6132. | ||
Mcquillan JS, Infante HG, Stokes E, Shaw AM. Silver nanoparticle enhanced silver ion stress response in Escherichia coli K12. Nanotoxicology. 2012;6:857–866. | ||
Radzig MA, Nadtochenko VA, Koksharova OA, Kiwi J, Lipasova VA, Khmel IA. Antibacterial effects of silver nanoparticles on gram-negative bacteria: influence on the growth and biofilms formation, mechanisms of action. Colloids Surf B Biointerfaces. 2013;102:300–306. | ||
Dibrov P, Dzioba J, Gosink KK, Häse CC. Chemiosmotic mechanism of antimicrobial activity of Ag(+) in Vibrio cholerae. Antimicrob Agents Chemother. 2002;46(8):2668–2670. | ||
Sondi I, Salopek-Sondi B. Silver nanoparticles as antimicrobial agent: a case study on E. coli as a model for Gram-negative bacteria. J Colloid Interface Sci. 2004;275(1):177–182. | ||
Lok CN, Ho CM, Chen R, et al. Proteomic analysis of the mode of antibacterial action of silver nanoparticles. J Proteome Res. 2006;5(4):916–924. | ||
Li WR, Xie XB, Shi QS, Zeng HY, Ou-Yang YS, Chen YB. Antibacterial activity and mechanism of silver nanoparticles on Escherichia coli. Appl Microbiol Biotechnol. 2010;85(4):1115–1122. | ||
Ramalingam B, Parandhaman T, das SK. Antibacterial effects of biosynthesized silver nanoparticles on surface ultrastructure and nanomechanical properties of Gram-negative bacteria viz. Escherichia coli and Pseudomonas aeruginosa. ACS Appl Mater Interfaces. 2016;8(7):4963–4976. | ||
ISO/IEC, 2017. General requirements for the competence of testing and calibration laboratories. International Standard, reference number ISO/IEC 17025:2017(E). Geneva, Switzerland: International Organization for Standardization. | ||
Bondarenko O, Rahman PK, Rahman TJ, Kahru A, Ivask A. Effects of rhamnolipids from Pseudomonas aeruginosa DS10-129 on luminescent bacteria: toxicity and modulation of cadmium bioavailability. Microb Ecol. 2010;59(3):588–600. | ||
Ivask A, Rõlova T, Kahru A. A suite of recombinant luminescent bacterial strains for the quantification of bioavailable heavy metals and toxicity testing. BMC Biotechnol. 2009;9:41. | ||
ISO. Water quality – Kinetic determination of the inhibitory effects of sediment, other solids and coloured samples on the light emission of Vibrio fischeri (kinetic luminescent bacteria test). International Standard, reference number ISO 21338:2010(E). International Organization for Standardization, Geneva, Switzerland. 2010. | ||
Kurvet I, Ivask A, Bondarenko O, Sihtmäe M, Kahru A. LuxCDABE – transformed constitutively bioluminescent Escherichia coli for toxicity screening: comparison with naturally luminous Vibrio fischeri. Sensors. 2011;11(8):7865–7878. | ||
MS Excel macro Regtox; 2009. Vindimian E. Available from: http://www.normalesup.org/~vindimian/en_index.html. Accessed September 7, 2018. | ||
Suppi S, Kasemets K, Ivask A, et al. A novel method for comparison of biocidal properties of nanomaterials to bacteria, yeasts and algae. J Hazard Mater. 2015;286:75–84. | ||
Helander IM, Mattila-Sandholm T. Fluorometric assessment of gram-negative bacterial permeabilization. J Appl Microbiol. 2000;88(2):213–219. | ||
Daugelavičius R, Gaidelytė A, Cvirkaitė-Krupovič V, Bamford DH. On-line monitoring of changes in host cell physiology during the one-step growth cycle of Bacillus phage Bam35. J Microbiol Methods. 2007;69(1):174–179. | ||
Daugelavičius R, Buivydas A, Senčilo A, Bamford DH. Assessment of the activity of RND-type multidrug efflux pumps in Pseudomonas aeruginosa using tetraphenylphosphonium ions. Int J Antimicrob Agents. 2010;36(3):234–238. | ||
Blinova I, Niskanen J, Kajankari P, et al. Toxicity of two types of silver nanoparticles to aquatic crustaceans Daphnia magna and Thamnocephalus platyurus. Environ Sci Pollut Res Int. 2013;20(5):3456–3463. | ||
Aruoja V, Pokhrel S, Sihtmäe M, Mortimer M, Mädler L, Kahru A. Toxicity of 12 metal-based nanoparticles to algae, bacteria and protozoa. Environ Sci Nano. 2015;2(6):630–644. | ||
Bulich AA. A practical and reliable method for monitoring the toxicity of aquatic samples. Process Biochem. 1982;17:45–47. | ||
Elnabarawy MT, Robideau RR, Beach SA. Comparison of three rapid toxicity test procedures: Microtox,® polytox,® and activated sludge respiration inhibition. Toxicity Assessment. 1988;3(4):361–370. | ||
Ocaktan A, Yoneyama H, Nakae T. Use of fluorescence probes to monitor function of the subunit proteins of the MexA-MexB-oprM drug extrusion machinery in Pseudomonas aeruginosa. J Biol Chem. 1997;272(35):21964–21969. | ||
Nicholls D, Ferguson SJ. Bioenergetics. 4th ed. San Diego: Academic Press; 2013 eBook ISBN: 9780123884312. | ||
Ivask A, Juganson K, Bondarenko O, et al. Mechanisms of toxic action of Ag, ZnO and CuO nanoparticles to selected ecotoxicological test organisms and mammalian cells in vitro: a comparative review. Nanotoxicology. 2014;8(Suppl 1):57–71. | ||
Juganson K, Ivask A, Blinova I, Mortimer M, Kahru A. NanoE-Tox: New and in-depth database concerning ecotoxicity of nanomaterials. Beilstein J Nanotechnol. 2015;6:1788–1804. | ||
Bondarenko OM, Heinlaan M, Sihtmäe M, et al. Multilaboratory evaluation of 15 bioassays for (eco)toxicity screening and hazard ranking of engineered nanomaterials: FP7 project NANOVALID. Nanotoxicology. 2016;10(9):1229–1242. | ||
Slavin YN, Asnis J, Häfeli UO, Bach H. Metal nanoparticles: understanding the mechanisms behind antibacterial activity. J Nanobiotechnology. 2017;15(1):65. | ||
Dakal TC, Kumar A, Majumdar RS, Yadav V. Mechanistic Basis of Antimicrobial Actions of Silver Nanoparticles. Front Microbiol. 2016;7:1831. | ||
Temple LM, Sage AE, Schweizer HP, Phibbs PV. Carbohydrate Catabolism in Pseudomonas aeruginosa. In: Montie TC, editor. Pseudomonas. Biotechnology Handbooks. Vol. 10. Boston, MA: Springer; 1998:35–72. | ||
Kennedy AJ, Chappell MA, Bednar AJ, et al. Impact of organic carbon on the stability and toxicity of fresh and stored silver nanoparticles. Environ Sci Technol. 2012;46(19):10772–10780. | ||
Lemire JA, Harrison JJ, Turner RJ. Antimicrobial activity of metals: mechanisms, molecular targets and applications. Nat Rev Microbiol. 2013;11(6):371–384. | ||
Mcquillan JS, Shaw AM. Differential gene regulation in the Ag nanoparticle and Ag(+)-induced silver stress response in Escherichia coli: a full transcriptomic profile. Nanotoxicology. 2014;8 Suppl 1:177–184. | ||
Randall CP, Gupta A, Jackson N, Busse D, O’Neill AJ. Silver resistance in Gram-negative bacteria: a dissection of endogenous and exogenous mechanisms. J Antimicrob Chemother. 2015;70(4):1037–1046. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.