Back to Journals » Lung Cancer: Targets and Therapy » Volume 11
Patients with NSCLCs Harboring Internal Inversions or Deletion Rearrangements of the ALK Gene Have Durable Responses to ALK Kinase Inhibitors
Authors Schrock AB, Madison R , Rosenzweig M, Allen JM, Erlich RL, Wang SY, Chidiac T, Reddy VS, Riess JW, Yassa AE , Shakir A , Miller VA, Alexander BM , Venstrom J, McGregor K , Ali SM
Received 22 November 2019
Accepted for publication 24 March 2020
Published 17 April 2020 Volume 2020:11 Pages 33—39
DOI https://doi.org/10.2147/LCTT.S239675
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sai-Hong Ignatius Ou
Alexa B Schrock,1 Russell Madison,1 Mark Rosenzweig,2 Justin M Allen,2 Rachel L Erlich,2 Siao-Yi Wang,3 Tarek Chidiac,4 Vodur Suresh Reddy,5 Jonathan W Riess,6 Ahmet Ersin Yassa,6 Abdur Shakir,7 Vincent A Miller,1 Brian M Alexander,1 Jeffrey Venstrom,1 Kimberly McGregor,1 Siraj M Ali1
1Foundation Medicine, Department of Clinical Development, Cambridge, MA, USA; 2Foundation Medicine, Department of Translational Oncology and Clinical Reporting, Cambridge, MA, USA; 3Loyola University Medical Cancer, Department of Hematology and Oncology, Maywood, IL, USA; 4Zangmeister Cancer Center, Department of Hematology and Oncology, Columbus, OH, USA; 5Cancer Care Specialists, Department of Hematology and Oncology, Reno, NV, USA; 6UC Davis Comprehensive Cancer Center, Department of Hematology and Oncology Sacramento, CA, USA; 7Sarah Bush Lincoln Health System, Department of Medical Oncology, Mattoon, IL, USA
Correspondence: Alexa B Schrock
Foundation Medicine, 150 Second Street, Cambridge, MA 02141, USA
Tel +1617-418-2200
Email [email protected]
Background: ALK fusions are targetable drivers in non-small-cell lung cancer (NSCLC). However, patients with NSCLC harboring ALK rearrangements without a fusion partner identified in DNA have also been shown to respond to ALK inhibitors. We aimed to characterize complex ALK variants that may predict sensitivity to multiple approved ALK inhibitors.
Methods: Comprehensive genomic profiling (CGP) of DNA isolated from formalin‐fixed paraffin‐embedded (FFPE) tumor tissue or blood-based circulating tumor DNA was performed for 39,159 NSCLC patients during routine clinical care. For a subset of cases, RNA sequencing was performed, and prior ALK test results and clinical treatment information were collected from treating physicians.
Results: We queried the Foundation Medicine NSCLC database and identified ALK internal inversions, as well as internal deletions, as the sole ALK rearrangements in 6 (0.02%) and 3 (0.01%) of cases, respectively. In cases with ALK internal inversions, RNA testing identified an EML4-ALK fusion in 2/2 cases evaluated, and 3/3 patients treated with ALK inhibitors had durable responses. A single patient with an ALK internal deletion and clinical data available responded to multiple ALK inhibitors. RNA data available for a subset of non-NSCLC cases suggest that ALK internal deletions removing a portion of the N-terminus are drivers themselves and do not result in ALK fusions. Fluorescence in situ hybridization (FISH) results were inconsistent for both classes of DNA events.
Conclusion: Rare internal inversions of ALK appear to be indicative of ALK fusions, which can be detected in RNA, and response to ALK inhibitors in patients with NSCLC. In contrast, ALK internal deletions are not associated with ALK fusions in RNA but likely represent targetable drivers themselves. These data suggest that CGP of DNA should be supplemented with immunohistochemistry or RNA-based testing to further resolve these events and match patients to effective therapies.
Keywords: ALK rearrangement, inversion, deletion, genomic profiling, targeted therapy
Introduction
ALK gene fusions are known oncogenic drivers in non-small-cell lung cancer (NSCLC) and other tumor types, and are targetable with multiple FDA-approved ALK tyrosine kinase inhibitors (TKIs).1 ALK rearrangements, identified using fluorescence in situ hybridization (FISH), immunohistochemistry (IHC) or next-generation sequencing (NGS), typically result in the ALK kinase domain fused to a 5ʹ dimerization partner. Patients with NSCLC positive for these alterations have excellent response rates to ALK TKIs.2–4 The various accepted methods for detection of ALK rearrangements are generally concordant; however, previous studies have shown that cases negative by ALK FISH can be positive using NGS, particularly cases with complex DNA events, and that these patients respond to ALK TKIs.5,6 Given the efficacy of ALK TKIs as a class, deep understanding and exploration of these more complex ALK variants is warranted.
While the majority of ALK rearrangements detected using NGS are fusions with an identified 5ʹ partner, in a subset of cases DNA rearrangements are detected without evidence of a gene fusion. Case reports of NSCLCs with ALK rearrangements but no fusion partner detected in DNA have demonstrated ALK fusions in RNA and responses to ALK TKIs; however the literature remains relatively scant.7–9 In a subset of cases, the N-terminal domain of ALK is predicted to be separated from the kinase domain through rearrangement or alternative transcription, resulting in activation in the absence of a gene fusion.10,11 In this report we present multiple patients whose tumors harbor novel rearrangements where both detected breakpoints occur within the ALK gene, and who experienced durable responses to ALK TKIs.
Methods
Hybrid‐capture based comprehensive genomic profiling (CGP; FoundationOneCDx) was performed prospectively for 39,159 NSCLC patients on formalin‐fixed paraffin‐embedded (FFPE) tumor tissue or circulating tumor DNA (ctDNA) submitted during routine clinical care in a Clinical Laboratory Improvement Amendments‐certified, College of American Pathologists‐accredited, New York State‐regulated reference laboratory (Foundation Medicine Inc., Cambridge, MA). DNA (>50ng) was extracted from FFPE NSCLC specimens; NGS was performed on hybridization‐captured, adaptor ligation‐based libraries to high, uniform coverage (>500x) for all coding exons of 236–405 cancer‐related genes plus selected introns.12 Additionally, since May 2016, hybrid-capture based NGS was performed on ctDNA.13 Two 10-mL aliquots of peripheral whole blood were collected, a double-spin protocol was used to isolate plasma, and 50–100ng of ctDNA was extracted to create adapted sequencing libraries before hybrid-capture and sample-multiplexed sequencing of 62–70 genes plus selected introns to >5000x unique coverage. All ALK exons were baited; dedicated intron baiting was included for ALK introns 18 and 19 in ctDNA and intron 19 in tissue. DNA and RNA CGP (FoundationOneHeme) was performed on selected samples where indicated as assay previously described.14 Approval for this study, including a waiver of informed consent and a Health Insurance Portability and Accountability Act waiver of authorization, was obtained from the Western Institutional Review Board (protocol no. 20152817).
Results
The index patient is a 50-year-old female never smoker diagnosed with stage III lung adenocarcinoma in 2015. ALK FISH testing, as well as ROS1 FISH and EGFR mutation testing, was negative. She received carboplatin/pemetrexed/bevacizumab followed by nivolumab and discontinued both due to toxicity. The treating physician then ordered CGP of a right lung core biopsy which showed an ALK rearrangement (intron 17/19 breakpoints) predicted to result in an internal inversion. No alterations in other known drivers were detected. Subsequent ALK IHC testing was positive and the patient began treatment with crizotinib. Due to intolerability, after less than 2 months she was switched to alectinib with excellent partial response (PR) lasting 26 months (Figure 1). At progression on alectinib no resistance alterations within ALK were identified, although an inactivating STK11 alteration was acquired, and the patient was recently switched to lorlatinib.
To assess the frequency and diversity of this class of ALK rearrangements we queried the Foundation Medicine database of 33,601 tissue and 4679 ctDNA samples from 39,159 NSCLC patients, which were submitted for CGP during clinical care (July 2015 - January 2019). DNA rearrangements with both breakpoints within ALK predicted to result in inversion of a portion of the N-terminus leaving the kinase domain (exons 20–29) intact were found in 25 cases (0.06%; 24 tissue and 1 ctDNA sample). In most cases (76%, 19/25), the inversion co-occurred with another ALK rearrangement, typically an EML4-ALK fusion, but in 6 cases the inversion was the only ALK alteration identified in DNA. All 6 cases were lung adenocarcinomas, 5/6 patients were female, and the median patient age was 60.5 years (range 23–82) (Table 1). There were no other known driver alterations in 5/6 cases; the sixth case, which was known to be treatment naïve, had a co-occurring EGFR L858R mutation (mutant allele frequency 30%). The 5ʹ ALK breakpoint was in introns 8–17 and the 3ʹ ALK breakpoint was in intron 18 (n=1) or intron 19 (n=5). Two cases were evaluated for ALK rearrangements in RNA and in both harbored an EML4-ALK RNA fusion.
 |
Table 1 Clinical and Genomic Characteristics of NSCLC Cases with ALK Internal Rearrangements |
Two additional patients with ALK internal inversions in DNA had clinical follow up data available. Case 2 is that of a 23-year-old male never-smoker diagnosed with stage IV lung adenocarcinoma in 2018. He had a right frontal lobe brain lesion resection, consistent with metastatic adenocarcinoma. CGP was ordered on a left lung biopsy, which showed an ALK rearrangement (intron 8/19 breakpoints) predicted to result in an internal inversion. ALK FISH testing on the lung biopsy was positive. Follow up DNA and RNA CGP on the resected brain tissue were performed showing the same ALK internal inversion in DNA but an EML4-ALK fusion in RNA. The patient began treatment with alectinib and underwent stereotactic radiosurgery (SRS) to his brain lesion approximately 1 month later. The main mass in his left lung mass measured 7.2x2.5cm at diagnosis and decreased to 3.7x2.5cm at 5 weeks and 2.7x1.5cm at 14 weeks after initiation of alectinib and no other metastatic sites outside the brain were identified. A brain MRI 3.5 months after treatment initiation showed a new 11mm lesion and SRS was repeated. He remains on alectinib for 7 months with an ongoing response in the lung.
Case 3 is that of a 78-year-old female never-smoker diagnosed with stage IV lung adenocarcinoma in 2018. CGP was performed on a cell pellet from pleural fluid collected at diagnosis, which detected an ALK rearrangement (intron 15/19 breakpoints) predicted to result in an internal inversion. The patient received chemotherapy and immunotherapy for approximately 2 months with some intolerance and no evidence of response, before beginning alectinib. Subsequent ALK FISH testing was negative and alectinib was interrupted for approximately 2 weeks. DNA and RNA CGP were performed on the original pleural fluid cell pellet and showed the same ALK internal inversion in DNA but an EML4-ALK fusion in RNA. Alectinib was resumed and the patient remains on therapy for 12 months with a complete response.
We also identified 8 NSCLC cases (0.02%) with DNA rearrangements with both breakpoints within the ALK gene predicted to result in deletion of a portion of the N-terminal region (Table 1). In 3/8 cases, no co-occurring ALK rearrangement or other drivers were detected. Clinical follow up data was available for 1 of these patients. Case 4 is that of a 66-year old female who was diagnosed with Stage IIIA lung adenocarcinoma in 2016. She initially underwent tri-modality therapy including cisplatin/etoposide plus radiation therapy followed by R middle and lower lobectomy with lymph node dissection. As both the 4R node and primary tumor were ALK positive by FISH, she was started on crizotinib for residual unresectable disease. Treatment was discontinued after 3 months due to toxicity and imaging demonstrated no evidence of disease. She was subsequently found to have liver lesions 1 year later and CGP of a liver metastasis detected an ALK rearrangement (intron 12/19 breakpoints) predicted to result in an internal deletion. She was restarted on crizotinib with a PR and remained on treatment for 16 months until progression of liver lesions. She was switched to brigatinib but discontinued due to pneumonitis and subsequently began treatment with alectinib, ongoing for 10 months with a PR.
Discussion
Following a dramatic prolonged response to alectinib in a patient with NSCLC harboring a previously uncharacterized ALK rearrangement, we queried a large genomic database of over 39,000 NSCLCs and found that ALK internal inversion rearrangements are rare (<0.1%) but recurrent events, sometimes difficult to detect using FISH but likely clinically actionable. These inversions involve a subset of exons 1–19 and typically include classic intron 19 breakpoints. They are often detected with other co-occurring ALK rearrangements but can occur as the sole ALK DNA event. FISH testing was available for 3/6 such cases and was positive in 1 instance (intron 8/19 breakpoints) but negative in the other 2 (intron 15/19 and 17/19 breakpoints), suggesting that detection by FISH may depend on the size of a given inversion. Importantly, subsequent analyses revealed that in 2/2 cases with only an ALK internal inversion in DNA, RNA testing detected a classic EML4-ALK fusion. ALK internal inversions as the sole ALK rearrangement were also detected in 5 additional cases in the Foundation Medicine database of ~170,000 non-NSCLC tumor samples; however, none of these cases had a canonical intron 19 breakpoints or RNA data available, so the functional status of these events is unclear.
A plausible biological explanation for our observation that 2/2 cases with ALK DNA internal inversions harbor RNA-only ALK fusions is that these inversions are not driver alterations per se but are indicators of more extensive rearrangement events affecting the ALK locus. Because typical commercial NGS sequencing technologies use short sequencing reads to obtain raw sequencing data, which is then computationally assembled and mapped to the reference genome, the full extent of a complex DNA rearrangement can be difficult to capture. Thus, DNA rearrangements might sometimes appear as partner-less events or events of unclear biological significance, such as ALK internal inversions, while RNA sequencing of these cases reveals in-frame fusions. This phenomenon has been reported previously using other NGS platforms, suggesting a general limitation of the technology.15,16
An important implication of this finding is that for cases where NGS DNA sequencing uncovers rearrangements of unclear significance in known oncogenes, orthogonal testing, such as with RNA or protein-based assays should be considered to ascertain whether the detected rearrangements are incompletely-captured oncogenic fusions or likely non-productive events.
In the case of the index patient, she responded to alectinib for 26 months, and at progression CGP of ctDNA showed the initial ALK intron 17/19 deletion and an ALK rearrangement with an intron 18 breakpoint, which was unlikely to have been detected initially because the tissue CGP assay utilized initially lacks dedicated intron 18 baiting. Without RNA evidence for this case it is not possible to assess directly, but given the excellent clinical response, we speculate that an ALK fusion would likely be detected in RNA. No ALK resistance mutations were detected at progression on alectinib; however, an acquired STK11 inactivating truncation was identified. STK11 alterations have been identified as resistance mechanisms to immune checkpoint inhibitors in NSCLC,17 but have not been described as acquired alterations driving resistance to ALK TKIs. However, in this case it’s conceivable that the STK11 alteration could be modulating the host anti-tumor response.
We also identified rare ALK N-terminal deletions and similar events have resulted in ALK activation in preclinical models.10,11 In 3/8 cases the deletion was the sole ALK rearrangement in DNA, and FISH testing available for 2/3 cases and was negative in 1 case (intron 3/19 breakpoints) and positive in the other (intron 12/19 breakpoints). Although no RNA data was available for any of the 5 NSCLC cases with ALK deletions, we also identified ALK internal deletions in 11 cases in the Foundation Medicine database of ~170,000 non-NSCLC tumor samples. In 3/3 of these cases with RNA data available (rhabdomyosarcoma with deletion of exons 2–17, unknown primary malignant neoplasm with deletion exons 3–18 and undifferentiated sarcoma with deletion of exons 2–16) the DNA-identified ALK internal deletion was confirmed in RNA, but none of these cases had an ALK fusion detected in RNA. Our report herein of a patient with NSCLC positive for an ALK internal deletion who responded to crizotinib, and then alectinib, provides preliminary validation of preclinical data showing that ALK deletions within the extracellular domain are activating and suggests that such deletions may define a novel class of clinically targetable driver alterations in NSCLC.
Overall this work provides preliminary clinical evidence for targetability of 2 distinct rare rearrangement classes that are detected as variants affecting the N-terminus of ALK. Although these events are uncommon, it’s possible that incomplete intron baiting leads to an underrepresentation in overall frequency. Limitations of this study include lack of clinical treatment data for a significant subset of patients, as well as incomplete FISH, IHC, and RNA sequencing data. However, available data suggest that ALK internal inversions detected in DNA correlate with the identification of ALK fusions in RNA (2/2 cases). Therefore, these events may be indicative of more extensive ALK rearrangements, and 100% (3/3) of patients with these events responded to ALK TKIs. Internal deletions in contrast do not appear to result in gene fusions (3/3 non-NSCLC cases with available RNA data were negative for ALK fusion) but do appear to be driver events with sensitivity to ALK TKIs. FISH results were mixed for both classes of rearrangements. This work suggests that CGP to identify these alterations, as well as supplemental RNA or IHC testing, is warranted to allow these patients to be matched to highly effective approved targeted therapies or enroll in clinical trials.
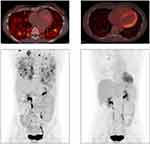 |
Figure 1 Response to alectinib in index case with ALK internal inversion. PET-CT scans just prior to initiation of alectinib (left) PET-CT scans after 3 months of treatment with alectinib (right). |
Consent for Publication
We confirm written informed consent for the case details to be published has been obtained for all patients described in this study.
Disclosure
ABS, RM, MR, JA, RLE, VAM, BA, JV, and KM are employees at Foundation Medicine, Inc., a wholly-owned subsidiary of Roche, and have Roche stock ownership. SMA was an employee at Foundation Medicine, Inc., at the time the study was conducted. JWR reports personal fees from Takeda, Celgene, and Abbvie; advisory board for Boehringer Ingelheim, Loxo Oncology, Spectrum Pharmaceuticals; grants from Merck, and grants provided to institution from Novartis and AstraZeneca, outside the submitted work. VAM is stockholder for Revolution Medicines and Mirati Therapeutics, outside the submitted work. In addition, he has a patent USPO85014313 issued to Sloan Kettering Institute for Cancer Research. The authors report no other conflicts of interest in this work.
References
1. Hallberg B, Palmer RH. Mechanistic insight into ALK receptor tyrosine kinase in human cancer biology. Nat Rev Cancer. 2013;13(10):685–700. doi:10.1038/nrc3580
2. Peters S, Camidge DR, Shaw AT, et al. Alectinib versus crizotinib in untreated ALK-positive non-small-cell lung cancer. N Engl J Med. 2017;377(9):829–838. doi:10.1056/NEJMoa1704795
3. Solomon BJ, Besse B, Bauer TM, et al. Lorlatinib in patients with ALK-positive non-small-cell lung cancer: results from a global Phase 2 study. Lancet Oncol. 2018;19(12):1654–1667. doi:10.1016/S1470-2045(18)30649-1
4. Camidge DR, Kim HR, Ahn M-J, et al. Brigatinib versus crizotinib in ALK-positive non-small-cell lung cancer. N Engl J Med. 2018;379(21):2027–2039. doi:10.1056/NEJMoa1810171
5. Ali SM, Hensing T, Schrock AB, et al. Comprehensive genomic profiling identifies a subset of crizotinib-responsive alk-rearranged non-small cell lung cancer not detected by fluorescence in situ hybridization. Oncologist. 2016;21(6):762–770. doi:10.1634/theoncologist.2015-0497
6. Li W, Zhang J, Guo L, Chuai S, Shan L, Ying J. Combinational analysis of FISH and immunohistochemistry reveals rare genomic events in ALK fusion patterns in NSCLC that responds to crizotinib treatment. J Thorac Oncol. 2017;12(1):94–101. doi:10.1016/j.jtho.2016.08.145
7. Peled N, Palmer G, Hirsch FR, et al. Next-generation sequencing identifies and immunohistochemistry confirms a novel crizotinib-sensitive ALK rearrangement in a patient with metastatic non-small-cell lung cancer. J Thorac Oncol. 2012;7(9):e14–e16. doi:10.1097/JTO.0b013e3182614ab5
8. Zhao R, Zhang J, Han Y, et al. Clinicopathological features of ALK expression in 9889 cases of non-small-cell lung cancer and genomic rearrangements identified by capture-based next-generation sequencing: a chinese retrospective analysis. Mol Diagn Ther. 2019;23(3):395–405. doi:10.1007/s40291-019-00389-y
9. Ou S-HI, Lee TK, Young L, et al. Dual occurrence of ALK G1202R solvent front mutation and small cell lung cancer transformation as resistance mechanisms to second generation ALK inhibitors without prior exposure to crizotinib. Pitfall of solely relying on liquid re-biopsy? Lung Cancer Amst Neth. 2017;106:110–114. doi:10.1016/j.lungcan.2017.02.005
10. Wiesner T, Lee W, Obenauf AC, et al. Alternative transcription initiation leads to expression of a novel ALK isoform in cancer. Nature. 2015;526(7573):453–457. doi:10.1038/nature15258
11. Cazes A, Louis-Brennetot C, Mazot P, et al. Characterization of rearrangements involving the ALK gene reveals a novel truncated form associated with tumor aggressiveness in neuroblastoma. Cancer Res. 2013;73(1):195–204. doi:10.1158/0008-5472.CAN-12-1242
12. Frampton GM, Fichtenholtz A, Otto GA, et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol. 2013;31(11):1023–1031. doi:10.1038/nbt.2696
13. Clark TA, Chung JH, Kennedy M, et al. Analytical validation of a hybrid capture-based next-generation sequencing clinical assay for genomic profiling of cell-free circulating tumor DNA. J Mol Diagn. 2018;20(5):686–702. doi:10.1016/j.jmoldx.2018.05.004
14. He J, Abdel-Wahab O, Nahas MK, et al. Integrated genomic DNA/RNA profiling of hematologic malignancies in the clinical setting. Blood. 2016;127(24):3004–3014. doi:10.1182/blood-2015-08-664649
15. Benayed R, Offin M, Mullaney K, et al. High yield of RNA sequencing for targetable kinase fusions in lung adenocarcinomas with no mitogenic driver alteration detected by DNA sequencing and low tumor mutation burden. Clin Cancer Res. 2019;25(15):4712–4722. doi:10.1158/1078-0432.CCR-19-0225
16. Davies KD, Le AT, Sheren J, et al. Comparison of molecular testing modalities for detection of ROS1 rearrangements in a cohort of positive patient samples. J Thorac Oncol. 2018;13(10):1474–1482. doi:10.1016/j.jtho.2018.05.041
17. Skoulidis F, Goldberg ME, Greenawalt DM, et al. STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov. 2018;8(7):822–835. doi:10.1158/2159-8290.CD-18-0099
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
