Back to Journals » Cancer Management and Research » Volume 12
Overexpression of FAM234B Predicts Poor Prognosis in Patients with Luminal Breast Cancer
Authors Lyu L, Wang M, Zheng Y , Tian T, Deng Y , Xu P , Lin S , Yang S, Zhou L , Hao Q, Wu Y, Dai Z , Kang H
Received 2 September 2020
Accepted for publication 12 November 2020
Published 3 December 2020 Volume 2020:12 Pages 12457—12471
DOI https://doi.org/10.2147/CMAR.S280009
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eileen O'Reilly
Lijuan Lyu,1,2,* Meng Wang,1,* Yi Zheng,1,2,* Tian Tian,1 Yujiao Deng,1,2 Peng Xu,1,2 Shuai Lin,1 Si Yang,1,2 Linghui Zhou,1,2 Qian Hao,1,2 Ying Wu,1,2 Zhijun Dai,2 Huafeng Kang1
1Department of Oncology, The 2nd Affiliated Hospital of Xi’an Jiaotong University, Xi’an 710004, People’s Republic of China; 2Department of Breast Surgery, The First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou 310003, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Huafeng Kang
Department of Oncology, The 2nd Affiliated Hospital of Xi’an Jiaotong University, West 5th Road, Xincheng District, Xi’an 710004, People’s Republic of China
Email [email protected]
Zhijun Dai
Department of Breast Surgery, The First Affiliated Hospital, College of Medicine, Zhejiang University, No. 79, Qingchun Road, Hangzhou 310003, People’s Republic of China
Email [email protected]
Background: Family with sequence similarity 234 member B (FAM234B), a protein-coding gene, is mainly expressed in brain tissues. Its clinical significance and biological function in tumors, especially in breast cancer (BC), have not been elucidated.
Methods: We firstly investigated the expression pattern of FAM234B at the mRNA and protein levels using Oncomine, TCGA portal, GEPIA, TIMER, HPA, and UALCAN databases, then applied bc-GenExMiner to assess the associations between expression level of FAM234B and clinicopathological features of BC. Besides, we also verified the expression of FAM234B expression in clinical BC samples using qRT-PCR. Subsequently, GEPIA, bc-GenExMiner, and TIMER databases were used to analyze the prognostic significance of FAM234B in all BC and different molecular subtypes. Finally, we conducted co-expression analysis and gene set enrichment analysis (GSEA). Additionally, we explored the regulatory mechanism of FAM234B in BC.
Results: Both bioinformatics analysis and experimental verification confirmed that the FAM234B expression was significantly higher at the mRNA and protein levels in luminal BC tissues than in adjacent normal tissues. High FAM234B expression was significantly correlated with older age, estrogen receptor-positive, progesterone receptor-positive, human epidermal growth factor receptor 2-negative, wild-type p53, low Nottingham prognostic index, low Scarff-Bloom-Richardson grade, lymph node metastasis positivity, and high tumor stage. Moreover, survival analysis indicated that high FAM234B expression was significantly related to a worse prognosis in patients with luminal BC. GSEA indicated that FAM234B was positively related to membrane transport process and negatively associated with immune response function. Besides, mechanism exploration indicated that pseudogene HTR7P1 might act as endogenous RNA to compete with has-miR-1271-5p or has-miR-381-3p for binding to FAM234B, thereby upregulating the expression of FAM234B in luminal BC.
Conclusion: Our results suggest that FAM234B may be a candidate therapeutic target or prognostic marker for luminal breast cancer.
Keywords: FAM234B, luminal breast cancer, HTR7P1, prognosis
Introduction
Breast cancer (BC) is one of the three most common cancers worldwide.1 Among females, BC is the leading cause of cancer-related deaths in the vast majority of countries.2,3 According to estrogen or progesterone receptor (ER or PR) expression and human epidermal growth factor receptor 2 (HER2) gene amplification, BC is classified into several molecular subtypes, including luminal, HER2-enriched, and basal-like.4,5 Different molecular subtypes have different clinical outcomes and treatment regimens. The luminal subtype is the most common, accounting for approximately 75% of all BC cases.6 Although great advances in early detection, surgical procedures, adjuvant endocrine therapy, and chemotherapy have substantially improved clinical outcomes in luminal BC, relapses and resistance to endocrine therapy are still important issues, leading to a low overall survival (OS) of patients with luminal BC.6,7 Since the disease remains a significant global health burden,8,9 the identification of specific and sensitive molecular biomarkers that may serve as therapeutic targets or prognostic indicators for patients with luminal BC is an important research goal.
Family with sequence similarity 234 member B (FAM234B), also known as KIAA1467, is a protein-coding gene located on chromosome 12. FAM234B serves as integral component of the membrane and is expressed in many cell types and organs, especially in brain tissues.10 Diseases associated with FAM234B include neurodevelopmental disorders and recurrent childhood high hyperdiploid acute lymphoblastic leukemia.11,12 Nevertheless, the clinical significance and underlying roles of FAM234B in solid tumors are not well studied. Thus, we analyzed the expression profile, prognostic value, biological functions, and molecular mechanisms of FAM234B in BC through the combination of bioinformatics analysis and experimental verification.
Materials and Methods
Expression Analysis Using a Bioinformatics Approach
The FAM234B mRNA expression profile was evaluated in samples of 20 cancer types and matched non-tumor samples using Oncomine (https://www.oncomine.org).13 P-values < 0.0001, fold change values of > 2.0, and genes ranking in the top 10% were set as thresholds. Then, FAM234B mRNA expression levels in BC were validated using data from TCGA portal (http://tumorsurvival.org/index.html),14 GEPIA (http://gepia.cancer-pku.cn/),15 TIMER (https://cistrome.shinyapps.io/timer/),16 and bc-GenExMiner (http://bcgenex.centregauducheau.fr/BC-GEM/)17 databases. Moreover, the protein expression of FAM234B in BC was explored using the Human Protein Atlas (HPA, https://www.proteinatlas.org/)18 and CPTAC data from the UALCAN (http://ualcan.path.uab.edu/analysis.html)19 database. In addition, mRNA expression level of HTR7P1 in BC were evaluated using GEPIA.15
Associations Between FAM234B or HTR7P1 Expression Levels and Clinicopathological Features of BC
The bc-GenExMiner (http://bcgenex.centregauducheau.fr/BC-GEM/)17 tool was applied to analyze the relationships between FAM234B or HTR7P1 mRNA expression levels and clinicopathological features of BC, including age, ER, PR, HER2, p53, Scarff-Bloom-Richardson (SBR) grades, Nottingham prognostic index (NPI), and molecular subtype. Additionally, we also analyzed the relationship between mRNA expression of FAM234B or HTR7P1 and tumor stages by GEPIA.15 The distribution of FAM234B or HTR7P1 mRNA expression levels across molecular subtypes was evaluated using UCSC Xena (http://xena.ucsc.edu/)20 and TCGA portal.14 Finally, CPTAC data from UALCAN19 was used to explore the protein expression of FAM234B in different subtypes.
Survival Analysis of FAM234B and HTR7P1
GEPIA,15 TIMER,16 and bc-GenExMiner21 databases were used to evaluate the effects of FAM234B mRNA expression levels on survival of patients with all BC and different molecular subtypes. The prognostic values of HTR7P1 in different subtypes were also evaluated using bc-GenExMiner21 database.
Co-Expressed Genes of FAM234B and Gene Set Enrichment Analysis
LinkedOmics (http://www.linkedomics.org/)22 was used for obtaining co-expression genes of FAM234B via the LinkFinder module and for performing gene set enrichment analysis (GSEA) via the LinkInterpreter module. A heat map of the top 50 positively or negatively correlated genes was generated. The correlation was evaluated by Spearman test.
Correlation Analysis of FAM234B with HTR7P1
Pearson’s pairwise correlation analysis of FAM234B with HTR7P1 in BC and different molecular subtypes was performed using bc-GenExMiner.17 Then, TIMER16 was used to verify the correlations based on Spearman correlation coefficients.
Prediction of Candidate MicroRNAs of FAM234B and HTR7P1
To better understand the molecular mechanism of HTR7P1 regulating FAM234B, we first explored the interaction molecules of HTR7P1 and FAM234B using RNAInter (http://www.rna-society.org/rnainter/home.html)23 database. Then, the subcellular localization of HTR7P1 was explored by its sequence extracted from the National Center for Biotechnology Information using IncLocator (http://www.csbio.sjtu.edu.cn/bioinf/lncLocator/).24 We applied miRanda (http://www.microrna.org/)25 and starBase (http://starbase.sysu.edu.cn/index.php)26 to determine potential microRNAs (miRNAs) binding to HTR7P1 3ʹ-UTR. Subsequently, potential binding miRNAs of FAM234B 3ʹ-UTR were predicted using TargetScanHuman7.2 (http://www.targetscan.org/),27 starBase,26 and miRWalk (http://mirwalk.umm.uni-heidelberg.de/).28 Then, we analyzed the potential miRNAs using a Venn diagram. The expression level of the candidate miRNA in BC was detected using dbDEMC2 (https://www.picb.ac.cn/dbDEMC/).29 The correlations of miRNAs with HTR7P1 and FAM234B were analyzed using starBase.26
Gathering of Clinical Samples and Ethics Statement
A total of 120 patients suffering from primary BC were recruited from the Second Affiliated Hospital of Xi’an Jiaotong University (Xi’an, China). All the patients were pathologically diagnosed, did not exhibit distant metastasis, and did not receive any other treatment before surgery. Samples were immersed in RNAlater reagent and further stored at −80°C. The clinicopathological parameters of patients were obtained from their medical records. The study was approved by the Ethics Committee of Xi’an Jiaotong University, and each patient gave an informed consent.
RNA Extraction and qRT-PCR Analysis
Total RNA was isolated from the tissue samples using Trizol reagent (Invitrogen, USA). Then, the RNA was reverse-transcribed into cDNA using PrimeScript TM RT reagent Kit (TAKARA, Japan) according to the manufacturer’s protocol. The levels of FAM234B and HTR7P1 were detected by quantitative reverse transcription-polymerase chain reaction (qRT-PCR) using the SYBR® Premix Ex Taq TM II kit (Takara, Japan) on ABI StepOne Real-Time PCR system. The primers for FAM234B, HTR7P1, and the internal control GAPDH are listed in Supplementary Table 1. The relative expression of FAM234B and HTR7P1 was quantified using 2−ΔCt values.
Statistical Analysis
The difference in the expression of FAM234B or HTR7P1 between two groups for our clinical samples was examined by Mann–Whitney test. The association of FAM234B or HTR7P1 expression with BC clinical characteristics was assessed by chi-square test. Spearman correlation coefficients were calculated to evaluate the association between FAM234B and HTR7P1. All the statistical analyses were conducted with IBM SPSS Statistics 18.0 software. P-value < 0.05 was considered statistically significant.
Results
High FAM234B Expression Indicated Poor Prognosis in Luminal BC Patients
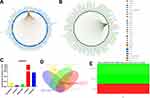
The expression profile of FAM234B in human cancer was first determined using microarray data by ONCOMINE database (Supplementary Figure 1A). FAM234B mRNA expression levels were highest in BC tissues in the Bittner multi-cancer dataset (Supplementary Figure 1B). Thus, we next focused on the role of FAM234B in BC. In the Curtis BC dataset, the mRNA level of FAM234B in BC tissues was highest in mucinous breast carcinoma, lowest in medullary breast carcinoma, and obviously higher than that in normal breast tissues (Supplementary Figure 1C and D, p < 0.001). Interestingly, compared with basal-like BC tissues, the FAM234B mRNA expression was significantly upregulated in luminal-like BC in the Farmer Breast 4 dataset (Supplementary Figure 1E, p < 0.001). We further examined FAM234B mRNA levels using RNA-seq data from TCGA via TCGA portal, GEPIA, TIMER, and bc-GenExMiner databases. As demonstrated in Figures 1A–D and 2A–C, consistent with the results of the ONCOMINE analysis, the mRNA levels of FAM234B were higher in luminal BC tissues and lower in basal-like BC tissues than that in normal breast tissues (p < 0.001). For further validation, we also detected the expression of FAM234B in clinical BC samples. As shown in Figure 1E, FAM234B expression in BC tissues was markedly higher than that in normal tissues (p = 0.027). Moreover, compared with adjacent-normal tissues, the mRNA levels of FAM234B were obviously upregulated in luminal BC tissues (Figure 2D, p = 0.0006) and downregulated in triple-negative BC (TNBC) tissues (Figure 2D, p = 0.0409). Using combination bioinformatics analysis and experimental validation, we demonstrated that FAM234B mRNA was significantly overexpressed in luminal BC. Next, we probed the protein expression pattern of FAM234B in BC using HPA and UALCAN. As depicted in Figure 3A, FAM234B was not expressed in normal breast tissues and was expressed at weak and moderate intensity in BC tissues, and located in cytoplasm. Similarly, a CPTAC analysis showed that the protein expression levels of FAM234B differed significantly between BC tissues and normal breast tissues (Figure 3B, p < 0.001). Moreover, CPTAC analysis also indicated that the protein expression of FAM234B was obviously higher in luminal BC than in TNBC (Figure 3C, p = 0.015). All evidence taken together, our results showed that FAM234B was upregulated at the mRNA and protein levels in luminal BC.
To better understand the clinical significance and function of FAM234B in BC, we assessed the relationship between FAM234B expression and clinical outcomes using GEPIA, TIMER, and bc-GenExMiner databases. First, the effect of FAM234B mRNA expression on survival in all BC patients was evaluated using GEPIA and TIMER databases. High FAM234B mRNA expression was significantly related to a worse OS (Figure 4A1 and B1, p < 0.05) in all BC patients. Subgroup analyses were subsequently performed to assess the prognostic value of FAM234B for different molecular subtypes. An analysis of TCGA data of GEPIA and TIMER revealed that increased FAM234B mRNA levels predicted worse survival in BC patients with luminal subtype (Figure 4A2 and B3, p < 0.05), but there was no significant correlation between FAM234B expression and prognosis in BC patients with HER2-enriched and basal-like BC (Figure 4A3, A4 and B2, B4). As determined using the bc-GenExMiner database, high FAM234B mRNA levels were obviously associated with a shorter OS in BC patients with luminal A and luminal B subtypes (Figure 4C1 and C2) but with a longer OS in BC patients with the HER2-enriched subtype (Figure 4C3). However, no significant correlation was found between FAM234B expression and prognosis in patients with basal-like BC (Figure 4C4).
Association of FAM234B Expression Levels with Clinicopathological Features of BC Patients
We further investigated the correlations between the expression level of FAM234B and BC clinicopathological parameters using bc-GenExMiner. FAM234B mRNA expression levels were positively related to age, with upregulated expression in the older group (Figure 5A and B, p < 0.0001). FAM234B mRNA expression levels were significantly higher in the ER-positive group and PR-positive group than in the corresponding negative groups (Figure 5C and D, p < 0.0001), and the opposite trend was observed for HER2 status (Figure 5E, p < 0.0001). Patients with BC harboring wild-type p53 displayed higher FAM234B levels than those of patients harboring mutant p53 (Figure 5F, p < 0.0001). Not surprisingly, compared with levels in TNBC and basal-like BC, the mRNA levels of FAM234B were clearly higher in the not-basal-like BC and non-TNBC group (Figure 5G, p < 0.0001). However, the SBR grade and NPI were negatively related to mRNA levels of FAM234B (Figure 5H and I, p < 0.0001). FAM234B mRNA expression was remarkably correlated with tumor stages, and the highest FAM234B mRNA expression was found in stage X (Figure 5J, p < 0.001).
Subsequently, we assessed the associations between FAM234B expression and BC clinicopathological parameters in clinical samples, including age, menopause status, tumor size, lymph node metastasis, TNM stage, ER status, PR status, HER2 status, and Ki67 expression level. BC patients were divided into two groups (high expression vs. low expression) using median value as the cut-off. As shown in Table 1 and Supplementary Figure 2, the increased expression of FAM234B was correlated with lymph node metastasis-positive, ER-positive, PR-positive, HER2-negative, and luminal subtype, whereas FAM234B did not show correlation with age, menopause status, tumor size, TNM stage, and Ki67 level.
 |
Table 1 Correlations of FAM234B Expression with Breast Cancer Clinical Features |
Co-Expressed Genes of FAM234B and Functional Enrichment Analysis
The identification of related genes will contribute to our understanding of the underlying roles of FAM234B in the development and progression of BC. We thus obtained the correlated genes of FAM234B by LinkFinder module using LinkedOmics database (Figure 6A). The heatmap of the top 50 positively and negatively correlated genes of FAM234B are presented in Figure 6B and C. Next, the gene set enrichment analysis (GSEA) of all significantly correlated genes was conducted via LinkInterpreter module. As shown in Figure 6D, we observed that FAM234B was positively related to the membrane transport and cytoskeleton formation process, such as peroxisome organization, microtubule-based movement, vesicle-mediated transport to the plasma membrane, cilium organization, peroxisomal transport, vacuolar transport, and so on, and negatively associated with immunity-related biological function, such as adaptive immune response, response to chemokine, interleukin-1 production, response to interferon-gamma, humoral immune response, lymphocyte-mediated immunity, natural killer cell and T cell activation, and so on.
HTR7P1 Was the Most Significantly Correlated with FAM234B, and High HTR7P1 Expression Predicted Poor Survival Rate in Luminal BC Patients
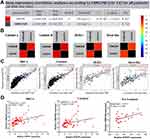
As depicted in Figure 6A, HTR7P1 (5-hydroxytryptamine receptor 7 pseudogene 1) had the most correlation with FAM234B. They were located on chr12p13.1, and HTR7P1 was located upstream of FAM234B (Figure 7A). Specially, a correlation analysis using bc-GenExMiner database revealed that correlation of FAM234B with HTR7P1 varied among various molecular subtypes, with higher correlation coefficient in luminal A and luminal B subtypes (Figure 7B and Supplementary Figure 3, p < 0.001). Similarly, we observed that the correlation between FAM234B and HTR7P1 was the most significant in luminal BC by TIMER database (Figure 7C, p < 0.001). We also found that HTR7P1 and FAM234B have a significant correlation in luminal subtype (p < 0.05), whereas no significant correlation was found in not-luminal subtype using clinical BC samples (Figure 7D, p > 0.05). Based on the above findings, we speculated that HTR7P1 expression in BC varied among molecular subtypes.
Not surprisingly, although HTR7P1 expression was not statistically different between BC and normal tissues (Supplementary Figure 4A and B, p > 0.05), its expression in luminal subtype was obviously higher than that in normal tissues (Figure 8A–D, p < 0.001). What is more, increased HTR7P1 mRNA level indicated worse prognosis in luminal A and luminal B subtypes (Figure 9A and B, p < 0.05), whereas it predicted better survival rate in HER2-enriched subtype (Figure 9C, p < 0.001). However, no significant correlation was found between HTR7P1 expression and prognosis in patients with basal-like BC (Figure 9D, p > 0.05). Subsequently, we analyzed associations of HTR7P1 mRNA expression with clinicopathological features of BC. The results from bc-GenExMiner database showed that HTR7P1 mRNA expression was obviously higher in older, ER-positive, PR-positive, HER2-negative, wild-type p53, not-basal-like and not-TNBC, low SBR grade, low NPI, and high-stage BC patients (Supplementary Figure 5, p < 0.05). Likewise, as shown in Table 2 and Supplementary Figure 6, the expression of HTR7P1 was correlated with ER status, PR status, and molecular subtype using clinical samples (p < 0.05).
 |
Table 2 Correlations of HTR7P1 Expression with Breast Cancer Clinical Features |
HTR7P1 Functions as an Competing Endogenous RNA of has-miR-1271-5p or has-miR-381-3p to Upregulate FAM234B
The pseudogene HTR7P1 acts as a special long non-coding RNA (lncRNA) and is located upstream of FAM234B. According to the above results, high HTR7P1 and FAM234B expression indicated poor prognosis in luminal BC. We thus assumed that HTR7P1 may cis-regulate the expression of FAM234B. The interaction network of the top 100 interactions of HTR7P1 and FAM234B are shown in Figure 10A and B. Subcellular localization of pseudogene HTR7P1 determined the underlying mechanisms. lncLocator predicted that HTR7P1 was mainly located in the cytosol but also distributed in the exosome (Figure 10C), which suggested that HTR7P1 regulated FAM234B expression more likely in a competing endogenous RNA (ceRNA) way. As shown in Figure 10D, after taking the intersection of the prediction results from four databases, there were three candidate miRNAs (has-miR-1271-5p, has-miR-381-3p, and has-miR-330). We further analyzed their expression in BC using dbDEMC. Compared with normal tissues, has-miR-1271-5p and has-miR-381-3p were downregulated (Figure 10E and Supplementary Table 2, p < 0.01), whereas has-miR-330 was upregulated in BC tissues (Figure 10E). Correlation analysis using starBase also confirmed a significantly negative correlation of expression of has-miR-1271-5p or has-miR-381-3p with HTR7P1 and FAM234B (Supplementary Figure 7A–D). These results demonstrated that HTR7P1 might serve as ceRNA to cis-regulate FAM234B expression by sponging has-miR-1271-5p or has-miR-381-3p in luminal BC.
Discussion
BC is a heterogeneous disease with distinct histological, molecular, and clinical phenotypes.30 Molecular stratification based on gene expression profiles has shown that BC could be divided into so-called intrinsic subtypes (luminal A and B, HER2-enriched, and basal-like).31 These molecular subtypes differ clearly with respect to incidence, prognosis, and response to treatments. Over 70% of BC cases belong to the two luminal categories. Patients with luminal subtypes often respond to “endocrine” therapies targeting ER.32 Unfortunately, many patients show intrinsic or acquired resistance to these therapies over time.33 In fact, more than 30% of ER-positive cancers do not respond to endocrine therapy, exhibiting intrinsic resistance. Another 30–40% of tumors initially respond to endocrine therapy and eventually develop resistance.34 Therefore, the current focus of research is the identification of new molecular targets or adjuvant endocrine therapies to overcome resistance in patients with luminal BC.
Several luminal subtype-specific genes have been identified, including NUP43,35 FANCM,36 SYCP2,37 and NCOA5.38 FAM234B is a recently identified gene in the FAM234 family. Previous studies have reported that it is mainly expressed in the nervous system.11 However, few studies have evaluated the role of FAM234B in human cancers. Results of bioinformatics analysis showed that FAM234B mRNA expression is highest in BC tissues among various tumor types, and its mRNA and protein levels are higher in BC tissues than in matched normal tissues, which was also verified by qRT-PCR for clinical samples. In particular, compared with normal breast tissues, the expression of FAM234B was increased in luminal BC but decreased in basal-like BC. qRT-PCR for clinical samples also demonstrated a high expression level of FAM234B in luminal BC tissues when compared with normal tissues. Subsequently, survival analysis indicated that high FAM234B levels are associated with a poor prognosis in patients with luminal BC. Positive associations between FAM234B expression and age, ER status, PR status, and tumor stage were found. Conversely, HER2 status, p53 status, NPI, SBR grade, basal-like status, and TNBC status were negatively related to FAM234B expression. Likewise, we also found that high mRNA expression of FAM234B was correlated with lymph node metastasis, ER-positive, PR-positive, HER2-negative, and luminal subtype by analyzing the qRT-PCR for clinical samples. Different from the results of bioinformatic analysis, no significant correlation between FAM234B expression and age and stage was found in our experimental verification using clinical samples. This may be due to the small sample size and the selection bias of the cohort of the present study. Additionally, FAM234B did not show correlation with menopause status, tumor size, and Ki67 level. These findings suggested that FAM234B functions as an oncogene and overexpression of FAM234B may be a promising prognostic biomarker in luminal BC.
Co-expression analysis facilitates understanding of the biological functions and molecular mechanisms of FAM234B; we obtained the positively and negatively correlated genes of FAM234B using Linkedomics database. Among them, genes in the ER-alpha pathway (ESR1, GATA3, FOXA1, and XBP1) were significantly positively correlated with FAM234B expression,39–41 which can explain the high expression of FAM234B in luminal subtypes. Estrogen is a key regulator of mammary gland development, breast carcinogenesis, and progression.42,43 The ER signaling pathway is the most promising target to date for the clinical treatment of ER-positive BC, and ERs are ligand-dependent transcription factors that regulate genes involved in cell proliferation, differentiation, apoptosis, and cell migration.44 The dysregulation of these signaling pathways can lead to the initiation, progression, and invasion of breast tumors.45 Therefore, we hypothesized that FAM234B is directly related to ER-mediated pathways and functions in luminal BC via these genes. Besides, GSEA showed that these positively correlated genes were mainly enriched in the membrane transport process, such as peroxisome organization, microtubule-based movement, vesicle-mediated transport to the plasma membrane, cilium organization, peroxisomal transport, vacuolar transport, and so on. We found that FAM234B was a single-pass membrane protein and was located in golgi apparatus, cytoplasm, cytoskeleton, microtubule organizing center, and membrane using UniProt (https://www.uniprot.org/uniprot/) database, which can explain that FAM234B was positively correlated with membrane transport process. Specially, GSEA showed that these negatively associated genes with FAM234B were mainly enriched in immunity-related biological functions, such as adaptive immune response, response to chemokine, interleukin-1 production, response to interferon-gamma, humoral immune response, lymphocyte-mediated immunity, natural killer cell and T cell activation, and so on. A large-scale quantitative phosphoproteomic analysis of TCR signaling identified that FAM234B protein phosphorylation was associated with T cell activation in human T cell leukemia.46 At present, there are limited research reports on the correlation of FAM234B with immune response. Therefore, the relationship between FAM234B expression and immunity needs further research.
What makes FAM234B upregulated in luminal BC? Co-expression analysis also provided critical clues for investigating the regulatory mechanisms of FAM234B and showed that HTR7P1 was most significantly correlated with FAM234B. Moreover, correlation analysis suggested that, among various BC subtypes, FAM234B expression was the most significantly correlated with HTR7P1 in luminal BC, which was also verified by qT-PCR using clinical samples. Simultaneously, both bioinformatics analysis and experimental verification have confirmed that HTR7P1 expression was higher in luminal BC tissues than that in normal breast tissues. What is more, overexpression of HTR7P1 indicated shorter survival in luminal BC patients. HTR7P1 is a pseudogene that has not been reported in human tumors. Pseudogenes have long been considered junk genes. However, in recent years, more and more evidence has shown that pseudogenes play a key role in physiological and pathological processes in human diseases.47 The dysregulation of pseudogenes also have a lot to do with tumor.48,49 Pseudogenes, as a kind of lncRNA, serve as ceRNA for miRNAs or interacting with RNA-binding proteins to cis-regulate or trans-regulate the expression of their parent genes or other protein-coding genes, which depend on the subcellular localization of pseudogenes.50–52 The prediction of subcellular localization of HTR7P1 showed that it was mainly distributed in the cytosol, using IncLocator database. Therefore, we speculated that HTR7P1 might be likely to upregulate the FAM234B expression through the ceRNA mechanism. Further exploration found that has-miR-1271-5p or has-miR-381-3p might be candidate miRNAs competing with HTR7P1 for binding to FAM234B. We also observed a significantly negative correlation between the has-miR-1271-5p or has-miR-381-3p and HTR7P1 or FAM234B in BC samples.
In summary, the overexpressions of FAM234B and HTR7P1 predict a poor prognosis in patients with luminal BC. Pseudogene HTR7P1 might cis-regulate the expression of FAM234B via competing with has-miR-1271-5p or has-miR-381-3p for binding to FAM234B in luminal BC. Besides, GSEA indicated that FAM234B was positively related to the membrane transport and cytoskeleton formation process and negatively associated with immune response biological function. However, the biological functions and molecular mechanisms of FAM234B in luminal BC still require laboratory research support.
Abbreviations
BC, breast cancer; TNBC, triple-negative breast cancer; ER, estrogen receptor; PR, progesterone receptor; OS, overall survival; HER2, human epidermal growth factor receptor 2; SBR, Scarff-Bloom-Richardson; NPI, Nottingham prognostic index; GSEA, gene set enrichment analysis; FAM234B, family with sequence similarity 234 member B; HTR7P1, 5-hydroxytryptamine receptor 7 pseudogene 1.
Data Sharing Statement
The data that support the findings of this study are available and derived from the available resource in the public domain.
Original Publication Statement
We declare that there are no prior or duplicate publication or submission elsewhere of any part of the work.
Acknowledgments
We thank all members of our study team for their whole-hearted cooperation.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising, or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
The authors would like to acknowledge financial support for this work from National Natural Science Foundation of China, Grant/Award Number: 81471670.
Disclosure
The authors declare that there are no competing interests associated with the manuscript.
References
1. Harbeck N, Gnant M. Breast cancer. Lancet. 2017;389(10074):1134–1150. doi:10.1016/S0140-6736(16)31891-8
2. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
3. DeSantis CE, Ma J, Goding Sauer A, Newman LA, Jemal A. Breast cancer statistics, 2017, racial disparity in mortality by state. CA Cancer J Clin. 2017;67(6):439–448.
4. Waks AG, Winer EP. Breast cancer treatment: a review. JAMA. 2019;321(3):288–300. doi:10.1001/jama.2018.19323
5. Prat A, Karginova O, Parker JS, et al. Characterization of cell lines derived from breast cancers and normal mammary tissues for the study of the intrinsic molecular subtypes. Breast Cancer Res Treat. 2013;142(2):237–255. doi:10.1007/s10549-013-2743-3
6. Ignatiadis M, Sotiriou C. Luminal breast cancer: from biology to treatment. Nat Rev Clin Oncol. 2013;10(9):494–506. doi:10.1038/nrclinonc.2013.124
7. Alfarsi L, Johnston S, Liu DX, Rakha E, Green AR. Current issues with luminal subtype classification in terms of prediction of benefit from endocrine therapy in early breast cancer. Histopathology. 2018;73(4):545–558.
8. Li N, Deng Y, Zhou L, et al. Global burden of breast cancer and attributable risk factors in 195 countries and territories, from 1990 to 2017: results from the Global Burden of Disease Study 2017. J Hematol Oncol. 2019;12(1):140. doi:10.1186/s13045-019-0828-0
9. Yang J, Li Y, Liu Q, et al. Brief introduction of medical database and data mining technology in big data era. J Evid Based Med. 2020;13(1):57–69. doi:10.1111/jebm.12373
10. Nagase T, Kikuno R, Ishikawa K, Hirosawa M, Ohara O. Prediction of the coding sequences of unidentified human genes. XVII. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. DNA Res. 2000;7(2):143–150. doi:10.1093/dnares/7.2.143
11. Reuter MS, Tawamie H, Buchert R, et al. Diagnostic yield and novel candidate genes by exome sequencing in 152 consanguineous families with neurodevelopmental disorders. JAMA Psychiat. 2017;74(3):293–299. doi:10.1001/jamapsychiatry.2016.3798
12. Chen C, Bartenhagen C, Gombert M, et al. Next-generation-sequencing of recurrent childhood high hyperdiploid acute lymphoblastic leukemia reveals mutations typically associated with high risk patients. Leuk Res. 2015;39(9):990–1001. doi:10.1016/j.leukres.2015.06.005
13. Rhodes DR, Yu J, Shanker K, et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004;6(1):1–6. doi:10.1016/S1476-5586(04)80047-2
14. Xu S, Feng Y, Zhao S. Proteins with evolutionarily hypervariable domains are associated with immune response and better survival of basal-like breast cancer patients. Comput Struct Biotechnol J. 2019;17:430–440. doi:10.1016/j.csbj.2019.03.008
15. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–W102. doi:10.1093/nar/gkx247
16. Li T, Fan J, Wang B, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77(21):e108–e10. doi:10.1158/0008-5472.CAN-17-0307
17. Jezequel P, Frenel JS, Campion L, et al. bc-GenExMiner 3.0: new mining module computes breast cancer gene expression correlation analyses. Database. 2013;2013:bas060. doi:10.1093/database/bas060
18. Ponten F, Jirstrom K, Uhlen M. The human protein atlas–a tool for pathology. J Pathol. 2008;216(4):387–393. doi:10.1002/path.2440
19. Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–658. doi:10.1016/j.neo.2017.05.002
20. Goldman M, Craft B, Hastie M, et al. The UCSC xena platform for public and private cancer genomics data visualization and interpretation. bioRxiv. 2019;326470.
21. Jezequel P, Campone M, Gouraud W, et al. bc-GenExMiner: an easy-to-use online platform for gene prognostic analyses in breast cancer. Breast Cancer Res Treat. 2012;131(3):765–775. doi:10.1007/s10549-011-1457-7
22. Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2017;46(D1):D956–D963. doi:10.1093/nar/gkx1090
23. Lin Y, Liu T, Cui T, et al. RNAInter in 2020: RNA interactome repository with increased coverage and annotation. Nucleic Acids Res. 2020;48(D1):D189–D197. doi:10.1093/nar/gkz804
24. Cao Z, Pan X, Yang Y, Huang Y, Shen HB. The lncLocator: a subcellular localization predictor for long non-coding RNAs based on a stacked ensemble classifier. Bioinformatics. 2018;34(13):2185–2194. doi:10.1093/bioinformatics/bty085
25. Betel D, Wilson M, Gabow A, Marks DS, Sander C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 2008;36(Database issue):D149–D153. doi:10.1093/nar/gkm995
26. Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42(Database issue):D92–D97. doi:10.1093/nar/gkt1248
27. Agarwal V, Bell GW, Nam JW, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs. eLife. 2015;4. doi:10.7554/eLife.05005
28. Sticht C, De La Torre C, Parveen A, Gretz N. miRWalk: an online resource for prediction of microRNA binding sites. PLoS One. 2018;13(10):e0206239. doi:10.1371/journal.pone.0206239
29. Yang Z, Wu L, Wang A, et al. dbDEMC 2.0: updated database of differentially expressed miRNAs in human cancers. Nucleic Acids Res. 2017;45(D1):D812–D818. doi:10.1093/nar/gkw1079
30. Li J, Chen Z, Su K, Zeng J. Clinicopathological classification and traditional prognostic indicators of breast cancer. Int J Clin Exp Pathol. 2015;8(7):8500–8505.
31. Russnes HG, Lingjaerde OC, Borresen-Dale AL, Caldas C. Breast cancer molecular stratification: from intrinsic subtypes to integrative clusters. Am J Pathol. 2017;187(10):2152–2162. doi:10.1016/j.ajpath.2017.04.022
32. Abdel-Hafiz HA, Horwitz KB. Role of epigenetic modifications in luminal breast cancer. Epigenomics. 2015;7(5):847–862. doi:10.2217/epi.15.10
33. Badia E, Oliva J, Balaguer P, Cavailles V. Tamoxifen resistance and epigenetic modifications in breast cancer cell lines. Curr Med Chem. 2007;14(28):3035–3045. doi:10.2174/092986707782794023
34. Milani A, Geuna E, Mittica G, Valabrega G. Overcoming endocrine resistance in metastatic breast cancer: current evidence and future directions. World J Clin Oncol. 2014;5(5):990–1001. doi:10.5306/wjco.v5.i5.990
35. Tian C, Zhou S, Yi C. High NUP43 expression might independently predict poor overall survival in luminal A and in HER2+ breast cancer. Future Oncol. 2018;14(15):1431–1442. doi:10.2217/fon-2017-0690
36. Wang Y, Wang J, Long F, et al. Correlation of FANCM expression with clinical factors in luminal B breast cancer. Breast Cancer. 2018;25(4):431–437. doi:10.1007/s12282-018-0841-0
37. Wu C, Tuo Y. SYCP2 expression is a novel prognostic biomarker in luminal A/B breast cancer. Future Oncol. 2019;15(8):817–826. doi:10.2217/fon-2018-0821
38. Ye XH, Huang DP, Luo RC. NCOA5 is correlated with progression and prognosis in luminal breast cancer. Biochem Biophys Res Commun. 2017;482(2):253–256. doi:10.1016/j.bbrc.2016.11.051
39. Lacroix M, Leclercq G. About GATA3, HNF3A, and XBP1, three genes co-expressed with the oestrogen receptor-alpha gene (ESR1) in breast cancer. Mol Cell Endocrinol. 2004;219(1–2):1–7. doi:10.1016/j.mce.2004.02.021
40. Andres SA, Wittliff JL. Relationships of ESR1 and XBP1 expression in human breast carcinoma and stromal cells isolated by laser capture microdissection compared to intact breast cancer tissue. Endocrine. 2011;40(2):212–221. doi:10.1007/s12020-011-9522-x
41. Andres SA, Wittliff JL. Co-expression of genes with estrogen receptor-alpha and progesterone receptor in human breast carcinoma tissue. Horm Mol Biol Clin Investig. 2012;12(1):377–390. doi:10.1515/hmbci-2012-0025
42. Zhou Z, Qiao JX, Shetty A, et al. Regulation of estrogen receptor signaling in breast carcinogenesis and breast cancer therapy. Cell Mol Life Sci. 2014;71(8):1549.
43. Yager JD, Davidson NE. Estrogen carcinogenesis in breast cancer. N Engl J Med. 2006;354(3):270–282. doi:10.1056/NEJMra050776
44. Brisken C, O’Malley B. Hormone action in the mammary gland. Cold Spring Harb Perspect Biol. 2010;2(12):a003178. doi:10.1101/cshperspect.a003178
45. Harrell JC, Dye WW, Allred DC, et al. Estrogen receptor positive breast cancer metastasis: altered hormonal sensitivity and tumor aggressiveness in lymphatic vessels and lymph nodes. Cancer Res. 2006;66(18):9308–9315. doi:10.1158/0008-5472.CAN-06-1769
46. Mayya V, Lundgren DH, Hwang SI, et al. Quantitative phosphoproteomic analysis of T cell receptor signaling reveals system-wide modulation of protein-protein interactions. Sci Signal. 2009;2(84):ra46. doi:10.1126/scisignal.2000007
47. Pink RC, Wicks K, Caley DP, Punch EK, Jacobs L, Carter DR. Pseudogenes: pseudo-functional or key regulators in health and disease? RNA. 2011;17(5):792–798. doi:10.1261/rna.2658311
48. Hu X, Yang L, Mo YY. Role of Pseudogenes in tumorigenesis. Cancers. 2018;10(8). doi:10.3390/cancers10080256
49. Poliseno L, Marranci A, Pandolfi PP. Pseudogenes in human cancer. Front Med. 2015;2:68. doi:10.3389/fmed.2015.00068
50. Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465(7301):1033–1038. doi:10.1038/nature09144
51. An Y, Furber KL, Ji S. Pseudogenes regulate parental gene expression via ceRNA network. J Cell Mol Med. 2017;21(1):185–192. doi:10.1111/jcmm.12952
52. Tutar L, Ozgur A, Tutar Y. Involvement of miRNAs and pseudogenes in cancer. Methods Mol Biol. 2018;1699:45–66.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.