Back to Journals » OncoTargets and Therapy » Volume 10
Niclosamide inhibition of STAT3 synergizes with erlotinib in human colon cancer
Authors Shi L, Zheng H, Hu W, Zhou B, Dai X, Zhang Y, Liu Z, Wu X, Zhao C , Liang G
Received 5 December 2016
Accepted for publication 20 January 2017
Published 23 March 2017 Volume 2017:10 Pages 1767—1776
DOI https://doi.org/10.2147/OTT.S129449
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr XuYu Yang
Lingyi Shi,1,* Hailun Zheng,1,* Wanle Hu,2 Bin Zhou,2 Xuanxuan Dai,3 Yi Zhang,3 Zhiguo Liu,1 Xiaoping Wu,1 Chengguang Zhao,1 Guang Liang1
1Chemical Biology Research Center, School of Pharmaceutical Sciences, 2Department of Coloproctology, The Second Affiliated Hospital and Yuying Children’s Hospital, 3Department of Oncological Surgery, The First Affiliated Hospital, Wenzhou Medical University, Wenzhou, People’s Republic of China
*These authors contributed equally to this work
Abstract: Niclosamide, an anthelmintic drug approved by the US Food and Drug Administration against cestodes, is used to treat tapeworm infection. In this study, we show that niclosamide can potentially inhibit signal transducer and activator of transcription 3 (STAT3) in colon cancer cell lines. Combined inhibition of epidermal growth factor receptor and STAT3 by erlotinib and niclosamide synergistically induces apoptosis and antiproliferation in colon cancer cell lines. Our findings suggest that erlotinib and niclosamide combination provides an effective therapeutic approach to improving the prognosis of colon cancer.
Keywords: STAT3, EGFR, erlotinib, niclosamide, colon cancer
Introduction
Colon cancer is the third most common cancer worldwide and the second leading cause of cancer death in the US.1 The mortality rate of patients with colon cancer has decreased owing to improved detection, but the incidence of colon cancer continues to increase.2 Considerable progress in the development of chemotherapy for advanced colon cancer has been observed in the recent decade.3 The epidermal growth factor receptor (EGFR) has been reported as a critical therapeutic target for the treatment of colon cancer.4 However, an anti-EGFR agent used to treat colon cancer has exhibited moderate efficacy.5 The mechanisms of this resistance have yet to be elucidated. Nevertheless, the potential of EGFR inhibitors to induce activation of the EGFR-independent pathway is under consideration.6,7
Signal transducer and activator of transcription (STAT) proteins significantly influence diverse biological processes, including cell proliferation, differentiation, survival, and inflammatory response.8,9 One of the well-studied STAT members, STAT3, has been shown to be an important molecule in various malignancies and verified as an effective target for cancer therapy, including endometrial, cervical, breast, brain, prostate, and colon cancer.10–13 A number of studies have indicated that aberrant STAT3 activation contributes to cell proliferation, differentiation, migration, and survival.14,15
Niclosamide, particularly effective against cestodes, has been used to treat tapeworm infection for approximately 50 years.16 Finding a new use for traditional medicines is considerably easier than inventing a new drug because traditional medicines have known pharmacokinetics and have often been approved for human treatments.17 In the past 5 years, niclosamide has been identified as a potential anticancer agent targeting multiple signaling pathways (eg, NF-κB, ROS, Notch, Wnt/b-catenin, and mTORc1).18–21 Recently, several studies have reported that niclosamide exhibits antiproliferative activity in head and neck, ovarian, breast, and hematologic cancer.22–25 Till date, no studies demonstrating the therapeutic efficacy of niclosamide in colon cancer are available. Taking this background into account, the present study aimed to investigate the effects of niclosamide on colon cancer and the molecular mechanisms underlying its therapeutic action.
Our outcomes demonstrated that niclosamide acted as a potent STAT3 inhibitor in colon cancer cell lines. The combination of niclosamide and erlotinib induced apoptosis and antiproliferation in colon cancer cell lines as well as sensitization of colon cancer cells to erlotinib. On the basis of our findings, we suggest that combination of niclosamide and erlotinib can potentially improve the prognosis of colon cancer.
Materials and reagents
Cell culture and antibodies
Human colon cancer cell lines (HCT116, SW620, and HT29) were obtained from Cell Resources center of the Shanghai Institutes for Biological Sciences (Chinese Academy of Sciences, Shanghai, People’s Republic of China). The cells were routinely cultured in RPMI-1640 (Gibco/BRL; Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% fetal bovine serum (Hyclone, Logan, UT, USA) and 1% of antibiotic solution (100 units/mL penicillin and 100 μg/mL streptomycin) in a humidified atmosphere of 5% CO2 at 37°C.
Antibodies such as anti-BCL-2, anti-Bax, anti-cleaved PARP, anti-GAPDH, horseradish peroxidase (HRP)-conjugated goat anti-mouse IgG, and HRP-conjugated donkey anti-rabbit IgG were purchased from Santa Cruz Biotechnology (Dallas, TX, USA). Antibodies against Phospho-STAT3 and STAT3 were purchased from Cell Signaling Technology (Danvers, MA, USA).
Inhibitor drugs
Nifuroxazide, niclosamide, cryptotanshinone, and alantolactone were purchased from Selleck Chemicals (Houston, TX, USA). Nifuroxazide is an oral nitrofuran antibiotic not currently approved for use in the USA but is used elsewhere as an antidiarrheal agent.26 Niclosamide has been used to treat tapeworm infection and also as a molluscicide for water treatment in schistosomiasis control programs.27 Cryptotanshinone, a major lipophilic component isolated from Salvia miltiorrhiza Bunge, has been shown to possess chemotherapeutic properties against various types of cancer cells.28 Alantolactone is used as an insect repellent, antibacterial, and anticancer agent.29 The drug compounds were dissolved in sterile dimethyl sulfoxide (DMSO) to produce a 20 mM stock solution, which was then stored at −20°C and further diluted freshly with cell culture medium.
MTT assay
Cells were seeded into wells of a 96-well plate at 5×103 cells per well in 100 μL of the corresponding medium. The cells were then treated with drugs at different concentrations for 72 h. Subsequently, they were treated with a fresh solution of MTT (5 mg/mL) for 4 h at 37°C. The purple formazan crystals were finally solubilized with DMSO solution, and absorbance was recorded using a multi-well plate reader at 490 nm.
Western blot analysis
Cells were lysed in a lysis buffer containing a phosphatase inhibitor, and the lysates were clarified by centrifugation (12,000 rpm) at 4°C for 10 min. The supernatant was run on 10% and 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred to a polyvinylidene fluoride membrane. After being blocked with 5% nonfat dry milk in Tris Buffer Solution Tween for 1.5 h, membranes were incubated with a specific primary antibody of 1:1,000 dilution overnight and a HRP-conjugated secondary antibody of 1:3,000 dilution for 1 h. Immunoreactive bands were visualized using enhanced chemiluminescence reagent.
Molecular docking
As one of the most widely used computational approaches for structure-based drug design, molecular docking study was used to predict the binding pose of compound in STAT3 SH2-binding site by using the software AutoDock (version 4.2.6).30 The crystallographic coordinate for human STAT3 SH2 (Protein Data Bank [PDB] ID: 1BG1) was obtained from the PDB.31 Prior to docking, protein structures were prepared by removing water molecules and other ligands using PyMol software.32 A grid box size of 60×60×60 dimensions with a spacing of 0.375 Å between the grid points was implemented and covered almost the entire SH2-binding site. The grid parameter files were created setting up the map files directly. The Lamarckian genetic algorithm was applied to deal with the interactions of protein and inhibitors. The number of individuals in population was set to 300, and trials of 100 dockings and maximum number of energy evaluations were set as default along with other settings. AutoDockTools version 1.5.6 and PyMol were used to analyze the docking results.
Clonogenic assay
A total of 500 cells per well were seeded into a 6-well plate with 2 mL of RPMI-1640 and incubated overnight. The cells were then pretreated with nifuroxazide and erlotinib or DMSO for 8–12 h. After treatment, the cells were washed with phosphate buffer saline (PBS) twice and transferred to a fresh medium to grow for 7 days. Colonies were washed with PBS and then fixed with 4% methanol for 15 min at room temperature. The cells were washed with PBS twice and stained with 1% crystal violet (25% methanol) for 10 min at room temperature. Each experiment was conducted thrice.
Analysis of cell apoptosis
Cells (3×105) were seeded in 6-well plates and incubated overnight and then treated with nifuroxazide and erlotinib for 24 h. After treatment, the cells were harvested with trypsin and then washed with cold PBS twice. The cells were stained with Annexin V for 10 min under dark conditions and then with propidium iodide (PI) for 5 min. Apoptotic cells were counted using the FACS Calibur flow cytometer and quantified by flow cytometric analysis.
Statistical analyses
Data are represented as mean ± standard error of the mean of 3 independent experiments. Student’s t-test was performed to determine the statistical significance between 2 groups by using GraphPad Prism 6.0 (GraphPad Software, Inc., La Jolla, CA, USA). Differences between groups were analyzed by the log-rank test using GraphPad Prism 6.0. P<0.05 was considered statistically significant.
Results
Antiproliferative effects of niclosamide in human colon cancer cells
Nifuroxazide acts as a potent inhibitor of STAT3 signaling pathway in breast cancer cells, though it has little effect on cells lacking STAT3 activation.33 Niclosamide has recently been identified to target multiple signaling pathways (eg, NF-κB, ROS, Notch, and STAT3).16 Cryptotanshinone has previously been observed to possess the most powerful antibacterial, anti-inflammatory, and antitumor effect.34 Alantolactone has an inhibitory effect on cancer cells migration, invasion, adhesion, and colony formation.35 Therefore, we screened the antiproliferative effects of these 4 compounds in human colon cancer cells by MTT assay.
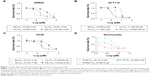
After the cells were treated with nifuroxazide, niclosamide, cryptotanshinone, and alantolactone for 72 h, MTT assay was employed. As shown in Figure 1, administration of these 4 compounds in a dose-dependent manner reduced viability in SW620 (Figure 1A), HCT116 (Figure 1B), and HT29 cells (Figure 1C). Among these compounds, niclosamide exhibited the most potent antiproliferative effect against all tested human colon cancer cell lines. After exposure of the cells to niclosamide for 72 h, the half maximal inhibitory concentration (IC50) in SW620, HCT116, and HT29 cell lines was found to be 2.9, 0.4, and 8.1 μM, respectively (Figure 1D). This suggests that niclosamide can inhibit the proliferation of human colon cancer cells.
Molecular docking between STAT3 and niclosamide
STAT3, a member of the STAT family of transcription factors, has recently been verified as an attractive therapeutic target and shown to be an important molecule in cancer therapy, including colon cancer.36 As the STAT3 SH2 domain is critical for the activation and biological function of STAT3,37 we investigated interactions between the compound and the SH2 domain of STAT3 by molecular docking analysis of the active compound niclosamide (Figure 2A). We applied PyMol software to remove water molecules and other ligands to prepare the STAT3 SH2 domain protein structures and used the software AutoDock (version 4.2.6) to predict the binding pose of compound in STAT3 SH2-binding site to identify whether compound niclosamide acts as a STAT3 inhibitor that targets the STAT3 SH2 domain.38 The molecular docking results indicated that niclosamide formed 4 hydrogen bonds with the side chain of ARG-595, SER-636, GLU-594, and LYS-591 as shown in Figure 2B. It was predicted that niclosamide could fit into the 2 major binding sites, the pTyr705 and the side pocket site, and so it could inhibit STAT3 phosphorylation. In summary, critical hydrogen bond interaction between niclosamide and STAT3 SH2 domain were predicted by a rational method combined with molecular docking. These methods clarified the interaction of niclosamide with STAT3. Further verification of co-crystallization of STAT3 and niclosamide is in progress.
Inhibition of STAT3 phosphorylation by niclosamide in colon cancer cells HCT116 and SW620
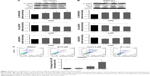
We performed Western blot analysis on colon cancer cell lines to examine whether niclosamide could inhibit STAT3. Western blot analysis indicated that niclosamide inhibited STAT3 phosphorylation (Figure 3). Notably, niclosamide inhibited STAT3 phosphorylation in a time- and dose-dependent manner in HCT116 and SW620 cell lines.
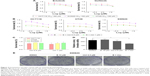
Sensitization of colon cancer cells to erlotinib by treatment of cells with niclosamide
Erlotinib or gefitinib, EGFR tyrosine kinase inhibitors, have been shown to benefit patients with non-small-cell lung cancer and pancreatic cancer. However, almost all patients develop a progressive disease during therapy. As shown in Figure 4A, after treatment of cells with erlotinib and gefitinib for 72 h, the IC50 in SW620, HCT116, and HT29 cells exceeded 60 μM. These results indicate that erlotinib and gefitinib could not inhibit the proliferation of SW620, HCT116, and HT29 cell lines. We thus examined the antiproliferative effects of erlotinib as well as erlotinib combined with niclosamide by MTT assay. As shown in Figure 4B, treatment of cells with niclosamide sensitized colon cancer cells to erlotinib, while niclosamide had no effect on SW620, HCT116, and HT29 cells at 0.5 μM and 1 μM concentrations (Figure 4C). To verify the anticancer properties of the combination, we performed colony-forming experiments. SW620 cells (5000) were seeded in a 6-well plate overnight and then treated with erlotinib (10 μM) or/and niclosamide (2.5 μM) for 1 week. We then removed the drugs and washed the cells with PBS twice. The cell colonies were stained with crystal violet and then counted. The combination of niclosamide and erlotinib suppressed the colony formation of colon cancer cells, as shown in Figure 4D and E. Colony formation assay demonstrated that the synergistic effect of niclosamide and erlotinib efficiently inhibits the growth of colon cancer cells.
Induction of apoptosis in colon cancer cells with combined niclosamide and erlotinib
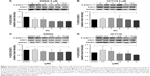
Compared with single-agent treatment, the combination of erlotinib and niclosamide enhanced the inhibition of cell viability in all tested cell lines (Figure 4B). We sought to investigate whether the combination of erlotinib and niclosamide could also induce cell apoptosis. We first employed Western blot analysis. HCT116 and SW620 cells were treated with erlotinib and niclosamide alone, or with both for 24 h. The combination significantly increased cleaved PARP and BAX, as shown in Figure 5A and B. BCL-2 expression was markedly decreased.
We also used flow cytometry to examine whether the combination induced cell apoptosis in SW620 cells. SW620 cells were treated with erlotinib (10 μM) or/and niclosamide (2.5 μM) for 24 h and then stained with Annexin V and PI for apoptosis analysis. The combination resulted in a significant increase in early (Annexin V+, PI−) and late apoptotic cells (Annexin V+, PI+), as shown in Figure 5C and D.
Discussion
Colon cancer is one of the most prevalent tumors worldwide.39 Despite conventional therapies such as surgery, radiation, and chemotherapy, the overall survival rate for colon cancer has not significantly improved.40 The human EGFR is overexpressed in various gastrointestinal cancer types, including 60%–80% of colon cancers, correlating with poor prognosis and early disease progression.41 In addition, cancer cell also upregulates other important downstream genes such as STAT3, which contributes to cell proliferation, cell survival, and angiogenesis in colon cancer.42 Therefore, more effective approaches need to be developed to improve the prognosis of colon cancer.
Here, our results showed that niclosamide reduces cell viability in a dose-dependent manner. This finding suggests that niclosamide could be effective against colon cancer. Several groups have independently revealed the activity of niclosamide against cancer cells;16,27,43 however, the precise mechanism underlying this antitumor activity has yet to be elucidated. Therefore, we first used molecular docking study to verify whether there is interaction between niclosamide and STAT3. In our study, we found critical hydrogen bond interaction between niclosamide and STAT3 SH2 domain. We further investigated the correlation between niclosamide and STAT3 in SW620 and HCT116 cell lines, and the results showed that niclosamide suppressed STAT3 phosphorylation in a time- and dose-dependent manner. Several studies have revealed that inhibition of EGFR by erlotinib induces activation of STAT3, which contributes to erlotinib resistance.44,45 Our data showed that when erlotinib and niclosamide were combined, they synergistically induced apoptosis of colon cancer cells. It has been reported that multiple signal transduction pathway inhibitors, including the mitogen-activated protein kinase, fibroblast growth factor receptor inhibitors, and chemotherapy drugs, can induce activation of STAT3 survival signaling pathway, leading to resistance.38,46 It is possible that, in addition to erlotinib, niclosamide may also sensitize these target drugs. Based on our findings, we conclude that niclosamide may represent a novel and more effective drug for colon cancer.
Drug development process from initial lead discovery to final medication is costly, lengthy, and incremental.33 Finding new uses for old or natural products is much easier and more economical than inventing a new drug from scratch. The present study demonstrates that niclosamide can inhibit the cell growth of colon cancer by inhibiting STAT3 phosphorylation. We also investigated the combined inhibition of EGFR and STAT3 by using erlotinib and niclosamide to synergistically repress colon cancer. This study was conducted only in vitro; some compounds may exhibit potent antitumor activities in vitro but show no antitumor activity in vivo. Therefore, further molecular investigations in preclinical and clinical settings are warranted to evaluate anticancer potential in a synergistic manner to substantiate niclosamide as an effective therapeutic candidate for colon cancer.
Acknowledgments
This research was supported by the National Natural Science Fund of China (21472142), the Zhejiang Provincial Natural Science Fund (LY16H160052, LY17H160050, and LY17H160055), Medical Scientific Research Fund of Zhejiang Province (2017192276), and Wenzhou Science and Technology Project (Y20160052). The authors thank Dr Jianzhang Wu for helpful discussions and assistance in writing the manuscript.
Disclosure
The authors report no conflicts of interest in this work.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66(1):7–30. | ||
Zahid A, Young CJ. How to decide on stent insertion or surgery in colorectal obstruction? World J Gastrointest Surg. 2016;8(1):84–89. | ||
Young JI, Mongoue-Tchokote S, Wieghard N, et al. Treatment and survival of small-bowel adenocarcinoma in the United States: a comparison with colon cancer. Dis Colon Rectum. 2016;59(4):306–315. | ||
de Castro-Carpeño J, Belda-Iniesta C, Casado Sáenz E, Hernández Agudo E, Feliu Batlle J, González Barón M. EGFR and colon cancer: a clinical view. Clin Transl Oncol. 2008;10(1):6–13. | ||
Kuwai T, Nakamura T, Sasaki T, et al. Phosphorylated epidermal growth factor receptor on tumor-associated endothelial cells is a primary target for therapy with tyrosine kinase inhibitors. Neoplasia. 2008;10(5):489–500. | ||
Li R, You S, Hu Z, et al. Inhibition of STAT3 by niclosamide synergizes with erlotinib against head and neck cancer. PLoS One. 2013;8(9):e74670. | ||
Kang Y, Hu W, Bai E, et al. Curcumin sensitizes human gastric cancer cells to 5-fluorouracil through inhibition of the NFkB survival-signaling pathway. Onco Targets Ther. 2016;9:7373–7384. | ||
Yu H, Pardoll D, Jove R. STATs in cancer inflammation and immunity: a leading role for STAT3. Nat Rev Cancer. 2009;9(11):798–809. | ||
Shodeinde AL, Barton BE. Potential use of STAT3 inhibitors in targeted prostate cancer therapy: future prospects. Onco Targets Ther. 2012;5:119–125. | ||
Lee H, Deng J, Kujawski M, et al. STAT3-induced S1PR1 expression is crucial for persistent STAT3 activation in tumors. Nat Med. 2010;16(12):1421–1428. | ||
Lesina M, Kurkowski MU, Ludes K, et al. Stat3/Socs3 activation by IL-6 transsignaling promotes progression of pancreatic intraepithelial neoplasia and development of pancreatic cancer. Cancer Cell. 2011;19(4):456–469. | ||
Gao SP, Mark KG, Leslie K, et al. Mutations in the EGFR kinase domain mediate STAT3 activation via IL-6 production in human lung adenocarcinomas. J Clin Invest. 2007;117(12):3846–3856. | ||
Haura EB, Zheng Z, Song L, Cantor A, Bepler G. Activated epidermal growth factor receptor-Stat-3 signaling promotes tumor survival in vivo in non-small cell lung cancer. Clin Cancer Res. 2005;11(23):8288–8294. | ||
Crescenzo R, Abate F, Lasorsa E, et al; European T-Cell Lymphoma Study Group, T-Cell Project: Prospective Collection of Data in Patients with Peripheral T-Cell Lymphoma and the AIRC 5xMille Consortium “Genetics-Driven Targeted Management of Lymphoid Malignancies”. Convergent mutations and kinase fusions lead to oncogenic STAT3 activation in anaplastic large cell lymphoma. Cancer Cell. 2015;27(4):516–532. | ||
Wake MS, Watson CJ. STAT3 the oncogene – still eluding therapy? FEBS J. 2015;282(14):2600–2611. | ||
Pan JX, Ding K, Wang CY. Niclosamide, an old antihelminthic agent, demonstrates antitumor activity by blocking multiple signaling pathways of cancer stem cells. Chin J Cancer. 2012;31(4):178–184. | ||
Chong CR, Sullivan DJ Jr. New uses for old drugs. Nature. 2007;448(7154):645–646. | ||
Jin Y, Lu Z, Ding K, et al. Antineoplastic mechanisms of niclosamide in acute myelogenous leukemia stem cells: inactivation of the NF-kappaB pathway and generation of reactive oxygen species. Cancer Res. 2010;70(6):2516–2527. | ||
Wang AM, Ku HH, Liang YC, Chen YC, Hwu YM, Yeh TS. The autonomous notch signal pathway is activated by baicalin and baicalein but is suppressed by niclosamide in K562 cells. J Cell Biochem. 2009;106(4):682–692. | ||
Osada T, Chen M, Yang XY, et al. Antihelminth compound niclosamide downregulates Wnt signaling and elicits antitumor responses in tumors with activating APC mutations. Cancer Res. 2011;71(12):4172–4182. | ||
Balgi AD, Fonseca BD, Donohue E, et al. Screen for chemical modulators of autophagy reveals novel therapeutic inhibitors of mTORC1 signaling. PLoS One. 2009;4(9):e7124. | ||
Zou M, Zhang X, Xu C. IL6-induced metastasis modulators p-STAT3, MMP-2 and MMP-9 are targets of 3,3′-diindolylmethane in ovarian cancer cells. Cell Oncol (Dordr). 2016;39(1):47–57. | ||
Eiring AM, Page BD, Kraft IL, et al. Combined STAT3 and BCR-ABL1 inhibition induces synthetic lethality in therapy-resistant chronic myeloid leukemia. Leukemia. 2015;29(3):586–597. | ||
Liu J, Chen X, Ward T, Pegram M, Shen K. Combined niclosamide with cisplatin inhibits epithelial-mesenchymal transition and tumor growth in cisplatin-resistant triple-negative breast cancer. Tumour Biol. 2016;37(7):9825–9835. | ||
Jin B, Wang C, Li J, Du X, Ding K, Pan J. Anthelmintic Niclosamide Disrupts the Interplay of p65 and FOXM1/β-catenin and Eradicates Leukemia Stem Cells in Chronic Myelogenous Leukemia. Clin Cancer Res. 2017;23(3):789–803. | ||
Zhu Y, Ye T, Yu X, et al. Nifuroxazide exerts potent anti-tumor and anti-metastasis activity in melanoma. Sci Rep. 2016;6:20253. | ||
Ye T, Xiong Y, Yan Y, et al. The anthelmintic drug niclosamide induces apoptosis, impairs metastasis and reduces immunosuppressive cells in breast cancer model. PLoS One. 2014;9(1):e85887. | ||
Zhu Z, Zhao Y, Li J, et al. Cryptotanshinone, a novel tumor angiogenesis inhibitor, destabilizes tumor necrosis factor-α mRNA via decreasing nuclear-cytoplasmic translocation of RNA-binding protein HuR. Mol Carcinog. 2016;55(10):1399–1410. | ||
Khan M, Yi F, Rasul A, et al. Alantolactone induces apoptosis in glioblastoma cells via GSH depletion, ROS generation, and mitochondrial dysfunction. IUBMB Life. 2012;64(9):783–794. | ||
Morris GM, Huey R, Lindstrom W, et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem. 2009;30(16):2785–2791. | ||
Becker S, Groner B, Müller CW. Three-dimensional structure of the Stat3beta homodimer bound to DNA. Nature. 1998;394(6689):145–151. | ||
Alexander N, Woetzel N, Meiler J. Bcl::Cluster: a method for clustering biological molecules coupled with visualization in the Pymol Molecular Graphics System. IEEE Int Conf Comput Adv Bio Med Sci. 2011;2011:13–18. | ||
Yang F, Hu M, Lei Q, et al. Nifuroxazide induces apoptosis and impairs pulmonary metastasis in breast cancer model. Cell Death Dis. 2015;6:e1701. | ||
Yu HJ, Park C, Kim SJ, Cho NP, Cho SD. Signal transducer and activators of transcription 3 regulates cryptotanshinone-induced apoptosis in human mucoepidermoid carcinoma cells. Pharmacogn Mag. 2014;10(Suppl 3):S622–S629. | ||
Chun J, Li RJ, Cheng MS, Kim YS. Alantolactone selectively suppresses STAT3 activation and exhibits potent anticancer activity in MDA-MB-231 cells. Cancer Lett. 2015;357(1):393–403. | ||
Zhao C, Xiao H, Wu X, et al. Rational combination of MEK inhibitor and the STAT3 pathway modulator for the therapy in K-Ras mutated pancreatic and colon cancer cells. Oncotarget. 2015;6(16):14472–14487. | ||
Zhao C, Li H, Lin HJ, Yang S, Lin J, Liang G. Feedback activation of STAT3 as a cancer drug-resistance mechanism. Trends Pharmacol Sci. 2016;37(1):47–61. | ||
van Kessel KE, Zuiverloon TC, Alberts AR, Boormans JL, Zwarthoff EC. Targeted therapies in bladder cancer: an overview of in vivo research. Nat Rev Urol. 2015;12(12):681–694. | ||
Chorost MI, Datta R, Santiago RC, et al. Colon cancer screening: where have we come from and where do we go? J Surg Oncol. 2004;85(1):7–13. | ||
Azcárate-Peril MA, Sikes M, Bruno-Bárcena JM. The intestinal microbiota, gastrointestinal environment and colorectal cancer: a putative role for probiotics in prevention of colorectal cancer? Am J Physiol Gastrointest Liver Physiol. 2011;301(3):G401–G424. | ||
Pohl M, Stricker I, Schoeneck A, et al. Antitumor activity of the HER2 dimerization inhibitor pertuzumab on human colon cancer cells in vitro and in vivo. J Cancer Res Clin Oncol. 2009;135(10):1377–1386. | ||
Wen W, Wu J, Liu L, et al. Synergistic anti-tumor effect of combined inhibition of EGFR and JAK/STAT3 pathways in human ovarian cancer. Mol Cancer. 2015;14:100. | ||
Sack U, Walther W, Scudiero D, et al. Novel effect of antihelminthic Niclosamide on S100A4-mediated metastatic progression in colon cancer. J Natl Cancer Inst. 2011;103(13):1018–1036. | ||
Wang X, Goldstein D, Crowe PJ, et al. Overcoming resistance of targeted EGFR monotherapy by inhibition of STAT3 escape pathway in soft tissue sarcoma. Oncotarget. 2016;7(16):21496–21509. | ||
Kraljevic S, Stambrook PJ, Pavelic K. Accelerating drug discovery. EMBO Rep. 2004;5(9):837–842. | ||
Sakamoto KM, Grant S, Saleiro D, et al. Targeting novel signaling pathways for resistant acute myeloid leukemia. Mol Genet Metab. 2015;114(3):397–402. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.