Back to Journals » Infection and Drug Resistance » Volume 14
Molecular Epidemiology and Mechanisms of Carbapenem-Resistant Acinetobacter baumannii Isolates from ICU and Respiratory Department Patients of a Chinese University Hospital
Received 29 December 2020
Accepted for publication 6 February 2021
Published 25 February 2021 Volume 2021:14 Pages 743—755
DOI https://doi.org/10.2147/IDR.S299540
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Sahil Khanna
Bin Liu,1 Lei Liu2
1Department of Respiratory and Critical Care Medicine, The First Affiliated Hospital of Harbin Medical University, Harbin, People’s Republic of China; 2Department of Respiratory Medicine, The Fourth Affiliated Hospital of Harbin Medical University, Harbin, People’s Republic of China
Correspondence: Lei Liu Email [email protected]
Background: The objective of our study is to estimate the differences in molecular epidemiology and resistance mechanisms in carbapenem-resistant Acinetobacter baumannii (CRAB) isolates from the ICU and respiratory department(RD) in Fourth Affiliated Hospital of Harbin Medical University.
Methods: Carbapenemase genes associated with carbapenem resistance were studied by polymerase chain reaction(PCR). Genotyping was analyzed using multi-locus sequence typing (MLST) and pulsed field gel electrophoresis (PFGE).
Results: Sixty non-duplicate CRAB isolates from the ICU and RD (n=30, respectively) were collected. All of CRAB strains were not resistant to colistin (0%). The CRAB strains from the ICU were significantly more resistant to tigecycline and cefoperazone/sulbactam compared with the RD (23.3% vs 0%, P=0.03; 53.3% % vs 23.3%, P=0.01, respectively). PCR detection of genes associated with CRAB revealed that the ratio in both the ICU and the RD of blaVIM-2, blaIMP-4, blaNDM-1, blaOXA-23, ampC, and mutation of CarO were present in 23.3% vs 0% (P=0.01), 40% vs 10% (P=0.02), 20% vs 0% (P=0.02), 80% vs 56.7%, 16.7% vs 13.3% and 86.7% vs 60% (P=0.04), respectively. Seven genotypes were detected by the PFGE in the RD and the ICU, respectively. Genotype I was significantly more frequent in the ICU compared with the RD (63.3% vs 36.6%, P=0.03). MLST showed that there were 10 ST genotypes in the RD and four in the ICU, but ST92 in both groups was 33.3% vs 63.3% (P=0.03), respectively.
Conclusion: There are differences in molecular epidemiology and resistance mechanisms in the CRAB isolates between the ICU and RD.
Keywords: Acinetobacter baumannii, carbapenem resistance, molecular epidemiology, resistance mechanisms
Background
Acinetobacter baumannii (A. baumannii or AB) has emerged as a very important pathogen of nosocomial infections and has been related to high morbidity and mortality in the past decade.1,2 Carbapenems were the first choice in the treatment of A. baumannii infection due to their previously high antimicrobial activity and low toxicity.3 However, A. baumannii has developed a resistance to this drug, resulting in a central global health threat. A rising incidence of carbapenem-resistant A. baumannii (CRAB) in intra-hospital and inter-hospital dissemination has appeared in the intensive care unit (ICU)4–6 of the chosen hospital. Recent studies have reported that CRAB strains are major causative organisms in hospital-acquired infections.7,8 It has been revealed that 65% of A. baumannii infections in the United States and Europe were caused by CRAB.9,10 Simultaneously, the infection rate of CRAB in China has reached 62%.11
CRAB is chiefly in connection with the emergence of class D carbapenem-hydrolyzing-lactamases and infrequently in connection with metallo-β-lactamases.12 There are the blaOXA-23-like gene that is mainly related to CRAB.13 The previous literature revealed the spread of resistant blaOXA-23-producing AB clones in 18 Chinese provinces.14 In addition, the resistance mechanism of CRAB was also bound with mutation in the CarO gene and efflux pump.15 However, there is extremely limited data about the molecular epidemiology and resistance mechanisms of CRAB in Harbin, China. Furthermore, no data are available on the epidemiological characteristics of CRAB isolates from the ICU and respiratory department.
Multi-locus sequence typing (MLST), pulsed field gel electrophoresis (PFGE) and polymerase chain reaction(PCR) are the most frequently and widely used methods for the characterization of the molecular epidemiology study of A. baumannii.16
The main objectives of our study are to estimate the differences in clinical characteristics, antimicrobial resistance profiles, molecular epidemiology, and resistance mechanisms in CRAB isolates from the ICU and respiratory department using MLST, PCR and PFGE in a university teaching hospital in China’s northeastern region. These will provide a theoretical basis for nosocomial infection control and prevention of CRAB.
Methods
The fourth Affiliated Hospital of Harbin Medical University, located in northeastern China, a tertiary teaching hospital with 2600 beds in use, including a 30-bed medical ICU and a 120-bed respiratory department.
Bacterial Isolates and Identification
During the period of July 2019 to January 2020, 60 non-duplicate CRAB strains from the ICU and respiratory department, 30 isolates, respectively, were collected. The clinical samples were specifically isolated for this research, all patients provided written informed consent. The method for identifying A. baumannii is to detect blaoxa-51-like gene of A. baumannii complex by PCR. The CRAB was defined as all the analyzed AB strains, whether in the ICU or respiratory department, were resistant to imipenem and meropenem. Clinical isolates from the samples including respiratory tract, urinary tract, blood, and wound surface samples were obtained. Carbapenemases were screened by the Modified Hodge test and the results were interpreted according to the Clinical and Laboratory Standards Institute (CLSI) 2018.17 Isolates identification, the minimum inhibitory concentration (MIC) and antimicrobial susceptibility testing of CRAB were performed using the VITEK 32 system (bioMerieux, France).
Antimicrobial Sensitivity and Resistance
The in vitro activity of regularly utilized antimicrobial agents was determined by broth microdilution in line with the guidelines of the CLSI.17 The interpretive standards for the MIC of the antibiotics were determined based on the CLSI breakpoints.17 Susceptibility testing of antimicrobials was performed using the broth microdilution method. The results were interpreted according to the breakpoints suggested by the CLSI.17 The seven types of antibiotic agents tested in this study were the extended-spectrum cephalosporins (ceftazidime), carbapenems (imipenem and meropenem), quinolones (ciprofloxacin), b-lactam/b-lactamase inhibitor combinations (piperacillin/tazobactam and cefoperazone/sulbactam), aminoglycosides (gentamicin and amikacin), glycopeptides (colistin), and tetracyclines (tigecycline). Colistin was examined in line with the recommendations of the CLSI.17
With regard to tigecycline, susceptibility/resistance breakpoints were interpreted according to the European Committee on Antimicrobial Susceptibility Testing criteria.18,19 Escherichia coli ATCC 25,922 and A. baumannii ATCC 19,606 were used as reference strains.
PCR Detection of Carbapenemases and Outer Membrane Protein of Carbapenem-Associated Resistance OMP (CarO)
Genomic DNA extractions were implemented according to a reformative protocol as, reported in the literature.20,21 First, the CRAB isolates were hatched on Mueller–Hinton agar plates all night at 37°C. Second, a loop full of each pure isolate was introduced in 300 μL of sterile distilled water and boiled for 10 minutes. After five minutes of centrifugation (6000 r.p.m.), the supernatant was kept at −20°C as a DNA template for PCR amplification. The existence of carbapenemase-encoding genes, including blaNDM, blaSPM, blaAIM, blaKPC, blaVIM, blaIMP, blaOXA-23-like, and bla-OXA51-like and genes related to carbapenem resistance, including CarO and ampC were detected by PCR assays.21,22 The total information of the PCR primers utilized in our study is shown in Table 1. The detected data were analyzed by applying Bioedit software, and the findings were contrasted by network blast software.27
 |
Table 1 Information Concerning the Primers Used in This Study |
Molecular Typing by MLST and PFGE
MLST
MLST was performed as previously described.28 The isolates were assigned to sequence types by using tools on the CRAB strains and MLST database (http://pubmlst.org/abaumannii/).
PFGE
Chromosomal DNA was prepared from CRAB strains and digested with ApaI (TaKaRa Bio) as previously reported.29 The PFGE results were interpreted by the criteria according to the reported literature.30
Statistical Analysis
Analyses were performed using SPSS 17.0 (SPSS Inc., Chicago, IL) for all data. Comparative analyses were executed by the Chi-square or Fisher exact tests for categorical variables, and by the t-test for continuous variables. All examinations were two-tailed, and P<0.05 was considered to indicate statistical significance.
Results
In total, 60 non-duplicate CRAB clinical isolates from the ICU and respiratory department, including 30 isolates, respectively, were collected between July 2019 and January 2020. All patients with the CRAB infection from the ICU and respiratory department had nosocomial infections. The rates of mechanical intervention were 70% (21/30) in the respiratory department and 100% (30/30) in the ICU. CRAB clinical isolates from the samples including the respiratory tract (81.7%, 49/60), blood (10%, 6/60), and wound surfaces (8.3%, 5/60) were obtained. Attributable mortality from the CRAB infection was 6.67% (2/30) in the respiratory department, and 23.33% (7/30) in the ICU; no significantly statistical difference was observed (P=0.15). Detailed information of all patients is provided in Table 2.
 |
Table 2 Summarized Demographic Data for All Patients from Respiratory Department and ICU (n=60) |
Antimicrobial Resistance Testing
Antibiotic susceptibility testing revealed that all the analyzed CRAB clinical strains, whether in the ICU or respiratory department, were resistant to imipenem (100%) and meropenem (100%), and all the analyzed CRAB clinical strains were not resistant to colistin (0%). Of all the CRAB clinical strains from the ICU and respiratory department (n=30, respectively), all the analyzed CRAB clinical strains from the respiratory department were resistant to tigecycline (0%), seven isolates were resistant to tigecycline from the ICU (23.3%), and a statistically significant difference was observed (P=0.03); seven isolates were resistant to cefoperazone/sulbactam from the respiratory department (23.3%), 16 isolates were resistant to cefoperazone/sulbactam from the ICU (53.3%), and a statistically significant difference was detected (P=0.01). The resistance rate was in the range of 43.3–70% for the other antimicrobials, including piperacillin/tazobactam, gentamicin, amikacin, ceftazidime, and ciprofloxacin; there was no statistically significant change between the ICU and respiratory department. Overall, colistin was the most effective antibiotic to the CRAB isolates, whether from the ICU or respiratory department. Antibiotic resistance and susceptibility profiles of the CRAB clinical isolates are shown in Table 3.
 |
Table 3 Antimicrobial Resistance and Susceptibility Profiles of CRAB Clinical Isolates Obtained from Patients at Respiratory Department and ICU |
Detection of Genes Associated with Carbapenem Resistance
The Modified Hodge test of 60 CRAB strains was positive. Of the various drug resistance genotypes tested, all of CRAB isolates from the ICU revealed that blaVIM-2, blaIMP-4, blaNDM-1, blaOXA-23, ampC, and mutation of CarO were present in 23.3% (7/30), 40% (12/30), 20% (6/30), 80% (24/30), 16.7% (5/30) and 86.7% (26/30) of the CRAB strains, respectively. Similarly, the CRAB isolates from the respiratory department indicated that blaVIM-2, blaIMP-4, blaNDM-1, blaOXA-23, ampC and mutation of CarO were present in 0% (0/30), 10% (3/30), 0% (0/30), 56.7% (17/30), 13.3% (4/30) and 60% (18/30) of the CRAB strains, respectively. A statistically significant difference was observed in blaVIM-2, blaIMP-4, blaNDM-1, and mutation of CarO between the respiratory department and the ICU (P=0.01, P=0.02, P=0.02, P=0.04, respectively). These results are shown in Table 4, Figures 1–3.
 |
Table 4 PCR Detection of Genes Associated with CRAB Isolates in Respiratory Department and ICU |
 |
Figure 1 Detection of IMP genes of carbapenem-resistant Acinetobacter baumannii clinical isolates by PCR. |
 |
Figure 2 Detection of VIM genes of carbapenem-resistant Acinetobacter baumannii clinical isolates by PCR. |
 |
Figure 3 Detection of NDM genes of carbapenem-resistant Acinetobacter baumannii clinical isolates by PCR. |
Typing by PFGE
CRAB isolates were genotyped by PFGE. The results showed that CRAB isolates could be divided into 10 clones (A-J). The 30 CRAB isolates in the respiratory department were distributed in seven genotypes (A, C, D, E, H, I and J), and the 30 CRAB isolates in the ICU were distributed in seven genotypes (A, B, D, E, G, H and I). The bacterial numbers of genotype I were the most popular, accounting for 50% (30/60), including 36.6% (11/30) from the respiratory department and 63.3% (19/30) from the ICU, respectively. However, a statistically significant difference was observed in genotype I between the ICU and respiratory department (P=0.03). Among type I, the I5 subtype is the most popular. These results are shown in Figure 4.
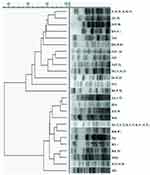 |
Figure 4 Dendrogram displaying the genetic relatedness of 60 CRAB strains, based on pulse field gel electrophoresis results. |
PFGE analysis of the CRAB strains isolated from the respiratory department and the ICU revealed that there was no significant difference in the genotypes. In the same homology analysis, the CRAB strains isolated from the respiratory department and the ICU showed cross coincidence distribution, and there was no obvious clumped distribution.
Molecular Typing by MLST
We used MLST technology to genotype the CRAB clinical isolates in the respiratory department and the ICU. The results showed that there were 10 ST genotypes in the respiratory department, among which the ST92 genotype was the most popular, accounting for 33.3% (10/30). The ST175 and ST298 genotypes accounted for 20.0% (6/30) and 13.3% (4/30), respectively. There were four genotypes in the ICU, among which the ST92 genotype was the most popular, accounting for 63.3% (19/30). The genotypes ST111 and ST244 accounted for 20.0% (6/30) and 10.0% (3/30), respectively. However, a statistically significant difference was observed in the most popular ST92 genotype between the respiratory department and the ICU (P=0.03). These results are shown in Table 5.
 |
Table 5 The MLST Results of CRAB in Respiratory Department and ICU |
Discussion
The CRAB is an urgent problem that needs to be solved. From the Taiwan Nosocomial Infection Surveillance System (2003–2012), the proportion of AB isolates resistant to carbapenem increased from 17.2% to 72.8%.31 There are extremely limited antibiotics that can be selected. In our study, colistin was found to be the most effective antibiotic assessed. Some drug sensitivity results are inconsistent with individual reports,32,33 and these differences could be explained by the differences in the habit, frequency, and region of the antibiotics use. A reported study showed that invasive procedures, ICU duration, recent surgery, and exposure to broad-spectrum antibiotics are risk factors for colonization or infection by CRAB.34 The ICU has been seen as an epicenter of the epidemiology of CRAB.35 In our study, we detected some major risk factors among inpatients, and medical intervention seems to be a major risk factor. Moreover, the predominant clinical picture of CRAB infection among patients was mostly respiratory. So far, some studies reported clonal outbreaks of the CRAB isolates that were related to environmental reservoirs36 to the extent that the ICU settings had to be completely shut down to terminally disinfect and control the epidemic.35,37 However, the departments in different hospitals have different circumstances, complicated situations and other unpredictable factors, such as department design, hospital economy, antibiotic management policy, and department management.
The AB isolates developed resistance to carbapenem by complex mechanisms. Carbapenemase is a key factor responsible for CRAB infection. In our study, some common carbapenemases were detected in the CRAB strains, including blaOXA-23, blaIMP, blaNDM, and blaVIM. The findings from our study, carrying the blaOXA-23 gene, was the main cause of the CRAB resistance phenotype, which is in line with previous studies.38–40 Recent reports have demonstrated that the VIM type is widespread in the Middle East region among CRAB.41 However, blaOXA-23-like still remains the most prevalent all over the world.42,43 The blaOXA-23-producing strains were popular in the respiratory department and the ICU, and blaOXA-23 can be inserted into plasmids and chromosomes. Our findings are similar to the reported literature.44 Similarly, this appearance made the control of CRAB infection further complicated. However, the spread of blaOXA-23 in the ICU should receive more attention than the respiratory department. New Delhi metallo-β-lactamase (NDM) was first identified in 2008 in India, and the NDM-producing CRAB isolate was first reported in 2011 in China.45 Currently, blaNDM has been identified as having low spread.46 Of the antibiotics detected, NDM‑positive CRAB strains in our study were identified that were extremely drug‑resistant and only sensitive to colistin and partial strains were susceptible to tigecycline. A laboratory in Argentina demonstrated that the outer membrane protein is encoded by the CarO gene, and when there is an insertion mutation or any other mutation, the CarO gene makes it off and thus the strain becomes resistant to certain drugs.15 In our research, all the CRAB isolates from the ICU revealed that mutation of CarO was present in 86.7% (26/30), and 60% (18/30) from the respiratory department, a statistically significant difference was observed. This is consistent with the reported literature.47 The above results indicate that the main mechanisms of CRAB between the ICU and the respiratory department may be the cooperation of blaOXA-23, blaIMP, and mutation of CarO.
PFGE analysis of CRAB strains isolated from the respiratory department and the ICU revealed that there was no significant difference in the classification. In the same homology analysis, CRAB strains isolated from the respiratory department and the ICU showed cross coincidence distribution, and there was no obvious clumped distribution. There was on the different floor and building between respiratory department and ICU, which suggested that the results of this molecular typing of CRAB strains from the respiratory department and the ICU were representative of the epidemic type of CRAB strains from the respiratory department and ICU in our hospital.
MLST analyses indicated that ST92 was the most prominent sequence type, which is completely concordant with previous observations that ST92 was the dominant clone of AB isolates in mainland China.48 Our findings showed that although there are some advantages of CRAB strains in the respiratory department, the resistant genotype diversity of the CRAB strains still exists. However, it is the vast majority of epidemic CRAB strains in the ICU that suggest there is a high risk of an outbreak in the ICU; therefore, the ICU setting should pay more attention to key monitoring and infection control measures. Some of our results are similar to the reported literature.44,45 Furthermore, the MLST analysis still revealed that some epidemic clones of CRAB from the ICU and respiratory department in this study were ST111 and ST175, which differed from the results obtained in other regions in China.47 These differences may be related to regional differences, carbapenem management, regional diseases, and infection control. These differences may also be explained as there are some endemic clones that are restricted to specific geographical regions. Our results indicated that ICU was more inclined to clonal transmission, however, respiratory department tended to be genetic diverse.
To our knowledge, the main advantage of our study consists of the first report of the molecular mechanisms and epidemiology profiles of CRAB from the respiratory department and the ICU in northeastern China, which permits the exact analysis differences between the respiratory department and the ICU in our hospital. In addition, obtaining a more effective solution for a better controlling incidence of CRAB in our hospital, which means heterogeneous measures can be taken in different departments from one hospital. However, there are a few limitations to our research. First, the environmental strains were not further obtained from the respiratory department and the ICU in our hospital; hence, this present study is absent from the identification of strain sources. This is also the direction of our next work. Second, there was a lack of monitoring for all patients who were recently admitted to the respiratory department and the ICU. Consequently, it was difficult to distinguish all patients between new infection cases obtaining CRAB in the respiratory department and the ICU and previously infected cases. Ultimately, due to limited funds and experimental conditions, some resistant genes associated with carbapenem-resistance, such as efflux pumps, have not been detected, which leads to the fact that we cannot understand the whole mechanism of CRAB infection.
Conclusion
In conclusion, this is the first report of the molecular mechanisms and epidemiology profiles of CRAB from the respiratory department and the ICU in northeastern China. Our findings suggest that there are partial differences in clinical characteristics, antimicrobial resistance profiles, molecular epidemiology, and resistance mechanisms in CRAB isolates between the ICU and respiratory department. The blaOXA-23-producing CRAB strains were popular in the respiratory department and the ICU. The MLST analysis revealed that the major epidemic clone of CRAB from the ICU was ST92. The dissemination of blaOXA-23-producing CRAB strains in the ICU should receive more attention than in the respiratory department, and an effective nosocomial infection control measure should be urgently required in the ICU. The heterogeneous control measures of CRAB infection can be taken in the ICU and respiratory department from one hospital.
Abbreviations
AB, Acinetobacter baumannii; CRAB, carbapenem-resistant Acinetobacter baumannii; ICU, intensive care units; RD, respiratory department; PCR, polymerase chain reaction; MLST, multi-locus sequence typing; PFGE, pulsed field gel electrophoresis; CLSI, Clinical and Laboratory Standards Institute; CarO, carbapenem-associated resistance OMP; MIC, minimum inhibitory concentration.
Data Sharing Statement
Availability of data and material need to be approved by the Microbiology Department of the Fourth Affiliated Hospital of Harbin Medical University.
Ethics Approval and Consent to Participate
This study has been reviewed and approved by the Research Ethics Committee of the Fourth Affiliated Hospital of Harbin Medical University (no. 2019-127). The clinical samples were specifically isolated for this research, all of the patients provided written informed consent. All relevant ethical safeguards have been met in line with the Declaration of Helsinki.
Consent for Publication
All authors have been personally and actively involved in substantive work leading to the report, and will hold themselves jointly and individually responsible for its content.
Acknowledgments
We would like to acknowledge the work of the Fourth Affiliated Hospital of the Harbin Medical University’s microbiology laboratory.
Funding
There is no funding to report.
Disclosure
The authors have no conflict of interests to declare.
References
1. Perez F, Hujer AM, Hujer KM, Decker BK, Rather PN, Bonomo RA. Global challenge of multidrug-resistant Acinetobacter baumannii. Antimicrob Agents Chemother. 2007;51(10):3471–3484. doi:10.1128/AAC.01464-06
2. Giamarellou H, Antoniadou A, Kanellakopoulou K. Acinetobacter baumannii: a universal threat to public health? Int J Antimicrob Agents. 2008;32(2):106–119. doi:10.1016/j.ijantimicag.2008.02.013
3. Kim YJ, Kim SI, Hong K-W, Kim YR, Park YJ, Kang M-W. Risk factors for mortality in patients with carbapenem-resistant Acinetobacter baumannii bacteremia: impact of appropriate antimicrobial therapy. J Korean Med Sci. 2012;27(5):471–475. doi:10.3346/jkms.2012.27.5.471
4. Nhu NTK, Lan NPH, Campbell JI, et al. Emergence of carbapenem-resistant Acinetobacter baumannii as the major cause of ventilator-associated pneumonia in intensive care unit patients at an infectious disease hospital in southern Vietnam. J Med Microbiol. 2014;63(Pt10):1386–1394. doi:10.1099/jmm.0.076646-0
5. Qureshi ZA, Hittle LE, O’Hara JA, et al. Colistin-resistant Acinetobacter baumannii: beyond carbapenem resistance. Clin Infect Dis. 2015;60(9):1295–1303. doi:10.1093/cid/civ048
6. Rafei R, Dabboussi F, Hamze M, et al. Molecular analysis of Acinetobacter baumannii strains isolated in Lebanon using four different typing methods. PLoS One. 2014;9(12):e115969. doi:10.1371/journal.pone.0115969
7. Evans BA, Hamouda A, Amyes SGB. The rise of carbapenem-resistant Acinetobacter baumannii. Curr Pharm Des. 2013;19(2):223–238. doi:10.2174/138161213804070285
8. Higgins PG, Dammhayn C, Hackel M, Seifert H. Global spread of carbapenem-resistant Acinetobacter baumannii. J Antimicrob Chemother. 2010;65(2):233–238. doi:10.1093/jac/dkp428
9. Lee CR, Lee JH, Park M, et al. Biology of acinetobacter baumannii: pathogenesis, antibiotic resistance mechanisms, and prospective treatment options. Front Cell Infect Microbiol. 2017;7:55.
10. Piana A, Palmieri A, Deidda S, et al. Molecular typing of XDR Acinetobacter baumannii strains in an Italian ICU. Epidemiol Prev. 2015;39(4Suppl 1):129–133.
11. Liu X, Zhao M, Chen Y, Bian X, Li Y, Shi J. Synergistic killing by meropenem and colistin combination of carbapenem-resistant Acinetobacter baumannii isolates from Chinese patients in an in vitro pharmacokinetic/pharmacodynamic model. Int J Antimicrob Agents. 2016;48(5):559–563. doi:10.1016/j.ijantimicag.2016.07.018
12. Martinez T, Martinez I, Vazquez GJ, Aquino EE, Robledo IE. Genetic environment of the KPC gene in Acinetobacter baumannii ST2 clone from Puerto Rico and genomic insights into its drug resistance. J Med Microbiol. 2016;65(8):784. doi:10.1099/jmm.0.000289
13. Higgins PG, Pérezllarena FJ, Zander E, Fernández A, Bou G, Seifert H. OXA-235, a novel class D β-lactamase involved in resistance to carbapenems in Acinetobacter baumannii. Antimicrob Agents Chemother. 2013;57(5):2121–2126. doi:10.1128/AAC.02413-12
14. Liu L, Ji S, Ruan Z, et al. Dissemination of blaOXA-23 in Acinetobacter spp. in China: main roles of conjugative plasmid pAZJ221 and transposon Tn2009. Antimicrob Agents Chemother. 2015;59(4):1998–2005. doi:10.1128/AAC.04574-14
15. Mussi MA, Limansky AS, Viale AM. Acquisition of resistance to carbapenems in multidrug-resistant clinical strains of Acinetobacter baumannii: natural insertional inactivation of a gene encoding a member of a novel family of β-barrel outer membrane proteins. Antimicrob Agents Chemother. 2005;49(4):1432–1440. doi:10.1128/AAC.49.4.1432-1440.2005
16. Tomaschek F, Higgins PG, Stefanik D, Wisplinghoff H, Seifert H, Gao F. Head-to-head comparison of two Multi-Locus Sequence Typing (MLST) schemes for characterization of Acinetobacter baumannii outbreak and sporadic isolates. PLoS One. 2016;11(4):e0153014. doi:10.1371/journal.pone.0153014
17. Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 28th Ed. CLSI Supplement M100. Wayne, PA: Clinical and Laboratory Standards Institute; 2018.
18. Deng M, Zhu M-H, Li J-J, et al. Molecular epidemiology and mechanisms of tigecycline resistance in clinical isolates of Acinetobacter baumannii from a Chinese University Hospital. Antimicrob Agents Chemother. 2014;58(1):297–303. doi:10.1128/AAC.01727-13
19. Zhao Y, Hu K, Zhang J, et al. Outbreak of carbapenem-resistant Acinetobacter baumannii carrying the carbapenemase OXA-23 in ICU of the eastern Heilongjiang Province, China. BMC Infect Dis. 2019;19(1):452. doi:10.1186/s12879-019-4073-5
20. Pournaras S, Gogou V, Giannouli M, et al. Single-locus-sequence-based typing of blaOXA-51-like genes for rapid assignment of Acinetobacter baumannii clinical isolates to international clonal lineages. J Clin Microbiol. 2014;52:1653–1657. doi:10.1128/JCM.03565-13
21. Woodford N, Ellington MJ, Coelho JM, et al. Multiplex PCR for genes encoding prevalent OXA carbapenemases in Acinetobacter spp. Int J Antimicrob Agents. 2006;27(4):351–353. doi:10.1016/j.ijantimicag.2006.01.004
22. Karah N, Jolley KA, Hall RM, Uhlin BE, Gao F. Database for the ampC alleles in Acinetobacter baumannii. PLoS One. 2017;12(5):e0176695. doi:10.1371/journal.pone.0176695
23. Mirshekar M, Shahcheraghi F, Azizi O, Solgi H, Badmasti F. Diversity of class 1 integrons, and disruption of carO and dacD by insertion sequences among Acinetobacter baumannii Isolates in Tehran, Iran. Microb Drug Resist. 2018;24(4):359–366. doi:10.1089/mdr.2017.0152
24. Piran A, Shahcheraghi F, Solgi H, Rohani M, Badmasti F. A reliable combination method to identification and typing of epidemic and endemic clones among clinical isolates of Acinetobacter baumannii. Infect Genet Evol. 2017;54:501–507. doi:10.1016/j.meegid.2017.08.018
25. El-Shazly S, Dashti A, Vali L, Bolaris M, Ibrahim AS. Molecular epidemiology and characterization of multiple drug-resistant (MDR) clinical isolates of Acinetobacter baumannii. Int J Infect Dis. 2015;41:42–49. doi:10.1016/j.ijid.2015.10.016
26. Sun F, Ou Q, Wang Q, et al. The resistance and transmission mechanism of Acinetobacter baumannii isolates in a tertiary care hospital, China. J Chemother. 2016;28(6):
27. Turton JF, Ward ME, Woodford N, et al. The role of ISAba1 in expression of OXA carbapenemase genes in Acinetobacter baumannii. FEMS Microbiol Lett. 2010;258(1):72–77. doi:10.1111/j.1574-6968.2006.00195.x
28. Bartual SG, Seifert H, Hippler C, et al. Development of a multilocus sequence typing scheme for characterization of clinical isolates of Acinetobacter baumannii. J Clin Microbiol. 2005;43(9):4382–4390. doi:10.1128/JCM.43.9.4382-4390.2005
29. Durmaz R, Otlu B, Koksal F. The optimization of a rapid pulsed-field gel electrophoresis protocol for the typing of Acinetobacter baumannii, Escherichia coli and Klebsiella spp. Jpn J Infect Dis. 2009;62(5):372–377.
30. Tenover FC, Arbeit RD, Goering RV, et al. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33(9):2233. doi:10.1128/JCM.33.9.2233-2239.1995
31. Chang -Y-Y, Chen H-P, Lin C-W, et al. Implementation and outcomes of an antimicrobial stewardship program: effectiveness of education. J Chin Med Assoc. 2017;80(6):353–359. doi:10.1016/j.jcma.2016.09.012
32. Warde E, Davies E, Ward A. Control of a multidrug-resistant Acinetobacter baumannii outbreak. Br J Nurs. 2019;28(4):242–248. doi:10.12968/bjon.2019.28.4.242
33. Muñoz-Price LS, Carling P, Clearly T, et al. Control of a two-decade endemic situation with carbapenem-resistant Acinetobacter baumannii: electronic dissemination of a bundle of interventions. Am J Infect Control. 2014;42:466–471. doi:10.1016/j.ajic.2013.12.024
34. Yamada K, Yanagihara K, Araki N, et al. Clinical characteristics of tertiary hospital patients from whom Acinetobacter calcoaceticus-Acinetobacter baumannii complex strains were isolated. Intern Med. 2012;51(1):51–57. doi:10.2169/internalmedicine.51.6018
35. Molter G, Seifert H, Mandraka F, et al. Outbreak of carbapenem-resistant Acinetobacter baumannii in the intensive care unit: a multi-level strategic management approach. J Hosp Infect. 2016;92(2):194–198. doi:10.1016/j.jhin.2015.11.007
36. Silva GJD, Domingues S. Insights on the horizontal gene transfer of carbapenemase determinants in the opportunistic pathogen Acinetobacter baumannii. Microorganisms. 2016;4(3):29. doi:10.3390/microorganisms4030029
37. Tsiatsiou O, Iosifidis E, Katragkou A, et al. Successful management of an outbreak due to carbapenem-resistant Acinetobacter baumannii in a neonatal intensive care unit. Eur J Pediatr. 2015;174(1):65–74. doi:10.1007/s00431-014-2365-8
38. Zowawi HM, Sartor AL, Sidjabat HE, et al. Molecular epidemiology of carbapenem-resistant Acinetobacter baumannii isolates in the gulf cooperation council states: dominance of OXA-23-type producers. J Clin Microbiol. 2015;53(3):896. doi:10.1128/JCM.02784-14
39. He C, Xie Y, Zhang L, et al. Increasing imipenem resistance and dissemination of the ISAba1-associated blaOXA-23 gene among Acinetobacter baumannii isolates in an intensive care unit. J Med Microbiol. 2011;60:337–341. doi:10.1099/jmm.0.022681-0
40. Martínez P, Mattar S. Imipenem-resistant Acinetobacter baumannii carrying the ISAba1-bla OXA-23,51 and ISAba1-bla ADC-7 genes in Monteria, Colombia. Braz J Microbiol. 2012;43:
41. Shahcheraghi F, Abbasalipour M, Feizabadi M, Ebrahimipour G, Akbari N. Isolation and genetic characterization of metallo-β-lactamase and carbapenamase producing strains of Acinetobacter baumannii from patients at Tehran hospitals. Iran J Microbiol. 2011;3:68.
42. Mugnier PD, Poirel L, Naas T, Nordmann P. Worldwide dissemination of the blaOXA-23 carbapenemase gene of Acinetobacter baumannii. Emerg Infect Dis. 2010;16:35. doi:10.3201/eid1601.090852
43. Thomas CM, Nielsen KM. Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat Rev Microbiol. 2005;3(9):711–721. doi:10.1038/nrmicro1234
44. Hua X, Zhou Z, Yang Q, et al. Evolution of Acinetobacter baumannii in vivo: international clone II, more resistance to ceftazidime, mutation in ptk. Front Microbiol. 2017;8:1256.
45. Savard P, Gopinath R, Zhu W, et al. First NDM‑positive Salmonella sp. strain identified in the United States. Antimicrob Agents Chemother. 2011;55:
46. Armin S, Fallah F, Azimi L, et al. Warning: spread of NDM-1 in two border towns of Iran. Cell Mol Biol. 2018;64:125–129. doi:10.14715/cmb/2018.64.10.20
47. Xiao S-Z, Chu H-Q, Han L-Z, et al. Resistant mechanisms and molecular epidemiology of imipenem-resistant Acinetobacter baumannii. Mol Med Rep. 2016;14(3):2483–2488. doi:10.3892/mmr.2016.5538
48. Huang L, Sun L, Yan Y. Clonal spread of carbapenem resistant Acinetobacter baumannii ST92 in a Chinese Hospital during a 6-year period. J Microbiol. 2013;51(1):113–117. doi:10.1007/s12275-013-2341-4
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
