Back to Journals » Infection and Drug Resistance » Volume 11
Molecular characteristics and comparative genomics analysis of a clinical Enterococcus casseliflavus with a resistance plasmid
Authors Yin M, Jiang Y, Qian C, Wu F, Ying Y, Wu C, Li P, Ying J, Li K, Xu T, Bao Q , Sun C
Received 14 July 2018
Accepted for publication 3 September 2018
Published 5 November 2018 Volume 2018:11 Pages 2159—2167
DOI https://doi.org/10.2147/IDR.S180254
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eric Nulens
Min Yin,1,* Yi Jiang,1,* Changrui Qian,1 Fei Wu,1 Yuanyuan Ying,1 Chongyang Wu,1 Peizhen Li,1 Jun Ying,1 Kewei Li,1 Teng Xu,1 Qiyu Bao,1 Caixia Sun2
1Department of Microbiology and Immunology, School of Laboratory Medicine and Life Sciences/Institute of Biomedical Informatics, Wenzhou Medical University, Wenzhou, Zhejiang 325035, China; 2Nursing Department, The First Affiliated Hospital of Wenzhou Medical University, Wenzhou, Zhejiang 325000, China
*These authors contributed equally to this work
Purpose: The aim of this work was to investigate the molecular characterization of a clinical Enterococcus casseliflavus strain with a resistance plasmid.
Materials and methods: En. casseliflavus EC369 was isolated from a patient in a hospital in southern China. The minimum inhibitory concentration was found by means of the agar dilution method to determine the antimicrobial susceptibilities of the strains. Whole-genome sequencing and comparative genomics analysis were performed to analyze the mechanism of antibiotic resistance and the horizontal gene transfer of the resistance gene-related mobile genetic elements.
Results: En. casseliflavus EC369 showed resistance to erythromycin, kanamycin, and streptomycin, but was susceptible to vancomycin, ampicillin, and streptothricin and other antimicrobials. There were six resistance genes (aph3′, ant6, bla, sat4, and two ermBs) carried by a transposon identified on the plasmid pEC369 and a complete resistance gene cluster of vancomycin and a tet (M) gene encoded on the chromosome. This is the first complete plasmid sequence reported in clinically isolated En. casseliflavus. The plasmid with the greatest sequence identity with pEC369 was the plasmid of Enterococcus sp. FDAARGOS_375, followed by the plasmids of Enterococcus faecium strains F12085 and pRE25, whereas the sequence with the greatest identity to the resistance genes carrying a transposon of pEC369 was on the chromosome of Staphylococcus aureus strain GD1677.
Conclusion: The resistance profiles of En. casseliflavus EC369 might contribute to the resistance genes encoded on the plasmid. The fact that the most similar sequence to the transposon carrying resistance genes of pEC369 was encoded in the chromosome of a S. aureus strain provides insights into the mechanism of dissemination of multidrug resistance between bacteria of different species or genera through horizontal gene transfer.
Keywords: Enterococcus casseliflavus, antimicrobial resistance, transposon, molecular characteristics, comparative genomics analysis
Introduction
Enterococci are gram-positive and facultative anaerobic organisms. They can grow under 6.5% NaCl concentrations and in a high-pH environment, and hydrolyze bile-esculin and L-pyrrolidonyl-B-naphthylamide.1 Enterococci are usually characterized by individual, paired, or short-chain gram-positive catalase-negative cocci.2 The Enterococcus genus was considered to belong to Lancefield group D Streptococcus; however, DNA homology studies have suggested that it is a distinct genus. To date, >40 Enterococcus species have been described and constitute a widespread group of bacteria.3 Enterococci have usually been found in the intestines of humans and animals, on the surfaces of plants, and in dairy products.4 They can persist in the environment because of their ability to survive under a wide range of harsh conditions, for instance, drying, extreme temperatures, high osmolarity, and the presence of disinfectants.5 Moreover, enterococci are also used in food production as probiotic products to monitor fecal contamination.6
Enterococcus spp. are an increasingly common cause of nosocomial infections, with Enterococcus faecalis and Enterococcus faecium accounting for the majority of human enterococcus infections.7 Other Enterococcus spp., including Enterococcus casseliflavus, have also been shown to be pathogenic to humans.8 En. casseliflavus was given species status in 1984.9 It can be motile and produces a yellow pigment.10 However, nonpigmented and nonmotile strains may also occur.11 The species En. casseliflavus was formerly thought to be associated primarily with vegetation.12 McGowan found that 9 of 27 (33.3%) tomato samples harbored Enterococcus spp.13 En. casseliflavus is among the normal flora in human and animal gastrointestinal tracts, unlike other enterococci, such as En. faecium and En. faecalis, which are the predominant conditionally pathogenic bacteria that cause hospital-acquired infections. En. casseliflavus is not frequently isolated from clinical specimens, with the rates being <1.3%.14 It is an opportunistic pathogen, which means it targets persons who are immunocompromised or chronically ill and is sometimes nosocomially acquired.15 Although the infection of En. casseliflavus is not common, it can be seriously invasive if infection occurs. Recently, En. casseliflavus has been increasingly implicated in infections and hospital outbreaks.16 En. casseliflavus is associated with a wide variety of invasive infections in humans, such as endocarditis, bacteremia, endophthalmitis, and spontaneous bacterial peritonitis.17–19
Enterococcus possesses intrinsic or acquired resistance to antimicrobials.20 The acquired resistance is usually mediated by mobile genetic elements such as insertion elements, transposons, and bacteriophages, which carry a pool of resistance genes and transmit the resistance between the bacteria of different species or genera via horizontal gene transfer.21 Enterococci also possess an inherent resistance to antimicrobial agents, for instance, low to moderate levels of resistance to aminoglycosides and low levels of resistance to vancomycin.22 It is possible that enterococci have the potential for resistance to virtually all clinically useful antibiotics. The current study was performed to introduce the molecular characteristics of a clinical En. casseliflavus with a plasmid carrying several resistance genes. Genome analysis has illuminated the extent of resistance genes related to the mobile genetic elements and the evolution of antibiotic resistance.
Materials and methods
Bacterial strain
En. casseliflavus EC369 was isolated from a bile specimen of a patient in Lishui Hospital, Zhejiang, China. The strain was identified using the Vitek-60 microorganism autoanalysis system (BioMerieux Corporate, Craponne, France). Further verification was performed using homologous comparisons of the sequences of 16S rRNA genes and the whole-genome sequence of the bacteria from the National Center for Biotechnology Information (NCBI) nucleotide database by BlastN and BlastP programs (http://www.ncbi.nlm.nih.gov). The bacteria and plasmids used in this work are listed in Table 1.
  | Table 1 Strains and plasmids used in this work Abbreviations: CHL, chloramphenicol; GM, gentamicin; ORFs, open reading frames; r, resistance; RF, rifampin; AP, ampcillin. |
Antimicrobial susceptibility testing
The minimum inhibitory concentrations (MICs) of the antibiotics for the bacteria were determined by the agar dilution method in accordance with the guidelines of the Clinical and Laboratory Standards Institute (CLSI document M100-S27, 2017). In the evaluation of MICs, resistance to streptothricin was set at ≥1,024 µg/mL, according to Werner et al.23 The MIC was defined as the lowest concentration producing no visible growth. Escherichia coli ATCC 25922 and En. faecalis ATCC 29212 served as the quality control strains for the antimicrobial susceptibility tests.
Whole-genome sequencing
Bacterial DNA was extracted using the Generay Genomic DNA Miniprep kit (Shanghai Generay Biotech Co., Ltd, Shanghai, China) from a single colony subcultured in brain heart infusion broth at 37°C for 16 hours, following the manufacturer’s instructions. Genomic DNA was sequenced with Illumina HiSeq-2500 and Pacific Bioscience sequencers at Annoroad Genomics Technology Co., Ltd (Beijing, China). Reads derived from the HiSeq-2500 sequencing were initially assembled de novo with the SOAPdenovo software to obtain contigs of the genome sequences. Pacific Bioscience sequencing reads of ~10–20 kb in length were mapped onto the primary assembly to scaffold the contigs. The gaps were filled either by remapping the short reads from HiSeq-2500 sequencing or by PCR product sequencing of the gap. Glimmer (http://ccb.jhu.edu/software/glimmer) was used to predict protein-coding genes with potential open reading frames (ORFs) >150 bp. GView was used to construct the basic genome features.24 BlastX (https://blast.ncbi.nlm.nih.gov) was used to annotate the predicted protein-coding genes against a nonredundant protein database with an e-value threshold of 1e–5.
Cloning experiments
The resistance gene sequences were PCR amplified, and the PCR products were then eluted from agarose gel and ligated into suitable vectors (pUCP20, pUCP24, or pAM401). The ORFs of the resistance genes were ligated into the pUCP20 or pUCP24. The resistance genes with the predicted promoter regions (http://www.softberry.com/berry.phtml?topic=bprom&group=programs&subgroup=gfindb) were ligated into pAM401. We designed the primers (Table 2) by using Primer Premier 5.0 and then synthesized them at Shanghai Sunny Biotechnology Co., Ltd (Shanghai, China). The recombinant plasmids (pUCP20-ORFs) were transformed into Es. coli DH5α via the calcium chloride method, and the bacterial colonies were grown on Luria-Bertani agar plates supplemented with ampicillin (100 µg/mL). The recombinant plasmids (pUCP24-ORFs) were transformed into Es. coli DH5α using the same method, but were grown on Luria-Bertani agar plates supplemented with gentamicin (20 µg/mL). Additionally, the recombinant plasmids (pAM401-ORFs) were transformed into En. faecalis JH2-2 by electrotransformation, and the bacterial colonies were grown on brain heart infusion agar plates supplemented with chloramphenicol (16 µg/mL). The recombinant plasmids (pUCP20-ORFs and pUCP24-ORFs) were extracted and digested with KpnI and BamHI (Takara Biomedical Technology, Beijing Co., Ltd, Beijing, China) and the recombinant plasmids (pAM401-ORFs) were digested with XbaI and BamHI (TaKaRa) to confirm the insert size, and then, the orientation and frame of the ORFs were further verified by sequencing with an ABI 3730 automated sequencer (Thermo Fisher Scientific, Waltham, MA, USA).
  | Table 2 Primers used in this study Note: aThe underlines represent the restriction enzyme sites and their protective bases; bThe primers with the predicted promotor regions. |
Conjugation experiments
En. faecalis JH2-2 was used as the recipient in the conjugation experiments to detect the transferable characteristics of the conjugative plasmid of the donor En. casseliflavus EC369 by filter mating, which allows tight cell-to-cell contact, as previously described.25,26 The transconjugants were selected on brain heart infusion plates supplemented with 25 µg/mL rifampicin (Sinopharm Chemical Reagent Co., Ltd, Shanghai, China), 50 µg/mL fusidic acid (Sinopharm Chemical Reagent Co., Ltd), and 512 µg/mL kanamycin (Sinopharm Chemical Reagent Co., Ltd). The plasmid DNA was extracted from the transconjugant (pEC369/En. faecalis JH2-2) and verified by PCR of the resistance genes and PCR products sequencing.
Collection and processing of the plasmids and the resistance gene-related sequences
The plasmid genome sequences for comparative genomics analysis in this study were selected based on the whole-genome sequence (pEC369) comparison against the whole-genome sequences available in the NCBI nucleotide database with coverage of >33%. The accession numbers of the plasmids were CP023514 (a plasmid from Enterococcus sp. FDAARGOS_375), X92945 (pRE25 from En. faecalis), and KY579372 (the plasmid of En. faecium strain F12085).
For the comparative genomics analysis of the resistance gene-related transposons, similar sequences were also obtained from the NCBI nucleotide database using the resistance gene-encoded region of pEC369 as the query. The resulting sequence was filtered and the sequence with an identity of >95% and coverage of >80% was retained. Moreover, we searched a plasmid pVEF1 (39 kb in length) that was free of the resistance gene related transposon, but shared about 15.6kb sequence with pEC369 which accounted for 40% (15.6/39) of the genome of pVEF1. The accession number of the plasmid was AM296544 (pVEF1 from En. faecium). Orthologous groups of genes from the candidate sequences were identified using BlastP and Paranoid.27 The sequence retrieval, statistical analysis, and other bioinformatics tools used in this study were applied with Python and Biopython scripts.28
Results and discussion
General features of the En. casseliflavus EC369 genome
En. casseliflavus EC369, producing yellow pigment on the plate, had the typical biological characteristics of a gram-positive Enterococcus. 16S ribosomal RNA gene homology analysis showed that the genes sharing the greatest nucleotide sequence identities to that of En. casseliflavus EC369 were from En. casseliflavus LMG10745 (NR114778) and two Enterococcus gallinarum strains (LMG 13129, NR104559; NBRC 100675, NR113964). They all showed sequence identities of 99%. The genome sequences sharing the highest identities to that of En. casseliflavus EC369 were from En. casseliflavus EC20 (CP004856, 98.0%) and Enterococcus sp. FDAARGOS_375 (CP023515, 98.0%). Therefore, we finally grouped the strain into a species of En. casseliflavus and named it En. casseliflavus EC369.
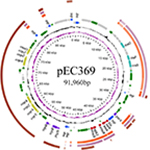
The whole genome of En. casseliflavus EC369 consisted of a chromosome (CP032739) of 3.58 Mb in length encoding 3,333 ORFs and a circular plasmid (pEC369, CP032740) of 91,960 bp in length encoding 95 ORFs. Of the ORFs encoded on the plasmid, 65% (62/95) were predicted to encode proteins with known functions, including a transposon carrying six antimicrobial genes (aph3′, ant6, bla, sat4, and two ermB genes), two clusters of copper resistance genes, and so on (Table 3; Figure 1). At present, there are ~29 genome sequences of En. casseliflavus available in the NCBI GenBank. Most of these 29 genome sequences are the incomplete genome sequences. In addition to the complete genome sequence of En. casseliflavus EC369 in this work, only one strain, En. casseliflavus EC20 (CP004856), had the complete genome sequence. Even though a variety of complete plasmid sequences have been reported in other species of the Enterococcus, such as En. faecium and En. faecalis, no complete plasmid sequence, however, from En. casseliflavus was available in the database.
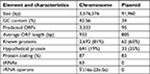  | Table 3 General features of Enterococcus casseliflavus EC369 Abbreviation: ORFs, open reading frames. |
The resistance genes and their functions in En. casseliflavus EC369
En. casseliflavus, similar to most enterococci, showed resistance to a variety of antibiotics, such as glycopeptides, aminoglycosides, macrolides, tetracycline, and beta-lactams.29–31 The resistance mechanisms are related to both intrinsic and acquired resistance genes. The results of MIC detection of several antibiotics showed that it was resistant to kanamycin, streptomycin, and erythromycin and susceptible to vancomycin, ampicillin, streptothricin, and other antimicrobials (Table 4). Six resistance genes (aph3¢, ant6, sat4, bla, and two ermBs with the same sequences) were identified on the plasmid pEC369. Among them, the resistance genes aph3¢, ant6, and ermB were functional, but sat4 and bla were not (the cloned genes with or without the promoter region). The cloned ORFs of the ant6 and aph3¢ genes exhibited 2- and 5-fold increases in the MIC levels to streptomycin and kanamycin, respectively, compared with those of the controls (DH5α or DH5α carrying the vector pUCP20). The gene ermB exhibited 4-fold increases in the MIC levels to erythromycin. When the ORFs of the resistance genes (ant6, aph3′, and ermB) were cloned with their predicted promoter regions and transformed into the En. faecalis JH2-2 recipients, they exhibited at least 4-fold increases in MIC levels to streptomycin, kanamycin, and erythromycin, respectively, compared with those of the control pAM401/JH2-2 (Table 4). The results of the function detection of the cloned resistance genes were in accordance with the resistance phenotypes of the host strain En. casseliflavus EC369.
It has been reported that En. casseliflavus is intrinsically resistant to vancomycin at low levels, and that resistance is mediated by the vanC-2-type genes.32 Some enterococci strains showed high-level resistance to vancomycin and teicoplanin (MIC >256 mg/L), coinciding with the presence of the vanA gene.33 The mechanisms of glycopeptide resistance in enterococci have been sufficiently investigated.34 To date, nine vancomycin resistance-related genes (vanA, vanB, vanC, vanD, vanE, vanG, vanL, vanM, and vanN) have been identified, with the acquired genotype vanA being the most common, followed by vanB. VanA is related to resistance to both vancomycin and teicoplanin, whereas VanB is resistant only to vancomycin. VanC is usually encoded by the vanC1 and vanC2/3 genes, which are intrinsic to En. gallinarum and En. casseliflavus, respectively.35 This characteristic can be used for species identification.36
The vanC-2 containing vancomycin resistance gene cluster of En. casseliflavus consists of five genes. The first three genes of the cluster, vanC-2, vanXYC-2, and vanTC-2, are essential for vancomycin resistance. vanc-2 encodes a D-Ala:D-Ser ligase, vanXYC-2 encodes a protein possessing both D, D-dipeptidase and d, D-carboxypeptidase activities, and vanTC-2 encodes a serine racemase. Expression of the resistance genes is controlled by a two-component regulatory system that is present downstream of vanT C-2 consisting of a response regulator, VanRC-2, and a histidine kinase, VanSC-2.37 In this work, a cluster of five vancomycin resistance-related genes (vanC-2, vanXY, vanT, vanR, and vanS) were all identified in the chromosome genome. However, EC369 showed a low resistance level to vancomycin with an MIC of 1 µg/mL. We may think that the vancomycin resistance genes in En. casseliflavus EC369 are not functional and the reason for this remains a question.
The transposon carrying multiple resistance genes on a conjugative plasmid
The acquisition of foreign resistance genes in Enterococcus is often related to transfer of the resistance plasmids that carry antibiotic resistance genes. Studies have reported that plasmids are abundant in enterococci, as illustrated by the finding of one to seven plasmids in 88 out of 93 En. faecium isolates and high resistance plasmid carriage rates observed in En. casseliflavus isolates.38,39 En. casseliflavus once showed high-level resistance to vancomycin due to the acquisition of plasmid pIP218 carrying the vanA gene.40 The plasmid-mediated aminoglycoside-modifying enzyme gene, aph (2”)-Ie, appeared in a strain of high-level gentamicin-resistant En. casseliflavus.41 A conjugative plasmid with transposon Tn6000 related to the tetracycline resistance gene was identified in En. casseliflavus 664.1H1.42 In this work, a cluster of resistance genes including two aminoglycoside resistance genes (aph(3’)-III and ant(6)-Ia) and two erythromycin resistance genes ermB was identified encoded on the plasmid. The plasmid (pEC369) was a conjugative plasmid and could be successfully transferred into the recipient cell through conjugation. The MIC levels of the transconjugant to the antibiotics detected were similar to those of the donor cell En. casseliflavus EC369 (Table 4). It demonstrated that the resistance activities of En. casseliflavus EC369 to the antibiotics streptomycin, kanamycin, and erythromycin were mainly related to the genes encoded in the conjugative plasmid pEC369 (Table 3).
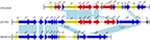
Resistance genes can translocate among plasmids or chromosomes through the mobile genetic elements such as transposons, insertion elements, integrons, and phages. The resistance genes (aph3¢, ant6, bla, sat4, and two ermB genes) encoded on pEC369 are carried by a transposon of ~7 kb in length. This transposon is characterized by two copies of a 5 bp direct repeat “GTGAT” that precisely borders the transposase gene and the peptide-binding protein gene (Figure 2). Further comparative genomics analysis demonstrated that the sequence with the highest identity to the transposon-carrying resistance genes of pEC369 was located on the chromosome of Staphylococcus aureus strain GD1677 (CP019595), and the sequence with the greatest identity to the upstream and downstream sequences of the resistance genes carrying the transposon of pEC369 was pVEF1 (AM296544), a plasmid in En. faecium. pVEF1 was 39 kb in length and 52 kb smaller than pEC369, and the region similar to the flanking sequences of the transposon of pEC369 was 15.6 kb in length, consisting of 40% (15.6/39 kb) of the pVEF1 genome. The transposon of pEC369 might have been fromed on the basis of the initial transposon of tnp-bla-sat4-ant6-ermB by the transposase (encoded by the tnp gene) of the transposon which captured the other two resistance genes (aph3’ and ermB) and the peptide-binding protein gene (orf E) (Figure 2).
Comparative genomics analysis of the plasmid
Comparative genomics analysis showed that the plasmid with the greatest sequence similarity to pEC369 was a plasmid (CP023514) from Enterococcus sp. FDAARGOS_375. It was 148 kb in length and 57 kb larger than pEC369. The two plasmids had nearly 38 kb (41%, 38/91.96) similar sequences. The similar regions were mainly located between 50 and 80 kb of the pEC369 and included two clusters of copper resistance genes (located in two regions of 57.5–63 kb and 71.5–77.7 kb, respectively), but the plasmid was free of the drug resistance-related transposon. Two other plasmids with higher similarities to pEC369 were pRE25 (X92945, 50 kb) and the plasmid of En. faecium strain F12085 (KY579372, 73 kb), and both had ~30 kb (33%, 30/91.96) sequences similar to that of pEC369. The regions of the two plasmids pRE25 and the plasmid of En. faecium strain F12085, similar to that of pEC369, were different from the plasmid from Enterococcus sp. FDAARGOS_375. The two plasmids shared nearly the same similar region, which spanned 10–45 kb of the pEC369, including the sequences of the drug resistance-related transposon and the genes related to conjugation (Figure 1). Although the plasmids with the greatest sequence similarity were the above three plasmids, the sequence most similar to that of resistance genes (transposon) was on the S. aureus chromosome, which suggested that exchanges of resistance genes may occur between these bacteria.
Conclusion
In this work, we sequenced the whole genome of the clinical isolate En. casseliflavus EC369. In addition to a resistance gene cluster of vancomycin, the genome encoded seven other resistance genes, of which a tet (M) was encoded in the chromosome and six resistance genes (aph3¢, ant6, bla, sat4, and two ermBs) were carried by a transposon encoded on the plasmid pEC369, which conferred the host high level of resistance to erythromycin, kanamycin, and streptomycin. Although the plasmid with the greatest sequence identity to pEC369 was the plasmid of Enterococcus sp. FDAARGOS_375, followed by the plasmids from the En. faecium strains, the sequence with the greatest identity to the resistance genes carrying the transposon of pEC369 was on the chromosome of S. aureus strain GD1677. This result demonstrates the potential for the spread of multiresistant mobile genetic elements within bacteria of different species or genera, which poses significant challenges for successful clinical treatment and infection control strategies. Therefore, the study of enterococcal biology and genetics is necessary and will undoubtedly contribute to our understanding of bacterial resistance dissemination.
Acknowledgments
We thank Professor Jianzhong Shen of the College of Veterinary Medicine, China Agricultural University for his kind donation of the strain Enterococcus faecalis JH2-2. This work was funded by grants from the Natural Science Foundation of Zhejiang Province (LY14C060005 and LQ17H190001), the Science and Technology Foundation of Zhejiang Province, China (2015C33196), and the National Natural Science Foundation of China (81401702, 81501808 and 81501780).
Disclosure
The authors report no conflicts of interest in this work.
References
Hollenbeck BL, Rice LB. Intrinsic and acquired resistance mechanisms in Enterococcus. Virulence. 2012;3(5):421–569. | ||
Castillo-Rojas G, Mazari-Hiríart M, Ponce de León S, et al. Comparison of Enterococcus faecium and Enterococcus faecalis strains isolated from water and clinical samples: antimicrobial susceptibility and genetic relationships. PLoS One. 2013;8(4):e59491. | ||
Yuen GJ, Ausubel FM. Enterococcus infection biology: lessons from invertebrate host models. J Microbiol. 2014;52(3):200–210. | ||
Pasquaroli S, di Cesare A, Vignaroli C, Conti G, Citterio B, Biavasco F. Erythromycin- and copper-resistant Enterococcus hirae from marine sediment and co-transfer of erm(B) and tcrB to human Enterococcus faecalis. Diagn Microbiol Infect Dis. 2014;80(1):26–28. | ||
Kuch A, Willems RJ, Werner G, et al. Insight into antimicrobial susceptibility and population structure of contemporary human Enterococcus faecalis isolates from Europe. J Antimicrob Chemother. 2012;67(3):551–558. | ||
Palmer KL, Godfrey P, Griggs A, et al. Comparative genomics of enterococci: variation in Enterococcus faecalis, clade structure in E. faecium, and defining characteristics of E. gallinarum and E. casseliflavus. MBio. 2012;3(1):e00318–00311. | ||
De Niederhäusern S, Bondi M, Anacarso I, et al. Antibiotics and heavy metals resistance and other biological characters in enterococci isolated from surface water of Monte Cotugno Lake (Italy). J Environ Sci Health A Tox Hazard Subst Environ Eng. 2013;48(8):939–946. | ||
Agudelo Higuita NI, Huycke MM. Enterococcal disease, epidemiology, and implications for treatment. In: Gilmore MS, Clewell DB, Ike Y, Shankar N, editors. Enterococci: From Commensals to Leading Causes of Drug Resistant Infection. Boston; 2014. 1–35. | ||
Liu Y, Wang Y, Dai L, Wu C, Shen J. First report of multiresistance gene cfr in Enterococcus species casseliflavus and gallinarum of swine origin. Vet Microbiol. 2014;170(3-4):352–357. | ||
Facklam RR, Collins MD. Identification of Enterococcus species isolated from human infections by a conventional test scheme. J Clin Microbiol. 1989;27(4):731–734. | ||
Pappas G, Liberopoulos E, Tsianos E, Elisaf M. Enterococcus casseliflavus bacteremia. Case report and literature review. J Infect. 2004;48(2):206–208. | ||
Mundt JO, Graham WF. Streptococcus faecium var. casselifavus, nov. var. J Bacteriol. 1968;95(6):2005–2009. | ||
Micallef SA, Goldstein RE, George A, et al. Diversity, distribution and antibiotic resistance of Enterococcus spp. recovered from tomatoes, leaves, water and soil on U.S. Mid-Atlantic farms. Food Microbiol. 2013;36(2):465–474. | ||
Liu Y, Liu K, Lai J, Wu C, Shen J, Wang Y. Prevalence and antimicrobial resistance of Enterococcus species of food animal origin from Beijing and Shandong Province, China. J Appl Microbiol. 2013;114(2):555–563. | ||
Abamecha A, Wondafrash B, Abdissa A. Antimicrobial resistance profile of Enterococcus species isolated from intestinal tracts of hospitalized patients in Jimma, Ethiopia. BMC Res Notes. 2015;8:213. | ||
Tan CK, Lai CC, Wang JY, et al. Bacteremia caused by non-faecalis and non-faecium enterococcus species at a Medical center in Taiwan, 2000 to 2008. J Infect. 2010;61(1):34–43. | ||
Narciso-Schiavon JL, Borgonovo A, Marques PC, et al. Enterococcus casseliflavus and Enterococcus gallinarum as causative agents of spontaneous bacterial peritonitis. Ann Hepatol. 2015;14(2):270–272. | ||
Choi SH, Lee SO, Kim TH, et al. Clinical features and outcomes of bacteremia caused by Enterococcus casseliflavus and Enterococcus gallinarum: analysis of 56 cases. Clin Infect Dis. 2004;38(1):53–61. | ||
Khurana RN, Leder HA, Nguyen QD, Do DV. Enterococcus casseliflavus endophthalmitis associated with a horse tail injury. Arch Ophthalmol. 2009;127(11):1547–1552. | ||
Rubinstein E, Keynan Y. Vancomycin-resistant enterococci. Crit Care Clin. 2013;29(4):841–852. | ||
Garcia-Ramon DC, Luque-Navas MJ, Molina CA, del Val C, Osuna A, Vilchez S. Identification, sequencing and comparative analysis of pBp15.S plasmid from the newly described entomopathogen Bacillus pumilus 15.1. Plasmid. 2015;82:17–27. | ||
Arias CA, Murray BE. The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol. 2012;10(4):266–278. | ||
Werner G, Hildebrandt B, Witte W. Linkage of erm(B) and aadE-sat4-aphA-3 in multiple-resistant Enterococcus faecium isolates of different ecological origins. Microb Drug Resist. 2003;9(Suppl 1):9–16. | ||
Petkau A, Stuart-Edwards M, Stothard P, van Domselaar G. Interactive microbial genome visualization with GView. Bioinformatics. 2010;26(24):3125–3126. | ||
He T, Shen Y, Schwarz S, et al. Genetic environment of the transferable oxazolidinone/phenicol resistance gene optrA in Enterococcus faecalis isolates of human and animal origin. J Antimicrob Chemother. 2016;71(6):1466–1473. | ||
Laverde Gomez JA, van Schaik W, Freitas AR, et al. A multiresistance megaplasmid pLG1 bearing a hylEfm genomic island in hospital Enterococcus faecium isolates. Int J Med Microbiol. 2011;301(2):165–175. | ||
Remm M, Storm CE, Sonnhammer EL. Automatic clustering of orthologs and in-paralogs from pairwise species comparisons. J Mol Biol. 2001;314(5):1041–1052. | ||
Cock PJ, Antao T, Chang JT, et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics. 2009;25(11):1422–1423. | ||
Monticelli J, Knezevich A, Luzzati R, di Bella S. Clinical management of non-faecium non-faecalis vancomycin-resistant enterococci infection. Focus on Enterococcus gallinarum and Enterococcus casseliflavus/flavescens. J Infect Chemother. 2018;24(4):237–246. | ||
Lebreton F, Willems RJL, Gilmore MS. Enterococcus diversity, origins in nature, and gut colonization. In: Gilmore MS, Clewell DB, Ike Y, Shankar N, editors. Enterococci: From Commensals to Leading Causes of Drug Resistant Infection. Boston; 2014. 1–59. | ||
Jia W, Li G, Wang W. Prevalence and antimicrobial resistance of Enterococcus species: a hospital-based study in China. Int J Environ Res Public Health. 2014;11(3):3424–3442. | ||
Iweriebor BC, Obi LC, Okoh AI. Macrolide, glycopeptide resistance and virulence genes in Enterococcus species isolates from dairy cattle. J Med Microbiol. 2016;65(7):641–648. | ||
Haenni M, Saras E, Châtre P, et al. vanA in Enterococcus faecium, Enterococcus faecalis, and Enterococcus casseliflavus detected in French cattle. Foodborne Pathog Dis. 2009;6(9):1107–1111. | ||
Yang JX, Li T, Ning YZ, et al. Molecular characterization of resistance, virulence and clonality in vancomycin-resistant Enterococcus faecium and Enterococcus faecalis: a hospital-based study in Beijing, China. Infect Genet Evol. 2015;33:253–260. | ||
Moura TM, Cassenego AP, Campos FS, et al. Detection of vanC1 gene transcription in vancomycin-susceptible Enterococcus faecalis. Mem Inst Oswaldo Cruz. 2013;108(4):453–456. | ||
Britt NS, Potter EM. Clinical epidemiology of vancomycin-resistant Enterococcus gallinarum and Enterococcus casseliflavus bloodstream infections. J Glob Antimicrob Resist. 2016;5:57–61. | ||
Dutta I, Reynolds PE. The vanC-3 vancomycin resistance gene cluster of Enterococcus flavescens CCM 439. J Antimicrob Chemother. 2003;51(3):703–706. | ||
Radu S, Toosa H, Rahim RA, et al. Occurrence of the vanA and vanC2/C3 genes in Enterococcus species isolated from poultry sources in Malaysia. Diagn Microbiol Infect Dis. 2001;39(3):145–153. | ||
Palmer KL, Kos VN, Gilmore MS. Horizontal gene transfer and the genomics of enterococcal antibiotic resistance. Curr Opin Microbiol. 2010;13(5):632–639. | ||
Dutka-Malen S, Blaimont B, Wauters G, Courvalin P. Emergence of high-level resistance to glycopeptides in Enterococcus gallinarum and Enterococcus casseliflavus. Antimicrob Agents Chemother. 1994;38(7):1675–1677. | ||
Qu TT, Zhang Y, Yu YS, Chen YG, Wei ZQ, Li LJ. A novel plasmid-mediated aminoglycosides-modifying enzyme gene, aph (2’’)-Ie, in a strain of high-level gentamicin resistant Enterococcus casseliflavus. Zhonghua Yi Xue Za Zhi. 2006;86(9):596–599. | ||
Brouwer MSM, Mullany P, Roberts AP. Characterization of the conjugative transposon Tn6000 from Enterococcus casseliflavus 664.1H1 (formerly Enterococcus faecium 664.1H1). FEMS Microbiol Lett. 2010;67(1):76. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.