Back to Journals » OncoTargets and Therapy » Volume 13
MOB1 Inhibits Malignant Progression of Colorectal Cancer by Targeting PAK2
Authors Liu J, Shi Z, Ma Y, Fu L, Yi M
Received 11 March 2020
Accepted for publication 3 August 2020
Published 3 September 2020 Volume 2020:13 Pages 8803—8811
DOI https://doi.org/10.2147/OTT.S253470
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Jianmin Xu
Jie Liu,1 Zhitao Shi,2 Yunyun Ma,1 Liang Fu,1 Man Yi1
1Department of Proctology, Affiliated Traditional Chinese Medicine Hospital, Xinjiang Medical University, Urumqi, People’s Republic of China; 2Department of General Surgery, Affiliated Traditional Chinese Medicine Hospital, Xinjiang Medical University, Urumqi, People’s Republic of China
Correspondence: Jie Liu
Department of Proctology, Affiliated Traditional Chinese Medicine Hospital, Xinjiang Medical University, Urumqi 830000, People’s Republic of China
Tel +8613999188169
Email [email protected]
Objective: We aimed at studying the mechanism of MOB1 inhibiting the proliferation and metastasis of colorectal cancer (CRC), to provide a new guidance for the early diagnosis and treatment of CRC.
Methods: MOB1 expression level in 68 pairs of CRC tissues and adjacent ones was detected by quantitative real-time polymerase chain reaction (qRT-PCR) analysis, and the associations between the expression level of MOB1 and the clinicopathological indicators as well as the prognosis of CRC patients were analyzed. After constructing CRC cell lines that stably overexpressing or silencing MOB1, the changes of cell proliferation and metastasis ability were examined by Cell Counting Kit (CCK-8) and Transwell assay. In addition, the interaction between MOB1 and PAK2 and how the these two genes affect the biological functions of CRC cell lines were investigated by luciferase assay, qRT-PCR and Western Blot experiments.
Results: Our data showed that MOB1 expression level in CRC tissues was remarkably lower than that in adjacent ones. In comparison to patients of the group of high MOB1 expression, these patients of low MOB1 expression group showed higher incidence of distant or lymph node metastasis and lower survival rate. Cell functional experiments revealed that overexpression of MOB1 markedly attenuated the proliferation and migration ability of CRC cell lines compared to the NC group; In contrast, knockdown of MOB1 enhanced the above-mentioned cell abilities compared to anti-NC group. Luciferase assay verified an interaction between MOB1 and PAK2; and Western blot analysis showed a negative correlation between the expression of the MOB1 and PAK2 protein levels in CRC tissues. Subsequently, we demonstrated that MOB1 interacted with PAK2 to regulate its expression and affected the proliferation and migration capacity of CRC cell lines in vitro.
Conclusion: In summary, the lowly expressed MOB1 in CRC tissues and cell lines may accelerate the proliferation and migration through modulating PAK2 expression.
Keywords: MOB1, PAK2, CRC, malignant progression
Introduction
Colorectal cancer (CRC) is a common malignant disease of the digestive system, with the number of new cases reaches 930,000 worldwide each year and 160,000 in China. So far, it has become the third most common cancer in China and the fourth leading cause of cancer death in the world.1–4 According to cancer statistics, approximately 147,950 individuals will be diagnosed with CRC and 53,200 will die from the disease, including 17,930 cases and 3640 deaths in individuals aged younger than 50 years in the United States in 2020.2,4 As we know, the development of CRC follows multiple pathways: benign adenomatous polyps that start in the inner wall of the colorectal, gradually develop into advanced adenomas and carcinomas in situ, invasive carcinomas and eventually distant metastases develop into advanced tumors.5,6 Studies have shown that the survival time of CRC patients with tumor metastasis significantly decreased, and the five-year survival time was only about 14%, which seriously affected the prognosis of CRC patients.7,8 Therefore, we need thoroughly identify the specific mechanism of CRC development and metastasis to provide some clues for exploring better therapies and improving patients’ prognosis.9,10
MOB1 (Mps One binder 1), a member of a protein family that are highly conserved from yeast to humans, has been found to be involved in the mitotic process of yeast and cell survival, proliferation, differentiation and organ formation through LATS/NDR kinase activity.11,12 Studies have proved that MOB1 can effectively promote the elongation of neuronal axons by regulating the axon skeleton, namely microtubules and microfilaments.13 In addition, the deletion of MOB1 gene in lung tissues can induce dephosphorylation of LATS1, stabilize YAP1 protein, and prevent the differentiation of bronchoalveolar stem cells into bronchial epithelial cells.14 Silencing the warts/lats gene in the brain of Drosophila can stable Yorkie/YAP protein and reduce neural stem cell differentiation, suggesting that LATS kinase is capable of promoting YAP phosphorylation and accelerating neural stem cell differentiation.15 Recent studies have suggested that MOB1 may be engaged in the regulation of proliferation, apoptosis, migration and invasion of pancreatic cancer, breast cancer and other malignant tumor cells.16,17 Therefore, the study of MOB1 in CRC not only helps to clarify the pathogenesis of this tumor but also provides valuable targets for its diagnosis and treatment.
In this study, bioinformatics analysis revealed a potential interaction between MOB1 and PAK2, which is a downstream effector of GTPase in the Rho family and plays an essential part in modulating cell proliferation, cell movement and cell apoptosis. The dysfunction of PAK2 can lead to various diseases, including tumors. Therefore, in this study, we detected the differential expressions of MOB1 and PAK2 in CRC, and explored the impact of MOB1 on biological functions of CRC cell lines.
Methods
Patients and CRC Samples
Tumor tissue specimens and adjacent ones of 68 CRC patients undergoing radical surgery were collected. All subjects had not received any radiotherapy or chemotherapy before surgery. Among them, there were 48 patients with Adenocarcinoma; 12 patients with adenosquamous carcinoma; 6 patients with squamous cell carcinoma; and 2 patients with undifferentiated carcinoma. CRC pathological classification and staging criteria are implemented in accordance with the International Union Against Cancer (UICC) CRC staging criteria. Patients and their families signed informed consent. This study was approved by the ethics committee of Affiliated Traditional Chinese Medicine Hospital, Xinjiang Medical University. This study complies with the Helsinki Declaration Clinical Practice Guidelines.
Cell Culture
Human colon cancer cell lines (HT-29, HCT-116, HCT-8, Caco-2, SW620) and normal human intestinal epithelial cell line (FHC) were provided by ATCC (Manassas, VA, USA). HT-29 and HCT-8 were cultured with Dulbecco’s modified eagle medium (DMEM) (Thermo Fisher Scientific, Waltham, MA, USA) containing 10% fetal bovine serum (FBS) (Gibco, Rockville, MD, USA), penicillin (100 U/mL), and streptomycin (100 ug/mL), while HCT-116, SW620 and Caco-2 were cultured with Roswell Park Memorial Institute 1640 (RPMI 1640) medium. All cells were cultured in a 37°C cell incubator with 5% CO2.
Transfection
To regulate MOB1 and PAK2 expression, oligonucleotides and plasmids were constructed. The following siRNAs targeting MOB1 were designed by RiboBio (Guangzhou, China). Fulllength MOB1 was cloned into the pEX-3 (GenePharma, Shanghai, China) overexpression vector. The PAK2 gene was cloned into pLVX plasmids and siRNA. The oligonucleotides and plasmids were transfected into cells with Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. These cells were collected 48 hours later for cell function experiments.
Cell Counting Kit-8 (CCK-8) Test
Cells were collected 48 h after transfection and seeded into 96-well plates (3000 cells/well). Cell Counting Kit-8 (CCK-8) assay was conducted based on instructions.
Transwell Assay
Cell migration capacity was tested using a 24-well plate cell pre-coated or not coated with matrix gel according to the manufacturer’s instructions.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
One milliliter of TRIzol (Invitrogen, Carlsbad, CA, USA) was used to lyse the cells to extract total RNA from the CRC tissue and cell lines. Real-time PCR was performed according to the instructions of SYBR® Premix Ex Taq ™ (Takara, Tokyo, Japan) kit, with glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as internal reference. The following primers were used in the qRT-PCR reaction: MOB1: forward, 5′-AAAATTTTCATATAAGTCCCGGCCA-3′; reverse, 5′-GGGAGTTGCCTTAGTAGGCG-3′; PAK2: forward: 5ʹ-TGCCCGGGGCCATTTCATAAT-3ʹ; reverse: 5ʹ-GCCTCCAGTGCTAAAGATGGT-3ʹ; GAPDH: forward, 5′-GGTGAAGGTCGGAGTCAACG-3′; reverse, 5′-CCATGTAGTTGAGGTCAATGAAG-3′.
Western Blot
Cells were lysed, shaken on ice for 30 minutes, and centrifuged at 4°C, 14,000 × g for 15 minutes. Total protein concentration was calculated by bicinchoninic acid (BCA) protein assay kit (Beyotime, Shanghai, China) to detect the expression level of MOB1 from the CRC tissue and cell lines. The extracted proteins were separated on a 10% Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gel and then transferred to a polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA, USA). Western blot analysis was carried out based on standard procedures. Primary antibodies against MOB1, PAK2 and GAPDH, and the secondary antibodies were provided by Cell Signaling Technology (Danvers, MA, USA).
Dual-Luciferase Reporter Assay
The transcription factor expression plasmid to be tested was co-transfected with the reporter plasmid into the CRC cell lines. The luciferase activity was measured using a luciferase reporter kit (Promega, Madison, WI, USA).
Statistical Analysis
GraphPad Prism 5 V5.01 software (La Jolla, CA, USA) was used for statistical analysis. Differences between two groups were analyzed by using the Student’s t-test. Comparison between multiple groups was done using one-way ANOVA test followed by Post Hoc Test (Least Significant Difference). Each experiment was repeated at least three independent experiments. P<0.05 indicated the significant difference.
Results
Low Expression of MOB1 in CRC
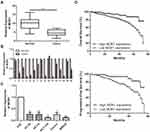
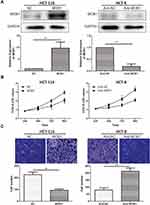
QRT-PCR results detected a significant reduction in MOB1 expression in the collected CRC tumor tissue samples as compared to paracancerous ones (Figure 1A, 1B). Meanwhile, compared with FHC, MOB1 was also remarkably lowly expressed in CRC cell lines (Figure 1C), suggesting that MOB1 may serve as a tumor suppressor gene in CRC cells.
MOB1 Expression Was Correlated with Pathological Staging and Metastasis Incidence of CRC Patients
We divided the 68 pairs of tumor tissue collected from CRC patients into high and low MOB1 expression group, and further explored the association of MOB1 with gender, age, pathological stage and incidence of metastasis of CRC patients. Table 1 indicates that compared with high-expressed MOB1, low-expressed MOB1 showed a positive correlation with metastasis incidence and pathological stage of CRC patients, but not with the other two indexes, suggesting that MOB1 may act as a new biological indicator to predict the malignant progress of CRC. In addition, Kaplan–Meier survival curve showed that disease-free survival or overall survival of CRC patients with low MOB1 expression was remarkably lower than those with high MOB1 expression (Figure 1D).
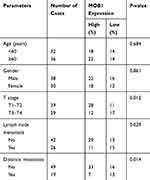 |
Table 1 Association of MOB1 Expression with Clinicopathologic Characteristics of Colorectal Cancer |
MOB1 Inhibited Proliferation and Migration of CRC Cell Lines
To specify the impact of MOB1 on CRC cell functions, we constructed MOB1 knockdown and overexpression lentiviral vectors in CRC cell lines HCT-8 and HCT-116, respectively, and verified the transfection efficiency by qRT-PCR and Western Blot experiments (Figure 2A). Subsequently, CCK8 assay showed that the proliferation rate of over-expressing MOB1 in colorectal cancer cell lines was about 2.4 times that of the control group; however, the proliferation rate of knockdown of MOB1 in colorectal cancer cell lines was about 0.3 times that of the control group (Figure 2B). Transwell experiments demonstrated that over-expressing MOB1 significantly reduced the number of perforating cells in the Transwell chamber; and the opposite results were observed after knockdown of MOB1 (Figure 2C). The above observations suggest that MOB1 is capable of suppressing the migration and proliferation capacities of CRC cells.
MOB1 Was Bound to PAK2
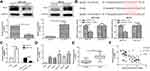
We then examined the changes in PAK2 mRNA and protein expression level after upregulation/downregulation of MOB1 in HCT-116/HCT-8 cell lines, respectively. As a result, PAK2 level was markedly increased after knockdown of MOB1, and the converse result was obtained after overexpression of MOB1 (Figure 3A). Further, luciferase assay was conducted and the binding relationship between PAK2 and MOB1 were verified (Figure 3B). Subsequently, MOB1 expression was detected after constructing PAK2 knockdown and overexpression vectors in CRC cell lines HCT-8 and HCT-116. It was found that knockdown of PAK2 significantly increased MOB1 mRNA expression (Figure 3C) while overexpression of PAK2 decreased the expression of it (Figure 3D). In addition, PAK2 also showed a significant increased expression in CRC tissues in comparison to paracancerous ones (Figure 3E). Finally, in CRC tissues, a negative correlation between above two proteins was shown in Figure 3F.
MOB1 Modulated PAK2 to Influence Proliferation and Migration of CRC Cell Lines
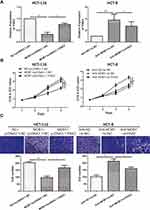
To further explore how MOB1 interacts with PAK2 and affects the malignant progression of CRC, we co-transfected MOB1 and PAK2 knockdown vectors in CRC cell lines. qRT-PCR results revealed that overexpression/knockdown of PAK2 reversed the decreased/increased the expression level of PAK2 induced by overexpression/knockdown of MOB1 (Figure 4A). Additionally, it was found by CCK-8 and Transwell experiments that silencing PAK2 reversed the promotion effect of MOB1 knockdown on the proliferation and migration ability of CRC cell lines, while overexpression of PAK2 led to an opposite result (Figure 4B and C).
Discussion
CRC is a heterogeneous disease induced by multiple causes which have not been clearly identified.1,2 At present, P53 gene mutation, KRAS gene mutation, APC gene mutation, MMR gene inactivation and other abnormalities are known to lead to the occurrence of colorectal cancer, which to a large extent provides people with a certain understanding of the molecular mechanism of the occurrence and development of colorectal cancer, and also provides a scientific basis for the clinical treatment of colorectal cancer.18,19 The basic treatment for CRC patients includes surgical treatment, neoadjuvant radiotherapy, adjuvant chemotherapy, etc., among which, surgical treatment is the most effective, yet approximately 40–50% of CRC patients develop distant metastasis at the time of diagnosis.6,7 Radical surgical treatment is not suitable for CRC patients with distant metastasis, leading to only 10% of 5-year survival rate.7,8 Tumor metastasis is a complex process involving multiple steps and factors, which is mostly responsible for the poor prognosis of CRC patients.3–6 In recent years, there have been numerous reports on CRC metastasis, but no effective treatment has been found. Therefore, it is necessary to study the molecular mechanism of CRC metastasis in depth and search for effective target molecules for CRC metastasis.9,10
Previous studies have reported that MOB1 expression might have great influence on the progression of some malignant tumors such as pancreatic cancer and breast cancer; however, whether it could affect the development of CRC still has not been studied.16,17 In this study, MOB1 expression was found remarkably upregulated in CRC tissues and cell lines. In addition, we found that patients with low MOB1 expression had higher metastasis incidence and pathological stage and lower progression-free survival rate, suggesting a worse prognosis. Meanwhile, experiments in vitro revealed that MOB1 was able to suppress the proliferation and migratory ability of CRC cells.
In this study, we predicted a potential binding relationship between MOB1 and PAK2 by bioinformatics analysis and further confirmed it by luciferase assay. Subsequently, qRT-PCR assay detected an increased expression of PAK2 in both CRC cell lines and tissues, suggesting a negative correlation between MOB1 and PAK2. Therefore, we suspected that MOB1 might inhibit the malignant progression of CRC through the modulation of PAK2. Subsequently, cell reverse experiments suggested that PAK2 silencing/overexpression could counteract the influence of MOB1 knockdown/overexpression on the migration and proliferation of CRC, revealing that the interaction between the MOB1 and PAK2 could be closely implicated in the occurrence of CRC.
Conclusions
In conclusion, the above data suggest that MOB1 promotes the malignant progression of CRC via regulating PAK2, and that MOB1/PAK2 crosstalk may play a key role in the proliferation and metastasis of CRC.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Gu MJ, Huang QC, Bao CZ, et al. Attributable causes of colorectal cancer in China. Bmc Cancer. 2018;18(1):38. doi:10.1186/s12885-017-3968-z
2. Brody H. Colorectal cancer. Nature. 2015;521(7551):S1. doi:10.1038/521S1a
3. Zhu J, Tan Z, Hollis-Hansen K, Zhang Y, Yu C, Li Y. Epidemiological trends in colorectal cancer in China: an Ecological Study. Dig Dis Sci. 2017;62(1):235–243. doi:10.1007/s10620-016-4362-4
4. Siegel RL, Miller KD, Sauer AG, et al. Colorectal cancer statistics, 2020. CA Cancer J Clin. 2020;70(3):145–164. doi:10.3322/caac.21601
5. Rodriguez-Salas N, Dominguez G, Barderas R, et al. Clinical relevance of colorectal cancer molecular subtypes. Crit Rev Oncol Hematol. 2017;109:9–19. doi:10.1016/j.critrevonc.2016.11.007
6. Gupta R, Sinha S, Paul RN. The impact of microsatellite stability status in colorectal cancer. Curr Probl Cancer. 2018;42(6):548–559. doi:10.1016/j.currproblcancer.2018.06.010
7. Jin M, Frankel WL. Lymph node metastasis in colorectal cancer. Surg Oncol Clin N Am. 2018;27(2):401–412. doi:10.1016/j.soc.2017.11.011
8. Huang D, Sun W, Zhou Y, et al. Mutations of key driver genes in colorectal cancer progression and metastasis. Cancer Metastasis Rev. 2018;37(1):173–187. doi:10.1007/s10555-017-9726-5
9. Yiu AJ, Yiu CY. Biomarkers in colorectal cancer. Anticancer Res. 2016;36(3):1093–1102.
10. Zhang X, Sun XF, Cao Y, et al. CBD: a biomarker database for colorectal cancer. Database (Oxford). 2018;2018. doi:10.1093/database/bay046
11. Quan M, Chen Z, Jiao F, et al. Lysine demethylase 2 (KDM2B) regulates hippo pathway via MOB1 to promote pancreatic ductal adenocarcinoma (PDAC) progression. J Exp Clin Cancer Res. 2020;39(1):13. doi:10.1186/s13046-019-1489-0
12. Song J, Wang T, Chi X, et al. Kindlin-2 inhibits the hippo signaling pathway by promoting degradation of MOB1. Cell Rep. 2019;29(11):3664–3677. doi:10.1016/j.celrep.2019.11.035
13. Lobo GP, Fulmer D, Guo L, et al. The exocyst is required for photoreceptor ciliogenesis and retinal development. J Biol Chem. 2017;292(36):14814–14826. doi:10.1074/jbc.M117.795674
14. Otsubo K, Goto H, Nishio M, et al. MOB1-YAP1/TAZ-NKX2.1 axis controls bronchioalveolar cell differentiation, adhesion and tumour formation. Oncogene. 2017;36(29):4201–4211. doi:10.1038/onc.2017.58
15. Kulaberoglu Y, Lin K, Holder M, et al. Stable MOB1 interaction with hippo/MST is not essential for development and tissue growth control. Nat Commun. 2017;8(1):695. doi:10.1038/s41467-017-00795-y
16. Chen M, Zhang H, Shi Z, et al. The MST4-MOB4 complex disrupts the MST1-MOB1 complex in the hippo-YAP pathway and plays a pro-oncogenic role in pancreatic cancer. J Biol Chem. 2018;293(37):14455–14469. doi:10.1074/jbc.RA118.003279
17. Turunen SP, von Nandelstadh P, Ohman T, et al. FGFR4 phosphorylates MST1 to confer breast cancer cells resistance to MST1/2-dependent apoptosis. Cell Death Differ. 2019;26(12):2577–2593. doi:10.1038/s41418-019-0321-x
18. Markowitz SD, Bertagnolli MM. Molecular origins of cancer: molecular basis of colorectal cancer. N Engl J Med. 2009;361(25):2449–2460. doi:10.1056/NEJMra0804588
19. Ng C, Li H, Wu W, Wong SH, Yu J. Genomics and metagenomics of colorectal cancer. J Gastrointest Oncol. 2019;10(6):1164–1170. doi:10.21037/jgo.2019.06.04
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.