Back to Journals » OncoTargets and Therapy » Volume 14
MiRNA-223-3p Affects Mantle Cell Lymphoma Development by Regulating the CHUK/NF-ƘB2 Signaling Pathway
Authors Yuan J, Zhang Q, Wu S, Yan S, Zhao R, Sun Y, Tian X, Zhou K
Received 22 September 2020
Accepted for publication 29 January 2021
Published 2 March 2021 Volume 2021:14 Pages 1553—1564
DOI https://doi.org/10.2147/OTT.S283486
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Arseniy Yuzhalin
Jingjing Yuan, Qing Zhang, Shengsheng Wu, Suran Yan, Ran Zhao, Yajuan Sun, Xiaoxu Tian, Keshu Zhou
Department of Hematology, Affiliated Cancer Hospital of Zhengzhou University, Henan Cancer Hospital, Zhengzhou, People’s Republic of China
Correspondence: Keshu Zhou
Department of Hematology, Affiliated Cancer Hospital of Zhengzhou University, Henan Cancer Hospital, Zhengzhou, People’s Republic of China
Tel/Fax +86-371-65587513
Email [email protected]
Background: Mantle cell lymphoma (MCL) is an aggressive malignancy that accounts for 5– 10% of non-Hodgkin’s lymphoma. MiRNA-223-3p has been demonstrated to be down-regulated in MCL and is a useful prognostic factor. However, little is known about underlying molecular mechanism of miRNA-233-3p in MCL.
Methods: The expression levels of miRNA-223-3p and CHUK mRNA in MCL cells were detected by real-time quantitative PCR (RT-qPCR). The effects of miRNA-223-3p/CHUK overexpression/knockdown on MCL cell proliferation and apoptosis were measured by CCK-8 assay and annexin V PE/7-AAD-based flow cytometry/TUNEL assay, respectively. A nude mouse subcutaneous xenograft model was used to further evaluate the potential effects in vivo. Dual-luciferase reporter assay was used to verify the inhibitory effect of miRNA-223-3p on CHUK. Furthermore, the regulatory function of miRNA-223-3p on the CHUK/NF-ƘB2 axis was assessed by RT-qPCR, western blot and immunofluorescence.
Results: In the present study, miRNA-223-3p overexpression inhibited proliferation and accelerated apoptosis of MCL cells in vitro and in vivo. The results of Luciferase reporter assay showed that CHUK was a direct target of miRNA-223-3p in HEK293T cells. Furthermore, the results of RT-qPCR, western blot confirmed that CHUK was targeted and negatively regulated by miRNA-223-3p for repressing NF-ƘB2 pathway activation in MCL cells. Importantly, CHUK overexpression promoted proliferation and suppressed apoptosis of MCL cells, whereas CHUK knockdown reversed down-regulated miRNA-223-3p -accelerated cell proliferation in vitro.
Conclusion: In conclusion, miRNA-223-3p affects MCL development by regulating the CHUK/NF-ƘB2 signaling pathway, which is crucial to provide a novel therapeutic strategy.
Keywords: mantle cell lymphoma, miRNA-223-3p, CHUK, NF-ƘB2 signaling pathway, disease progression
Introduction
MCL is a subtype of non-Hodgkin’s lymphoma (NHL) which is derived from naïve CD5+ B cells originating in the mantle zone around reactive germinal centers. MCL accounts for 5–10% of NHL.1 Though advances in chemical compound, immunotherapy and targeted therapy have achieved,2–11 most of MCL patients relapse and die from the disease finally. Therefore, new therapeutic targets are eagerly required for MCL patients.
 |
Table 1 Primers Used in This Study |
MicroRNAs(miRNAs) are small non-coding RNAs of approximately 22 nucleotides which could regulate gene expression through hybridization with a target mRNA sequence, thus resulting in translational repression or mRNA degradation.12 Expression profiling studies have detected specific miRNAs expression “signatures” in various human cancers, especially B cell tumors.13 MiRNA-223-3p has been demonstrated to be down-regulated in MCL and is a useful prognostic factor.14 However, the underlying molecular mechanism remains not entirely clear.
In this study, we investigated molecular mechanism of miRNA-223-3p affecting MCL development. Up-regulation of miRNA-223-3p in MCL cells lead to growth failure and cell apoptosis. NF-ƘB non-classical signaling pathways are abnormally activated in MCL.15 Li et al reported that miRNA-223-3p affect macrophage differentiation by regulating CHUK expression.16 On this basis, we further found that miRNA-223-3p affected MCL development by regulating the CHUK/NF-ƘB2 signaling pathway. These findings define a mechanism for a tumor suppressor role of miRNA-223-3p in MCL and foster its translational research as a therapeutic target.
Materials and Methods
MCL Patient Samples
Tumor tissues were collected from 14 newly diagnosed MCL patients who underwent lymph modes biopsy between 2017 and 2020 at Institute of Hematology and Blood Disease Hospital of Chinese Academy of Medical Sciences and Peking Union Medical College and embedded in paraffin wax. This study was approved by ethics committee of Institute of Hematology and Blood Disease Hospital of Chinese Academy of Medical Sciences and Peking Union Medical College and conducted according to ethical guidelines of 1975 Declaration of Helsinki. Written informed consent was obtained from each patient.
Cell Lines
MCL cell lines Granta-519 and Jeko-1 (American Type Culture Collection, USA) were all cultured in RPMI-1640 (Gibco, New York, USA) supplemented with 10% fetal calf serum (Thermo Fisher Scientific, Massachusetts, USA) at 37°C with 5% CO2.
RT-qPCR
RNA extraction was performed using TRIzol (Invitrogen, California, USA) according to the manufacturer’s instructions. cDNA was synthetized by reverse transcription using an ABI 2720 thermal cycler (ABI Biosystems, Massachusetts, USA) according to the manufacturer’s instructions (M-MLV-RTase, Promega, Madison, USA). The cDNA product was detected using a 2X ChamQ Universal SYBR qPCR Master Mix (Vazyme, Nanjing, China) with an ABI 7500 Real-Time PCR System (ABI Biosystems, Massachusetts, USA). The cycling parameters were 95 °C for a 30s hot start followed by 40 cycles of 95 °C for 10s and 60°C for 30s. The relative mRNA (Candidate gene/GAPDH) or miRNA expressions (Candidate gene/U6) were determined using the 2−ΔCt method. The relative mRNA or miRNA fold change was determined using the 2−ΔΔCt method. All primers are listed in Table 1.
Western Blot Analysis
Granta-519 cells were incubated with CD40L (2 μg/mL)(CST, #32,621) for 4h, Jeko-1 cells were incubated with CD40L (2 μg/mL) for 24h. Whole-cell lysates were prepared in RIPA buffer (Beijing Dingguo, Shanghai, China) with protease inhibitor, quantified, and loaded onto SDS-PAGE. After electrophoresis, proteins in the gel were transferred to a nitrocellulose membrane and incubated with primary antibodies at 4°C overnight. Western blot was conducted using antibodies specific for CHUK (Santa Cruz Biotechnology, sc-71,333), NF-ƘB2 (CST, #3017) and β-actin (Proteintech, 66,009-1-Ig), followed with horseradish peroxidase-labeled secondary antibody (Santa Cruz Biotechnology, sc-2004/sc-2005). Signals were detected by chemiluminescence (ECL Western Blotting Detection Reagents, GE Healthcare, Buckinghamshire, England) and visualized using G:BOX Chemi GelDocumentation System (Syngene, Frederick, MD, USA).
Lentiviral Transduction
The lentiviral vectors were purchased from Shanghai Genechem Company Ltd., China. A scramble siRNA (5ʹ-TTC TCC GAA CGT GTC ACG T-3ʹ) was used as the negative control (NC). The siRNA targeting sequence of miRNA-223-3p was 5ʹ-TGG GGT ATT TGA CAA ACT GAC A-3ʹ. The siRNA targeting sequence of CHUK gene was 5ʹ-GCA AAT GAG GAA CAG GGC AAT-3ʹ. CHUK over-expression plasmid was constructed by subcloning CHUK cDNA into GV492 plasmid. MiRNA-223-3p over-expression plasmid was constructed by subcloning pre-miRNA into GV646 plasmid (Shanghai Genechem Company Ltd., Shanghai, China). We used puromycin (2 μg/mL) to select infected cells. For CHUK promoter reporter plasmid, a sequence containing miRNA-223-3p binding sites (Chr10:100,188,675–100,188, 681 strand (-)) were inserted into PGL3-promoter plasmid before SV40 promoter.
Cell Proliferation Assay
Cell proliferation was detected by Cell Counting Kit-8 assay (Solarbio, Beijing, China). Infected MCL cells were plated in 96-well plates at 5×10^4 cells per well, respectively. The cells were incubated overnight before experimentation. After 10 μL of CCK-8 reagent was added to each well at 0, 24, 48, 72, 96 and 120h, the plate was incubated for 4 h. The optical density of viable cells was measured at 450 nm using a microplate reader (BioTek, Vermont, USA).
Cell Apoptosis Assay
Cell apoptosis was detected by Annexin V PE/7-AAD apoptosis detection kit (Solarbio, Beijing, China). Infected MCL cells were seeded in 6-well plates at 2×10^6 cells per well. Cell apoptosis was detected after 120 h. Briefly, cells were collected and washed by phosphate-buffered saline, and apoptosis was evaluated according to the kit protocol. Although standard flow cytometry apoptosis assays use co-staining with propidium iodide, we analyzed annexin V PE/7-AAD since the cells were transfected with an EGFP-expressing vector and the excitation wavelengths of propidium iodide and EGFP overlap.
Immunofluorescence Staining
Granta-519 cells were incubated with CD40L (2 μg/mL)(CST, #32,621) for 4h, Jeko-1 cells were incubated with CD40L (2 μg/mL) for 24h. Cells were fixed with 4% paraformaldehyde, permeabilized with 0.5% Triton X-100 and blocked with 2% BSA. Then, cells were probed with anti-RelB antibodies (Abcam, ab33907). After washing 3 times with TBST (1% Tween-20 in TBS), the cells were incubated with Alexa Fluor 594-conjugated secondary antibodies (Abways, AB0152). Then, they were washed 3 times with TBST, and DAPI solution was added to stain the cell nuclei. Images were obtained using a confocal microscope (Zeiss).
TUNEL Staining
TUNEL (terminal deoxynucleotidyl transferase dUTP nick-end labeling) staining was performed using a TUNEL kit (Roche, Basel, Switzerland). TUNEL+ cells were counted with ImageJ software.
Luciferase Reporter Assay
Cells cultured in 24-well plates were transfected with promoter reporter construct or mutant variants and overexpression vector. A Renilla luciferase–containing plasmid, which is driven by thymidine kinase promoter, was always included in transfection to control transfection efficiency. Luciferase activity was determined using a dual-luciferase reporter assay system following the manufacturer’s instructions (Promega, Madison, WI).
Xenograft Mouse Model Assay
Five-week-old female BALB/c athymic nude mice (Charles River) were injected with 2 × 107 Jeko-1 cells that had been transfected with e-GFP or miRNA-223-3p overexpression lentiviral vector subcutaneously in the flank. When the tumor approached 20–30 mm3, mice were divided randomly into groups of seven. Tumor volume was measured with a caliper every 3 days, and calculated with the formula: V=0.5× length×width2. The subcutaneous tumors were harvested and weighted at the 34th day after implantation. All experimental protocols were approved by Zhengzhou University Institutional Review Board and carried out in accordance with the National Institutes of Health guide for the care and use of Laboratory animals.
Statistical Analysis
All experiments were performed at least three times independently, unless otherwise specified. Statistical analyses were performed using GraphPad Prism. Correlations were analyzed by Spearman correlation. T-tests were used to determine significance, with a p-value of <0.05 to indicate a significant difference. Data are presented as means±SD.
Results
MiRNA-223-3p Affected MCL Development in vitro
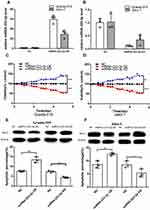
We explored miRNA-223-3p biological function through lentivirus-mediated overexpression and knockdown of miRNA-223-3p in Granta-519 and Jeko-1 cells. The lentivirus infection efficacy was above 95% for miRNA-223-3p OE, miRNA-223-3p KD and NC lentivirus, which ensure the synchronization of all the following experiments (Supplementary Figure 1A). MiRNA-223-3p levels were assessed by qPCR. The results showed that miRNA-223-3p OE lentivirus infected cultures exhibited significantly increased miRNA-223-3p expression compared with cells infected with NC lentivirus in Granta-519 (p<0.01) and Jeko-1 (p<0.05) (Figure 1A); miRNA-223-3p KD lentivirus infected cultures exhibited significantly decreased miRNA-223-3p expression compared with cells infected with NC lentivirus in Granta-519 (p<0.001) and Jeko-1 (p<0.05) cells (Figure 1B).
Granta-519 and Jeko-1 cells infected with miRNA-223-3p OE lentivirus, miRNA-223-3p KD lentivirus or NC lentivirus were seeded in 96-well plates. Cell growth was monitored by CCK-8 assays every day for 5 days, while cell apoptosis was measured by FACS analysis of Annexin V-PE/7-AAD-stained cells after 5 days. The results showed that overexpression and knockdown of miRNA-223-3p respectively decreased and increased cell viability compared with the control group in Granta-519 (miRNA-223-3p OE:p<0.001; miRNA-223-3p KD:p<0.001) and Jeko-1 (miRNA-223-3p OE:p<0.001; miRNA-223-3p KD:p<0.01) cells after 5 days (Figure 1C and D). FACS analysis of Annexin V-PE/7-AAD-stained cells demonstrated that apoptotic percentages of miRNA-223-3p OE cells (Granta-519:p<0.01; Jeko-1:p<0.05) and miRNA-223-3p KD cells (Granta-519:p<0.01;Jeko-1:p<0.05) undergoing were respectively higher and lower compared with that of NC cells after 5 days. Western blot analysis showed that anti-apoptotic protein Bcl-2 expressions were respectively decreased and increased in miRNA-223-3p OE and miRNA-223-3p KD cells compared with the control group (Figure 1E and F).
MiRNA-223-3p Down-Regulated CHUK Expression and Repressed NF-ƘB2 Pathway Activation
The above results indicated that miRNA-223-3p play an important role in MCL development in vitro. However, the mechanisms underlying miRNA-223-3p-mediated MCL development are still unclear.
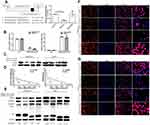
NF-ƘB2 signaling pathway is abnormally activated in MCL.15 Previous studies reported that miRNA-223-3p affects macrophage differentiation through regulating the expression level of CHUK,16 which is important NF-ƘB2 signaling molecule. Interestingly, we found that CHUK 3ʹUTR region harbor miRNA-223-3p binding sites (Chr10: 100,188,675–100,188,681 strand (-)) (Supplementary Figure 2). In addition, miRNA-223-3p expression was inversely correlated with CHUK mRNA expression (rs=−0.7758, p<0.01) or the level of NF-ƘB2 p100 processing into p52 (rs=−0.5956, p<0.05) in MCL patient tissues (Figure 2D). We then hypothesized that miRNA-223-3p plays a critical role in the development of MCL through regulating CHUK. The effect of miRNA-223-3p on the regulation of CHUK transactivity is examined. MiRNA-223-3p was overexpressed in HEK293T cells by lentivirus-mediated overexpression. Wild type CHUK reporter activity was significantly decreased in miRNA-223-3p OE cells, whereas no significant changes were found in corresponding mutant type luciferase activity (Figure 2A). Importantly, CHUK mRNA levels were significantly decreased in miRNA-223-3p OE cells (Granta-519: p<0.01; Jeko-1:p<0.001) and increased in miRNA-223-3p KD cells (Granta-519:p<0.001;Jeko-1:p<0.01) compared with the control group (Figure 2B). The similar trend on CHUK protein levels was observed as on mRNA levels by Western blot analysis in Granta-519 and Jeko-1 cells (Figure 2C). These results further suggest the specific effect of miRNA-223-3p on CHUK transactivity and translation activity.
Besides, we further examined whether miRNA-223-3p affected NF-ƘB2 pathway activation. Overexpression of miRNA-223-3p decreased the protein expression level of CHUK and phospho-CHUK, the level of NF-ƘB2 p100 processing to p52 and nuclear translocation of RelB, while knockdown of miRNA-223-3p increased the protein expression level of CHUK and phospho-CHUK, the level of NF-ƘB2 p100 processing to p52 and nuclear translocation of RelB in Granta-519 and Jeko-1 cells after NF-ƘB2 pathway stimulation with CD40L which signified that miRNA-223-3p repressed the NF-ƘB2 pathway activation (Figure 2E–G).
Altering CHUK Affected MCL Development in vitro
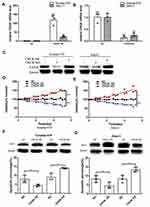
Based on above data, we probed CHUK biological function through protein overexpression and knockdown. We did lentivirus-mediated overexpression and knockdown of CHUK in Granta-519 and Jeko-1 cells. The lentivirus infection efficacy is above 95% for CHUK OE, CHUK KD and NC lentivirus (Supplementary Figure 1B). CHUK levels were accessed by qPCR and Western blot. The results showed that CHUK OE lentivirus infected cultures exhibited significantly increased CHUK mRNA expression compared with cells infected with NC lentivirus in Granta-519 (p<0.01) and Jeko-1 (p<0.01) (Figure 3A); CHUK KD lentivirus infected cultures exhibited significantly decreased CHUK mRNA expression compared with cells infected with NC lentivirus in Granta-519 (p<0.01) and Jeko-1 (p<0.01) (Figure 3B). The similar trend on CHUK protein levels was observed as on mRNA levels by Western blot analysis in Granta-519 and Jeko-1 cells (Figure 3C).
Granta-519 and Jeko-1 cells infected with CHUK OE lentivirus, CHUK KD lentivirus or NC lentivirus were seeded in 96-well plates. Cell growth and apoptosis were measured as mentioned above. The results showed that overexpression and knockdown of CHUK respectively increased and decreased cell viability compared with the control group after 5 days in Granta-519 (CHUK OE: p<0.001; CHUK KD:p<0.001) and Jeko-1 (CHUK OE: p<0.05; CHUK KD:p<0.001) cells (Figure 3D and E). The analysis of Annexin V-PE/7-AAD-stained cells demonstrated that apoptotic percentages of CHUK OE cells (Granta-519:p<0.05; Jeko-1:p<0.05) and CHUK KD cells (Granta-519:p<0.05;Jeko-1:p<0.05) were respectively lower and higher compared with that of NC cells after 5 days. Western blot analysis showed that anti-apoptotic protein Bcl-2 expressions were respectively increased and decreased in CHUK OE cells and CHUK KD cells compared with the control group (Figure 3F and G, table 1).
MiRNA-223-3p Affected MCL Development by Regulating the CHUK/NF-ƘB2 Signaling Pathway
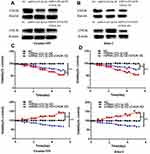
To identifying whether miRNA-223-3p affects MCL development through CHUK/NF-ƘB2 signaling pathway, we then silenced CHUK in miRNA-223-3p KD cells and up-regulated CHUK in miRNA-223-3p OE cells, which were identified by Western blot (Figure 4A and B). Figure 4C and D showed that knockdown CHUK inhibited cell proliferation led by miRNA-223-3p KD in Granta-519 and Jeko-1 cells, while up-regulated CHUK accelerated cell proliferation led by miRNA-223-3p OE in Granta-519 and Jeko-1 cells.
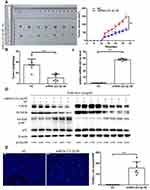
MiRNA-223-3p Affected MCL Development in vivo
To investigate whether miRNA-223-3p affected MCL cells proliferation and apoptosis in vivo, we injected nude mice with Jeko-1 cells transfected with NC or miRNA-223-3p overexpression lentiviral vector subcutaneously. The results of tumor growth curves revealed that miRNA-223-3p overexpression dramatically impaired tumor growth in nude mice (p<0.001, Figure 5A and B). The miRNA-223-3p content and protein expression level of CHUK, phospho-CHUK and NF-ƘB2 in tumor tissues were respectively detected by qPCR and Western blot. It was observed that the level of miRNA-223-3p in tumor tissues that developed by injecting miRNA-223-3p OE Jeko-1 cells was significantly higher than that in the control group (p<0.001, Figure 5C). MiRNA-223-3p overexpression decreased the protein expression level of CHUK, phospho-CHUK and the level of NF-ƘB2 p100 processing to p52, which proved that miRNA-223-3p repressed the NF-ƘB2 pathway activation in tumor tissues (Figure 5D). The results of TUNEL staining suggested that miRNA-223-3p overexpression remarkably increased apoptotic cell number compared with the control group in vivo (p<0.001, Figure 5E).
Discussion
The key findings of this study are that increased expression of miRNA-223-3p inhibited proliferation and accelerated apoptosis of MCL cells in vitro and in vivo. MiRNA-223-3p regulates MCL development by the CHUK/NF-ƘB2 signaling pathway. This study provides new sights and strong evidences that the miRNA-223-3p/CHUK/NF-ƘB2 signaling pathway plays an important role in MCL development.
Wong et al reported that miRNA-223-3p increases apoptosis and decreases cell viability, proliferation of hepatocellular carcinoma cells by targeting STMN1, a key microtubule-regulatory protein that directly participates in cellular proliferation and cell cycle.17 In addition to solid tumors, miRNA-223-3p is up-regulated in nodal marginal zone lymphoma and inhibits LMO2 expression. Moreover, our previous studies demonstrated that miRNA-223-3p is down-regulated in MCL compared with healthy donors and is associated with poor survival in MCL patients.14 However, the specific mechanism is not explored thoroughly.
Activating NF-ƘB2 signaling pathway maintains cell proliferation and protects cells from apoptosis. NF-ƘB2 is widely used as a gene regulator to control cell proliferation and cell survival in eukaryotic cells. Besides, it also regulates anti-apoptotic genes and thus eliminate caspase family activity, which is the core of most apoptosis. Studies showed that NF-ƘB2 signaling pathway is abnormally activated in MCL.14
Although the therapeutic effect of MCL is improved with the development of treatment strategy, emerging new therapies have intolerable adverse effects including cytopenia and infection.14,18–27 Besides, recurrence and drug resistance are main problems. Hence, it is meaningful and important to identify the specific pathogenesis of MCL thoroughly.
The fact that miRNA−223−3p plays a critical role in the pathogenesis of MCL has been gradually recognized. Our previous studies demonstrated that miRNA−223−3p is repressed and is associated with poor prognosis in MCL by targeting SOX11.14 In the present study, we found that down-regulation of miRNA-223-3p accelerated proliferation and inhibited apoptosis of MCL cells by upregulating CHUK and thus abnormally activating the non-classical NF-ƘB signaling pathway. These findings were in consistent with the previous study and further clarify the underlying molecular mechanism of miRNA-223-3p affecting MCL development thoroughly.
Aberrant versions of miRNA have been implicated in not only tumor development,28–31 but also drug resistance.32–36 Therefore, further studies are needed for the function of miRNA-223-3p in drug resistance and corresponding mechanisms in MCL.
In summary, our results suggest that miRNA-223-3p plays an important role in MCL development by regulating the CHUK/NF-ƘB2 signaling pathway. Downregulation of miRNA-223-3p accelerated proliferation and inhibited apoptosis of MCL cells by up-regulating CHUK and thus abnormally activating the non-classical NF-ƘB2 signaling pathway. Thorough understanding of the precise role of miRNA-223-3p in MCL provides the opportunity to develop a novel therapeutic strategy by upregulating expression of miRNA-223-3p in MCL cells.
Funding
This study was supported by grants from National Natural Science Foundation of China (81470336), Natural Science Foundation of Henan Province (182300410347).
Disclosure
The authors report no conflicts of interest.
References
1. Velders GA, Kluin-Nelemans JC, De Boer CJ, et al. Mantle-cell lymphoma: a population-based clinical study. J Clin Oncol. 1996;14(4):1269–1274. doi:10.1200/JCO.1996.14.4.1269
2. Gauthier J, Maloney DG. Allogeneic transplantation and chimeric antigen receptor-engineered T-Cell therapy for relapsed or refractory mantle cell lymphoma. Hematol Oncol Clin North Am. 2020;34(5):957–970. doi:10.1016/j.hoc.2020.06.010
3. Bond DA, Maddocks KJ. Current role and emerging evidence for bruton tyrosine kinase inhibitors in the treatment of mantle cell lymphoma. Hematol Oncol Clin North Am. 2020;34(5):903–921. doi:10.1016/j.hoc.2020.06.007
4. Ruan J. Approach to the initial treatment of older patients with mantle cell lymphoma. Hematol Oncol Clin North Am. 2020;34(5):871–885. doi:10.1016/j.hoc.2020.06.005
5. Guy D, Kahl BS. Initial and consolidation therapy for younger patients with mantle cell lymphoma. Hematol Oncol Clin North Am. 2020;34(5):861–870. doi:10.1016/j.hoc.2020.06.004
6. Sawalha Y, Bond DA, Alinari L. Evaluating the therapeutic potential of zanubrutinib in the treatment of relapsed/refractory mantle cell lymphoma: evidence to date. Onco Targets Ther. 2020;13:6573–6581. doi:10.2147/OTT.S238832
7. Santambrogio E, Novo M, Rota-Scalabrini D, Vitolo U. Chemotherapy combinations for B-cell lymphoma and chemo-free approach in elderly patients: an update on best practice. Expert Rev Hematol. 2020;13(8):851–869.
8. Rule S, Johns S. Advances in classification and treatment of non-hodgkin lymphoma: mantle cell. Cancer J. 2020;26(4):348–356. doi:10.1097/PPO.0000000000000462
9. Wang X, Li W, Wang X, et al. The effectiveness and harms of bortezomib in combination with chemotherapy for mantle cell lymphoma: a protocol for systematic review and meta-analysis. Medicine (Baltimore). 2020;99(28):e20961. doi:10.1097/MD.0000000000020961
10. Li P, Dong N, Zeng Y, et al. Chimeric antigen receptor T-cell therapy: a promising treatment modality for relapsed/refractory mantle cell lymphoma. Front Med. 2020;14(6):811–815.
11. Villa D, Sehn LH, Savage KJ, et al. Bendamustine and rituximab as induction therapy in both transplant-eligible and -ineligible patients with mantle cell lymphoma. Blood Adv. 2020;4(15):3486–3494. doi:10.1182/bloodadvances.2020002068
12. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–297. doi:10.1016/S0092-8674(04)00045-5
13. Calin GA, Croce CM. MicroRNA-cancer connection: the beginning of a new tale. Cancer Res. 2006;66:7390–7394. doi:10.1158/0008-5472.CAN-06-0800
14. Zhou K, Feng X, Wang Y, et al. miR-223 is repressed and correlates with inferior clinical features in mantle cell lymphoma through targeting SOX11. Exp Hematol. 2018;58(27–34.e1). doi:10.1016/j.exphem.2017.10.005
15. Rahal R, Frick M, Romero R, et al. Pharmacological and genomic profiling identifies NF-κB-targeted treatment strategies for mantle cell lymphoma. Nat Med. 2014;20(1):87–92. doi:10.1038/nm.3435
16. Li T, Morgan MJ, Choksi S, Zhang Y, Kim YS, Liu ZG. MicroRNAs modulate the noncanonical transcription factor NF-κB pathway by regulating expression of the kinase IKKα during macrophage differentiation. Nat Immunol. 2010;11(9):799–805. doi:10.1038/ni.1918
17. Wong WL, Lung WM, Law TY, Lai BS, Wong N. MicroRNA-223 is commonly repressed in hepatocellular carcinoma and potentiates expression of stathmin1. Gastroenterology. 2008;135(1):257–269. doi:10.1053/j.gastro.2008.04.003
18. Wang M, Munoz J, Goy A, et al. KTE-X19 CAR T-cell therapy in relapsed or refractory mantle-cell lymphoma. N Engl J Med. 2020;382(14):1331–1342. doi:10.1056/NEJMoa1914347
19. Phillips T, Barr PM, Park SI, et al. A Phase 1 trial of SGN-CD70A in patients with CD70-positive diffuse large B cell lymphoma and mantle cell lymphoma. Invest New Drugs. 2019;37(2):297–306. doi:10.1007/s10637-018-0655-0
20. Davids MS, Kim HT, Nicotra A, et al. Umbralisib in combination with ibrutinib in patients with relapsed or refractory chronic lymphocytic leukaemia or mantle cell lymphoma: a multicentre phase 1-1b study. Lancet Haematol. 2019;6(1):e38–e47. doi:10.1016/S2352-3026(18)30196-0
21. Gressin R, Daguindau N, Tempescul A, et al. A Phase 2 study of rituximab, bendamustine, bortezomib and dexamethasone for first-line treatment of older patients with mantle cell lymphoma. Haematologica. 2019;104(1):138–146. doi:10.3324/haematol.2018.191429
22. Salles G, Gopal AK, Minnema MC, et al. Phase 2 study of daratumumab in relapsed/refractory mantle-cell lymphoma, diffuse large B-cell lymphoma, and follicular lymphoma. Clin Lymphoma Myeloma Leuk. 2019;19(5):275–284. doi:10.1016/j.clml.2018.12.013
23. Martin P, Ruan J, Furman R, et al. A Phase I trial of palbociclib plus bortezomib in previously treated mantle cell lymphoma. Leuk Lymphoma. 2019;60(12):2917–2921. doi:10.1080/10428194.2019.1612062
24. Jerkeman M, Eskelund CW, Hutchings M, et al. Ibrutinib, lenalidomide, and rituximab in relapsed or refractory mantle cell lymphoma (PHILEMON): a multicentre, open-label, single-arm, phase 2 trial. Lancet Haematol. 2018;5(3):e109–e116. doi:10.1016/S2352-3026(18)30018-8
25. Matsumoto Y, Kobayashi T, Shimura Y, et al. Combined rituximab, bendamustine, and dexamethasone chemotherapy for relapsed or refractory indolent B-cell non-hodgkin lymphoma and mantle cell lymphoma: a multicenter Phase II study. Int J Hematol. 2019;110(1):77–85. doi:10.1007/s12185-019-02650-w
26. Hermine O, Hoster E, Walewski J, et al. Addition of high-dose cytarabine to immunochemotherapy before autologous stem-cell transplantation in patients aged 65 years or younger with mantle cell lymphoma (MCL Younger): a randomised, open-label, Phase 3 trial of the European Mantle Cell Lymphoma Network. Lancet. 2016;388(10044):565–575. doi:10.1016/S0140-6736(16)00739-X
27. Tessoulin B, Bouabdallah K, Burroni B, et al. Safety and efficacy of temsirolimus in combination with three different immuno-chemotherapy regimens in relapse and refractory mantle cell lymphoma, final results of the T3 phase IB trial of the LYSA. Ann Hematol. 2020;99(8):1771–1778. doi:10.1007/s00277-020-04159-3
28. Pulikkan JA, Viola D, Peramangalam PS, et al. Cell-cycle regulator E2F1 and microRNA-223 comprise an autoregulatory negative feedback loop in acute myeloid leukemia. Blood. 2010;115:1768–1778. doi:10.1182/blood-2009-08-240101
29. Yang W, Lan X, Li D, Li T, Lu S. MiR-223 targeting MAFB suppresses proliferation and migration of nasopharyngeal carcinoma cells. BMC Cancer. 2015;15(1):461. doi:10.1186/s12885-015-1464-x
30. Dong W, Li B, Wang J, Song Y, Zhang Z, Fu C. MicroRNA-337 inhibits cell proliferation and invasion of cervical cancer through directly targeting specificity protein 1. Tumour Biol. 2017;39(6):101042831771132. doi:10.1177/1010428317711323
31. Zuo XL, Chen ZQ, Wang JF, Wang JG, Cai J. miR-337-3p suppresses the proliferation and invasion of hepatocellular carcinoma cells through targeting JAK2. Am J Cancer Res. 2018;8(4):662–674.
32. Yu FY, Yao HR, Zhu PC, et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell. 2007;131:1109–1123. doi:10.1016/j.cell.2007.10.054
33. Pan YZ, Gao WQ, Yu AM. MicroRNAs regulate CYP3A4 expression via direct and indirect targeting. Drug Metab Dispos. 2009;37:2112–2117. doi:10.1124/dmd.109.027680
34. Xia L, Zhang D, Du R, et al. miR-15b and miR-16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells. Int J Cancer. 2008;123(2):372–379. doi:10.1002/ijc.23501
35. Bai H, Xu R, Cao Z, Wei D, Wang C. Involvement of miR-21 in resistance to daunorubicin by regulating PTEN expression in the leukaemia K562 cell line. FEBS Lett. 2011;585(2):402–408. doi:10.1016/j.febslet.2010.12.027
36. Jeon HM, Sohn YW, Oh SY, Oh SY, Kim H. ID4 imparts chemoresistance and cancer stemness to glioma cells by derepressing miR-9*-mediated suppression of SOX2. Cancer Res. 2011;71(9):3410–3421. doi:10.1158/0008-5472.CAN-10-3340
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.