Back to Journals » OncoTargets and Therapy » Volume 13
MiR-486-5p Act as a Biomarker in Endometrial Carcinoma: Promotes Cell Proliferation, Migration, Invasion by Targeting MARK1
Authors Zheng X , Xu K, Zhu L, Mao M, Zhang F , Cui L
Received 21 January 2020
Accepted for publication 23 April 2020
Published 28 May 2020 Volume 2020:13 Pages 4843—4853
DOI https://doi.org/10.2147/OTT.S246841
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr William C. Cho
Xiaojiao Zheng, Kejun Xu, Linyan Zhu, Meiya Mao, Fubin Zhang, Lining Cui
Department of Obstetrics and Gynecology, Ningbo First Hospital, Ningbo, Zhejiang 315035, People’s Republic of China
Correspondence: Lining Cui
Department of Obstetrics and Gynecology, Ningbo First Hospital, 59 Liuting Street, Haishu District, Ningbo, Zhejiang 315035, People’s Republic of China
Email [email protected]
Background: miRNA expression acts as a potential biomarker in many diseases including endometrial carcinoma (EC). miR-486-5p dysregulation is observed in several tumor types, but the roles of miR-486-5p in EC are hardly ever studied.
Objective: This study aimed to analyze the expression profile of miR-486-5p in tumor tissues and serum samples of patients with EC and explore the target prediction, function analysis and validation in immortal cell lines.
Patients and Methods: A total of 42 freshly paired EC tissues, the corresponding adjacent non-neoplastic tissues and serum samples were also collected from patients with EC, and 42 matched normal serum samples were included as control group. The level of miR-486-5p expression was tested by quantitative real-time polymerase chain reaction (qRT-PCR). Cell proliferation was determined by colony formation assay and CCK-8 assay. Furthermore, functional evaluation of miR-486-5p on migration was performed by wound-healing assay and invasion was estimated by transwell invasion assay. qRT-PCR, luciferase reporter assay and Western blotting (WB) were performed to verify the targeting of MARK1 by miR-486-5p.
Results: miR-486-5p was significantly up-regulated in EC tissues and serum samples, promoting the proliferation, migration and invasive activities of EC cells by targeting MARK1.
Conclusion: These data indicated miR-486-5p as a novel molecular biomarker for diagnosing and treating EC, and MARK1 might act as a critical and functional target of miR-486-5p with the implications on cell proliferation, migration and invasiveness of EC tumor cells.
Keywords: endometrial carcinoma, miR-486-5p, MARK1, proliferation, migration, invasion
Introduction
Endometrial carcinoma (EC) is one of the most common gynecological malignancies in women, with more than 80000 newly diagnosed patients in Europe yearly.1 EC is dichotomized into two types based on estrogen levels, clinical and histopathological behavior as less aggressive and estrogen-dependent type I, highly aggressive and estrogen-independent type II.2 Hysterectomy and bilateral salpingo-oophorectomy, chemotherapy and radiation are considered as effective treatment strategies for EC; however, it demonstrated an extremely poor prognosis. So, novel screening methods for pre-cancerous and early disease are warranted. The molecular changes that underlie EC initiation and progression are poorly understood, and so it is imperative to find key signaling pathways for novel diagnostic, prognostic and treatment strategies.3
MicroRNAs (miRNAs) are a class of short 18-25-nucleotide noncoding RNAs that post-transcriptionally regulate gene expression and are considered important in many fundamental biological functions.4,5 Microenvironment that favors replication and growth of miRNAs expression is regarded as an important factor. This is because they bind to partially complementary sequences that target the 3ʹ-untranslated region (3ʹ-UTR) of mRNA and modulate gene expression, resulting in either mRNA degradation or translation repression or both, which therefore influences the biological function.5,6 miRNAs are new biomarkers in the pathogenesis and progression of different types of cancers.7,8 Increasing evidence has demonstrated the important link between miRNAs and cancer, in which miRNAs act as oncogene or anti-oncogene virtually in all tumors including gynecological cancers. So, these may become specific therapeutic targets in the future.8,9
Evidence has revealed that miRNA-486-5p functions both as an oncogene and tumor suppressor in multiple tumor types. It is downregulated in non-small cell lung cancer,10 esophageal squamous cell carcinoma,11 breast cancer,12,13 gastric adenocarcinoma14 and colorectal carcinogenesis.15 However, miR-486-5p was upregulated in cervical cancer.16 Also, other researches indicated reduced miR-486-5p expression in renal cell carcinoma17,18 and chronic myeloid leukemia.19 Disordered expression of miR-486-5p was associated with the occurrence and development of various cancers, functions including proliferation, migration and invasion. To date, the clinical significance and the exact mechanism of miR-486-5p of EC are not fully elucidated.
We herein analyzed the expression of miR-486-5p in EC clinical samples and cell lines, investigated the roles of miR-486-5p in the regulation of EC cell proliferation, migration and invasion, and explored the underlying mechanisms regulated by miR-486-5p. Also, the potential roles of miR-486-5p in EC were investigated. More importantly, MARK1 that acts as a specific key regulator of microtubule dynamics was confirmed as a functional target of miR-486-5p.
Patients and Methods
Patients and Specimen
Forty-two matched specimens of Endometrial carcinoma and adjacent normal tissues were collected from patients enrolled at Ningbo First Hospital from 2013 to 2018. Endometrial carcinoma patients who had undergone surgery written informed consent. Endometrial carcinoma serum samples were obtained from Endometrial carcinoma patients who had provided the tissues, 42 control normal serum samples were also collected from healthy donors who came from a checkup. Forty-two samples of Endometrial carcinoma were obtained from patients who were without chemotherapy or radiotherapy before surgery, in addition, subgrouped into stage (I~IV) group according to the Federation International FIGO criteria. Tissue samples after surgical removal and serum samples after centrifugal collect were both frozen immediately in liquid nitrogen and stored at −80°C. Clinical parameters including age, grade, FIGO stage, estrogen receptor (ER) status, and lymph node metastasis. The data are illustrated in Table 1.
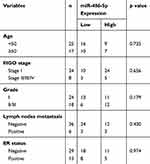 |
Table 1 Relationship Between Expression of miR-486-5p and Basic Clinical Characteristics in Tumor Tissues from Endometrial Carcinoma Patients |
Healthy donors provided written informed consent, in accordance with the Declaration of Helsinki, all tissue and serum samples were collected and all procedures were performed in accordance with the China Ethical Review Committee and Ningbo First Hospital Ethical Committee (Petition Number: 2018-R061).
Cell Culture
The endometrial carcinoma cell lines ECC-1 and HEC-1-B were purchased from Boster Company (Boster, Wuhan, China). Human endometrial carcinoma cell lines AN3CA and Ishikawa were obtained from Shanghai Fuxiang Biotechnology Company (Fuxiang, Shanghai, China). Cells were cultured under condition in a humidified incubator at 37°C containing 5% CO2 in RPMI1640 (ECC-1, Ishikawa), DMEM (AN3CA) and MEM (HEC-1-B) supplemented with 10% fetal bovine serum (FBS)(Gibco, Auckland, NZ).
Cell Transfection
Ishikawa cells were transfected with miR-486-5p mimics or negative control and HEC-1-B were transfected with miR-486-5p inhibitor or negative control (GenePharma, Shanghai, China) for 24 or 48 h using Lipofectamine™ 2000 (Invitrogen, Carlsbad, USA) in accordance with the manufacturer’s instructions. Primers used in the study are shown in Table 2.
 |
Table 2 Primers Used in the Study |
Cell Proliferative Capacity Assay
Cell Counting Kit-8 (CCK-8) assay: Following transfection, cells were seeded into 96-well culture plates (2x103 cells/well). At different time points, 10 µL CCK-8 solution was added to each well and then incubated at 37°C for another 1–2 h (Dojindo, Kumamoto, Japan). The absorbance at 450 nm (A450) was detected using a microplate reader.
For the colony formation assay, following transfection, cells were seeded at 500 cells/well in six-well culture plates. Cells were washed thrice with phosphate-buffered saline (PBS), fixed with paraformaldehyde (4%) for 30 min, and then stained with crystal violet (0.1%) for 20 min after 10–14 days at 37°C following inoculation. Any colony with more than 50 cells was counted under a microscope. The rate of colony formation was calculated according to method: (number of colonies/number of cells inoculated) ×100%.
Cell Cycle Analysis
For the different phases of cell-cycle analysis, transfected cells were plated in a six-well transwell plates (5 × 104 cells/well). Cell-cycle progression was determined using a Cell Cycle Detection Kit (MultiSciences, China). After transfection for 48 h, the cells were washed with PBS and collected by centrifugation, and then treated with 10 μL permeabilization solution and 1mL DNA-staining solution. Finally, incubated for 30 min in the dark at room temperature and analyzed via flow cytometry (BD Biosciences, USA).
Wound-Healing Assay
A wound scratch assay was used to determine the migratory ability, following transfection, cells were seeded in 6-well plates and reached confluence. Cell monolayers were scratched using sterile 200 micropipette tips. The cells were rinsed three times with PBS and afterward incubated in a 0.1%FBS medium. Images of the wound scratches were captured at each time at 0, 24 and 48 h hours. Wound closure was calculated via measuring areas migrated by cells.
Cell Invasion Assay
Following transfection, the serum-free cells (8×104 cells) in serum-free DMEM medium were plated into a 24-well transwell plate into Matrigel-coated. Matrigel (BD Biosciences, Franklin Lakes, USA) was diluted (1:10) with standard medium. 500 µL Standard medium supplemented with 10% FBS was placed in the lower chambers. Over a time period of 48 h, the chamber was fixed with formalin for 30 min and stained with crystal violet (Beyotime Institute Biotech, Shanghai, China) for 30 min at room temperature, then photographed under a microscope at 200 × magnification and calculated the number of penetrating cells (Leica, Wetzlar, Germany).
RNA Extraction and Quantitative Real-Time PCR
Total RNA from cultured cells, frozen serum and tissue specimens were isolated using TRizol reagent (Invitrogen, Carlsbad, USA) according to the manufacturer’s instructions. The quality of the RNA was measured using Nanodrop. qRT-PCR assays were used to detect mRNA expression using SYBR Green Kit (TaKaRa, Dalian, China) and miRNA expression using SYBR Green miRNA Kit (GenePharma, Shanghai, China) according to the manufacturer’s instructions. The U6 small nuclear RNA and gapdh mRNA were used as the references for quantification. qRT-PCR amplification was performed by using an ABI StepOne real-time PCR system. The primers used in the study for amplifying are shown in Table 2.
Luciferase Reporter Assay
To certify directly binding of miR-486-5p to the 3ʹ-untranslated region (UTR), the wildtype (WT) or mutant reporter plasmid vector and up or up negative control were co-transfected in Ishikawa cells using Lipofectamine™ 2000 (Invitrogen, Carlsbad, USA) in 24-well plates. After transfection 48 h, the cells were lysed. Subsequently, luciferase activity was measured using a Dual Luciferase Reporter Assay Kit (Promega, USA) according to the manufacturer’s instructions. Data were normalized to the intensities of Renilla luminescence, and the results were presented as relative to luciferase activity.
Western Blotting
The cells were harvested and lysed in RIPA buffer (Solarbio, USA) containing protease inhibitors. The concentration of protein samples was tested using the bicinchoninic acid protein kit (beyotime, Shanghai, China) 0.10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) was employed to separate equal amounts of the soluble proteins, then transferred to polyvinylidene difluoride membranes (Millipore, Billerica, MA, USA) and blocked in 5% non-fat milk at room temperature for 1 h. Membranes were incubated with rabbit monoclonal anti-MARK1 antibodies (1:1000 diluted) from Cell Signaling Technology (Beverly, MA, USA) and rabbit polyclonal anti-actin antibodies (1:1000 diluted) from Santa Cruz Biotechnology (Santa Cruz, CA, USA) overnight at 4°C, subsequently 1 h incubation with appropriate goat anti-rabbit IgG-H&L secondary antibodies (1:2000 diluted) from Abcam (Cambridge, MA, USA). The protein bands were visualized using the enhanced chemiluminescence (ECL) detection solution.
Statistical Analysis
Statistical analysis of all data was analyzed by software Statistical Package for Social Sciences (SPSS) version 22.0 (SPSS Inc., Chicago, IL, USA), performed by software GraphPad Prism version 7.0 (GraphPad, San Diego, USA). Student’s t-test was calculated to compare when two groups, and one-way analysis of variance (ANOVA) to compare three or more groups, and clinical data were assessed using chi-square test. A P-value of less than 0.05 was considered as significantly difference.
Results
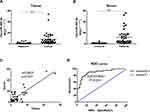
There are clearly higher levels of miR-486-5p were observed in tissue samples of EC when compared with control, as well as serum samples. As shown in Figure 1(A), miR-486-5p was significantly overexpressed EC tissues than in matched adjacent tissues (***P<0.001). As shown in Figure 1(B), the expression of miR-486-5p was significantly increased in serum samples of EC patients when compared with controls (***P<0.001). Figure 1(C) shows a close correlation of miR-486-5p expression in between the patients’ serum and tissues (***P<0.001). Figure 1(D) reveals significant results of receiver operating characteristic (ROC) analysis, indicating that the expression of serum miR-486-5p assay in EC patients was significant when compared to healthy donors (***P<0.001).
The relationship between miR-486-5p expression level and age, grade, FIGO stage, estrogen receptor (ER) status, and lymph node metastasis was analyzed in EC. Nevertheless, no association was shown between the basic clinical characteristics and the expression level of miR-486-5p in tumor tissues (Table 1).
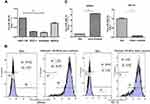
To determine the relative expression of miR-486-5p in four EC cell lines, quantitative RT-PCR analysis was performed via small RNA fractions. As shown in Figure 2A, the relative expression level of miR-486-5p remained the highest in HEC-1-B cells and lowest in Ishikawa cells (***P<0.001). Next, miR-486-5p transfection efficiency was determined after transfection for 24 h, and the cells were analyzed by flow cytometry. Figure 2B shows overexpression of miR-486-5p in Ishikawa cells and low expression of miR-486-5p in HEC-1-B cells. These data suggested that 96.97% of Ishikawa cells carried FAM-labeled miR-486-5p, and 95.98% of HEC-1-B cells carried FAM-labeled miR-486-5p. Furthermore, the levels of miR-486-5p were determined by qRT-PCR (Figure 2C). The levels of mRNA expression of miR-486-5p were significantly increased in miR-486-5p-transfected Ishikawa cells when compared to those with mock-transfected cells (***P<0.001). The mRNA expression levels of miR-486-5p were significantly decreased in miR-486-5p-transfected HEC-1-B cells when compared with mock-transfected cells (**P<0.05).
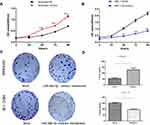
To determine the possible role of miR-486-5p in tumor cellular proliferation, cell viability was examined through CCK-8 assay. Figure 3(A and B) demonstrated that Ishikawa cells transfected with miR-486-5p mimic had enhanced proliferation at 48, 72 and 96 h when compared to the control groups (***P<0.001), and HEC-1-B cells transfected with miR-486-5p inhibitors inhibited the cell growth at 48, 72 and 96 h when compared to the control groups (**P<0.01, ***P<0.001). The colony formation assay was performed to determine the possible role of miR-486-5p in tumor cell proliferation. As shown in Figure 3(C and D), cell viability was detected through colony formation assay. Overexpression of miR-486-5p has significantly promoted the clonogenicity of Ishikawa cells (*P<0.05). The downregulated expression of miR-486-5p has significantly reduced the colony formation rate of HEC-1B cells (*P<0.05).
The abnormal expression of miR-486-5p has changed the G1 phase of cell cycle in tumor cells. Previous studies have shown that proliferation control is primarily achieved in the G1 phase of the cell cycle in cancer cells.20 Figure 4(A–C). Ishikawa cells transfected with miR-486-5p mimics showed a marked increase in S phase cells, and so the G1 phase cells were decreased (*P < 0.05). Figure 4(D–F). HEC-1-B cells transfected with miR-486-5p inhibitors after 48 h resulted in significant increase of G1 phase cells, and so the S phase cells were decreased (*P < 0.05).
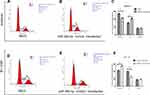 |
Figure 4 MiR-486-5p changes G1 to S phase transition of cell cycle of EC cells. (A–F) Cell cycle changes detected by flow cytometry. Data are expressed as the mean ± SD. **P < 0.01, ***P< 0.001. |
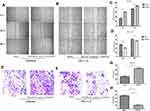
To investigate the possibility that the upregulated expression of miR-486-5p in Ishikawa cells promotes migration and invasion, while its downregulated expression in HEC-1-B cells suppressed their migration and invasion. Figure 5(A–D), wound-healing assay. Representative images show in Figure 5(A and B). As shown in Figure 5(A and C), the scratched width of Ishikawa cells transfected with miR-486-5p mimics was remarkably reduced after 24 h and 48 h of transfection when compared with negative control group, and so the migratory ability was remarkably promoted (**P<0.01). As shown in Figure 5(B and D), the scratched width of HEC-1-B cells transfected with miR-486-5p inhibitors was significantly wider after 24 h and 48 h when compared with negative control group, and so the migratory ability was shown to be significantly inhibited (**P<0.01, ***P<0.001); Figure 5(E–H), through matrigel invasive assay. As shown in Figure 5(E and G), the representative images showed that the cell invasion of Ishikawa cells transfected with miR-486-5p mimics was significantly increased when compared with negative control group, and miR-486-5p expression upregulation promoted the invasiveness of Ishikawa cells (**P<0.01). As shown in Figure 5(F and H), decreased cell invasiveness of HEC-1-B cells transfected with miR-486-5p inhibitors was observed when compared with negative control group, and so downregulated miR-486-5p expression inhibited HEC-1-B cell invasiveness (***P<0.001).
To investigate the molecular mechanisms related to miR-486-5p functions in EC, its potential targets were predicted by bioinformatics programs[miRanda (http://www.miRanda.org), TargetScan (http://www.targetscan.org) and miRWalk (http://www.miRWalk.org)], and selected showed association with the phenotype of miR-486-5p. MARK1 was shown to be the target gene whether regulated by miR-486-5p in EC for target validation (Figure 6). Figure 6(A) presents a layout of luciferase reporter constructs, ie, double–stranded oligonucleotides were bound to a putative miR-486-5p site within MARK1 mRNA 3ʹ-UTR. To explore whether miR-486-5p directly targets MARK1, luciferase reporter assays were performed to examine the direct binding sites of miR-486-5p in its 3ʹUTR, Figure 6(B). The luciferase activity of MARK1_wt with miR-486-5p mimic showed significant reduction (~50%) (**P < 0.01), indicating that miR-486-5p targets MARK1 through miR-486-5p binding sites within the MARK1 3ʹUTR. To experimentally explore whether bona fide targeting of MARK1 by miR-486-5p results from miRNA-induced mRNA degradation, the Ishikawa cells were transfected with miR-486-5p mimics and HEC-1-B cells with miR-486-5p inhibitors, Figure 6(C). qPCR analysis was performed to evaluate MARK1 mRNA levels of transfected cells. The results revealed that transfection with miR-486-5p mimics has significantly modified MARK1 downtranscript levels in Ishikawa cells (**P < 0.01), and miR-486-5p inhibitors have significantly modified MARK1 uptranscript levels in HEC-1-B cells (**P < 0.01), suggesting the involvement of a biotranslational process within mRNA expression. Figure 6(D). Ishikawa cells transfected with miR-486-5p mimics and HEC-1-B cells transfected with miR-486-5p inhibitors were assayed 48 h after transfection by Western blotting, the level of β-actin served as a loading control, MARK1 protein expression was shown to be changed.
Discussion
The present study demonstrated that miRNAs are considered as diagnostic and prognostic biomarkers in EC owing to their entanglement in neoplastic initiation and progression.21 An increasing trend of miR-486-5p was discovered in several types of cancers, and is considered to be involved in various bio-effects such as transcriptional and epigenetic regulations. We herein showed that the (i) expression of miR-486-5p in both serum and tumor tissue was elevated in EC patients; (ii) miR-486-5p is a putative binder of MARK1-3ʹ UTR; (iii) manipulation of miR-486-5p levels can affect the expression levels of MARK1; and (iv) suppression of miR-486-5p expression has significantly attenuated proliferation, colony formation, migration, and invasion of an EC cell line (HEC-1-B) in vitro. Overexpression of miR-486-5p in another EC cell line (Ishikawa) stimulated cell proliferation, colony formation, migration, and invasion in vitro. Collectively, there is no clear conclusion on the association of miR-486-5p and EC to the best of our knowledge. Our results for the first time showed that miR-486-5p regulated tumor progression by regulating the function of MARK1 in EC.
Currently, there are several up or down-regulated miRNAs in EC that are considered to influence cancer initiation, pathogenesis and progression,21 and so miRNAs are being considered as new candidate therapeutic targets. The relationship between miRNAs and carcinogenesis is a key aspect in demonstrating the epigenetic effects.22 Tumor proliferation and metastases are the leading causes of death in EC. However, the molecular mechanism that underlies the miRNAs still remains inconclusive. So far, the miRNAs have simultaneously functioned in the modification of multiple targets, and miR-486-5p can specifically target various genes in the involvement of tumor progressions such as fibrillin-1 in papillary thyroid carcinoma,23 PTEN in cervical cancer,16 OLFM4 in gastric cancer14 and PIK3R1 in nonsmall cell lung cancer.24 Our previous study has remarkably demonstrated that overexpression of miR-486-5p promoted proliferation, migration and invasion in cervical cancer.16 Furthermore, miR-486-5p acts as an oncogene in chronic myeloid leukemia progenitors and up-regulation of miR-486-5p expression has significantly promoted the growth, survival and drug sensitivity. However, overexpression of miR-486-5p has effectively inhibited cell proliferation and invasion and successfully induced apoptosis in vitro in non-small cell lung cancer.24 Therefore, miR-486-5p function in cells promotes or inhibits proliferation and showed association with a wide spectrum of targets. This paper provides results that miR-486-5p-MARK1 signaling pathway is the functional mechanism by which miR-486-5p promotes EC.
There are four MARK isoforms (MARK1-4), and they play several roles in regulating cell polarization and microtubule dynamics, which in turn are associated with cell movement in cancer and its development, promotion of cell migration and invasiveness of carcinoma cells.25 Among the multiple molecular mechanisms that are altered in human cancers, those that are involved in cell division cycle control are believed to be the main for the process of oncogenesis.26 Our results showed the complex bio-effects of MARK1 on cell growth, migration and invasive pathways mediated by miR-486-5p, which were finally connected to each other. Like other protein kinases, MARKs play a role in various signaling pathways and both are regulated by multiple mechanisms, and also through MAP phosphorylation or non-canonical pathways, promoting Wnt activation27 and LKB1 signaling28 that might benefit cancer research.
MARK1 binds to other “activating” proteins or membrane-bound targets. The gain of function of this MAP in epithelial cells is regulated by MARK1 by inducing microtubule dynamics29,30 and by binding to the microtubule lattice, which can stabilize/destabilize the microtubules through dephosphorylation/phosphorylation cycles. Recently, MARK1 functions as a tumor suppressor in several types of cancers. microRNA-23a31 and miR-125a-5p25 promotes cancer cell migration and proliferation by targeting at MARK1, and our study revealed that miR-486-5p acts as an oncogene in the progression of EC by influencing MARK1.
In the present study, the miR-486-5p expression levels were markedly varied between EC and normal tissues. A larger sample size to further verify the function in EC is warranted in the future. Collectively, our results for the first time showcased that miR-486-5p regulated tumor progression through MARK1 function regulation. There are many factors that are related to the regulation of functional dynamics of EC, and whether any other signaling pathways played a role via miR-486-5p in EC was not determined. MARK1 induced the regulation of G1/S transition and the invasion system. Additional analyses are required to precisely define on how miR-486-5p induced MARK1 activation and how MARK1 regulated signaling of cell proliferation, and invasion. To address these questions and to further understand the miR-486-5p-mediated MARK1 signaling, further research should be conducted.
Conclusions
In summary, miR-486-5p functions as potential tumor stimulative factors for endometrial carcinoma progression. And MARK1 was identified as a specific target for miR-486-5p playing important roles in malignant transformation.MiR-486-5p expression inversely correlated with MARK1 expression, leads to downregulation of MARK1 consequently influencing cell proliferation, migration and invasion, suggesting a potential role for diagnosis and treatment in endometrial carcinoma tumorigenesis.
Acknowledgments
Project supported by the Ningbo Health Branding Subject Fund (PPXK2018-04), the Medical and Health Plan of Zhengjiang (grant number 2019KY166), and the Natural Science Foundation of Ning Bo (2019A610306).We would like to thank the Central Laboratory of Ningbo First Hospital.
Disclosure
Xiaojiao Zheng and Kejun Xu are co-first authors. The authors report no conflicts of interest in this work.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. doi:10.3322/caac.21387
2. Bokhman JV. Two pathogenetic types of endometrial carcinoma. Gynecol Oncol. 1983;15(1):10–17. doi:10.1016/0090-8258(83)90111-7
3. Bansal N, Yendluri V, Wenham RM. The molecular biology of endometrial cancers and the implications for pathogenesis, classification, and targeted therapies. Cancer Control. 2009;16(1):8–13. doi:10.1177/107327480901600102
4. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–233. doi:10.1016/j.cell.2009.01.002
5. Pasquinelli AE. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet. 2012;13(4):271–282. doi:10.1038/nrg3162
6. Hwang HW, Mendell JT. MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer. 2006;94(6):776–780. doi:10.1038/sj.bjc.6603023
7. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6(11):857–866. doi:10.1038/nrc1997
8. Perge P, Nagy Z, Igaz I, Igaz P. Suggested roles for microRNA in tumors. Biomol Concepts. 2015;6(2):149–155. doi:10.1515/bmc-2015-0002
9. Kanekura K, Nishi H, Isaka K, Kuroda M. MicroRNA and gynecologic cancers. J Obstet Gynaecol Res. 2016;42(6):612–617. doi:10.1111/jog.12995
10. Wang J, Tian X, Han R, et al. Downregulation of miR-486-5p contributes to tumor progression and metastasis by targeting protumorigenic ARHGAP5 in lung cancer. Oncogene. 2014;33(9):1181–1189. doi:10.1038/onc.2013.42
11. Yi Y, Lu X, Chen J, et al. Downregulated miR-486-5p acts as a tumor suppressor in esophageal squamous cell carcinoma. Exp Ther Med. 2016;12(5):3411–3416. doi:10.3892/etm.2016.3783
12. Tahiri A, Leivonen SK, Luders T, et al. Deregulation of cancer-related miRNAs is a common event in both benign and malignant human breast tumors. Carcinogenesis. 2014;35(1):76–85. doi:10.1093/carcin/bgt333
13. Li H, Mou Q, Li P, et al. MiR-486-5p inhibits IL-22-induced epithelial-mesenchymal transition of breast cancer cell by repressing Dock1. J Cancer. 2019;10(19):4695–4706. doi:10.7150/jca.30596
14. Oh HK, Tan AL, Das K, et al. Genomic loss of miR-486 regulates tumor progression and the OLFM4 antiapoptotic factor in gastric cancer. Clin Cancer Res. 2011;17(9):2657–2667. doi:10.1158/1078-0432.CCR-10-3152
15. Liu X, Chen X, Zeng K, et al. DNA-methylation-mediated silencing of miR-486-5p promotes colorectal cancer proliferation and migration through activation of PLAGL2/IGF2/beta-catenin signal pathways. Cell Death Dis. 2018;9(10):1037. doi:10.1038/s41419-018-1105-9
16. Li C, Zheng X, Li W, Bai F, Lyu J, Meng QH. Serum miR-486-5p as a diagnostic marker in cervical cancer: with investigation of potential mechanisms. BMC Cancer. 2018;18(1):61. doi:10.1186/s12885-017-3753-z
17. Goto K, Oue N, Shinmei S, et al. Expression of miR-486 is a potential prognostic factor after nephrectomy in advanced renal cell carcinoma. Mol Clin Oncol. 2013;1(2):235–240. doi:10.3892/mco.2012.46
18. He Y, Liu J, Wang Y, et al. Role of miR-486-5p in regulating renal cell carcinoma cell proliferation and apoptosis via TGF-beta-activated kinase 1. J Cell Biochem. 2019;120(3):2954–2963. doi:10.1002/jcb.26900
19. Wang LS, Li L, Li L, et al. MicroRNA-486 regulates normal erythropoiesis and enhances growth and modulates drug response in CML progenitors. Blood. 2015;125(8):1302–1313. doi:10.1182/blood-2014-06-581926
20. Sclafani RA, Holzen TM. Cell cycle regulation of DNA replication. Annu Rev Genet. 2007;41:237–280. doi:10.1146/annurev.genet.41.110306.130308
21. Umene K, Banno K, Kisu I, et al. New candidate therapeutic agents for endometrial cancer: potential for clinical practice (review). Oncol Rep. 2013;29(3):855–860. doi:10.3892/or.2013.2221
22. Toh TB, Lim JJ, Chow EK. Epigenetics in cancer stem cells. Mol Cancer. 2017;16(1):29.
23. Ma X, Wei J, Zhang L, et al. miR-486-5p inhibits cell growth of papillary thyroid carcinoma by targeting fibrillin-1. Biomed Pharmacother. 2016;80:220–226. doi:10.1016/j.biopha.2016.03.020
24. Tian F, Wang J, Ouyang T, et al. MiR-486-5p serves as a good biomarker in nonsmall cell lung cancer and suppresses cell growth with the involvement of a target PIK3R1. Front Genet. 2019;10:688. doi:10.3389/fgene.2019.00688
25. Natalia MA, Alejandro GT, Virginia TJ, Alvarez-Salas LM. MARK1 is a novel target for miR-125a-5p: implications for cell migration in cervical tumor cells. Microrna. 2018;7(1):54–61. doi:10.2174/2211536606666171024160244
26. Manning AL, Dyson NJ. pRB, a tumor suppressor with a stabilizing presence. Trends Cell Biol. 2011;21(8):433–441. doi:10.1016/j.tcb.2011.05.003
27. Wallingford JB, Habas R. The developmental biology of Dishevelled: an enigmatic protein governing cell fate and cell polarity. Development. 2005;132(20):4421–4436. doi:10.1242/dev.02068
28. Goodwin JM, Svensson RU, Lou HJ, Winslow MM, Turk BE, Shaw RJ. An AMPK-independent signaling pathway downstream of the LKB1 tumor suppressor controls Snail1 and metastatic potential. Mol Cell. 2014;55(3):436–450. doi:10.1016/j.molcel.2014.06.021
29. Hammond JW, Huang CF, Kaech S, Jacobson C, Banker G, Verhey KJ. Posttranslational modifications of tubulin and the polarized transport of kinesin-1 in neurons. Mol Biol Cell. 2010;21(4):572–583. doi:10.1091/mbc.e09-01-0044
30. Witte H, Bradke F. The role of the cytoskeleton during neuronal polarization. Curr Opin Neurobiol. 2008;18(5):479–487. doi:10.1016/j.conb.2008.09.019
31. Tang X, Yang M, Wang Z, Wu X, Wang D. MicroRNA-23a promotes colorectal cancer cell migration and proliferation by targeting at MARK1. Acta Biochim Biophys Sin (Shanghai). 2019;51(7):661–668. doi:10.1093/abbs/gmz047
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.