Back to Journals » OncoTargets and Therapy » Volume 13
MiR-483 Targeted SOX3 to Suppress Glioma Cell Migration, Invasion and Promote Cell Apoptosis
Authors Lu S, Yu Z, Zhang X, Sui L
Received 30 November 2019
Accepted for publication 1 February 2020
Published 9 March 2020 Volume 2020:13 Pages 2153—2161
DOI https://doi.org/10.2147/OTT.S240619
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Gaetano Romano
Shujing Lu,1 Zhengyang Yu,2 Xia Zhang,2 Lingling Sui2
1Department of Critical Care Medicine, Liaocheng Third People’s Hospital, Liaocheng 252000, Shandong, People’s Republic of China; 2Department of Internal Neurology, Liaocheng Third People’s Hospital, Liaocheng 252000, Shandong, People’s Republic of China
Correspondence: Lingling Sui
Department of Internal Neurology, Liaocheng Third People’s Hospital, No. 62 Weiyu Road, Liaocheng 252000, Shandong, People’s Republic of China
Tel +86 635-8385120
Email [email protected]
Objective: Glioma is the most common malignant brain tumor that has high aggressiveness. The aim of this study was to investigate the potential therapeutic targets for gliomas.
Materials and Methods: Real-time quantitative polymerase chain reaction (RT-qPCR) was employed to calculate the expression of miRNA and genes. The connection between the expression of miR-483 and patients’ overall survival rate was evaluated using Kaplan–Meier analysis. In addition, the underlying mechanism was detected using luciferase assay.
Results: The expression level of miR-483 was significantly decreased in glioma tissue samples and cell lines, compared to the adjacent tissues and normal cell lines. Downregulation of miR-483 or upregulation of SOX3 was associated with overall survival of glioma patients. Additionally, overexpression of miR-483 promotes cell invasion and migration and inhibits apoptosis. In addition, miR-483 directly targeted to SOX3, and the expression of miR-483 has a negative correlation with SOX3 in glioma tissues. SOX3 reversed partial functions of miR-483 on cell migration, invasion, and promoted cell apoptosis in glioma.
Conclusion: MiR-483 inhibited glioma cell migration, invasion, and promoted glioma cell apoptosis by targeting SOX3. MiR-483 maybe acted as a potential target for the treatment of glioma.
Keywords: miR-483, SOX3, glioma, apoptosis, invasion
Introduction
Glioma, the most prevalent and aggressive primary malignant brain tumor, accounts for almost 28% of all central nervous system tumors according to the statistics of the American Brain Tumor Registry (CBTRUS).1,2 Currently, clinical treatment of glioma includes surgery, radiation therapy, chemotherapy; however, the prognosis of glioma is still poor, and the recurrence rate is quite high after initial treatment.3,4 Therefore, it is important to look for potential therapeutic targets for gliomas to improve current diagnostic and therapeutic status.
MicroRNA (miRNA), a type of single-strand noncoding RNA approximately 22 nt in length, regulates the expression of a number of genes in human and other organisms by interacting with target mRNA 3ʹ-UTR and interrupting protein generating.5,6 Increasing studies have indicated that miRNAs were involved in the process of glioma, including proliferation, cell cycle and apoptosis.7,8 As a tumor suppressor gene, miR-483 can impact multiple tumors, which are generally downregulated in tumors and exerts inhibiting tumor efficacy. For instance, miR-483 regulated oxaliplatin resistance through targeting FAM171B in human colorectal cancer.9 Similarly, miR-483 decreases the radiosensitivity of nasopharyngeal carcinoma cells by targeting DAPK1.10 Besides, a research demonstrated that miR-483 suppressed cell proliferation via oncogene HDAC8 in human triple-negative breast cancer.11 In addition, miR-483 inhibited cell proliferation, invasion in gastric cancer via OGT.12 However, the roles of miR-483 in the recurrence and metastasis of glioma remain unknown.
SRY-box transcription factor 3 (SOX3) encodes a member of the SRY-related HMG-box (SOX) family of transcription factors, which involved in the regulation of embryonic development and in the determination of the cell fate.13,14 In recent years, increasing evidence has been provided to suggest the involvement of SOX3 in tumorigenesis. SOX3 acted as an oncogene by promoting cells proliferation, migration, and invasion, while restrained apoptosis and adhesion of ovarian cancer.15 Ectopic expression of Sox3 causes oncogenic transformation of chicken embryo fibroblasts.16 Overexpression of SOX3 in ESCC cells could significantly promote the proliferation, invasion, migration and tube formation of lymphatic endothelial cells.17 In this study, we found that miR-483 was low expressed and SOX3 was overexpressed in glioma. MiR-483 suppressed cell migration, invasion and promoted cell apoptosis of LN229 cells via targeting SOX3.
Materials and Methods
Patients and Samples
A collection of 40 glioma patients were selected from Liaocheng Third People’s Hospital and obtained pairs of glioma tissue samples and corresponding adjacent tissue samples. No radiotherapy or chemotherapy was performed before the surgery. The study was approved by the Ethics Committee of Liaocheng Third People’s Hospital and written informed consent was obtained from each patient. All the experiments were performed in accordance with the Declaration of Helsinki and subsequent updates.
Cell Culture
The normal-immortalized gliocyte HEB and two glioma cells LN18 and LN229 were obtained from the American Type Culture Collection (ATCC, Rockville, USA). All the cells were cultured in Roswell Park Memorial Institute 1640 (RPMI 1640) medium (Gibico, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS; Thermo Fisher Scientific, Waltham, MA, USA), penicillin (100 U/mL) and streptomycin (100 μg/mL), which were maintained at 37°C in an incubator with a humidified atmosphere with 5% CO2.
Cell Transfection
LN229 cells were seeded in 6-well plates and cultured to a density of 70%. The miR-483 mimic, miR-483 inhibitor and negative control were purchased from GenePharma, while pcDNA3.1-SOX3 and pcDNA3.1-NC were purchased from RiboBio (Guangzhou, China). Their sequences were as follows: miR-483 mimic: sense 5ʹ-UCACUCCUCUCCUCCCGUCUU-3ʹ, antisense 5ʹ-GACGGGAGGAGAGGAGUGAUU-3ʹ; NC mimic: sense 5ʹ-UUCUCCGAACGUGUCACGUTT-3ʹ, antisense 5ʹ-ACGUGACACGUUCGGAGAATT-3ʹ; miR-483 inhibitor: 5ʹ-AAGACGGGAGGAGAGGAGUGA-3ʹ; NC inhibitor: 5ʹ-CAGUACUUUUGUGUAGUACAA-3ʹ. Cells were transfected with Lipofectamine 3000 (Invitrogen; Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol.
Luciferase Reporter Assays
To investigate that miR-483 directly targeting to the mRNA 3ʹ-untranslated region (3ʹ-UTR) of SOX3, the putative binding site on SOX3 mRNA was mutated using Quick-change site-directed mutagenesis kit (Stratagene, La Jolla, CA, USA), and both the wild type and the mutant were amplified and cloned into the pGL3 vector (Promega, Madison, WI, USA). The wild type or the mutant 3ʹ-UTR were co-transfected into LN229 cell with miR-483 mimic or negative control. After co-transfection for 48 h, luciferase activity was detected using a Dual-Luciferase reporter assay system (Promega, Madison, WI, USA).
RNA Extraction and Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR)
Pursuant to the command of the manufacturer, TRIzol reagent (Ambion, Thermo Fisher Scientific, Waltham, MA, USA) was used to extract total RNA. The reverse transcription system (Promega Corporation, Madison, WI, USA) was conducted to perform the reverse transcription of RNA to cDNA. According to the manufacturer’s protocol, RT-qPCR was then performed using SYBR Premix Ex Taq TM (Takara, Tokyo, Japan) on an Mx3000P real-time PCR system (Applied Biosystems; Thermo Fisher Scientific, Waltham, MA, USA). The expression of miR-483 and SOX3 mRNA were normalized using U6 and glyceraldehyde 3-phosphate dehydrogenase (GAPDH), respectively. The primer sequences for the detection of mRNA expression were as follows: miR-483 Forward: 5ʹ- CTCTCGTAACTGGCTAGTCAAGAGAGTCATTTGACTCGCA-3ʹ, Reverse: 5ʹ-CTGTACTCATGAGGTGGTTGACGCAGGAGG-3ʹ; U6 Forward: 5ʹ-CTCGCTTCGGCAGCACA-3ʹ, Reverse: 5ʹ-AACGCTTCACGAATTTGCGT-3ʹ; SOX3 Forward: 5ʹ-GACCTGTTCGAGAGAACTCATCA-3ʹ, Reverse: 5ʹ-CGGGAAGGGTAGGCTTATCAA-3ʹ; GAPDH Forward: 5ʹ-GTCTCCTCTGACTTCAACAGC-3ʹ and Reverse: 5ʹ-ACCACCCTGTTGCTGTAGCCAA-3ʹ.
Western Blot Analysis
Radioimmunoprecipitation assay (RIPA) Lysis Buffer (Sigma, St. Louis, MO, USA) containing 10% phenylmethylsulfonyl fluoride (PMSF) (Sigma, St. Louis, MO, USA) was conducted to lyse total proteins. All the protein separation was using SDS-PAGE and then the blots were electro-transferred to polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA, USA). After being blocked by 5% fat-free milk at ream temperature for 1 h, the membranes were incubated with primary antibodies. The primary antibodies were against SOX3 (1:1000; Abcam, Cambridge, MA, USA), BCL-2 (1:1000, Abcam), Bax (1:1000, Abcam), Caspase-3 (1:1000, Abcam). Subsequently, the blots were incubated by secondary anti-rabbit HRP-conjugated antibody (Cell Signaling, Danvers, MA, USA). Enhanced chemiluminescence detection Kit (ECL, Pharmacia Biotech, Arlington, USA) was applied to capture the protein signals.
Transwell Assay
The cell invasion ability was calculated by the transwell insert (8 μm membrane, Corning, Cambridge, MA, USA), which were placed in 24-well plate and formed up and low two chambers. The upper chamber was added 200 μL LN229 cells suspension that were suspended by serum-free medium, whereas the lower chamber was filled with 500 μL medium containing 15% FBS. After the cells were incubated 24 h at 37°C, the non-invaded cells were removed by cotton swabs, while the invaded cells were fixed using 4% paraformaldehyde and then stained by 10% crystal violet. Finally, a microscope (Olympus Corporation, Tokyo, Japan) was utilized to count the cells number.
Statistical Analysis
All data are presented as the mean ± standard deviation (SD) of at least three experimental repeats. SPSS software package v.9.05 (SPSS Inc. Chicago, IL, USA) and GraphPad Prism software (version 6.0; GraphPad Prism Software, San Diego, CA, USA) were used to carry out statistical analysis. Analyses were performed using Student’s t-test with GraphPad prism 7 software. Differences between two groups were analyzed by using the Student’s t-test. Comparison between multiple groups was done using one-way ANOVA test followed by Post Hoc Test (Least Significant Difference). P<0.05 was considered to indicate a statistically significant difference.
Results
MiR-483 Expression Level in Glioma Tissues and Cell Lines
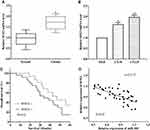
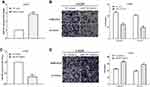
Previous studies have demonstrated the role of miR-483 in other cancers; thus, we first measured the expression of miR-483 by using RT-qPCR in 40 pairs of glioma and paracancerous tissues. As expected, miR-483 expression was significantly lower in tumor tissue samples than that of paracancerous tissues (P<0.05) (Figure 1A). Next, Kaplan–Meier analysis was used to calculate the overall survival, and we discovered that glioma patients with lower miR-483 level had a worse overall survival (P<0.05) (Figure 1B). Meanwhile, the expression of miR-483 was evaluated in two glioma cell lines LN18 and LN229 and a normal-immortalized gliocyte HEB by RT-qPCR. Not unfortunately, miR-483 was generally up-regulated in HEB cells compared with LN18 (P<0.05) and LN229 (P<0.01) (Figure 1C). LN229 cells had the lowest level of miR-483; therefore, LN229 was chosen as the experimental cell.
MiR-483 Inhibited LN229 Cell Migration, Invasion
To study the effects of miR-483 on glioma cells, miR-483 mimic or miR-483 inhibitor was transfected into LN229 cells, and the transfection efficiency was measured by RT-qPCR. The results showed a remarkable increased in miR-483 mimic-treated LN229 cells (P<0.05) (Figure 2A), while miR-483 was downregulated by treating miR-483 inhibitor (P<0.05) (Figure 2C). Transwell assay was used to detect cell migration and invasion after transfecting miR-483 mimic or miR-483 inhibitor. We found that overexpression of miR-483 could remarkably suppressed LN229 cell migration (P<0.05) and invasion (P<0.05) compared with the control groups (Figure 2B). In contrast, cell migration (P<0.05) and invasion (P<0.05) were enhanced by miR-483 inhibitor compared to the negative control (Figure 2D).
MiR-483 Promoted LN229 Cell Apoptosis in vitro
The effect of miR-483 on cell apoptosis was also examined. The apoptosis-related proteins such as Bcl-2, Bax and cleaved caspase-3 were assessed by RT-qPCR and Western blotting. And we found that upregulation of miR-483 enhanced the expression of Bcl-2, and reduced Bax and caspase-3 expression (Figure 3A–D). On the contrary, miR-483 inhibitor reduced Bcl-2 expression while promoted Bax and caspase-3 expression in LN229 cells (Figure 3E–H). All the results demonstrated that the apoptosis of LN229 cells was significantly repressed by miR-483.
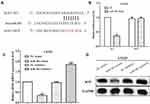
SOX3 Was a Predicted Target of miR-483 in LN229 Cells
To investigate the mechanism underlying miR-483 in glioma, TargetScan was applied to predict the target genes of miR-483, and we discovered SOX3 as a potential target gene (Figure 4A). In order to verify that miR-483 directly binding to the 3ʹuntranslated region (3ʹUTR), the presumptive binding site was mutated and followed calculated the luciferase activity. The Luciferase assay indicated that the luciferase activity of cells co-transfected with wild type SOX3 and miR-483 mimic was reduced, while no changes were observed in mutant SOX3 cells (Figure 4B). Furthermore, RT-qPCR and Western blotting were employed to measure the expression of SOX3 after altering miR-483 in LN229 cells. As we speculation, both the mRNA and protein levels of SOX3 was significantly decreased in miR-483 mimic group, whereas it was increased by miR-483 inhibitor (Figure 4C and D).
The Expression of SOX3 in Glioma Tissues and Cell Lines
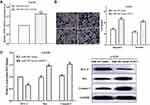
The expression of SOX3 was evaluated in glioma tissues and normal tissues by RT-qPCR. RT-qPCR result showed that the expression level of SOX3 in glioma was significantly higher compared with the normal tissues (P<0.05) (Figure 5A). The association between the expression of SOX3 and the overall survival of the glioma patients were calculated, and we found that upregulation of SOX3 predicted poor outcome (P<0.05) (Figure 5B). The mRNA level of SOX3 in cell lines was also detected, and we discovered that SOX3 expression was higher in glioma cell lines LN18 (P<0.05) and LN229 (P<0.05) than normal cell line HEB (Figure 5C). Subsequently, correlation analysis was performed to analyze the relationship between the expression of miR-483 and SOX3 in glioma tissues. Not unfortunately, the expression of miR-483 had a negative correlation with SOX3 expression (P<0.05) (Figure 5D).
Elevated Expression of SOX3 Reversed the Inhibitory Effect of Migration, Invasion and Cell Apoptosis of LN229 Cells by miR-483 Mimic
To validate that miR-483 works through SOX3, the changes in cell migration and invasion were evaluated under the co-transfection of miR-483 mimic and pcDNA3.1-SOX3 (Figure 6A). The results shown that upregulation of SOX3 significantly partially recovered the impaired effects of miR-483 mimic on the migratory and invasive ability in LN229 cells (P<0.01) (Figure 6B). Meanwhile, compared with only transfected with miR-483 mimic, the cell apoptosis ability was also inhibited after co-transfection with miR-483 and SOX3 by reducing BCL-2 expression, and enhanced Bax and Caspase-3 expression (P<0.05) (Figure 6C). Collectively, overexpression of SOX3 partially suppressed the suppressive roles of miR-483 on LN229 cell migration, invasion and inhibited cell apoptosis.
Discussion
Glioma is the most frequent primary intracranial tumor in the central nervous system. Due to the high invasiveness, the prognosis for patients remains poor.18 Thus, a better understanding of the molecular mechanisms of cancer development and progression, it is urgent to explore new markers for the diagnosis and treatment of glioma.
Increasing evidences have been demonstrated that miRNAs are abnormally expressed in many diseases, especially in cancer.19,20 In fact, it has been reported that a variety of miRNAs can function as biomarkers for the diagnosis and prognosis of human gliomas.21 In recent years, miR-483 has been demonstrated to be downregulated in colon cancer, hepatocellular carcinoma and gastric cancer,9,12,22 while it was upregulated in anaplastic thyroid cancer and Wilms’ Tumor,23,24 suggesting that the expression of miR-483 is distinct in different types of cancers. In this study, we found that miR-483 was low expressed in glioma tissues compared to the paracancerous normal tissues. Kaplan–Meier analysis demonstrated that downregulation of miR-483 has a negative connection with poor prognosis, which was opposite of the results in esophageal squamous cell carcinoma.25 A previous study has demonstrated that miR-483 inhibited cell proliferation and migration in colon cancer.26 Wang et al have indicated that miR-483 suppressed the proliferation of glioma cells.27 However, the effect of miR-483 in glioma was still unknown. Thus, to explore the functions of miR-483 in glioma, miR-483 mimic and miR-483 inhibitor were conducted to up- or down-regulate the expression of miR-483 in LN229 cells. In this study, we found that upregulation of miR-483-inhibited cell migration and invasion in LN229 cells, while downregulation of miR-483 promoted cell migration and invasion. Apoptosis is the orderly death of cells controlled by genes in order to maintain internal environment stability. BCL, Bax and Caspase-3 were cell apoptosis-related genes. In this study, we found that miR-483 overexpression suppressed cell apoptosis by upregulating BCL-2 expression and reducing the expression of Bax and Caspase-3, while low expression of miR-483 presents opposite results.
To explore the molecular mechanism, we used TargetScan to predict the target gene of miR-483, and we selected SOX3 as a candidate target gene. Luciferase assay revealed that miR-483 mimic reduced the luciferase activity of wild type SOX3. In addition, miR-483 mimic decreased the expression of SOX3, whereas miR-483 inhibitor enhanced it. MiR-483 had a negative connection with SOX3 in glioma tissues. All the results indicated that miR-483 directly targeted to the 3ʹUTR of SOX3, which was consistent with that in breast cancer.28 SOX3 was upregulated and played as an oncogene in several cancers, such as osteosarcoma and esophageal squamous cell carcinoma.17,29 We also discovered that SOX3 was overexpressed in glioma tissues and cell lines in comparison with normal tissues and normal-immortalized gliocyte HEB. Overexpression of SOX3 was associated with poor glioma overall survival, which was consistent with that in esophageal squamous cell carcinoma.30 Based on previous studies, SOX3 promoted the proliferation, viability, migration and invasion of glioblastoma cells.31 In this study, we found that overexpression of SOX3 reversed miR-483 mimic-induced cell migration, invasion and apoptosis. Therefore, miR-483 targeted SOX3 to inhibit glioma invasion and migration, and promote apoptosis in glioma.
Conclusions
Our study found that miR-483 directly regulated the expression of SOX3, inhibited invasion, migration and promotes apoptosis of glioma cells, suggesting that miR-483 can be a potential target for glioma treatment.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Wen PY, Huse JT. 2016 World Health Organization classification of central nervous system tumors. Continuum (Minneap Minn). 2017;23(6, Neuro–oncology):1531–1547. doi:10.1212/CON.0000000000000536
2. Stupp R, Mason WP, van den Bent MJ, et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 2005;352(10):987–996. doi:10.1056/NEJMoa043330
3. Weller M, Cloughesy T, Perry JR, Wick W. Standards of care for treatment of recurrent glioblastoma–are we there yet? Neuro Oncol. 2013;15(1):4–27. doi:10.1093/neuonc/nos273
4. Kumar AA, Abraham KA. Regression of recurrent high-grade glioma with temozolomide, dexamethasone, and levetiracetam: case report and review of the literature. World Neurosurg. 2017;108:911–990. doi:10.1016/j.wneu.2017.08.136
5. Zhang WC, Liu J, Xu X, Wang G. The role of microRNAs in lung cancer progression. Med Oncol. 2013;30(3):675. doi:10.1007/s12032-013-0675-8
6. Guz M, Rivero-muller A, Okon E, et al. MicroRNAs-role in lung cancer. Dis Markers. 2014;2014:218169. doi:10.1155/2014/218169
7. Carthew RW, Sontheimer EJ. Origins and Mechanisms of miRNAs and siRNAs. Cell. 2009;136(4):642–655. doi:10.1016/j.cell.2009.01.035
8. Luo H, Xu R, Chen B, et al. MicroRNA-940 inhibits glioma cells proliferation and cell cycle progression by targeting CKS1. Am J Transl Res. 2019;11(8):4851–4865.
9. Liang H, Xu Y, Zhang Q, et al. MiR-483-3p regulates oxaliplatin resistance by targeting FAM171B in human colorectal cancer cells. Artif Cells Nanomed Biotechnol. 2019;47(1):725–736. doi:10.1080/21691401.2019.1569530
10. Tian Y, Yan M, Zheng J, et al. miR-483-5p decreases the radiosensitivity of nasopharyngeal carcinoma cells by targeting DAPK1. Lab Invest. 2019;99(5):602–611. doi:10.1038/s41374-018-0169-6
11. Menbari MN, Rahimi K, Ahmadi A, et al. miR-483-3p suppresses the proliferation and progression of human triple negative breast cancer cells by targeting the HDAC8 oncogene. J Cell Physiol. 2019. doi:10.1002/jcp.29167
12. Yu FY, Zhou CY, Liu YB, Wang B, Mao L, Li Y. miR-483 is down-regulated in gastric cancer and suppresses cell proliferation, invasion and protein O-GlcNAcylation by targeting OGT. Neoplasma. 2018;65(3):406–414. doi:10.4149/neo_2018_170608N411
13. Schepers GE, Teasdale RD, Koopman P. Twenty pairs of sox: extent, homology, and nomenclature of the mouse and human sox transcription factor gene families. Dev Cell. 2002;3(2):167–170. doi:10.1016/s1534-5807(02)00223-x
14. Dong C, Wilhelm D, Koopman P. Sox genes and cancer. Cytogenet Genome Res. 2004;105(2–4):442–447. doi:10.1159/000078217
15. Yan Q, Wang F, Miao Y, et al. Sex-determining region Y-box3 (SOX3) functions as an oncogene in promoting epithelial ovarian cancer by targeting Src kinase. Tumour Biol. 2016;37(9):12263–12271. doi:10.1007/s13277-016-5095-x
16. Xia Y, Papalopulu N, Vogt PK, Li J. The oncogenic potential of the high mobility group box protein Sox3. Cancer Res. 2000;60(22):6303–6306.
17. Zheng YF, Li K, Cai QY, et al. The effect of high Sox3 expression on lymphangiogenesis and lymph node metastasis in esophageal squamous cell carcinoma. Am J Transl Res. 2017;9(6):2684–2693.
18. Pan Y, Yuan F, Li Y, Wang G, Lin Z, Chen L. Bromodomain PHDfinger transcription factor promotes glioma progression and indicates poor prognosis. Oncol Rep. 2019;41(1):246–256. doi:10.3892/or.2018.6832
19. Petrescu G, Sabo AA, Torsin LI, Calin GA, Dragomir MP. MicroRNA based theranostics for brain cancer: basic principles. J Exp Clin Cancer Res. 2019;38(1):231. doi:10.1186/s13046-019-1180-5
20. Si W, Shen J, Zheng H, Fan W. The role and mechanisms of action of microRNAs in cancer drug resistance. Clin Epigenetics. 2019;11(1):25. doi:10.1186/s13148-018-0587-8
21. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–233. doi:10.1016/j.cell.2009.01.002
22. Cao N, Mu L, Yang W, Liu L, Liang L, Zhang H. MicroRNA-298 represses hepatocellular carcinoma progression by inhibiting CTNND1-mediated Wnt/β-catenin signaling. Biomed Pharmacother. 2018;106:483–490. doi:10.1016/j.biopha.2018.06.135
23. Zhang X, Liu L, Deng X, et al. MicroRNA 483-3p targets Pard3 to potentiate TGF-beta1-induced cell migration, invasion, and epithelial-mesenchymal transition in anaplastic thyroid cancer cells. Oncogene. 2019;38(5):699–715. doi:10.1038/s41388-018-0447-1
24. Che G, Gao H, Tian J, Hu Q, Xie H, Zhang Y. MicroRNA-483-3p promotes proliferation, migration, and invasion and induces chemoresistance of Wilms’ tumor cells. Pediatr Dev Pathol. 2019;2118459181. doi:10.1177/1093526619873491
25. Xue L, Nan J, Dong L, et al. Upregulated miR-483-5p expression as a prognostic biomarker for esophageal squamous cell carcinoma. Cancer Biomarkers. 2017;19(2):193–197. doi:10.3233/CBM-160506
26. Niu ZY, Li WL, Jiang DL, Li YS, Xie XJ. Mir-483 inhibits colon cancer cell proliferation and migration by targeting TRAF1. Kaohsiung J Med Sci. 2018;34(9):479–486. doi:10.1016/j.kjms.2018.04.005
27. Wang L, Shi M, Hou S, et al. MiR-483-5p suppresses the proliferation of glioma cells via directly targeting ERK1. FEBS Lett. 2012;586(9):1312–1317. doi:10.1016/j.febslet.2012.03.035
28. Cui K, Zhang H, Wang GZ. MiR-483 suppresses cell proliferation and promotes cell apoptosis by targeting SOX3 in breast cancer. Eur Rev Med Pharmacol Sci. 2019;23(5):2069–2074. doi:10.26355/eurrev_201903_17248
29. Guo Y, Yin J, Tang M, Yu X. Downregulation of SOX3 leads to the inhibition of the proliferation, migration and invasion of osteosarcoma cells. Int J Oncol. 2018;52(4):1277–1284. doi:10.3892/ijo.2018.4278
30. Li K, Wang RW, Jiang YG, Zou YB, Guo W. Overexpression of Sox3 is associated with diminished prognosis in esophageal squamous cell carcinoma. Ann Surg Oncol. 2013;20 Suppl 3:S459–S466. doi:10.1245/s10434-012-2792-6
31. Marjanovic VJ, Drakulic D, Garcia I, et al. SOX3 can promote the malignant behavior of glioblastoma cells. Cell Oncol (Dordr). 2019;42(1):41–54. doi:10.1007/s13402-018-0405-5
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.