Back to Journals » OncoTargets and Therapy » Volume 11
miR-19b serves as a prognostic biomarker of breast cancer and promotes tumor progression through PI3K/AKT signaling pathway
Authors Li C, Zhang J, Ma Z, Zhang F, Yu W
Received 13 April 2018
Accepted for publication 26 May 2018
Published 16 July 2018 Volume 2018:11 Pages 4087—4095
DOI https://doi.org/10.2147/OTT.S171043
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Samir Farghaly
Chuansheng Li,1,* Jingwei Zhang,1,* Zhongliang Ma,2 Fan Zhang,1 Wenlong Yu1
1Department of General Surgery, Shanxian Central Hospital, Heze, Shandong, China; 2Department of Breast Surgery, The Affiliated Hospital of Qingdao University, Qingdao, Shandong, China
*These authors contributed equally to this work
Background: MicroRNAs (miRNAs) are involved in tumor progression of various human malignancies. MicroRNA-19b (miR-19b) has been described as serving a crucial role in tumorigenesis of breast cancer. The purpose of this study was to investigate the expression patterns, clinical value, and functional role of miR-19b in breast cancer.
Methods: Expression of miR-19b was estimated by quantitative real time PCR. Kaplan–Meier survival analysis and Cox regression assay were performed to explore the prognostic value of miR-19b. The functional role of miR-19b was verified using cell experiments.
Results: Upregulated miR-19b expression was observed in breast cancer tissues and cells compared with the controls (all P<0.05). The miR-19b expression was associated with distant metastasis and TNM stage. The survival curves showed that high miR-19b was correlated with poor overall survival of the patients (log-rank P=0.002). Furthermore, miR-19b was proven to be an independent prognostic factor for patients. By using miR-19b mimic and inhibitor, cell proliferation, migration, and invasion were enhanced by miR-19b overexpression but were suppressed by reduction of miR-19b (all P<0.05). Besides, PI3K/AKT was demonstrated to be activated by miR-19b in breast cancer cells.
Conclusion: The overexpression of miR-19b serves as a candidate prognostic biomarker of breast cancer and may be involved in the tumor progression through PI3K/AKT pathway.
Keywords: microRNA-19b, prognosis, progression, breast cancer
Introduction
Breast cancer is the most frequently diagnosed malignancy in women worldwide and ranks as the second leading cause of cancer death among females.1 Patients with breast cancer usually have some typical signs, such as a change in breast shape, a lump in the breast, fluid coming from the nipple or dimpling of the skin, which affect their mental and physical health.2 Several risk factors have been reported to be closely correlated with the occurrence of breast cancer, including obesity, alcohol abuse, lack of physical exercise, childbearing absence or family history.3 Although great efforts have been made regarding cancer treatment, the morbidity and mortality of breast cancer are increasing on a yearly basis.4 Currently, advanced therapeutic strategies are presented, such as operative surgery, chemotherapy, radiotherapy, and hormonotherapy, however, the prognosis and outcomes of patients with breast cancer remain dismal.5 Therefore, it is crucial to improve the prognosis to meet the clinical requirements for breast cancer.
MicroRNAs (miRNAs) represent a series of small and non-coding RNA molecules with important gene expression regulatory functions on post-transcriptional levels.6–8 By targeting the 3′ untranslated region (3′-UTR) of the targeted messenger RNAs (mRNAs), miRNAs are involved in the translation or degradation of the mRNAs, thus modulating expression of genes.9 Evidence has revealed that miRNAs serve pivotal roles in diverse biological processes, such as cell proliferation, differentiation, migration, invasion, and apoptosis.10,11 Moreover, miRNAs with abnormal expression levels are also involved in tumorigenesis and tumor progression.12 Also, aberrant miRNAs have attracted a lot of attention regarding their diagnostic and prognostic value for various malignancies.13,14 Thus, identification of novel miRNAs, which can be used for outcome prediction, is important for breast cancer prognosis. A study by Liu et al15 indicated that microRNA-19b (miR-19b) could promote tumorigenesis of breast cancer cells. However, data about the clinical significance of miR-19b have rarely been reported, until now.
In this study, we sought to assess the expression patterns of miR-19b in breast cancer tissues and cells, as well as its prognostic value for cancer patients. Additionally, the effects of miR-19b on biological behaviors of breast cancer cells were also analyzed.
Materials and methods
Patients and sample collection
This research was approved by the Ethics Committee of Shanxian Central Hospital. The experimental protocols were in accordance with the guidelines of the Ethics Committee of Shanxian Central Hospital. All patients signed informed consent. A total of 124 patients who were pathologically diagnosed as having breast cancer between October 2007 and February 2011 in Shanxian Central Hospital, were included in this study. None of the patients had received any preoperative therapy. One hundred and twenty-four paired breast cancer tissues and adjacent normal tissues were collected from the patients during surgery and snap frozen in liquid nitrogen for further analyses. The clinicopathological characteristics of the patients were recorded and listed in Table 1. Estrogen receptor (ER), progesterone receptor (PR), and HER-2 status were determined by immunohistochemistry analysis. After surgery, all patients were followed-up by means of a 5-year follow-up survey, during which the survival information of the patients was obtained by telephone.
  | Table 1 Association of miR-19b with the clinicopathological features of breast cancer patients |
Cell culture and transfection
Human breast cancer cell lines MCF-7, BT474, MDA-MB-231, T47D, and a normal breast epithelial cell line MCF-10A were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). The four breast cancer cell lines were cultured in DMEM supplemented with 10% fetal bovine serum (FBS), and the MCF-10A was maintained in RPMI 1640 medium supplemented with 10% FBS. All the cells were kept at 37°C in a humidified incubator with 5% CO2.
The breast cancer cells were seeded in 24-well plates and cultured overnight, then transfected with miR-19b mimic, miR-19b inhibitor or the corresponding negative controls of mimic and inhibitor (mimic NC and inhibitor NC) using Lipofectamine 2000 (Thermo Fisher Scientific, Waltham, MA, USA) following the manufacturer’s protocols. The miR-19b mimic, miR-19b inhibitor, and the NCs were purchased from Thermo Fisher Scientific.
RNA extraction and quantitative real time PCR (qRT-PCR)
Total RNA in the tissues and cells were isolated using Trizol Reagent (Thermo Fisher Scientific). The absorbance of RNA at 260 and 280 nm was measured to evaluate the quality and concentration of the RNA. Only the RNA with an OD absorption (A)260/A280 ratio of close to 2.0 was used in the subsequent analyses.
Single stranded cDNA was synthesized from the RNA using a PrimeScript RT reagent kit (TaKaRa, Kyoto, Japan) in compliance with the manufacturer’s instructions and then stored at −20°C. To estimate the expression of miR-19b, qRT-PCR was carried out using SYBR Green I Master Mix kit (Thermo Fisher Scientific) and 300 Real-Time PCR System (Thermo Fisher Scientific). In the reactions, U6 was used as the internal control gene. The final relative expression of miR-19b was calculated using 2−ΔΔCt method and normalized to U6.
MTT assay
The proliferation of breast cancer cells was investigated using MTT assay. The stably transfected cells were seeded in a 96-well plate at a density of 4×105 cells/well and incubated at 37°C in a humidified incubator with 5% CO2 for 3 days. An amount of 10 μL MTT (5 mg/mL) was added to each well every 24 hours and then incubated for a further 4 hours. After the incubation, the media were removed and 150 μL dimethylsulfoxide was added to the wells. The cell proliferation was evaluated by measuring the absorbance at 490 nm using a microplate reader. The MTT assay was repeated three times.
Transwell assay
Transwell assay was performed to examine the cell migration and invasion using Transwell chambers (8 μm pore size; Corning Incorporated, Corning, NY, USA). The membranes used for invasion analysis were coated with Matrigel (Corning Incorporated). The stably transfected cells (4×105 cells/well) were seeded into the upper chamber with serum-free medium. The lower chamber was full of medium supplemented with 10% FBS, which served as the chemo-attractant. The chambers were incubated at 37°C for 48 hours, and the number of cells in the lower chambers were counted using light microscopy.
Western blot analysis
The cells transfected with the corresponding vectors were lysed using RIPA buffer (Applygen, Beijing, China). The proteins in the lysates were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis and then transferred to polyvinylidene fluoride membranes (Corning Incorporated). The proteins were first blocked using 5% non-fat dry milk for 1 hour at room temperature and then incubated with primary antibodies (anti-PI3K, anti-p-PI3K, anti-AKT, anti-p-AKT, anti-CCND1) (Applygen) at 4°C overnight. The membranes were washed with tris buffered saline Tween, then incubated with secondary antibodies (Applygen) for 1 hour at room temperature. An electrochemiluminescence detection system was used to visualize the protein bands, and the protein expression levels were measured by Image-Pro Plus 6.0 software (Media Cybernetics, Rockville, MD, USA).
Statistical analysis
The differences between groups were analyzed using Student’s t-test or one-way analysis of variance (ANOVA). The relationship between expression of miR-19b and clinicopathological features was assessed using chi-squared test. Survival analysis was performed for breast cancer patients using Kaplan–Meier methods and log-rank test. Multivariate Cox regression analysis was carried out to confirm the prognostic value of miR-19b in cancer patients. All the statistical analyses were conducted using SPSS version 18.0 software (SPSS Inc., Chicago, IL, USA) and GraphPad Prism 5.0 software (GraphPad Software, Inc., La Jolla, CA, USA). Data used in these analyses were expressed as the mean ± SD. Statistical significance was set at P<0.05.
Results
Overexpression of miR-19b in breast cancer tissues and cell lines
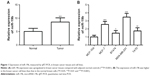
The expression of miR-19b measured by qRT-PCR was found to be upregulated in breast cancer tissues compared with the adjacent normal controls (P<0.001, Figure 1A). Moreover, miR-19b expression was further confirmed in breast cancer cell lines by qRT-PCR. As shown in Figure 1B, the expression levels of miR-19b in the four breast cancer cell lines (MCF-7, BT474, MDA-MB-231, and T47D) were also higher than that in the normal cell line, MCF-10A (all P<0.05).
Association of miR-19b expression with clinicopathological features of breast cancer patients
In addition to the expression patterns of miR-19b, the study also focused on its relationship with clinicopathological characteristics of patients. MiR-19b expression was classified into low expression group (n=56) and high expression group (n=68) based on the mean miR-19b expression level (8.523). According to the chi-squared test, miR-19b expression was found to be associated with distant metastasis (P=0.010) and TNM stage (P=0.004). However, no significant correlation was identified between miR-19b expression and other clinical parameters (all P>0.05, Table 1) such as age, tumor size, ER, PR, and HER-2 status.
Prognostic value of miR-19b expression for patients with breast cancer
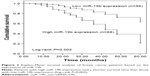
In the present study, the clinical significance of miR-19b in terms of breast cancer prognosis was investigated. The Kaplan–Meier survival curves, shown in Figure 2, demonstrate that patients with high miR-19b expression had poor overall survival compared to those with low miR-19b expression (log-rank P=0.002). In addition, the clinical parameters and miR-19b expression were included in the multivariate Cox analysis to determine their influence on overall survival of cancer patients. The analysis results indicated that miR-19b expression hazard ratio (HR =2.560, 95% CI =1.130–5.796, P=0.024) and TNM stage (HR =2.079, 95% CI =1.033–4.186, P=0.040) were two independent prognostic factors for breast cancer (Table 2).
  | Table 2 Multivariate Cox regression analysis for miR-19b in breast cancer patients |
Effects of miR-19b on cell proliferation, migration, and invasion in breast cancer cells
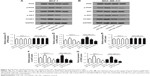
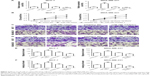
To examine the functional role of miR-19b in tumor progression of breast cancer, miR-19b mimic, inhibitor, and the corresponding NCs were transfected into MCF-7 and MDA-MB-231 cells, which had extremely high miR-19b expression, to modulate the expression of miR-19b in vitro. The expression of miR-19b in the transfected cells was verified by qRT-PCR and was demonstrated to be significantly increased in the cells transfected with miR-19b mimic but decreased in those with miR-19b inhibitor compared with the NCs (all P<0.001, Figure 3A). The cell proliferation of MCF-7 and MDA-MB-231 cells, measured by MTT, was proven to be markedly promoted in cells transfected with miR-19b mimic, and suppressed in the cells with miR-19b inhibitor (all P<0.05, Figure 3B). Transwell analysis was used to assess cell migration and invasion abilities. As shown in Figure 3C–F, overexpression of miR-19b by miR-19b mimic significantly increased cell migration and invasion (all P<0.05) but silencing of miR-19b by miR-19 inhibitor decreased the migration and invasion ability of both MCF-7 and MDA-MB-231 cells (all P<0.05).
MiR-19b regulated PI3K/AKT signaling pathway in breast cancer cells
To investigate the molecular mechanisms underlying the suppressor role of miR-19b in breast cancer, we studied the effects of miR-19b expression on PI3K/AKT signaling pathway, which is considered a crucial pathway during breast cancer development.16 According to Western blot analysis, the expression of PI3K, p-PI3K, AKT, and p-AKT was examined. Besides, protein expression of CCND1, a downstream effector of PI3K/AKT, was also analyzed. The analysis results shown in Figure 4 revealed that the protein expression of p-PI3K, p-AKT, and CCND1 in transfected MCF-7 and MDA-MB-231 cells was obviously elevated by miR-19b overexpression (all P<0.05) but was decreased by knockdown of miR-19b compared with the NCs (all P<0.05). No significant alteration was found for the expression of total PI3K and AKT (all P>0.05).
Discussion
Breast cancer currently remains a serious health burden among women worldwide. Despite great progress regarding various treatment options, improvements in prognosis and overall survival of patients suffering from breast cancer have been limited.17 Emerging research indicates that biomarkers used for diagnosis and prognosis exhibit high potential to improve outcomes of cancer patients.18–21 In recent studies, miRNAs have received more and more attention for their high diagnostic and prognostic significance in a variety of human malignancies.22 The aberrant expression of miRNAs has been detected in tumor samples, and usually has pivotal roles during tumor progression.23,24 In breast cancer, some functional miRNAs have also been identified. For instance, overexpression of miR-203 has been found in breast cancer tissues, which could significantly enhance cancer cell proliferation.25 Expression of miR-150 has been found to be decreased in triple-negative breast cancer tissues, and might suppress cell migration by downregulation of HMGA2.26 In addition, Li et al reported that miR-329 expression was downregulated in breast cancer tissues, and served as a candidate biomarker for breast cancer diagnosis and prognosis.27 All this previous research indicates that it is important to investigate the clinical value and functional role of miRNAs for the improvement of breast cancer treatment.
In the present study, we examined the expression patterns of miR-19b in breast cancer tissues and cell lines. The expression of miR-19b has never been studied in some human cancers. For example, Ohira et al28 reported that miR-19b expression was significantly increased in melanoma cells compared with normal melanocyte cells. The upregulated expression of miR-19b has also been identified in osteosarcoma cells, and exerted cancer-promoting effects in this malignancy.29 In breast cancer, the expression of miR-19b has also been investigated and was demonstrated to be elevated in tumor tissues compared with the normal controls.30 Similarly, the results of our research revealed that miR-19b expression, measured by qRT-PCR, was markedly higher in breast cancer tissues than in the matched normal tissues. The expression results in breast cancer cells were inconsistent with the tissues, which also showed increased miR-19b expression levels in cancer cell lines compared with the normal cells. Moreover, the expression of miR-19b was found to be associated with distant metastasis and TNM stage of breast cancer patients. Considering these results, miR-19b might serve as an oncogene, and play a crucial role during tumor development of breast cancer.
Based on the altered miR-19b expression in breast cancer, we further analyzed the clinical value of miR-19b in breast cancer prognosis. According to the Kaplan–Meier survival analysis, we observed that patients with high miR-19b expression had a shorter survival time than those with low miR-19b expression, suggesting that overexpression of miR-19b might be closely correlated with poor overall survival of breast cancer patients. Furthermore, multivariate Cox regression analysis results showed that the expression of miR-19b was an independent prognostic factor of patients with breast cancer. In addition to breast cancer, the prognostic significance of miR-19b has also been reported in other malignancies. A study by Wu et al31 indicated that upregulated expression of miR-19b was associated with poor survival and acted as a candidate prognostic biomarker in non-small-cell lung cancer. Besides, elevated miR-19b expression has also been demonstrated to be related to poor prognosis and disease recurrence in patients with myeloid leukemia.32
To uncover the functional role of miR-19b in the progression of breast cancer, the effects of miR-19b on breast cancer cell proliferation, migration, and invasion were explored by regulating miR-19b expression using miR-19b mimic and inhibitor. In two breast cancer cell lines, MCF-7 and MDA-MB-231, we found that the overexpression of miR-19b by miR-19b mimic could promote cell proliferation, migration, and invasion. Conversely, cell proliferation, migration, and invasion were all inhibited by reduction of miR-19b in breast cancer cells. Collectively, we considered that miR-19b could promote tumor progression in breast cancer.
The tumor-promoting activity of miR-19b has been shown in other human cancers. A study by Xu et al33 revealed that expression of miR-19b was upregulated in cervical cancer and was associated with enhanced cancer cell proliferation and invasion. The overexpression of miR-19b has also been reported to promote cell proliferation of prostate cancer cells by regulating PI3K/AKT pathway.34 However, the precise mechanisms of the role of miR-19b in breast cancer remain elusive. In the current work, we investigated the expression of proteins involved in PI3K/AKT pathway, which represents pivotal signaling in the pathogenesis of cancers.35,36 As shown by Western blot analysis, protein expression of p-PI3K, p-AKT, and CCND1 was upregulated in cells with overexpression of miR-19b, suggesting that PI3K/AKT signaling pathway might be activated by miR-19b (Figure 5). However, the detailed mechanisms underlying the relationship between miR-19b and PI3K/AKT pathway were not assessed in the present study and need to be investigated in further studies.
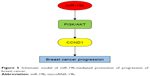  | Figure 5 Schematic model of miR-19b-mediated promotion of progression of breast cancer. |
Conclusion
Our data in this research revealed that upregulated expression of miR-19b serves as a candidate prognostic biomarker and is associated with distant metastasis and TNM stage in breast cancer. MiR-19b may promote cell proliferation, migration, and invasion via the PI3K/AKT pathway, and has the potential to be used in targeted therapy in breast cancer.
Disclosure
The authors report no conflicts of interest in this work.
References
Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108. | ||
Cedolini C, Bertozzi S, Londero AP, et al. Type of breast cancer diagnosis, screening, and survival. Clin Breast Cancer. 2014;14(4):235–240. | ||
Rudolph A, Chang-Claude J, Schmidt MK. Gene-environment interaction and risk of breast cancer. Br J Cancer. 2016;114(2):125–133. | ||
Jemal A, Center MM, Desantis C, Ward EM. Global patterns of cancer incidence and mortality rates and trends. Cancer Epidemiol Biomarkers Prev. 2010;19(8):1893–1907. | ||
Liu RZ, Garcia E, Glubrecht DD, et al. CRABP1 is associated with a poor prognosis in breast cancer: adding to the complexity of breast cancer cell response to retinoic acid. Mol Cancer. 2015;14:129. | ||
Adams BD, Kasinski AL, Slack FJ. Aberrant regulation and function of microRNAs in cancer. Curr Biol. 2014;24(16):R762–R776. | ||
Bhaskaran M, Mohan M. MicroRNAs: history, biogenesis, and their evolving role in animal development and disease. Vet Pathol. 2014;51(4):759–774. | ||
Dong Y, Chang C, Liu J, Qiang J. Targeting of GIT1 by miR-149* in breast cancer suppresses cell proliferation and metastasis in vitro and tumor growth in vivo. Onco Targets Ther. 2017;10:5873–5882. | ||
Liu B, Li J, Cairns MJ. Identifying miRNAs, targets and functions. Brief Bioinform. 2014;15(1):1–19. | ||
di Leva G, Garofalo M, Croce CM. MicroRNAs in cancer. Annu Rev Pathol. 2014;9:287–314. | ||
Song B, Lin HX, Dong LL, Ma JJ, Jiang ZG. MicroRNA-338 inhibits proliferation, migration, and invasion of gastric cancer cells by the Wnt/β-catenin signaling pathway. Eur Rev Med Pharmacol Sci. 2018;22(5):1290–1296. | ||
Quan J, Pan X, He T, et al. Tumor suppressor miR-211-5p is associated with cellular migration, proliferation and apoptosis in renal cell carcinoma. Exp Ther Med. 2018;15(4):4019–4028. | ||
Qin C, Huang RY, Wang ZX. Potential role of miR-100 in cancer diagnosis, prognosis, and therapy. Tumour Biol. 2015;36(3):1403–1409. | ||
Bertoli G, Cava C, Castiglioni I. MicroRNAs: New Biomarkers for Diagnosis, Prognosis, Therapy Prediction and Therapeutic Tools for Breast Cancer. Theranostics. 2015;5(10):1122–1143. | ||
Liu M, Yang R, Urrehman U, et al. MiR-19b suppresses PTPRG to promote breast tumorigenesis. Oncotarget. 2016;7(39):64100–64108. | ||
Nicolini A, Ferrari P, Kotlarova L, Rossi G, Biava PM. The PI3K-AKt-mTOR Pathway and New Tools to Prevent Acquired Hormone Resistance in Breast Cancer. Curr Pharm Biotechnol. 2015;16(9):804–815. | ||
Nagasawa S, Sedukhina AS, Nakagawa Y, et al. LSD1 overexpression is associated with poor prognosis in basal-like breast cancer, and sensitivity to PARP inhibition. PLoS One. 2015;10(2):e0118002. | ||
Esfahani M, Ataei N, Panjehpour M. Biomarkers for evaluation of prostate cancer prognosis. Asian Pac J Cancer Prev. 2015;16(7):2601–2611. | ||
Moch H, Srigley J, Delahunt B, et al. Biomarkers in renal cancer. Virchows Arch. 2014;464(3):359–365. | ||
Han SX, Zhou X, Sui X, et al. Serum dickkopf-1 is a novel serological biomarker for the diagnosis and prognosis of pancreatic cancer. Oncotarget. 2015;6(23):19907–19917. | ||
Li W, Wang Y, Zhang Q, et al. MicroRNA-486 as a Biomarker for Early Diagnosis and Recurrence of Non-Small Cell Lung Cancer. PLoS One. 2015;10(8):e0134220. | ||
Chen J, Wang X. MicroRNA-21 in breast cancer: diagnostic and prognostic potential. Clin Transl Oncol. 2014;16(3):225–233. | ||
Chen QG, Zhou W, Han T, et al. MiR-345 suppresses proliferation, migration and invasion by targeting Smad1 in human prostate cancer. J Cancer Res Clin Oncol. 2016;142(1):213–224. | ||
Xia M, Duan ML, Tong JH, Xu JG, Jg X. MiR-26b suppresses tumor cell proliferation, migration and invasion by directly targeting COX-2 in lung cancer. Eur Rev Med Pharmacol Sci. 2015;19(24):4728–4737. | ||
Muhammad N, Bhattacharya S, Steele R, Ray RB. Anti-miR-203 suppresses ER-positive breast cancer growth and stemness by targeting SOCS3. Oncotarget. 2016;7(36):58595–58605. | ||
Tang W, Xu P, Wang H, et al. MicroRNA-150 suppresses triple-negative breast cancer metastasis through targeting HMGA2. Onco Targets Ther. 2018;11:2319–2332. | ||
Li P, Dong J, Zhou X, et al. Expression patterns of microRNA-329 and its clinical performance in diagnosis and prognosis of breast cancer. Onco Targets Ther. 2017;10:5711–5718. | ||
Ohira T, Naohiro S, Nakayama Y, et al. miR-19b regulates hTERT mRNA expression through targeting PITX1 mRNA in melanoma cells. Sci Rep. 2015;5:8201. | ||
Li X, Wang FS, Wu ZY, Zy W, et al. MicroRNA-19b targets Mfn1 to inhibit Mfn1-induced apoptosis in osteosarcoma cells. Neoplasma. 2014;61(3):265–273. | ||
Zhao L, Zhao Y, He Y, Mao Y. miR-19b promotes breast cancer metastasis through targeting MYLIP and its related cell adhesion molecules. Oncotarget. 2017;8(38):64330–64343. | ||
Wu C, Cao Y, He Z, et al. Serum levels of miR-19b and miR-146a as prognostic biomarkers for non-small cell lung cancer. Tohoku J Exp Med. 2014;232(2):85–95. | ||
Zhang TJ, Lin J, Zhou JD, et al. High bone marrow miR-19b level predicts poor prognosis and disease recurrence in de novo acute myeloid leukemia. Gene. 2018;640:79–85. | ||
Xu XM, Wang XB, Chen MM, et al. MicroRNA-19a and -19b regulate cervical carcinoma cell proliferation and invasion by targeting CUL5. Cancer Lett. 2012;322(2):148–158. | ||
Tian L, Fang YX, Xue JL, Chen JZ. Four microRNAs promote prostate cell proliferation with regulation of PTEN and its downstream signals in vitro. PLoS One. 2013;8(9):e75885. | ||
Polivka J, Janku F. Molecular targets for cancer therapy in the PI3K/AKT/mTOR pathway. Pharmacol Ther. 2014;142(2):164–175. | ||
Danielsen SA, Eide PW, Nesbakken A, et al. Portrait of the PI3K/AKT pathway in colorectal cancer. Biochim Biophys Acta. 2015;1855(1):104–121. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.