Back to Journals » Cancer Management and Research » Volume 12
MicroRNA-155 Suppresses the Translation of p38 and Impairs the Functioning of Dendritic Cells in Endometrial Cancer Mice
Authors Jia J, Li X, Guo S, Xie X
Received 3 December 2019
Accepted for publication 31 March 2020
Published 30 April 2020 Volume 2020:12 Pages 2993—3002
DOI https://doi.org/10.2147/CMAR.S240926
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Antonella D'Anneo
Jianjun Jia,1 Xiaomao Li,2 Suiqun Guo,3 Xingmei Xie1
1Department of Obstetrics and Gynecology, The First Affiliated Hospital, Jinan University, Guangzhou 510632, People’s Republic of China; 2Department of Obstetrics and Gynecology, The Third Affiliated Hospital, Sun Yat-Sen University, Guangzhou 510632, People’s Republic of China; 3Department of Obstetrics and Gynecology, The Third Affiliated Hospital, Southern Medical University, Guangzhou 510632, People’s Republic of China
Correspondence: Jianjun Jia
Department of Obstetrics and Gynecology, The First Affiliated Hospital, Jinan University, Guangzhou 510632, People’s Republic of China
Tel/ Fax +86-020-38688359
Email [email protected]
Background: Dendritic cells (DCs) are reported to play an important role in activating the anti-tumor immune responses. p38 MAPK14 signaling plays an important role in controlling their activity. Here, we identified that miR-155 suppressed the translation of p38 and impaired the functioning of dendritic cells in endometrial cancer.
Methods: HEC1A endometrial cancer cell lines were used for the study which was transfected in the C57BL/6 mice. Murine bone marrow-derived dendritic cells (BMDCs) were isolated from the mice. Target prediction was done by TargetScan which was confirmed by RT-PCR analysis. The protein expression was carried by Western blot analysis. Levels of IL-12 were evaluated by ELISA. Mice injected with HEC1A cells were subjected to tumor challenge study.
Results: On screening the binding sites of p38 MAPK14 gene, miR-155 was found to bind the 3ʹUTR directly and blocked its translation. The levels of miR-155 were upregulated in dendritic cells and RAW264.7 cells, miR-155 showed inhibitory effect on expression levels of p38. In dendritic cells, miR-155 was found to regulate the expression of IL-12, also miR-155 inhibitor stimulated the differentiation of Th1 cells in mice induced with endometrial cancer. In dendritic cells, miR-155 inhibited the expression of p38 gene and decreased their ability to interfere in tumor growth.
Conclusion: The study concludes suppressive role of miR-155 in the process of dendritic cells mediated anti-tumor immunity, also inhibiting miR-155 provides a novel strategy for countering endometrial cancer.
Keywords: miR-155, dendritic cells, p38, IL-12, RAW264.7 cells
Introduction
Dendritic cells (DCs) or accessory cells are antigen presenting cells which have an exclusive property of activating the T cells through the antigens present on their surface.1 As DCs have their property due to presence of antigens on their surface they show increased levels of MHC class II molecules to exhibit them efficiently and then lead to activation of CD4+ and CD8+ T cells. In addition to this, DCs interact with natural killer cells (NK) and B cells to create a link between adaptive and innate immune systems,2,3 hence they are regarded as Prime activators of immune response and are actively associated with autoimmunity, inflammation and immune response in organ transplantation. Looking into their versatile potential, scientific community has focused on their ability to produce reactions in T and B cells. Recently, the anti-tumor functions of dendritic cells have gathered attention of scientific community.4 Dendritic cells have been evidenced to play potential role in immune response against tumors, whereas it has been found that tumors secrete soluble factors such as IL-10 and TGF-β for disrupting the differentiation of DCs and also suppress their ability to activate immune response to fight back which is very important barrier for treating the tumors.5,6 These tumor derived factors cause interruption in the regular functioning of DCs via activation of several pathways such as NF-κB, JAK/STAT and MAPK14. Importantly, MAPK14 pathway is confirmed to play important role in dysfunctioning of DCs,7 hence, interfering MAPK14 pathway can reduce the extent of damage on DCs mediated by cancer cells. p38 is crucial member of the MAPK14 family proteins, it regulates the various cell activities like transduction of signals, hence targeting the expression of p38 could be efficacious in DC mediated approaches for treating cancer.
Endometrial cancer (EC) is ranked as the 4th most common malignancy in females worldwide.7,8 EC is broadly divided into two subclasses, Type 1: endometrioid endometrial cancer and Type 2: non-endometrioid endometrial cancer.9 In endometrioid endometrial cancer, abnormal expression of phosphoinositide 3-kinase (PI3K) is the most common pathway. In addition to PI3K endometrioid endometrial cancer is accompanied with decreased levels of PTEN and alterations in PIK3R1, PIK3R2 and PIK3CA genes10.
Small coding RNAs also called as microRNAs (miRs) are distributed widely in various species and are involved in regulating the expression of genes associated with various pathological and physiological processes such as immunity.11 Earlier various miRs (miR-155, miR-146a and miR-142-3p) have been found to regulate the functioning of DCs via regulating the production of cytokines.12 As discussed earlier, DCs are involved in building immune responses via releasing suitable cytokines and inducing the differentiation of CD4+T cells.12 miRs may hence play important role for modifying DCs in improving the immune response against cancer.
MiR-155 also described as host gene was earlier reported to be involved in lymphoma.13 Beside its involvement in lymphoma, miR-155 is also involved in breast cancer, cardiovascular diseases, viral infections and some other solid tumors.14 It has been also reported that miR-155 has about 400 different gene targets.15 Here in the present work we found that miR-155 was expressed in DCs and could inhibit the tumor suppressing function of in them and could also bind to the 3ʹUTR region of MAPK14 mRNA leading to down-regulation in protein levels of MAPK14. We also evidenced that decreased levels of MAPK14 interfered with the dendritic cells derived synthesis of IL-12 and the differentiation of Dendritic cells driven Th17 cells.
Materials and Methods
Cell Lines, Experimental Mice and Isolation of Bone Marrow-Derived Dendritic Cells
We used HEC1A endometrial cancer cell lines, EM-E6 as normal endometrial cells, HEK 293T cells and RAW264.7 which are suitable for transfection host. The cells were supplied by the microbiology department of The First Affiliated Hospital, Jinan. The use of the cell lines received approval by the institutional research ethics committee of The First Affiliated Hospital, Jinan. All the cells were incubated and multiplied in DMEM media supplemented with FBS 10% at room temperature with 5% CO2.
C57BL/6 mice aging 6–7 weeks (Jackson Laboratories, Bar Harbor ME) were used for the study (n=8 in each group). The mice were kept under identical pathogen free conditions 1 week prior to their use. The animals were subjected to 12 hour dark and light cycle and were provided free access to food and water. The animal studies were approved from the Ethical clearance board of The First Affiliated Hospital, Jinan. The experiments adhered to guidelines laid by Chinese Academy of Medical Sciences’s Institute of Laboratory Animal Sciences (ILAS) in Beijing, China.
Murine bone marrow-derived dendritic cells (BMDCs) were isolated from the mice, which were sacrificed, the femur and tibia bones were flushed with phosphate buffer saline using a injection and the bone marrow cells were collected. The isolated cells were centrifuged at 600 rpm for 10 minutes and any erythrocytes were removed by incubating with lysis buffer of ammonium chloride for 5 minutes at 37°C. The bone marrow cells were rinsed with RPMI media added with fetal bovine serum (10%). These bone marrow cells were incubated for 5 days with Granulocyte-Macrophage Colony Stimulating Factor 500 U/mL and the BMDCs were obtained. Fluorescence activated cell sorting (FACS) was done and it was found that after 5 days the CD11c+ Dendritic cell population was about 75%.
Surface Staining and Intracellular Staining
The single cells were isolated from the mouse inguinal nodes, about 106 cells/well were maintained in 24 well plates. The cells were again stimulated for 3 hours using phorbol myristate acetate (50 ng/mL) (PMA), Ionomycin (1 μg/mL) and protein transport inhibitor (0.66 μL/mL). The intracellular and surface staining was done with an Immunophenotyping Kit (ThermoFisher USA) in accordance to suppliers instructions. The cells stained were evaluated using CellQuest software on BD-FACScaliber (BD bioscience USA). The staining experiments were done in replicates of three.
Bioinformatics Analysis for Prediction of Potential Binding Sites
For identifying the potential binding sites of miRNA, bioinformatics analysis was done by scanning the TargetScan data base using TargetScan 5.1.
Real-Time PCR (RT-PCR)
Total RNA was separated with the help of TRIzol reagent (ThermoFisher USA), about 1µg of RNA was subjected to reverse transcription using Reverse Transcription System (Promega, USA) following the supplied instructions. PCR analysis was done with the help of SYBR green RT-PCR master mix following the provided instructions and as per literatures published;16 the primers used are described in Table 1. The process of amplification was done by denaturation at 95°C for 10 minutes followed by 40 rounds of 95°C for 30 seconds, 72°C for 30 seconds and 58°C for 30 sec. GAPDH was selected as loading control for every analysis. RT-PCR was carried using iQ5 Real-Time PCR Systems (Bio-Rad), all the experiments were done in replicates of three.
 |
Table 1 Primers Used for RT-PCR Analysis |
Process of Transfection
The Oligonucleotide, RNA inhibitor and RNA mimics were obtained from RiboBio,China. The target cells received transfection with the respective Oligonucleotides (100 nm) using cell transfection kit (Qiagen USA) following the supplied instructions.
Western Blot Analysis
The cells were subjected to lysis using a lysis buffer, the resultant lysate was centrifuged at 12,000 rpm and then was boiled in sodium dodecyl sulfate. The proteins were resolved on poly acrylamide gel (10%) by electrophoresis in presence of SDS-PAGE and were then transferred on PVDF membranes, which were exposed to rabbit anti-beta-actin primary antibody (1:1000) followed by horseradish peroxidase conjugated anti-rabbit secondary antibodies (1:1000). The polypeptides were evaluated using an ECL reagent. The experiments were repeated three times.
Enzyme Linked Immunosorbent Assay (ELISA)
The cells were stimulated by LPS at low doses (1 μg/mL) for 24 hours, for levels of IL-12 in the cell supernatants; they were subjected to analysis using an ELISA kit (ThermoFisher USA) following the supplied instructions.
Luciferase Assay
The recombinant luciferase mRNAs was obtained by amplification of the 3ʹUTR region of mouse p38 MAPK14 and its mutant was done, the binding site of miR-155 was amplified by PCR with the help of modified primers (Forward 5′-CTCAGACTCGGTTAATGCTAATCGTGATAGG-3′, Reverse 5′-GCTGTGGCAGTGGAAGCGTGATTTATT-3′) for mutant and wild type 3ʹUTRs. The primers were cloned into the vector siCHECK-2 and CDS. The constructs were verified by sequencing. The normal endometrial cells EM-E6 were co-transfected using the prepared constructs which were combined with dsRNA control or dsRNA mimics of miR-155 with the help of Lipofectamine reagent. After 24 hours of transfection the cells were subjected to luciferase assay (Promega USA) following the supplied instructions.
Tumor Challenge Study
HEC1A cells were cultured and maintained as described earlier and were injected (1×106) into the C57BL/6 mice subcutaneously for developing tumors. The mice were housed under identical pathogen free conditions and were monitored for development of tumors regularly. The volumes of tumors were measured after 2 weeks of injecting cancer cells. The mice were injected with intra-tumoral injection of DC cells (1×106) to each mouse every day for next 7 days. Simultaneously, analgesics were administered (Buprenorphine 0.05 mg/kg) every 12 hours. The mice were euthanized and tumors were removed when the mouse showed signs of moribund state (signs such as: Hunched back, severe weight loss, severe mobility loss and piloerection). The time between grafting of cells till the moribund state was regarded as survival time. The size of tumors was measured by formula volume of tumor = ½ (L × W2), where W= width, L= Length.
Results
Target miRNAs of p38 MAPK14 Gene
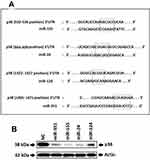
It was evidenced earlier that the DCs of cancer patients are non-functional, whereas the p38 MAP kinase cascade plays a potential role in regulating the functioning of DCs in tumor environments.17 In the process to find the miRs involved in regulation of p38 MAPK14 gene and therefore having the capacity to control the activity of DCs, we screened the possible miRs binding sites in the 3ʹUTR region of p38 mRNA bioinformatically using TargetScan (www.targetscan.org).15 The findings demonstrated that 4 miRs had binding sites for p38 gene which included miR-155, miR-24, miR-124 and miR-351 (Figure 1A). Ahead we studied whether these miRs could silence the target p38, for this we transfected these miRs mimics in the RAW264.7 cells which is established to be the cell line suitable for establishing intracellular signaling pathway.18–20 Followed by transfection, we measured the expression of p38 protein levels by Western blot analysis. The outcomes indicated significant downfall in the protein levels post transfection of miR-155, miR-351 and miR-24 (Figure 1B).
miR-155 Binds to the 3ʹUTR of p38 Gene and Blocks Its Translation
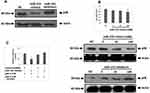
From the outcomes which suggested and validated about 3 miRs having suppressive effect on the expression levels of p38 gene, we deducted that all the three could act as regulators for DCs to alter the antitumor effects. In a study earlier miR-155 was found to express majorly in mouse DCs.21 In addition to this miR-155 has been identified to act as oncogene in breast cancer.22 Hence the present study was planned to evaluate the effect of miR-155 on DCs. Prior to this a study has suggested that miR-155 induces differentiation of RAW 264.7 cells to DC cells,23 hence we further studied the effect of miR-155 on the p38 gene utilizing RAW264.7 cells. At first the RAW264.7 cells were transfected with miR-155 mimics or inhibitor, followed by evaluation of protein levels of p38 gene by Western blot analysis. The outcomes clearly indicated that miR-155 mimic decreased the expression of p38 compared to control group, whereas the inhibitor increased the levels of p38 (Figure 2A). The level of expression of p38 were similar under different concentrations of miR-155 mimics when evaluated by RT-PCR, but when studied by Western blot technique, the levels were found to decrease with increasing concentration. But in contrast, when the dose of inhibitor increased the levels of p38 were elevated (Figure 2B). These findings demonstrated that miR-155 was involved in regulation of p38 gene at post transcriptional level via inhibiting the translation of mRNA rather than encouraging the degradation. As it was evidenced that miR-155 acted at post transcriptional level causing inhibition of p38 mRNA translation, in sight that some miRs can exert their effect via binding to the 3ʹUTR region of target mRNAs, we fabricated a recombinant mRNA for testing p38 mRNA as direct target of miR-155 by displacing the 3ʹUTR of luciferase reporter gene with that of p38 gene, mutant or wild type in the plasmid siCHECK2. The constructed plasmids were transfected in the HEK 293T cells followed my measurement of fluorescence intensity post 24 hours. The results showed that the recombinant luciferase gene having wild type p38 3ʹUTR was inhibited significantly by miR-155 mimics, whereas, the mutant showed normal expression (Figure 2C). These outcomes suggested that miR-155 halted the translation of p38 gene by directly binding to the 3ʹUTR region.
miR-155 Shows Expression and Functions in DCs
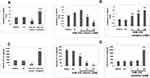
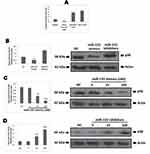
In order to clarify whether miR-155 caused differentiation of DCs because expression levels of miR-155 where found to be upregulated in DCs and RAW264.7 cells (Figure 3A). To illustrate the function of miR-155 in DCs, the dendritic cells were transfected with either miR-155 mimics/inhibitors and the protein levels of p38 were evaluated. It was observed that miR-155 showed an inhibitory effect on expression of p38 which is its target gene in presence of miR-155 mimics and the levels were increased in presence of miR-155 inhibitor (Figure 3B). This interconnection between protein levels of p38 and dose of miR-155 mimics/inhibitor further confirmed the effect by dose response studies (Figure 3C and D).
miR-155 Regulates the Expression of IL-12 in DCs
MAPK14 p38 is an important member in the MAPK family, it is reported that IL-12 activated p38 is necessary for IL-12 mediated expression and Th1 development in STAT-4 independent pathway.24 In context to DCs, inhibition of p38 MAPK14 has shown to cause increased production of IL-12 by stimulating macrophages and DCs.25 Looking into the influence of p38 on IL-12 as well as the pre-discussed relation between miR-155 and p38, made us to explore whether miR-155 could be involved in regulation of IL-12 production. The mice derived BMDCs were subjected to transfection with miR-155 inhibitor/mimics followed by treatment of LPS, the BMDCs were then collected for studying the mRNA levels of IL-12 using RT-PCR. The outcomes suggested that, the levels of IL-12 decreased significantly compared to control group under the influence of miR-155 mimics, whereas the levels increased when treated with inhibitor (Figure 4A). In addition to this, the effects of various doses of miR-155 inhibitor as well as mimics were also studied, which was confirmed by the changes in mRNA tendency (Figure 4B). From the findings it was confirmed that miR-155 exerted negative effect on protein levels of p38 and the inducing effect of p38 on levels of IL-12 which the miR-155 inhibitor and mimics would increase or decrease the mRNA concentration of IL-12 mRNA respectively. This was in agreement with our outcomes and also with previous reports which suggest that miR-155 exerts negative effect on IL-12 at transcriptional and post transcriptional level.26 To further confirm, enzyme linked immunosorbent assay (ELISA) was done for IL-12. We found very low levels of IL-12 protein in the supernatant of Dendritic cells in mimics treated group compared to control, whereas the inhibitor treated group showed significantly increased levels of IL-12 (Figure 4C). In addition to this, the dose response studies showed parallel results to RT-PCR except the results of inhibitor dose in the ELISA plot (Figure 4D). This phenomenon may be due to various processes which are responsible for production of mature RNA and functional proteins.
miR-155 Inhibitor Can Stimulate Th1 Cell Generation Through IL-12
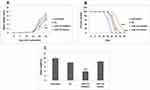
The correlation between miR-155 and IL-12 prompted us that miR-155 could participate in many processes involving IL-12 in them. In the immune system, IL-12 is involved in the process of differentiation of T cells into Th1cells.27 Also, IL-12 is reported to alter the development of Th1 involving TGF-β.28 Looking into this function, we speculated that miR-155 could also exert inhibitory effect on differentiation of Th1 cell which was proved experimentally. For in vivo studies we established HEC1A tumor model in C57BL/6 mice and then treated them with DC immunotherapy by injecting the bone marrow-derived dendritic cells pre-transfected with miR-155 mimics/inhibitor or negative control directly into the tumors of mice. After the experimental period was over (The mice showed signs of moribund state) the tumors were removed and subjected for detecting Th17, Th1 and Th2 cells by fluorescence activated cell sorting assay (FACS). The results suggested that Th1 cell population in the mice treated with inhibitor was significantly higher than the other 2 groups, suggesting potential ability of miR-155 inhibitor transfected Dendritic cells to induce the differentiation of Th1 cells in animal model of endometrial cancer.
miR-155 Reduces the Tumor-Suppressive Ability of Dendritic Cells
The main aim of the study was to identify miRs having ability to inhibit p38 gene which would help in developing a reasonable tumor therapy. Therefore the main focus of the study was to evaluate the effect of miR-155 transfected dendritic cells on HEC1A tumor cells induced C57BL/6 mice. The outcomes of our study suggested that miR-155 showed a tumor promoting effect and decreased the effectiveness of immunotherapy. It was observed that the miR-155 mimics treatment caused tumor with greatest size and highest weight (Figure 5A and C), developed at quickest rate (Figure 5A) and had least survival rate (Figure 5B) in all the dendritic cells treated mice. Opposite to this, the miR-155 inhibitor showed the most significant signs of its curative effect (Figure 5). It was also observed that the effect of negative control DCs increased when miR-155 inhibitor was included. These outcomes actuated us to conclude that in Dendritic cells, miR-155 could inhibit expression of p38 gene and decrease their ability to interfere in tumor growth, whereas miR-155 inhibitor could reverse this effect and could serve as a potential immunotherapeutic agent.
Discussion
Dendritic cells are also called as antigen presenting cells (APCs) which also link adaptive and innate immunity by stimulating the activation and differentiation of T cells. DCs show a range of receptors called as pattern recognition receptors which act by sensing the microbial pathogenic products and trigger the signaling pathway that lead to activation of DCs and the adaptive immune response.29 It has been reported that MAPK14, p38, JNK and ERK play an important role in regulating this process.30 Human efforts in these events may play an important part in enhancing the therapeutic effect of DCs in cancer patients. In the present work, the experiments were focused on p38 MAPK14 as it was very crucial in facilitating the functioning of DCs particularly for causing the release of IL-12 and differentiation of Th1 cells.
To study these targets of DCs we used a miR approach for which we screened the levels of miRs associated with p38. Looking into the outcomes of bioinformatics analysis by TargetScan and experimental outcomes we identified miR-155 which negatively regulated the protein levels of p38 by directly binding to the 3ʹUTR region of target mRNA. Our findings suggested that miR-155 exerted a negative effect on functioning of DCs via silencing p38 which was in agreement to earlier findings which confirmed that miR-155 silenced the LPS mediated activation of DCs via the p38 MAPK14 pathway.31
Over-expression of miR-155 by mimics showed inhibitory effect of expression on mRNA as well as protein levels of IL-12 in DCs whereas the inhibitor had reverse effect. This influence on the levels of IL-12 altered the Th1 cell population in the tumor micro-environment. This is an important phenomenon to Dendritic cells because initiation of Th1 response is crucial to them. This potential effect of miR-155 mimics and inhibitor on dendritic cells shows the importance of this miR in dendritic cells.
Consistent to our findings were the reports published earlier which suggested that inhibition of p38 showed suppression on expression of IL-12 in mice compared to wild type.24 Inhibitory effect of miR-155 to p38 in dendritic cells made miR-155 mimics a negative regulator and inhibitor as the positive regulator of DC-mediated differentiation of Th1 and IL-12 expression.
This influence of Th1 and IL-12 was a partial contribution of dendritic cells influenced by miR-155, their anti cancer effect which is the prime focus of our work. It is already established that IL-12 is a promising candidate for anti cancer therapy,32 this was in consistent to our present findings which confirmed higher tumor eradicating efficiency of miR-155 inhibited DCs having upregulated IL-12 and Th1.
It was observed that, miR-155 mimics transfected DCs showed lower therapeutic efficiency compared to normal group. It was also observed that the prime tumor suppressing ability of DCs was inhibited under the influence of miR-155. This suggested that blockade of miR-155 could serve a useful strategy for enhancing the anti tumor activity of dendritic cells whereas the miR-155 inhibitor could be a potential agent which would improve the potential of existing dendritic cells based therapies. In future more studies targeting different target genes of miR-155 can be done for treating cancers like endometrial cancer.
In conclusion, the present study established miR-155 as an endogenous miRNA of dendritic cells which could inhibit the translation of p38 mRNA and also down regulate the expression levels of IL-12 which in turn intervenes with development of Th1 cell in the tumor microenvironment. These findings confirm that miR-155 could be a novel target to be interfered, also blocking of miR-155 could be a potential strategy in improving the performance of dendritic cells based immunotherapy. Indeed, more preclinical studies in different animal models as well as clinical studies are still needed.
Acknowledgment
This work was supported by the Research Fund of Traditional Medicine Bureau of Guangdong Province (No.20201081).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Steinman RM. Decisions about dendritic cells: past, present, and future. Annu Rev Immunol. 2012;30(1):1–22. doi:10.1146/annurev-immunol-100311-102839
2. Dubois B, Bridon J-M, Fayette J, Barthélémy C, Banchereau J, Caux C. Dendritic cells directly modulate B cell growth and differentiation. J Leukoc Biol. 1999;66(2):224–230. doi:10.1002/jlb.66.2.224
3. Gerosa F, Baldani-Guerra B, Nisii C, Marchesini V, Carra G, Trinchieri G. Reciprocal activating interaction between natural killer cells and dendritic cells. J Exp Med. 2002;195(3):327–333. doi:10.1084/jem.20010938
4. Palucka K, Banchereau J. Cancer immunotherapy via dendritic cells. Nature Review Cancer. 2012;12(4):265–277. doi:10.1038/nrc3258
5. Kim R, Emi M, Tanabe K. Cancer immunosuppression and autoimmune disease: beyond immunosuppressive networks for tumour immunity. Immunol. 2006;119(2):254–264. doi:10.1111/j.1365-2567.2006.02430.x
6. Rabinovich GA, Gabrilovich D, Sotomayor EM. Immunosuppressive strategies that are mediated by tumor cells. Annu Rev Immunol. 2007;25(1):267–296. doi:10.1146/annurev.immunol.25.022106.141609
7. Siegel RL, Miller KD, Jemal A. Cancer statistics. CA Cancer J Clin. 2017;67(1):7–30. doi:10.3322/caac.21387
8. Kandoth C, Schultz N. The cancer genome atlas research network. integrated genomic characterization of endometrial carcinoma. Nature. 2013;497(7447):67–73. doi:10.1038/nature12113
9. Lax SF, Kurman RJ. A dualistic model for endometrial carcinogenesis based on immunohistochemical and molecular genetic analyses. Verh Dtsch Ges Pathol. 1997;81:228–232.
10. Cheung LWT, Hennessy BT, Li J. High frequency of PIK3R1 and PIK3R2 mutations in endometrial cancer elucidates a novel mechanism for regulation of pten protein stability. Cancer Discov. 2011;1(2):170–185. doi:10.1158/2159-8290.CD-11-0039
11. He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nature Review Gene. 2004;5(7):522–531. doi:10.1038/nrg1379
12. Turner ML, Schnorfeil FM, Brocker T. MicroRNAs regulate dendritic cell differentiation and function. J Immunol. 1950;2011(187):3911–3917.
13. Zhu FQ, Zeng L, Tang N, et al. MicroRNA-155 downregulation promotes cell cycle arrest and apoptosis in diffuse large b-cell lymphoma. Oncol Res. 2016;27;24(6):415–427. doi:10.3727/096504016X14685034103473
14. Faraoni I, Antonetti FR, Cardone J, Bonmassar E. miR-155 gene: a typical multifunctional microRNA. Biochim Biophys Acta. 2009;1792(6):497–505. doi:10.1016/j.bbadis.2009.02.013
15. Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120(1):15–20. doi:10.1016/j.cell.2004.12.035
16. Bustin SA, Benes V, Garson JA, et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009;55(4):611–622. doi:10.1373/clinchem.2008.112797
17. Wang S, Yang J, Qian J, Wezeman M, Kwak LW, Yi Q. Tumor evasion of the immune system: inhibiting p38 MAPK signaling restores the function of dendritic cells in multiple myeloma. Blood. 2006;107(6):2432–2439. doi:10.1182/blood-2005-06-2486
18. Xu J, Yue F, Wang J, Chen L, Qi W. High glucose inhibits receptor activator of nuclear factor-κB ligand-induced osteoclast differentiation via downregulation of v-ATPase V0 subunit d2 and Dendritic cell-specific transmembrane protein. Mol Med Rep. 2015;11(2):865–870. doi:10.3892/mmr.2014.2807
19. Lin CC, Liu CP, Hsieh FC, Lee CM, Wang WS. Antimicrobial susceptibility and clinical outcomes of Candida parapsilosis bloodstream infections in a tertiary teaching hospital in Northern Taiwan. J Microbiol Immunol Infection. 2015;48(5):552–558. doi:10.1016/j.jmii.2014.07.007
20. Yang R, Murillo FM, Cui H, Blosser R, Uematsu S, Takeda K. Papillomavirus-like particles stimulate murine bone marrow-derived dendritic cells to produce alpha interferon and Th1 immune responses via MyD88. J Virol. 2004;78(20):11152–11160. doi:10.1128/JVI.78.20.11152-11160.2004
21. Lippai D, Szabo G. Chapter 11-Prevention of Alcohol-Induced Inflammation of Murine Small Intestine by Microrna-155 Deficiency. In: Berczi I, editor. Insights to Neuroimmune Biology.
22. Mattiske S, Suetani RJ, Neilsen PM, Callen DF. The oncogenic role of miR-155 in breast cancer. Cancer Epidemiol Biomarkers Prev. 2012;21(8):1236–1243. doi:10.1158/1055-9965.EPI-12-0173
23. Ma YL, Ma ZJ, Wang M, Liao MY, Yao R, Liao YHM. MicroRNA-155 induces differentiation of RAW264.7 cells into dendritic-like cells. Int J Clin Exp Pathol. 2015;8(11):14050–14062.
24. Zhang S, Kaplan MH. The p38 mitogen-activated protein kinase is required for IL-12-induced IFN-gamma expression. J Immunol. 2000;165(3):1374–1380. doi:10.4049/jimmunol.165.3.1374
25. Yang Z, Zhang X, Darrah PA, Mosser DM. The regulation of Th1 responses by the p38 MAPK. J Immunol. 2010;185(10):6205–6213. doi:10.4049/jimmunol.1000243
26. Cui B, Liu W, Wang X, et al. Brucella omp25 upregulates miR-155, miR-21-5p, and miR-23b to inhibit interleukin-12 production via modulation of programmed death1 signaling in human monocyte/macrophages. Front Immunol. 2017;26(8):708. doi:10.3389/fimmu.2017.00708
27. Hsieh CS, Macatonia SE, Tripp CS, Wolf SF, O’Garra A, Murphy KM. Development of TH1 CD4+ T cells through IL-12 produced by Listeria-induced macrophages. Science. 1993;260(5107):547–549. doi:10.1126/science.8097338
28. Prochazkova J, Pokorna K, Holan V. IL-12 inhibits the TGF-β-dependent T cell developmental programs and skews the TGF-β-induced differentiation into a Th1-like direction. Immunobiol. 2012;217(1):74–82. doi:10.1016/j.imbio.2011.07.032
29. Iwasaki A, Medzhitov R. Regulation of adaptive immunity by the innate immune system. Science. 2010;327(5963):291–295. doi:10.1126/science.1183021
30. Huang G, Shi LZ, Chi H. Regulation of JNK and p38 MAPK in the immune system: signal integration, propagation and termination. Cytokine. 2009;48(3):161–169. doi:10.1016/j.cyto.2009.08.002
31. Ceppi M, Pereira PM, Dunand-Sauthier I, et al. MicroRNA155 modulates the interleukin1 signaling pathway in activated human monocyte-derived dendritic cells. Proc Natl Acad Sci U S A. 2009;106(8):2735–2740. doi:10.1073/pnas.0811073106
32. Colombo MP, Trinchieri G. Interleukin-12 in anti-tumor immunity and immunotherapy. Cytokine Growth Factor Rev. 2002;13(2):155–168. doi:10.1016/S1359-6101(01)00032-6
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.