Back to Journals » Infection and Drug Resistance » Volume 12
Mechanisms of azole resistance in clinical isolates of Candida glabrata from two hospitals in China
Authors Yao D, Chen J, Chen W , Li Z, Hu X
Received 18 January 2019
Accepted for publication 13 March 2019
Published 5 April 2019 Volume 2019:12 Pages 771—781
DOI https://doi.org/10.2147/IDR.S202058
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Joachim Wink
Dongting Yao*, Jia Chen*, Weiqin Chen, Zhen Li, Xiaobo Hu
Department of Laboratory Medicine, Longhua Hospital, Shanghai University of Traditional Chinese Medicine, Shanghai, People’s Republic of China
*These authors contributed equally to this work
Purpose: Candida glabrata has emerged as the second or third most common non-albicans species responsible for an increasing number of systemic infections. Moreover, its high-level of resistance to azole is associated with a high mortality rate. This study aimed to evaluate nosocomial infections and resistance characteristics of C. glabrata and to explore the mechanism of azole resistance in C. glabrata.
Patients and methods: Fifty-nine clinical C. glabrata isolates were collected from two hospitals in China. The susceptibility of the strains to antifungal agents was determined by both the ATB Fungus 3 strip and CLSI M27 broth microdilution method. Efflux of rhodamine 6G was examined to evaluate the effects of efflux pumps. The expression levels of CgCDR1, CgCDR2, CgSNQ2, CgERG11, and CgPDR1 were examined by real-time PCR. The sequences of CgERG11 and CgPDR1 were determined by PCR-based DNA sequencing.
Results: All 59 isolates of C. glabrata were susceptible to flucytosine and amphotericin B. Twelve (20.3%) isolates were determined to be fluconazole-resistant, whereas 13 (22.0%) and 27 (45.7%) isolates were categorized as non-wild-type for itraconazole and voriconazole, respectively. Efflux pumps in azole-resistant isolates showed stronger effects than those in azole-susceptible-dose dependent isolates, which is consistent with the significant upregulation of CgCDR1 and CgCDR2 (P<0.05), whereas no obvious differences were found for CgSNQ2, CgERG11, and CgPDR1 (P>0.05). Sequencing of CgERG11 showed no alteration favoring the hypothesis that CgERG11 is not involved in the azole resistance of C. glabrata. Four CgPDR1 missense mutations were found in azole-resistant isolates, of which the high frequency of the CgPDR1 mutation, A848V, has not been reported previously.
Conclusion: Efflux pump function is the main mechanism of resistance to fluconazole in our collected clinical isolates of C. glabrata, and further studies of the related gene disruption and genome-wide expression are needed to verify the function.
Keywords: Candida glabrata, antifungal susceptibility, azole, resistance mechanisms, efflux pump
Introduction
Candida species have become important opportunistic pathogens responsible for a growing number of candiduria and systemic infections.1 The morbidity and mortality caused by non-albicans Candida species are rising, although Candida albicans remains the commonest Candida species.2 Recent studies showed that C. glabrata has emerged as the second most common non-albicans species and causes fungemia in many countries,3,4 representing the highest proportion in Northern and Central Europe,5 and accounting for 21% of all Candida bloodstream isolates in the United States,6 18.1–40.7% in North America and 8.5–31.0% in northern Europe.7 A study of 814 yeast isolates in the China Hospital Invasive Fungal Surveillance Net (CHIF-NET) showed that Candida spp. isolates were broadly distributed and constituted 92.3% of blood culture isolates, while the C. glabrata species complex was the third most common non-albicans species in China.8 Fungal infections can be treated with several antifungal agents including fluconazole, itraconazole, and voriconazole, which are widely used for Candida treatment. However, C. glabrata exhibits intrinsically low susceptibility and can develop resistance frequently during prolonged exposure to the class of azole antifungals.9,10 Azole resistance among Candida spp. arises from the overexpression of efflux pumps, while up-regulation of transporters, overexpression of drug targets and mutations are increasingly becoming common, and alternative mechanisms, such as mitochondrial defects and biofilm formation, have also been recently documented. In C. glabrata, studies of azole-resistant isolates have demonstrated frequent constitutive overexpression of multidrug transporters of the ATP-binding cassette transporter. Several genes encoding these transporters contribute to this phenotype and have been identified, including C. glabrata CDR1 (CgCDR1), CgCDR2, and CgSNQ2.11,12 These transporters are regulated by the zinc finger transcription factor CgPdr1, which functionally resembles the Saccharomyces cerevisiae Pdr1 (ScPdrl) and ScPdr3 transcription factors.13 Vermitsky et al14 previously reported that expression of CgPDR1 was elevated in one of seven C. glabrata fluconazole resistant mutants. Compared to the wild type Pdr1, the strain showing a higher CgPDR1 expression had a single mutation, P927L, which caused azole resistance in the mutant strain. Additionally, azole resistance may be caused by increased expression of the gene coding for the azole target (CgERG11).15,16 In contrast, point mutations in CgERG11 that are involved in the resistance of C. glabrata to azoles have not been identified. The aim of the present study was to explore whether the molecular mechanisms described above, alone or in combination, are enough to fully interpret the phenotype of azole resistance in clinical isolates of C. glabrata or if other mechanisms are correlated with azole resistance. Thus, we set out to explore these mechanisms in 59 clinical isolates of C. glabrata collected in China by focusing on their efflux pumps, transporter and azole antifungal target enzyme.
Material and methods
Strains and medium
A total of 59 C. glabrata isolates were collected from the Renji hospital affiliated to Shanghai Jiao Tong University School of Medicine and Dongfang hospital affiliated to Shanghai Tongji University. The isolates were identified based on standard clinical microbiological methods, involving the assessment of the carbohydrate assimilation pattern (API 21C, Paris, France) and colony characteristics on chromogenic medium (Chromagar, Paris, France) and then confirmed by Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) (Skyray, JiangSu, China). All isolates were cultured on yeast extract-peptone-glucose agar and preserved in broth containing 20% glycerol at −80 °C. The study samples were collected from the samples that were received routinely in the laboratory. It was not collected separately for this project; hence, informed consent from the patients was not required, and the Research Ethics Committee of Longhua Hospital, Shanghai University of Traditional Chinese Medicine approved to conduct this study.
Antifungal susceptibility testing
The ATB Fungus 3 strip (bioMérieux, Marcy l’Etoile, France) was used to determine the antifungal susceptibility (flucytosine, amphotericin B, fluconazole, voriconazole, and itraconazole) of the 59 clinical isolates.17,18 Quality control was ensured by testing strains C. krusei ATCC 6258 and C. parapsilosis ATCC 22,019. Susceptibility to fluconazole was also assessed for all 59 C. glabrata strains using the broth microdilution method recommended by the Clinical and Laboratory Standards Institute (CLSI). The minimal inhibitory concentrations (MICs) were determined in RPMI 1640 buffered to a PH of 7.0 with an inoculum size of approximately 1.5×103 cells/mL. The MIC for amphotericin B and flucytosine was defined as the lowest concentration at which no visible growth was detected. Recently approved CLSI breakpoints were used:19 strains with fluconazole MICs≤32 μg/mL were defined as susceptible-dose dependent, while those with MICs≥64 μg/mL were defined as resistant. There were no clinical break points for voriconazole and itraconazole, and thus, species-specific epidemiological cut-off values (ECVs) were used to define isolates as wide-type (WT) or non-WT. The epidemiological cut off values (ECVs) for voriconazole and itraconazole were 0.25 μg/mL and 4 μg/mL, respectively.
Flow cytometric analysis of the efflux of rhodamine 6G
The efflux of rhodamine 6G (Sigma-Aldrich, St Louis, MO, USA), a fluorescent dye that uses the same membrane transporter as azoles in some Candida spp.,20 was measured to evaluate the activity of efflux pumps by flow cytometry, as described previously.21 Briefly, isolates cultured overnight were adjusted to a concentration of 5×107 cells/mL in 2 mL phosphate-buffered saline (PBS) and incubated at 30 °C for 4 h with shaking for deprivation. Rhodamine 6G was then added at a final concentration of 10 μM and shaken at 30 °C for 2 h. After washing twice with cold sterile PBS, the uptake of fluorescence by the cells was first measured at 535 nm with an LSRII flow cytometer (BD Biosciences, Franklin Lakes, NJ, USA). Glucose at a final concentration of 4 mM was added and shaken at 30 °C for another 2 h. The fluorescence of the cells was then evaluated. The fluorescence of cells incubated without rhodamine 6G served as a control. The data of geometric mean values presented correspond to fluorescence frequency distribution histograms (relative number of cells vs relative fluorescence intensity, expressed in arbitrary units on a logarithmic scale).
Quantitative real-time RT-PCR
Total RNA was extracted from isolates grown to the mid-exponential phase in YPD medium with the Yeast RNAiso Reagent Kit (TaKaRa, Shiga, Japan) according to the manufacturer’s instructions. Quantitative real-time–PCR of CgCDR1, CgCDR2, CgSNQ2, CgPDR1, and CgERG11 was performed with the SYBR Premix Ex Taq Kit (TaKaRa) and LightCycler 480 Real-Time PCR System (Roche, Basel, Switzerland), as described previously.22 All reactions were performed as follows: denaturation at 95 °C for 1 min, followed by 40 cycles that consisted of 15 s at 95 °C and 60 s at 60 °C. The primers used in the real-time RT–PCR are shown in Table 1. Quantification of each target gene was performed by the 2-△△CT method using the housekeeping gene ACT1 as a control.23 Each reaction was performed in triplicate.
 | Table 1 Primers used for real-time PCR |
Sequencing of CgERG11 and CgPDR1
The genomic DNA of azole-resistant and azole-susceptible-dose dependent isolates was extracted with the DNeasy plant mini kit (Qiagen, Hilden, Germany). Two pairs of oligonucleotide primers were designed according to the CgERG11 GenBank sequence (accession number L40389) to cover the whole gene (Table 2). CgPDR1 coding sequences (accession number AY700584) were amplified using 6 pairs of primers shown in Table 2. Amplification conditions were as follows: 5 min of denaturation at 94 °C, followed by 30 cycles consisting of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min, and finally 7 min of elongation at 72 °C. The PCR products were purified with the High Pure PCR product purification kit (Roche Diagnostics GmbH, Mannheim, Germany) and sequenced directly, as described previously.24
 | Table 2 Primers used for PCR |
Statistical analysis
Data were analyzed using SPSS version 13.0 software (SPSS, Inc., Chicago, IL, USA). Statistical significance was determined by independent samples t-test. Statistical significance was indicated when P<0.05.
Results
Antifungal susceptibilities of C. glabrata isolates
The ATB Fungus 3 strip was used to determine the antifungal susceptibility of the 59 clinical isolates. For fluconazole, the susceptibility of all the isolates was also detected by the broth microdilution method, and the results were consistent with those of the ATB fungus 3 strip. All the 59 isolates of C. glabrata, were found to be susceptible to amphotericin B and flucytosine. According to the CLSI interpretive criteria, 20.3% (12/59) of the isolates were determined to be fluconazole-resistant (MICs ≥64 μg/mL), with a high level of MICs ranging from 64 to 128 μg/mL. MIC50 and MIC90 were 4 μg/mL and 64 μg/mL for fluconazole respectively. After applying epidemiological cutoff values, 22.0% (13/59) and 45.7% (27/59) of the isolates were categorized as non-wild-type for itraconazole (>4 μg/mL) and voriconazole (>0.25 μg/mL), respectively. The results of antifungal susceptibility testing are shown in Table 3.
 | Table 3 Information and MIC values for the 59 Candida glabrata isolates |
Uptake and efflux of rhodamine 6G
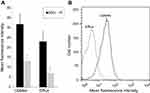
The accumulation and efflux of rhodamine 6G were quantitatively analyzed by flow cytometry. After 4 h of starvation and 2 h of incubation in PBS containing a fluorochrome, the mean fluorescence intensity of the cells was 13.16±6.26 (Table 4) in the azole-resistant group and 31.81±7.27 in the azole-susceptible-dose dependent group, with a significant difference between the two groups (Figure 1A). Moreover, after removing the free dye and incubating for another 1 h in PBS containing 4 mM glucose, the mean fluorescence intensity in the resistant isolate was significantly lower than that in the azole-susceptible-dose dependent isolate (Figure 1A), and the mean fluorescence intensity was 7.09±6.48 in the azole-resistant group and 23.09±5.61 in the azole- susceptible-dose dependent group. The visual variation of rhodamine 6G for azole-resistant isolate number 35 is presented in Figure 1B.
 | Table 4 Efflux pump activity of Candida glabrata |
Expression of CgCDR1, CgCDR2, CgSNQ2, CgERG11, and CgPDR1
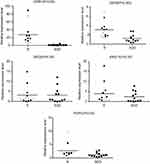
Quantitative RT–PCR experiments showed that the expression levels of CgCDR1 and CgCDR2 in the azole-resistant group were higher than those in the azole-susceptible-dose dependent group (P<0.05). On the contrary, there was no significant difference in the expression levels of CgSNQ2, CgERG11, and CgPDR1 between the azole-resistant and azole-susceptible-dose dependent isolates (P>0.05) (Figure 2).
Mutations in CgERG11
DNA sequence analysis of the CgERG11 open reading frame revealed 4 mutations (T1328C, T1394C, G1487A, and A1583G) in 24 isolates compared to the corresponding sequence in the GenBank database (GenBank accession number L40389). These point mutations did not change the amino acid sequence, indicating that this gene polymorphism was functionally silent. T1394C and A1583G were discovered in 11 azole-susceptible-dose dependent isolates and 7 azole-resistant isolates. T1328C was found in 9 azole-susceptible-dose dependent isolates and 4 azole-resistant isolates, while G1487A existed in only one azole-susceptible-dose dependent isolate.
Mutations in CgPDR1
DNA sequence analysis of the CgPDR1 open reading frame revealed 11 mutations in 24 isolates compared to the corresponding sequence in the GenBank database (GenBank accession number AY700584), of which 5 were synonymous mutations and the others were missense mutation sites. The 5 synonymous mutations were T309C, C837T, C1563T, A3156G, and T3228C, which only appeared in azole-susceptible-dose dependent isolates. For the 6 missense mutations, 2 were found in azole-susceptible-dose dependent isolates, which were A776G (E259G) and A1665C (E555D), while the other 4 missense mutations were found in azole-resistant isolates and were G1042T (G348C), A2290G (N764D), G2626A (D876N) (found in one isolate, respectively) and C2543T (A848V) (found in 8 isolates). Among the 4 missense mutations, G1042T (G348C) and A2290G (N764D) have been reported previously.25 Although G2626A (D876N) has also been reported, the amino acid mutation was different. C2543T (A848V) has not been reported and showed the highest frequency in this experiment (Table 5).
 | Table 5 Positions of polymorphic nucleotide sites in CgPDR1 |
Discussion
The mechanisms of Candida resistance against azole antifungal drugs include the following: 1) an increase in lanosterol 14α-demethylase, which increases the concentration of azole drug required to inhibit target enzyme production, 2) a decrease in the affinity between the azole drug and its target enzyme, 3) enhanced excretion effects against the drug, preventing the drug from collecting in the fungal cell, 4) inactivation of the sterol, Δ5, 6- desaturase, leading to the dysfunction of ergosterol synthesis.
Antifungal resistance can be acquired by C. glabrata during therapy.26 In addition, C. glabrata is intrinsically resistant or less susceptible to specific antifungals.27 One of the largest and a representative study that tested 1699 isolates of C. glabrata from two large antifungal surveys, the SENTRY Global Surveillance Program from 2006 to 2010 and the Centers for Disease Control and Prevention (CDC) population-based surveillance conducted in the Atlanta, GA, and Baltimore, MD, metropolitan areas between 2008 and 2010, showed that 162(9.7%) of the isolates were resistant to fluconazole, including 62 of 847 (7.3%) isolates from the SENTRY Program and 100 of 822 (12.2%) isolates from the CDC population-based surveillance.28 Data of the global SENTRY Program in 2010 and 2011 show 8.8% of C. glabrata isolates were resistant to fluconazole and that voriconazole was active against all Candida spp. except C. glabrata (10.5% non-WT).29 A study from China showed that resistance to fluconazole (MIC >32 μg/mL) was observed in 68 of 411 (16.5%) C. glabrata isolates from 11 hospitals participating in CHIF-NET (2010–2014), while MICs of >0.5 μg/mL for voriconazole and >2 μg/mL for itraconazole were observed for 28.7% and 6.8% of isolates, respectively.30 Data from CHIF-NET (2015) showed that 14 of 158 (8.9%) C. glabrata isolates were resistant to fluconazole (MICs≥64 μg/mL) and the non-WT rates of C. glabrata for voriconazole and itraconazole were 19.0% and 5.1%, respectively.31
In the present study, 20.3% (12/59) of the isolates were determined to be fluconazole-resistant (MICs≥64 μg/mL), and 22.0% (13/59) and 45.7% (27/59) of the isolates were categorized as non-wild-type for itraconazole (>4 μg/mL) and voriconazole (>0.25 μg/mL), respectively. According to the distribution of the strains, the number of C. glabrata isolates from the intensive care unit was the highest, accounting for 27.1% (16/59), followed by that from the neurosurgery ward (18.6%,11/59). This may be related to the frequent invasive operation and long-term bed rest of patients in these wards. Therefore, the prevention and control of nosocomial fungal infections in these wards should be strengthened, and attention should be paid to the examination of patients with high risk factors of invasive fungal infections. Narrow-spectrum antibiotics should be used according to the results of drug sensitivity to avoid unnecessary invasive operations, to reduce the incidence of nosocomial fungal infections. All the 59 clinical isolates of C. glabrata were genotyped by the MLST technique in our previous studies.32,33 Six ST genotypes (ST-7, ST-10, ST-3, ST-15, ST-43 and ST-55) were obtained with six housekeeping genes (FKS, LEU2, NMT1, TRP1, UGP1, and URA3) as target genes. The discriminatory power index was 0.470. Among these isolates, 43 were ST-7, accounting for 72.9% (43/59), and thus ST-7 was the dominant genotype. By comparing MLST genotype with the isolation sources, we found that, of the 11 strains of C. glabrata isolated from sterile sites (7 strains from blood and 4 strains from venous catheter, BALF, drain, and debridement, respectively), 7 isolates were ST-7, 2 were ST-55 and, the remaining 2 were ST-3 and ST-15. The sequence types (STs) were independent of isolation sources. Likewise, there was also no significant difference between different genotypes and drug susceptibility.
We attempted to evaluate different resistance mechanisms in the 59 clinical isolates of C. glabrata. Overexpression of the efflux pump increases drug efflux and prevents the drug from accumulating in the cell, leading to drug resistance. Because the same protein transports the azole drug and the fluorescent dye rhodamine 6G, we used the latter to detect the function of the efflux pump in C. glabrata. Our results showed that C. glabrata excreted significant amounts of rhodamine 6G, indicating that efflux pump function plays an important role in the drug resistance of C. glabrata. Two types of efflux pumps are present on the fungal cell membrane: the ATP-binding cassette transporter and major facilitator superfamily. CgCDR1, CgCDR2, and CgSNQ2 have been examined previously.34,35 Shin et al36 discovered 2 strains of C. glabrata isolated from the same patient; one was resistant to fluconazole and the other was sensitive to fluconazole. The mRNA expression of CgCDR1 and CgCDR2 in the resistant strain was 13.2-and 3.5-fold higher than that in the sensitive strain. Tumbarello et al37 also detected the mRNA expression of CgCDR1, CgCDR2, and CgSNQ2 in approximately 35 strains of C. glabrata, finding that 6 resistant strains were obviously upregulated, 8 susceptible- dose dependent strains were slightly upregulated, and the other sensitive strains showed no change. Our results also show that compared to that in the susceptible-dose dependent group, the expression of CgCDR1 and CgCDR2 in the resistant group was upregulated, while the expression of CgSNQ2 did not differ. In this study, comparison of the expression levels of CgCDR1 and CgCDR2 revealed the upregulation of CgCDR1 (3.4–89.6 fold) was more obvious than that of CgCDR2 (1.4–6.9 fold), which is consistent with the results of other studies. These findings support the view that CgCDR1 is more closely related to azole resistance than CgCDR2.38
Many studies have indicated that overexpression of CgERG11 causes C. glabrata to be resistant to fluconazole.39–41 However, in this study, we found that the expression difference between the resistant group and susceptible-dose dependent group was not significant. Sequencing of CgERG11 revealed 4 synonymous mutation sites, in which T1328C, T1394C, and A1583G appeared in both resistant and susceptible-dose dependent strains, while G1487A was only observed in susceptible-dose dependent strains. These 4 synonymous sites were also reported by Shen et al42 Additionally, G1487A was detected in their resistant strains. Thus, our result was conducive to the hypothesis that CgERG11 did not participate in the azole resistance of C. glabrata.
CgCDR1 and CgCDR2 have the same transcriptional regulatory factor, CgPdr1. Some CgPDR1 mutation promotes resistance to azole antibiotics by increasing the expression of CgCDR1 and CgCDR2 or enhancing the virulence of yeast and plays a central role in fluconazole resistance acquired by C. glabrata. Ferrari et al25 investigated the incidence of CgPDR1 mutations in a large collection of clinical isolates and identified 57 distinct single amino acid substitutions located at 50 locations, including the 3 mutations found in our research. Caudle et al43 reported upregulation of CgCDR1, CgCDR2, and CgSNQ2 and a mutation in CgPDR1 T2837C (L946S) in C. glabrata. They introduced this mutation into the sensitive strain and found that the MIC increased from 2 to 64 μg/mL. Tsai et al44 also introduced the gain of function mutation sites of CgPDR1 C2465T (P822L) from resistant C. glabrata into the sensitive strain and observed that the MIC was increased. Therefore, mutation in CgPDR1 is one mechanism leading to the resistance of C. glabrata. In the present study, we detected CgPDR1 mutations in 12 azole resistant clinical isolates, among which we found that only one isolate (isolate 21) did not have CgPDR1 mutation, while all the 11 remaining isolates carried a single missense mutation. Interestingly, the isolates number 21 showed a moderate upregulation of CgCDR1 (3.4-fold) while CgCDR2 was significantly upregulated (6.7-fold). The other 11 isolates carrying CgPDR1 mutations seemed to result in higher CgCDR1 expression levels, ranging from 7.5- to 89.6-fold increases. Compared with that of CgCDR1, the impact of CgPDR1 mutations on CgCDR2 was much weaker in these 11 isolates, with an increase of no more than 5.7-fold. Therefore, the CgPDR1 mutation has different regulatory effects on the expression of CgCDR1 and CgCDR2, and not all clinical isolates of C. glabrata showing azole resistance can be explained by amino acid substitutions in PDR1. Additionally, we found 4 missense mutation sites in only resistant strains, among which C2543T (A848V) and the other three were found in 8 of the 11 isolates, and one isolate, respectively. The high mutation frequency of A848V in our present study has not been reported previously, and thus, further studies of the related gene disruption and genome-wide expression may help to determine the regulation of these mutations in C. glabrata.
Conclusion
In summary, efflux pump function is the main mechanism of resistance to fluconazole in C. glabrata. Resistance development is a complex process regulated by multiple factors, and thus, cannot be explained by a single resistance gene or a single mechanism; therefore, further studies are needed to understand the relationship.
Acknowledgments
This study was supported by National Nature Science Foundation of China (Grant number 81803893).
Author contributions
All authors contributed to data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Perlin DS, Rautemaa-Richardson R, Alastruey-Izquierdo A. The global problem of antifungal resistance: prevalence, mechanisms, and management. Lancet Infect Dis. 2017;17(12):e383–e392. doi:10.1016/S1473-3099(17)30316-X
2. Sanguinetti M, Posteraro B, LassFlörl C. Antifungal drug resistance among Candida species: mechanisms and clinical impact. Mycoses. 2015;58(Suppl 2):2–13. doi:10.1111/myc.12330
3. Guinea J. Global trends in the distribution of Candida species causing candidemia. Clin Microbiol Infect. 2014;20(Suppl 6):5–10. doi:10.1111/1469-0691.12539
4. Xiao M, Fan X, Chen CA, et al. Antifungal susceptibilities of Candida glabrata species complex, Candida krusei, Candida parapsilosis species complex and Candida tropicalis causing invasive candidiasis in china: 3 year national surveillance. J Antimicrob Chemother. 2015;70(3):802–810. doi:10.1093/jac/dku460
5. Falagas ME, Roussos N, Vardakas KZ. Relative frequency of albicans and the various non-albicans Candida spp. among candidemia isolates from inpatients in various parts of the world: a systematic review. Int J Infect Dis. 2010;14(11):e954–e966. doi:10.1016/j.ijid.2010.04.006
6. Pfaller MA, Diekema DJ, Jones RN, et al. International surveillance of bloodstream infections due to Candida species: frequency of occurrence and in vitro susceptibilities to fluconazole, ravuconazole, and voriconazole of isolates collected from 1997 through 1999 in the SENTRY antimicrobial surveillance program. J Clin Microbiol. 2001;39(9):3254–3259.
7. Schwartz IS, Patterson TF. The emerging threat of antifungal resistance in transplant infectious diseases. Curr Infect Dis Rep. 2018;20(3):2. doi:10.1007/s11908-018-0608-y
8. Wang H, Xiao M, Chen SC, et al. In vitro susceptibilities of yeast species to fluconazole and voriconazole as determined by the 2010 national China Hospital Invasive Fungal Surveillance Net (CHIFNET) study. J Clin Microbiol. 2012;50:3952–3959. doi:10.1128/JCM.01263-11
9. Borst A, Raimer MT, Warnock DW, Morrison CJ, Arthington-Skaggs BA. Rapid acquisition of stable azole resistance by Candida glabrata isolates obtained before the clinical introduction of fluconazole. Antimicrob Agents Chemother. 2005;49(2):783–787. doi:10.1128/AAC.49.2.783-787.2005
10. Redding SW, Kirkpatrick WR, Saville S, et al. Multiple patterns of resistance to fluconazole in Candida glabrata isolates from a patient with oropharyngeal candidiasis receiving head and neck radiation. J Clin Microbiol. 2003;41(2):619–622.
11. Bennett JE, Izumikawa K, Marr KA. Mechanism of increased fluconazole resistance in Candida glabrata during prophylaxis. Antimicrob Agents Chemother. 2004;48(5):1773–1777.
12. Torelli R, Posteraro B, Ferrari S, et al. The ATP‐binding cassette transporter–encoding gene CgSNQ2 is contributing to the CgPDR1‐dependent azole resistance of Candida glabrata. Mol Microbiol. 2008;68(1):186–201. doi:10.1111/j.1365-2958.2008.06143.x
13. Vermitsky JP, Earhart KD, Smith WL, Homayouni R, Edlind TD, Rogers PD. Pdr1 regulates multidrug resistance in Candida glabrata: gene disruption and genome‐wide expression studies. Mol Microbiol. 2006;61(3):704–722. doi:10.1111/j.1365-2958.2006.05235.x
14. Vermitsky JP, Edlind TD. Azole resistance in Candida glabrata: coordinate upregulation of multidrug transporters and evidence for a Pdr1-like transcription factor. Antimicrob Agents Chemother. 2004;48(10):3773–3781. doi:10.1128/AAC.48.10.3773-3781.2004
15. Miyazaki H, Miyazaki Y, Geber A, et al. Fluconazole resistance associated with drug efflux and increased transcription of a drug transporter gene, PDH1, in Candida glabrata. Antimicrob Agents Chemother. 1998;42(7):1695–1701.
16. Vanden Bossche H, Marichal P, Odds FC, Le Jeune L, Coene MC. Characterization of an azole-resistant Candida glabrata isolate. Antimicrob Agents Chemother. 1992;36(12):2602–2610.
17. Li F, Wu L, Cao B, Zhang Y, Li X, Liu Y. Surveillance of the prevalence, antibiotic susceptibility, and genotypic characterization of invasive candidiasis in a teaching hospital in China between 2006 to 2011. BMC Infect Dis. 2013;13(1):353. doi:10.1186/1471-2334-13-353
18. Chen PL, Lo HJ, Wu CJ, et al. Species distribution and antifungal susceptibility of blood Candida isolates at a tertiary hospital in southern Taiwan, 1999–2006. Mycoses. 2011;54(4):e17–23. doi:10.1111/j.1439-0507.2009.01818.x
19.
20. Maesaki S, Marichal P, Vanden Bossche H, Sanglard D, Kohno S. Rhodamine 6G efflux for the detection of CDR1-overexpressing azole-resistant Candida albicans strains. J Antimicrob Chemother. 1999;44(1):27–31.
21. Peng Y, Dong D, Jiang C, Yu B, Wang X, Ji Y. Relationship between respiration deficiency and azole resistance in clinical, Candida glabrata. FEMS Yeast Res. 2012;12(6):719–727. doi:10.1111/j.1567-1364.2012.00821.x
22. Jiang C, Dong D, Yu B, et al. Mechanisms of azole resistance in 52 clinical isolates of Candida tropicalis in China. J Antimicrob Chemother. 2013;68(4):778–785. doi:10.1093/jac/dks481
23. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
24. Brun S, Bergès T, Poupard P, et al. Mechanisms of azole resistance in petite mutants of Candida glabrata. Antimicrob Agents Chemother. 2004;48(5):1788–1796.
25. Ferrari S, Ischer F, Calabrese D, et al. Gain of function mutations in CgPDR1 of Candida glabrata not only mediate antifungal resistance but also enhance virulence. PLoS Pathog. 2009;5(1):e1000268. doi:10.1371/journal.ppat.1000706
26. Lewis JS, Wiederhold NP, Wickes BL, Patterson TF, Jorgensen JH. Rapid emergence of echinocandin resistance in Candida glabrata resulting in clinical and microbiologic failure. Antimicrob Agents Chemother. 2013;57(9):4559–4561. doi:10.1128/AAC.01144-13
27. Arendrup MC, Patterson TF. Multidrug-resistant Candida: epidemiology, molecular mechanisms, and treatment. J Infect Dis. 2017;216(suppl 3):S445–S451. doi:10.1093/infdis/jix131
28. Pfaller MA, Castanheira M, Lockhart SR, Ahlquist AM, Messer SA, Jones RN. Frequency of decreased susceptibility and resistance to echinocandins among fluconazole-resistant bloodstream isolates of Candida glabrata. J Clin Microbiol. 2012;50(4):1199–1203. doi:10.1128/JCM.01263-11
29. Pfaller MA, Messer SA, Woosley LN, Jones RN, Castanheira M. Echinocandin and triazole antifungal susceptibility profiles for clinical opportunistic yeast and mold isolates collected from 2010 to 2011: application of new CLSI clinical breakpoints and epidemiological cutoff values for characterization of geographic and temporal trends of antifungal resistance. J Clin Microbiol. 2013;51(8):2571–2581. doi:10.1128/JCM.00308-13
30. Xin H, Meng X, Chen SCA, et al. Molecular epidemiology and antifungal susceptibility of Candida glabrata in China (august 2009 to July 2014): a Multi-center study. Front Microbiol. 2017;8:880. doi:10.3389/fmicb.2017.00880
31. Hou X, Xiao M, Wang H, et al. Profiling of PDR1 and MSH2 in Candida glabrata bloodstream isolates from a multi-center study in China. Antimicrob Agents Chemother. 2018;62(6):e00153–18. doi:10.1128/AAC.00153-18
32. Yao D, Ying C, Zheng B. Application evaluation of microsatellite polymorphism and multilocus sequence typing in genotyping of Candida glabrata. Lab Med. 2014;29(6):593–596.
33. Zheng B, Yao D, Ying C, Wang Y, Zhang H, Yang J. Epidemiology and drug-resistance analysis of nosocomial infection of Candida glabrata. Lab Med. 2012;27(6):461–466.
34. Ferrari S, Sanguinetti M, Torelli R, Posteraro B, Sanglard D, Cramer R. Contribution of CgPDR1-regulated genes in enhanced virulence of azole-resistant Candida glabrata. PLoS One. 2011;6(3):e17589. doi:10.1371/journal.pone.0017589
35. Ferrari S, Sanguinetti M, De Bernardis F, et al. Loss of mitochondrial functions associated with azole resistance in Candida glabrata results in enhanced virulence in mice. Antimicrob Agents Chemother. 2011;55(5):1852–1860. doi:10.1128/AAC.01271-10
36. Shin JH, Chae MJ, Song JW, et al. Changes in karyotype and azole susceptibility of sequential bloodstream isolates from patients with Candida glabrata candidemia. J Clin Microbiol. 2007;45(8):2389–2391. doi:10.1128/JCM.00381-07
37. Tumbarello M, Sanguinetti M, Trecarichi EM, et al. Fungaemia caused by Candida glabrata with reduced susceptibility to fluconazole due to altered gene expression: risk factors, antifungal treatment and outcome. J Antimicrob Chemother. 2008;62(6):1379–1385. doi:10.1093/jac/dkn381
38. Sanguinetti M, Posteraro B, Fiori B, Ranno S, Torelli R, Fadda G. Mechanisms of azole resistance in clinical isolates of Candida glabrata collected during a hospital survey of antifungal resistance. Antimicrob Agents Chemother. 2005;49(2):668–679. doi:10.1128/AAC.49.2.668-679.2005
39. Henry KW, Nickels JT, Edlind TD. Upregulation of ERG genes in Candida species by azoles and other sterol biosynthesis inhibitors. Antimicrob Agents Chemother. 2000;44(10):2693–2700.d
40. Niimi M, Nagai Y, Niimi K, et al. Identification of two proteins induced by exposure of the pathogenic fungus Candida glabrata to fluconazole. J Chromatogr B Analyt Technol Biomed Life Sci. 2002;782(1–2):245–252.
41. Samaranayake YH, Cheung BPK, Wang Y, Yau JY, Yeung KW, Samaranayake LP. Fluconazole resistance in Candida glabrata is associated with increased bud formation and metallothionein production. J Med Microbiol. 2013;62(Pt 2):303–318. doi:10.1099/jmm.0.044123-0
42. Shen Y, Lu H, Zhang Y. Analysis of ERG11 gene mutations in fluconazole-resistant Candida glabrata strains. Chin J Infect Dis. 2010;6(28):331–335.
43. Caudle KE, Barker KS, Wiederhold NP, Xu L, Homayouni R, Rogers PD. Genomewide expression profile analysis of the Candida glabrata Pdr1 regulon. Eukaryot Cell. 2011;10(3):373–383. doi:10.1128/EC.00073-10
44. Tsai HF, Sammons LR, Zhang X, et al. Microarray and molecular analyses of the azole resistance mechanism in Candida glabrata oropharyngeal isolates. Antimicrob Agents Chemother. 2010;54(8):3308–3317. doi:10.1128/AAC.00535-10
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.