Back to Journals » OncoTargets and Therapy » Volume 13
lncRNA UCA1 Contributes to 5-Fluorouracil Resistance of Colorectal Cancer Cells Through miR-23b-3p/ZNF281 Axis
Authors Xian Z, Hu B, Wang T, Zeng J, Cai J, Zou Q, Zhu P
Received 17 April 2020
Accepted for publication 22 June 2020
Published 31 July 2020 Volume 2020:13 Pages 7571—7583
DOI https://doi.org/10.2147/OTT.S258727
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjay Singh
Zhenyu Xian,1,* Bang Hu,2,* Ting Wang,1 Junyi Zeng,1 Jinlin Cai,1 Qi Zou,2 Peixuan Zhu3
1Graceland Medical Center, The Sixth Affiliated Hospital of Sun Yat-sen University, Guangzhou, Guangdong, People’s Republic of China; 2Department of Colorectal Surgery, The Sixth Affiliated Hospital of Sun Yat-sen University (Gastrointestinal and Anal Hospital), Guangzhou, Guangdong, People’s Republic of China; 3International Medical Center, Guangzhou General Hospital of Foresea Life Insurance, Guangzhou, Guangdong, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Peixuan Zhu
International Medical Center, Guangzhou General Hospital of Foresea Life Insurance, 703 Xinchengdadao, Zengcheng District, Guangzhou 511300, Guangdong, People’s Republic of China
Tel +86 13682275819
Email [email protected]
Purpose: The chemoresistance of 5-fluorouracil (5-FU) limited the application of chemotherapy in colorectal cancer (CRC) treatment. Herein, we aimed to uncover the potential mechanism behind the 5-FU resistance of CRC cells.
Methods: The abundance of long noncoding RNA urothelial carcinoma associated 1 (lncRNA UCA1), microRNA-23b-3p (miR-23b-3p) and zinc finger protein 281 (ZNF281) was measured by quantitative real-time polymerase chain reaction (qRT-PCR) in CRC tissues and cells. Western blot was conducted to examine autophagy-related proteins, apoptosis-associated proteins and ZNF281 in CRC tissues and cells. Cell counting kit-8 (CCK8) assay was performed to detect the viability and inhibitory concentration 50% (IC50) value of 5-FU of CRC cells. The apoptosis of CRC cells was measured by flow cytometry. The binding sites between miR-23b-3p and UCA1 or ZNF281 were predicted by miRcode and Starbase software, respectively, and the combination was confirmed by dual-luciferase reporter assay and RIP assay. Murine xenograft model was established to verify the role of UCA1 on the 5-FU resistance of CRC in vivo.
Results: The 5-FU resistance of CRC was positively related to the level of UCA1 and autophagy. UCA1 accelerated the 5-FU resistance of CRC cells through facilitating autophagy and suppressing apoptosis. MiR-23b-3p was a target of UCA1 in 293T and CRC cells. The knockdown of miR-23b-3p reversed the inhibitory effects of UCA1 interference on the 5-FU resistance and autophagy and the promoting impact on the apoptosis of CRC cells. ZNF281 could bind to miR-23b-3p in 293T cells. MiR-23b-3p elevated the 5-FU sensitivity through down-regulating ZNF281 in CRC cells. UCA1 interference enhanced the 5-FU sensitivity of CRC through miR-23b-3p/ZNF281 axis in vivo.
Conclusion: UCA1 mediated 5-FU resistance of CRC cells through facilitating autophagy and inhibiting apoptosis via miR-23b-3p/ZNF281 axis in vivo and in vitro.
Keywords: colorectal cancer, 5-FU, UCA1, miR-23b-3p, ZNF281, autophagy, apoptosis
Introduction
Chemotherapy and surgical resection are the primary treatment methods of colorectal cancer (CRC).1 5-fluorouracil (5-FU) is a common chemotherapeutic drug for CRC therapy. However, chemoresistance is an enormous challenge for the effective application of chemotherapeutic agents in CRC.
Long noncoding RNAs (lncRNAs) are long-chain RNAs, and they have been reported to regulate the levels of microRNAs (miRNAs) through serving as miRNAs sponges.2–6 Wang et al demonstrated that lncRNA urothelial carcinoma associated 1 (UCA1) facilitated the progression of bladder cancer, suggesting that UCA1 played an oncogenic role in bladder cancer.7 Accumulating articles have also confirmed the oncogenic functions of UCA1 in multiple cancers.8–11 For instance, Fang et al claimed that UCA1 contributed to multi-drug resistance of gastric cancer through sponging miR-27b.9 Han et al reported that the enrichment of UCA1 was positively related to tumor size and depth.11 Besides, Bian et al found that UCA1 facilitated CRC cell proliferation, and it also contributed to 5-FU resistance through miR-204-5p/CREB1.12 Nevertheless, the precise signal pathway of UCA1 in the chemoresistance of CRC is not fully addressed.
miRNAs participated in the proliferation, metastasis and apoptosis by negatively modulating the levels of their target messenger RNAs (mRNAs) or inhibiting their translation.13–16 Accumulating articles reported that miR-23b-3p played an anti-tumor role in a variety of cancers.17–20 For example, Kou et al demonstrated that the low expression of miR-23b was associated with the poor prognosis of CRC patients.20 Herein, we aimed to assess the function of miR-23b-3p in the chemoresistance of CRC.
Zinc finger protein 281 (ZNF281) mediates both transcriptional activation and suppression of its target genes.21 Hahn et al claimed that ZNF281 promoted the epithelial-mesenchymal transition (EMT), and it was positively regulated by SNAIL and negatively modulated by miR-34a.22 Besides, ZNF281 has been reported to accelerate the proliferation and metastasis of pancreatic cancer cells and CRC cells through Wnt/β-catenin signal pathway.23,24 However, the potential role of ZNF281 in the chemoresistance of CRC remains poorly understood.
In this study, we assessed the role of UCA1 in the chemoresistance of CRC. And then we explored the molecular mechanism by which UCA1 contributed to the 5-FU chemoresistance of CRC cells.
Patients and Methods
Patients
5-FU-resistant CRC patients (n=25) and 5-FU-sensitive CRC patients (n=25) who had not received surgery, chemotherapy or radiotherapy in The Sixth Affiliated Hospital of Sun Yat-sen University were recruited in this study. According to RECIST 1.1 criteria, CRC patients were divided into Sensitive-CRC group (containing Complete Response, Partial Response and Stable Disease) and Resistant-CRC group (Progressive Disease).25 This research had granted approval by the Ethics Committee of The Sixth Affiliated Hospital of Sun Yat-sen University. Patients whose tissues used in this study had provided written informed consents.
Cell Culture and Drug
Human CRC cell line SW480 and SW620 and human embryonic kidney cell line 293T were obtained from Bena Culture Collection (Beijing, China). All cells were cultivated with Dulbecco’s Modified Eagle Medium (DMEM; Gibco, Carlsbad, CA, USA) added with 10% fetal bovine serum (FBS; Gibco) and 10% penicillin (100 U/mL)-streptomycin (100 μg/mL) mixed solution in a 37°C, 5% CO2 humidified incubator.
5-FU was obtained from Sigma (St. Louis, MO, USA). The 5-FU-resistant CRC cell lines were built by treating normal SW480 and SW620 cells with increased concentrations (1 µg/mL, 2 µg/mL, 5 µg/mL, 10 µg/mL, 20 µg/mL, 40 µg/mL) of 5-FU. 1 µg/mL 5-FU was used for the maintenance of drug resistance in SW480/5-FU and SW620/5-FU cells.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
qRT-PCR was performed through using SYBR Green PCR Kit (Takara, Dalian, China) and the following primers (GeneCopoeia, Rockville, MD, USA). U6 or β-actin acted as the internal reference of UCA1, miR-23b-3p and ZNF281. Relative expression of the above genes was analyzed by 2−ΔΔCt method.26 The primers were listed as follows: UCA1 (Forward, 5ʹ-CTCTCCATTGGGTTCACCATTC-3ʹ; Reverse, 5ʹ-GCGGCAGGTCTTAAGAGATGAG-3ʹ), miR-23b-3p (Forward, 5ʹ-NBS]?>GAGCATCACATTGCCAGGG-3ʹ; Reverse, 5ʹ-GTGCAGGGTCCGAGGT-3ʹ), ZNF281 (Forward, 5ʹ-GAGGACACATAGTGGAGAAAAGCC-3ʹ; Reverse, 5ʹ-GTGGATCTAGACTAGGGCTGGAGTT-3ʹ), U6 (Forward, 5ʹ-CTCGCTTCGGCAGCACA-3ʹ; Reverse, 5ʹ-AACGCTTCACGAATTTGCGT-3ʹ), β-actin (Forward, 5ʹ-TGGAATCCTGTGGCATCCATGAAAC-3ʹ; Reverse, 5ʹ-ACGCAGCTCAGTAACAGTCCG-3ʹ).
Western Blot Assay
The proteins were isolated from CRC cells using RIPA lysis solution (Beyotime, Shanghai, China). The proteins were quantified through using a BCA detecting kit (Beyotime), and then the samples were subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gel and transferred onto a polyvinylidene fluoride (PVDF) membrane. After incubating with 5% non-fat milk for 1 h, the PVDF membrane was incubated with specific antibodies overnight. The antibodies against Bcl-2 associated X, apoptosis regulator (Bax; ab32503), Caspase-3 (ab13847), light chain 3 I/II (LC3I/II; ab51520), Beclin-1 (ab210498), p62 (ab155686), ZNF281 (ab101318) and β-actin (ab8226) were obtained from Abcam (Cambridge, MA, USA). The membrane was then incubated with a secondary antibody (ab205718, Abcam) for 1 h. The protein images were obtained through the enhanced chemiluminescent (ECL) system (Beyotime).
Cell Transfection
When CRC cells grew to 7080% confluence, transfection was carried out using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). Small interfering RNA negative control (si-NC), siRNA against UCA1 (si-UCA1), siRNA against ZNF281 (si-ZNF281), short hairpin RNA negative control (sh-NC) and shRNA against UCA1 (sh-UCA1) were purchased from Genepharma (Shanghai, China). miRNA-NC (miR-NC), miR-23b-3p, anti-miR-NC and miR-23b-3p inhibitor were obtained from Ribobio (Guangzhou, China).
Cell Counting Kit-8 (CCK8) Assay
CRC cells were cultured in 96-well plates to assess cell viability and inhibitory concentration 50% (IC50) value of 5-FU with Cell Counting Kit-8 (Sigma). Ten microliter CCK8 reagent was added to the wells of 96-well plates, and the CRC cells were incubated with CCK8 reagent for an additional 2 h. The optical density was measured by a microplate reader at 450 nm.
Cell Apoptosis Analysis
CRC cells were harvested using cold phosphate buffer saline (PBS) buffer and re-suspended in binding buffer. The cells were mixed with Annexin V and propidium iodide (PI; Solarbio, Beijing, China) for 10 min. Then the samples were analyzed by the flow cytometer (BD Biosciences, San Jose, CA, USA).
Dual-Luciferase Reporter Assay
miRcode online software was used for predicting the targets of lncRNA UCA1. The miR-23b-3p binding sites in UCA1 sequences (wild-type or mutant type) were embedded into the luciferase reporter vector. The luciferase activity was analyzed through the dual-luciferase reporter assay system (Promega, Madison, WI, USA). The targets of miR-23b-3p were predicted by Starbase software. The confirmation of the combination between miR-23b-3p and ZNF281 was conducted following the same approach.
RNA Immunoprecipitation (RIP) Assay
RIP buffer (Millipore, Billerica, MA, USA) was used to lyse CRC cells, and the cell extract was incubated with anti-Argonaute-2 (anti-Ago2; Millipore) or anti-Immunoglobulin G (anti-IgG; Millipore) beads. The immunoprecipitated RNAs were identified by qRT-PCR.
Murine Xenograft Assay
This protocol was permitted by the Animal Care and Use committee of The Sixth Affiliated Hospital of Sun Yat-sen University. All animal procedures were performed in accordance with the Guidelines for Care and Use of Laboratory Animals of “National Institutes of Health”. Nude mice were obtained from Orient Bio Inc (Seongnam, South Korea). SW480/5-FU cells stably expressing sh-NC or sh-UCA1 were inoculated to the flank of nude mice. When the average tumor size reached about 50 mm3, 30 mg/kg 5-FU was intraperitoneally injected into murine three times a week. The dimension of tumor was measured every 5 d. After 37-d inoculation, the nude mice were sacrificed and the tumors were excised and weighed.
Statistical Analysis
All data were displayed as mean ± standard deviation (S.D.). Student’s t-test was used to assess the differences between two groups. The correlation between the expression of miR-23b-3p and UCA1 or ZNF281 in 5-FU resistant-CRC patients was calculated by Spearman correlation coefficient. P<0.05 was considered to be statistically significant.
Results
5-FU Resistance of CRC is Related to the Abundance of UCA1 and Autophagy
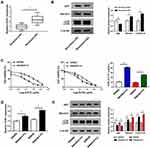
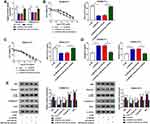
To illustrate the chemoresistance mechanism of CRC, we aimed to find crucial molecules involved in 5-FU resistance. As showed in Figure 1A, UCA1 was up-regulated in 5-FU resistant-CRC tissues compared with that in 5-FU sensitive-CRC tissues. Besides, we found that autophagy was promoted in 5-FU resistant-CRC tissues (Figure 1B). These findings demonstrated that the 5-FU resistance of CRC was associated with the level of UCA and autophagy. We established two 5-FU-resistant CRC cell lines (SW480/5-FU and SW620/5-FU) to further confirm our results. The IC50 values of 5-FU in SW480/5-FU and SW620/5-FU cell lines were notably elevated compared with their parental cell lines (SW480 and SW620) (Figure 1C), suggested that the establishment of 5-FU-resistant CRC cell lines was successful. As indicated in Figure 1D, the expression of UCA1 was elevated in the two 5-FU-resistant subclones compared with that in their parental cells. Additionally, the autophagy-related proteins (LC3II and Beclin1) were up-regulated in the two 5-FU-resistant subclones (Figure 1E), while p62 was down-regulated in SW480/5-FU and SW620/5-FU cells, suggesting that autophagy was accelerated in two 5-FU-resistant subclones compared with that in their parental cells.
UCA1 Contributes to 5-FU Resistance of CRC Cells Through Promoting Autophagy and Inhibiting Apoptosis
To explore the role of UCA1 in 5-FU resistance of CRC cells, we conducted loss-of-function experiments. We evaluated the knockdown efficiency of si-UCA1 in SW480/5-FU and SW620/5-FU cells. As mentioned in Figure 2A, the expression of UCA1 in SW480/5-FU and SW620/5-FU was prominently reduced by the transfection of si-UCA1. As indicated in Figure 2B and C, UCA1 knockdown decreased the viability and IC50 of 5-FU in SW480/5-FU and SW620/5-FU cells, demonstrating that UCA1 depletion elevated the 5-FU sensitivity of CRC cells. To investigate the mechanism by which UCA1 contributing to 5-FU resistance of CRC cells, we performed flow cytometry and Western blot assay. The apoptosis was notably facilitated in si-UCA1 transfected SW480/5-FU and SW620/5-FU cells compared with that in si-NC group (Figure 2D). Besides, Western blot assay showed that the abundance of Bax, Caspase-3 and p62 was enhanced (Figure 2E), while the enrichment of Beclin1 and LC3II was conspicuously decreased, suggesting that UCA1 intervention promoted the apoptosis while impeded the autophagy of SW480/5-FU and SW620/5-FU cells. Taken together, UCA1 enhanced the 5-FU resistance of CRC cells through promoting autophagy and restraining apoptosis.
miR-23b-3p Could Bind to UCA1 in CRC Cells
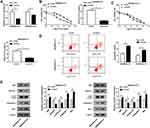
The enrichment of miR-23b-3p was lower in 5-FU resistant-CRC tissues than that in 5-FU sensitive-CRC tissues (Figure 3A). Apart from this, the level of miR-23b-3p was declined in 5-FU-resistant CRC cells (SW480/5-FU and SW620/5-FU) compared with that in SW480 and SW620 cells (Figure 3B). The level of miR-23b-3p was negatively correlated with the abundance of UCA1 in 5-FU resistant-CRC tissues (Figure 3C). miR-23b-3p was predicted as a target of UCA1 by miRcode software, and we confirmed the combination between miR-23b-3p and UCA1 in 293T cells by dual-luciferase reporter assay. Luciferase activity was conspicuously declined in 293T cells co-transfected with miR-23b-3p mimic and UCA1 WT (Figure 3D), while it has little impact in miR-23b-3p mimic and UCA1 MUT group, suggesting that miR-23b-3p was a direct target of UCA1 in 293T cells. Meanwhile, RIP assay also showed that UCA1 could bind to the RNA-induced silencing complex (RISC) in SW480/5-FU and SW620/5-FU cells, likely through the combination with miR-23b-3p (Figure 3E).
UCA1 Promotes the 5-FU Resistance of CRC Cells Through Sponging miR-23b-3p
To clarify the modulatory relationship between UCA1 and miR-23b-3p in CRC cells and whether miR-23b-3p was involved in UCA1-mediated 5-FU resistance of CRC cells, we transfected si-NC, si-UCA1, si-UCA1 + anti-miR-NC or si-UCA1 + miR-23b-3p inhibitor into SW480/5-FU and SW620/5-FU cells. As showed in Figure 4A, the level of miR-23b-3p was dramatically increased by si-UCA1 transfection, and the addition of miR-23b-3p inhibitor reversed this promoting effect, demonstrating that miR-23b-3p was inversely modulated by UCA1 in CRC cells. miR-23b-3p depletion abolished the inhibitory effect of UCA1 interference on the 5-FU resistance of SW480/5-FU and SW620/5-FU cells (Figure 4B and C). Besides, the addition of miR-23b-3p inhibitor counteracted the promoting effect of UCA1 depletion on the apoptosis and the suppressive effect on the autophagy of SW480/5-FU and SW620/5-FU cells (Figure 4D and E). Taken together, UCA1 contributed to 5-FU resistance of CRC cells through negatively regulating miR-23b-3p.
ZNF281 is a Direct Target of miR-23b-3p in CRC Cells
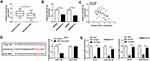
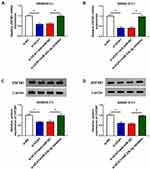
To illustrate the underlying mechanism by which miR-23b-3p elevating the 5-FU sensitivity of CRC cells, we aimed to investigate the downstream component of miR-23b-3p in CRC cells. As showed in Figure 5A and B, the mRNA and protein expression of ZNF281 was markedly up-regulated in 5-FU resistant-CRC tissues. Meanwhile, the abundance of ZNF281 mRNA and protein was notably increased in 5-FU-resistant CRC cells compared with that in their matching parental CRC cells (Figure 5C and D). As mentioned above, the expression of miR-23b-3p was lower in 5-FU resistant-CRC tissues, and we proposed the hypothesis that there was a significantly negative correlation between the levels of ZNF281 and miR-23b-3p in 5-FU resistant-CRC tissues. Subsequently, the result of the correlation analysis supported our hypothesis (Figure 5E). ZNF281 was predicted as a target of miR-23b-3p by Starbase software, and then dual-luciferase reporter assay verified the combination between ZNF281 and miR-23b-3p in 293T cells (Figure 5F). The luciferase activity was markedly reduced with the accumulation of miR-23b-3p in ZNF281 3ʹ UTR-WT group compared with that in ZNF281 3ʹ UTR-MUT group, suggesting that ZNF281 was a direct target of miR-23b-3p in 293T cells.
miR-23b-3p Elevates the 5-FU Sensitivity Through Negatively Regulating ZNF281 in CRC Cells
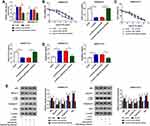
To explore whether ZNF281 could regulate the 5-FU resistance of CRC cells through serving as a downstream gene of miR-23b-3p, we transfected si-NC, si-ZNF281, si-ZNF281 + anti-miR-NC or si-ZNF281 + miR-23b-3p inhibitor into SW480/5-FU and SW620/5-FU cells. As mentioned in Figure 6A, the transfection of si-ZNF281 down-regulated the abundance of ZNF281, and the co-transfection of si-ZNF281 and miR-23b-3p inhibitor recovered the level of ZNF281 in SW480/5-FU and SW620/5-FU cells. The viability of SW480/5-FU and SW620/5-FU cells and IC50 value of 5-FU were down-regulated by si-ZNF281 transfection, and the addition of miR-23b-3p inhibitor recovered the viability and IC50 value of 5-FU in the two CRC cells (Figure 6B and C). Flow cytometry revealed that miR-23b-3p depletion abated the promoting effect of si-ZNF281 transfection on the apoptosis of SW480/5-FU and SW620/5-FU cells (Figure 6D). Moreover, we conducted Western blot assay to detect the proteins-associated with apoptosis and autophagy. As indicated in Figure 6E, ZNF281 interference elevated the abundance of Caspase-3, Bax and p62 and declined the expression of Beclin1 and LC3II, and the addition of miR-23b-3p inhibitor reversed the effects caused by ZNF281 depletion, suggesting that the depletion of miR-23b-3p diminished the promoting effect of ZNF281 silencing on the apoptosis and the inhibitory impact on the autophagy of CRC cells. These findings revealed that ZNF281 functioned as a downstream gene of miR-23b-3p to contribute to the 5-FU resistance of CRC cells.
ZNF281 is Modulated by UCA1/miR-23b-3p Axis in CRC Cells
To further explore the modulatory relationship among UCA1, miR-23b-3p and ZNF281 in SW480/5-FU and SW620/5-FU cells, we conducted the following experiments. SW480/5-FU and SW620/5-FU cells were transfected with si-NC, si-UCA1, si-UCA1 + anti-miR-NC or si-UCA1 + miR-23b-3p inhibitor. As showed in Figure 7A and B, UCA1 knockdown decreased the mRNA level of ZNF281 in SW480/5-FU and SW620/5-FU cells, and the addition of miR-23b-3p inhibitor recovered the abundance of ZNF281 mRNA. Meanwhile, the change of ZNF281 protein revealed the same trend to the change of ZNF281 mRNA in the above SW480/5-FU and SW620/5-FU cells (Figure 7C and D). Taken together, the level of ZNF281 was regulated by UCA1/miR-23b-3p signaling in CRC cells.
UCA1 Contributes to 5-FU Resistance of CRC in vivo
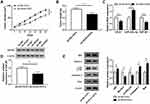
To validate the biological significance of UCA1 on the 5-FU resistance of CRC in vivo, we established the murine xenograft model using SW480/5-FU cells stably transfected with sh-NC or sh-UCA1. As mentioned in Figure 8A and B, tumor volume and weight were less in sh-UCA1 + 5-FU group than that in sh-NC + 5-FU group, suggesting that UCA1 depletion elevated the 5-FU sensitivity of CRC in vivo. The abundance of UCA1 was notably decreased in sh-UCA1 + 5-FU group (Figure 8C). UCA1 intervention enhanced the level of miR-23b-3p. Meanwhile, the abundance of ZNF281 mRNA and protein was prominently reduced in sh-UCA1 + 5-FU group compared with that in sh-NC + 5-FU group (Figure 8C and D). Additionally, pro-apoptotic proteins (Bax and Caspase-3) and p62 were up-regulated while the enrichment of Beclin1 and LC3II was conspicuous declined in sh-UCA1 + 5-FU group compared with that in sh-NC + 5-FU group (Figure 8E). Taken together, UCA1 promoted the 5-FU resistance of CRC through accelerating autophagy and inhibiting apoptosis via miR-23b-3p/ZNF281 axis in vivo.
Discussion
Accumulating studies have revealed the crucial role of UCA1 in multiple cancers, including bladder cancer, CRC and HCC.11,27,28 Han et al found that high expression of UCA1 was closely correlated with poor prognosis of CRC patients, and UCA1 was also found to accelerate the proliferation and cell cycle and suppress the apoptosis of CRC cells.11 UCA1 was also reported to promote the development of HCC via suppressing miR-216b and activating FGFR1/ERK signal axis.28 Furthermore, accumulating works have pointed out the functional association between UCA1 and chemoresistance in many cancers, including CRC.12,29,30 Zhang et al demonstrated that UCA1 enhanced the adriamycin (ADR) resistance of pediatric acute myeloid leukemia (AML) through promoting glycolysis via regulating miR-125a/hexokinase 2 signaling.30 Sun et al demonstrated that UCA1, CRNDE, H19 and HOTAIR were associated with Oxaliplatin or Irinotecan resistance in CRC cells according to the results of integrative bioinformatics analysis.31 Bian et al reported that UCA1 elevated the 5-FU resistance and proliferation of CRC cells through sponging miR-204-5p.12 Herein, we found that the abundance of UCA1 was enhanced in 5-FU resistant-CRC tissues and cells, suggesting that UCA1 played a pivotal role in the chemoresistance of CRC. Besides, autophagy was enhanced in 5-FU resistant-CRC tissues and cells. Autophagy has been reported to accelerate chemoresistance and the viability of cancer cells under diverse stresses.32–34 lncRNA H19 was found to enhance 5-FU resistance of CRC cells through accelerating SIRT1-mediated autophagy.32 Zhang et al demonstrated that miR-22 elevated the sensitivity of CRC cells to 5-FU through promoting apoptosis and restraining the autophagy.33 We proposed the hypothesis that UCA1 contributed to the 5-FU resistance of CRC through promoting autophagy.
We conducted knockdown experiments to assess the function of UCA1 in the 5-FU resistance of CRC cells. UCA1 depletion inhibited the viability of CRC cells treated with different concentrations of 5-FU compared with that in the control group. And the IC50 value was notably reduced by si-UCA1 transfection in CRC cells. Besides, UCA1 interference also accelerated the apoptosis and inhibited the autophagy of CRC cells. These findings suggested that UCA1 facilitated the 5-FU resistance of CRC cells through promoting autophagy and inhibiting apoptosis.
The target relationship between UCA1 and miR-23b-3p was predicted by miRcode software, and we confirmed the combination between UCA1 and miR-23b-3p in 293T cells and CRC cells by dual-luciferase reporter assay and RIP assay. MiR-23b-3p played an anti-tumor role in diverse cancers.20,35,36 For instance, miR-23b-3p hampered the proliferation and motility of cervical cancer cells through targeting c-Met.35 Kou et al reported that low abundance of miR-23b in plasma of CRC patients was related to the poor prognosis.20 Besides, Hu et al found that miR-23b-3p acted as the downstream target of MALAT1 to suppress autophagy-related chemoresistance.37 The knockdown of miR-23b-3p reversed the inhibitory effects of UCA1 depletion on the 5-FU resistance and autophagy and the promoting impact on the apoptosis of CRC cells, suggesting that miR-23b-3p functioned as a downstream component of UCA1 to elevate the 5-FU sensitivity of CRC cells.
ZNF281 has been reported to play an oncogenic role in non-small-cell lung cancer and CRC.24,38 ZNF281 promoted the proliferation and metastasis of CRC cells through regulating Wnt/β-Catenin signal pathway.24 Consistent with the above findings, we found that ZNF281 contributed to the 5-FU resistance of CRC cells. We found that the mRNA and protein expression of ZNF281 was conspicuously elevated in 5-FU resistant-CRC tissues and cells. Starbase software predicted that ZNF281 was a target of miR-23b-3p, and we verified the combination between ZNF281 and miR-23b-3p by dual-luciferase reporter assay. The addition of miR-23b-3p inhibitor counteracted the suppressive effects of ZNF281 depletion on the 5-FU resistance, viability and autophagy, and the promoting impact on the apoptosis of CRC cells. To further clarify the modulatory relationship between UCA1, miR-23b-3p and ZNF281 in CRC cells, we conducted the following experiments. The level of ZNF281 was declined with UCA1 depletion, and the addition of miR-23b-3p inhibitor recovered the expression of ZNF281 in SW480/5-FU and SW620/5-FU cells, demonstrating that ZNF281 was modulated by UCA1/miR-23b-3p axis in CRC cells.
We built the murine xenograft model using SW480/5-FU cells stably transfected with sh-UCA1 or sh-NC to assess the function of UCA1 in the 5-FU resistance of CRC in vivo. UCA1 depletion enhanced the 5-FU sensitivity of CRC tumor through promoting apoptosis and suppressing autophagy via miR-23b-3p/ZNF281 signaling in vivo.
In conclusion, UCA1/miR-23b-3p/ZNF281 axis was identified in CRC cells for the first time in this study. UCA1 contributed to the 5-FU resistance of CRC through promoting autophagy and suppressing apoptosis via miR-23b-3p/ZNF281 axis. Restoration of miR-23b-3p level might be an effective therapeutic strategy for CRC.
Disclosure
The authors declare that they have no conflicts of interest in this work.
References
1. Jonker FH, Tanis PJ, Coene PP, et al. Impact of neoadjuvant radiotherapy on complications after hartmann procedure for rectal cancer. Dis Colon Rectum. 2015;58:931–937. doi:10.1097/dcr.0000000000000432
2. Zhang L, He X, Jin T, et al. Long non-coding RNA DLX6-AS1 aggravates hepatocellular carcinoma carcinogenesis by modulating miR-203a/MMP-2 pathway. Biomed Pharmacother. 2017;96:884–891. doi:10.1016/j.biopha.2017.10.056
3. Wei H, Hu J, Pu J, et al. Long noncoding RNA HAGLROS promotes cell proliferation, inhibits apoptosis and enhances autophagy via regulating miR-5095/ATG12 axis in hepatocellular carcinoma cells. Int Immunopharmacol. 2019;73:72–80. doi:10.1016/j.intimp.2019.04.049
4. Cesana M, Cacchiarelli D, Legnini I, et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell. 2011;147:358–369. doi:10.1016/j.cell.2011.09.028
5. Kallen AN, Zhou XB, Xu J, et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell. 2013;52:101–112. doi:10.1016/j.molcel.2013.08.027
6. Wang K, Long B, Zhou LY, et al. CARL lncRNA inhibits anoxia-induced mitochondrial fission and apoptosis in cardiomyocytes by impairing miR-539-dependent PHB2 downregulation. Nat Commun. 2014;5:3596. doi:10.1038/ncomms4596
7. Wang F, Li X, Xie X, et al. UCA1, a non-protein-coding RNA up-regulated in bladder carcinoma and embryo, influencing cell growth and promoting invasion. FEBS Lett. 2008;582:1919–1927. doi:10.1016/j.febslet.2008.05.012
8. Huang J, Zhou N, Watabe K, et al. Long non-coding RNA UCA1 promotes breast tumor growth by suppression of p27 (Kip1). Cell Death Dis. 2014;5:e1008. doi:10.1038/cddis.2013.541
9. Fang Q, Chen X, Zhi X. Long non-coding RNA (LncRNA) Urothelial Carcinoma Associated 1 (UCA1) increases multi-drug resistance of gastric cancer via downregulating miR-27b. Med Sci Monit. 2016;22:3506–3513. doi:10.12659/msm.900688
10. Li JY, Ma X, Zhang CB. Overexpression of long non-coding RNA UCA1 predicts a poor prognosis in patients with esophageal squamous cell carcinoma. Int J Clin Exp Pathol. 2014;7:7938–7944.
11. Han Y, Yang YN, Yuan HH, et al. UCA1, a long non-coding RNA up-regulated in colorectal cancer influences cell proliferation, apoptosis and cell cycle distribution. Pathology. 2014;46:396–401. doi:10.1097/pat.0000000000000125
12. Bian Z, Jin L, Zhang J, et al. LncRNA-UCA1 enhances cell proliferation and 5-fluorouracil resistance in colorectal cancer by inhibiting miR-204-5p. Sci Rep. 2016;6:23892. doi:10.1038/srep23892
13. Croce CM, Calin GA. miRNAs, cancer, and stem cell division. Cell. 2005;122:6–7. doi:10.1016/j.cell.2005.06.036
14. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. doi:10.1016/s0092-8674(04)00045-5
15. Ambros V. The functions of animal microRNAs. Nature. 2004;431:350–355. doi:10.1038/nature02871
16. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–866. doi:10.1038/nrc1997
17. Salvi A, Sabelli C, Moncini S, et al. MicroRNA-23b mediates urokinase and c-met downmodulation and a decreased migration of human hepatocellular carcinoma cells. FEBS J. 2009;276:2966–2982. doi:10.1111/j.1742-4658.2009.07014.x
18. Loftus JC, Ross JT, Paquette KM, et al. miRNA expression profiling in migrating glioblastoma cells: regulation of cell migration and invasion by miR-23b via targeting of Pyk2. PLoS One. 2012;7:e39818. doi:10.1371/journal.pone.0039818
19. Li W, Liu Z, Chen L, et al. MicroRNA-23b is an independent prognostic marker and suppresses ovarian cancer progression by targeting runt-related transcription factor-2. FEBS Lett. 2014;588:1608–1615. doi:10.1016/j.febslet.2014.02.055
20. Kou CH, Zhou T, Han XL, et al. Downregulation of miR-23b in plasma is associated with poor prognosis in patients with colorectal cancer. Oncol Lett. 2016;12:4838–4844. doi:10.3892/ol.2016.5265
21. Wang ZX, Teh CH, Chan CM, et al. The transcription factor Zfp281 controls embryonic stem cell pluripotency by direct activation and repression of target genes. Stem Cells. 2008;26:2791–2799. doi:10.1634/stemcells.2008-0443
22. Hahn S, Jackstadt R, Siemens H, et al. SNAIL and miR-34a feed-forward regulation of ZNF281/ZBP99 promotes epithelial-mesenchymal transition. EMBO J. 2013;32:3079–3095. doi:10.1038/emboj.2013.236
23. Qian Y, Li J, Xia S. ZNF281 promotes growth and invasion of pancreatic cancer cells by activating Wnt/beta-catenin signaling. Dig Dis Sci. 2017;62:2011–2020. doi:10.1007/s10620-017-4611-1
24. Qin CJ, Bu PL, Zhang Q, et al. ZNF281 regulates cell proliferation, migration and invasion in colorectal cancer through wnt/beta-catenin signaling. Cell Physiol Biochem. 2019;52:1503–1516. doi:10.33594/000000104
25. Wang D, Yang L, Yu W, et al. Colorectal cancer cell-derived CCL20 recruits regulatory T cells to promote chemoresistance via FOXO1/CEBPB/NF-κB signaling. J Immunother Cancer. 2019;8:215–219. doi:10.1186/s40425-019-0701-2
26. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi:10.1006/meth.2001.1262
27. Wang XS, Zhang Z, Wang HC, et al. Rapid identification of UCA1 as a very sensitive and specific unique marker for human bladder carcinoma. Clin Cancer Res. 2006;12:4851–4858. doi:10.1158/1078-0432.ccr-06-0134
28. Wang F, Ying HQ, He BS, et al. Upregulated lncRNA-UCA1 contributes to progression of hepatocellular carcinoma through inhibition of miR-216b and activation of FGFR1/ERK signaling pathway. Oncotarget. 2015;6:7899–7917. doi:10.18632/oncotarget.3219
29. Wang H, Guan Z, He K, et al. LncRNA UCA1 in anti-cancer drug resistance. Oncotarget. 2017;8:64638–64650. doi:10.18632/oncotarget.18344
30. Zhang Y, Liu Y, Xu X. Knockdown of LncRNA-UCA1 suppresses chemoresistance of pediatric AML by inhibiting glycolysis through the microRNA-125a/hexokinase 2 pathway. J Cell Biochem. 2018;119:6296–6308. doi:10.1002/jcb.26899
31. Sun F, Liang W, Qian J. The identification of CRNDE, H19, UCA1 and HOTAIR as the key lncRNAs involved in oxaliplatin or irinotecan resistance in the chemotherapy of colorectal cancer based on integrative bioinformatics analysis. Mol Med Rep. 2019;20:3583–3596. doi:10.3892/mmr.2019.10588
32. Wang M, Han D, Yuan Z, et al. Long non-coding RNA H19 confers 5-Fu resistance in colorectal cancer by promoting SIRT1-mediated autophagy. Cell Death Dis. 2018;9:1149. doi:10.1038/s41419-018-1187-4
33. Zhang H, Tang J, Li C, et al. MiR-22 regulates 5-FU sensitivity by inhibiting autophagy and promoting apoptosis in colorectal cancer cells. Cancer Lett. 2015;356:781–790. doi:10.1016/j.canlet.2014.10.029
34. Yu X, Shi W, Zhang Y, et al. CXCL12/CXCR4 axis induced miR-125b promotes invasion and confers 5-fluorouracil resistance through enhancing autophagy in colorectal cancer. Sci Rep-UK. 2017;7:42226. doi:10.1038/srep42226
35. Campos-Viguri GE, Peralta-Zaragoza O, Jiménez-Wences H, et al. MiR-23b-3p reduces the proliferation, migration and invasion of cervical cancer cell lines via the reduction of c-Met expression. Sci Rep-UK. 2020;10:3256. doi:10.1038/s41598-020-60143-x
36. Yang T, He X, Chen A, et al. LncRNA HOTAIR contributes to the malignancy of hepatocellular carcinoma by enhancing epithelial-mesenchymal transition via sponging miR-23b-3p from ZEB1. Gene. 2018;670:114–122. doi:10.1016/j.gene.2018.05.061
37. YiRen H, YingCong Y, Sunwu Y, et al. Long noncoding RNA MALAT1 regulates autophagy associated chemoresistance via miR-23b-3p sequestration in gastric cancer. Mol Cancer. 2017;16:174. doi:10.1186/s12943-017-0743-3
38. Xue YB, Ding MQ, Xue L, et al. CircAGFG1 sponges miR-203 to promote EMT and metastasis of non-small-cell lung cancer by upregulating ZNF281 expression. Thorac Cancer. 2019;10:1692–1701. doi:10.1111/1759-7714.13131
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.