Back to Journals » OncoTargets and Therapy » Volume 13
LncRNA SNHG15 Contributes to Immuno-Escape of Gastric Cancer Through Targeting miR141/PD-L1
Authors Dang S , Malik A, Chen J, Qu J, Yin K, Cui L, Gu M
Received 15 April 2020
Accepted for publication 1 August 2020
Published 24 August 2020 Volume 2020:13 Pages 8547—8556
DOI https://doi.org/10.2147/OTT.S251625
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjay Singh
Shengchun Dang,1,2 Abdul Malik,1 Jixiang Chen,1 Jianguo Qu,1 Kai Yin,1 Lei Cui,1 Min Gu3
1Department of General Surgery, Affiliated Hospital of Jiangsu University, Zhenjiang, Jiangsu Province 212001, People’s Republic of China; 2Department of General Surgery, Pucheng Hospital, Weinan, Shaanxi Province 715500, People’s Republic of China; 3Department of Oncology, Zhenjiang Hospital of Traditional Chinese and Western Medicine, Zhenjiang, Jiangsu 212001, People’s Republic of China
Correspondence: Min Gu
Department of Oncology, Zhenjiang Hospital of Traditional Chinese and Western Medicine, Zhenjiang, Jiangsu 212001, People’s Republic of China
Tel/Fax +86-511-88820988
Email [email protected]
Introduction: Long non-coding RNAs (lncRNAs) have been demonstrated to participate in many biological processes and severs as important regulators during the progression of gastric cancer.
Methods: Here, we introduced human lncRNA SNHG15 which was highly expressed in gastric cancer and cells. Interestingly, the expression of SNHG15 was correlated with programmed cell death ligand 1 (PD-L1), which promotes the resistance of gastric cancer cells to immune responses. Meanwhile, SNHG15 downregulation suppressed the expression of PD-L1 and resistance of immune responses.
Results: Further, our results suggested that SNHG15 acted as a competing endogenous RNA (CeRNA) to sponge miR-141, which was downregulated in gastric cancers and negatively correlated to PD-L1.
Conclusion: Our results suggested that SNHG15 improved the expression of PD-L1 by inhibiting miR-141, which in turn promoted the resistance of stomach cancer cells to the immune responses.
Keywords: lncRNA SNHG15, gastric cancer, PD-L1, immuno-escape, miR-141
Introduction
Gastric cancer is a common malignant tumor with poor prognosis in the world which has becoming a serious threat to human health.1 According to Jacques Ferlay et al, there were about 951,000 new cases of gastric cancer worldwide in 2012 and about 723,000 deaths from gastric cancer, which were ranking 5th in malignant tumor morbidity and 3rd in mortality.2 More than 70% of new cases of stomach cancer occur in developing countries.1–4 However, based on current therapeutic strategies, it is difficult to be cured for advanced gastric cancer. Therefore, a better understanding of molecular mechanism is important to improve the early diagnosis and targeted therapy for gastric cancer.
Long non-coding RNAs (lncRNAs) are RNA molecules with more than 200 nucleotides in length which has a limit or no protein-coding ability.5 Over the past decade, a series of studies have shown that IncRNA can be served as tumor marker for cancers.6,7 Abnormally expressed lncRNAs are suggested play an important regulatory role in the development of cancer which some of them were functionally characterized to be involved in multiple cellular biological processes, such as cell proliferation, invasion, apoptosis, cancer metastasis and even tumor genesis.8,9 Several lncRNAs have been reported to be related to gastric cancer.8 Some lncRNAs such as TINCR, TUSC7, UCA1 mediate cell proliferation and invasion.8,10,11 Other lncRNAs including CCAT1, GAPLINC, GAS5, H19 play oncogenic or tumor suppressor roles.12–15 The human SNHG15 gene is an 837-bp-long transcript without protein-coding which located on chromosome 7p13.16 It has been reported that SNHG15 is up-regulated in GC tissues, breast cancer, papillary thyroid carcinoma, colon cancer, hepatocellular carcinoma, Osteosarcoma, etc.16–21 However, the function of SNHG15 in gastric cancer is still unclear.
Here in this study, the biological function of lncRNA SNHG15 and gastric cancer was investigated. First, we examined the expression of SNHG15 in gastric cancer tissue and found that its expression was significantly up-regulated, and the same result was also found in gastric cancer cells. This phenomenon indicates that SNHG15 may play an important role in gastric cancer. Interestingly, the expression of SNHG15 was correlated with programmed cell death ligand 1 (PD-L1) which promotes the resistance of gastric cancer cells to immune responses. Meanwhile, overexpression of SNHG15 improved the expression of PD-L1 and thus promoted immune escape of HGC-27 cells while knocking down SNHG15 showed the opposite results. Further studies had found that the expression of miR-141 was negatively correlated with the expression of SNHG15 and PD-L1. The target relationship between SNHG15 and miR-141, PD-L1 and miR-141 were also verified. At last, our work demonstrated that LncRNA SNHG15 contributes to immuno-escape of gastric cancer cells through regulating the inhibition of PD-L1 by miR141. At the same time, it also provides a new therapeutic target and direction for the diagnosis and treatment of gastric cancer.
Methods
Sample Collection
Nine gastric cancer samples were obtained from patients who were confirmed by two pathologists by histopathological diagnosis in the Affiliated Hospital of Jiangsu University. Primary cancer tissues and paired adjacent non-cancerous tissues were immediately frozen in liquid nitrogen after excision and stored at −80°C before RNA extraction. The study was approved by the Research Ethics Committee of the Affiliated Hospital of Jiangsu University, China. Informed consents were obtained from all patients.
Cell Culture and Transfection
The gastric mucosal cell line GES-1 and human gastric adenocarcinoma cancer cell line HGC-27 were obtained from the Chinese Academy of Sciences Committee on Type Culture Collection cell bank (Shanghai, China). Cells were cultured using RPMI-1640 medium (Gibco, USA) supplemented with 10% fetal bovine serum (Gibco, USA) which maintained at 37°C in a humidified 5% CO2 incubator.
For transfection, cells were transfected with the plasmid of SNHG15 over-expression, siRNA against SNHG15 (Sense: 5′- UCA AAC UUG CUC AAU UAA GGU-3′, Anti-sence:3′- CUU AAU UGA GCA AGU UUG AAA-5′), miR-141 mimic (Guide strand: 5ʹ-UAA CAC UGU CUG GUA AAG AUG G-3ʹ, Passenger strand: 5ʹ-CAU CUU CCA GUA CAG UGU UGG A-3ʹ), miR-141 inhibitor (5ʹ-CCA UCU UUA CCA GAC AGU GUU A-3ʹ), plasmid of PD-L1 or their negative corresponding controls (purchased from Synthgene, China) using transfection reagent Lipofectamine 2000 (Invitrogen, CA, USA) according to the manufacturer’s instructions.
RNA Isolation, qRT-PCR Analysis
Trizol reagent (Invitrogen, Shanghai, China) was used to extract total RNA from tissues and cells. The expression levels of SNHG15 were analyzed by HiScript II One Step qRT-PCR SYBR Green Kit (Vazyme, Nanjing, China) with the GAPDH gene as a standard control. The primer sequences used are as follows: SNHG15 (Forward: 5′-CAACCATAGCGGTGCAACTGTGC-3′, Reverse: 5ʹ-GTACTGAACGTTGAACCAAGTCGG-3′); GAPDH (Forward: 5′-CAGTGCCAGCCTCGTCTAT-3′, Reverse: 5ʹ-CTTCTGACACCTACCGGGGA-3′). For the detection of miR-141 expression, extracted RNA was reverse-transcribed using the TaqMan MicroRNA Reverse Transcription Kit and miRNA-specific stem-loop primers (Applied Biosystems, USA). The reaction conditions were 16 °C for 30 min, 42 °C for 30 min, 85 °C for 5 min, and 4 °C until the end of the reaction. Real-time PCR assays of the transcribed cDNA were performed using the TaqMan MicroRNA assays (Applied Biosystems, USA). The reaction conditions were 95 °C for 10 min, followed by 40 cycles at 95 °C for 15 s, and 60 °C for 1 min. The reactions were carried out using 0.5 μg of RNA extracted from cells and U6 was used as internal control.
Luciferase Constructs and Transfection
According to the information of TargetScan, the potential target of miR-141 was predicted. The sequences of SNHG15 that could bind to miR-141 were partly mutated and inserted into the reporter plasmid in order to identify the binding specificity. The WT/MUT plasmids, miR-141 mimic and its control were co-transfected into HGC-27 cells. After 48 h of transfection, the relative luciferase activities were detected by the Dual-Glo Luciferase Assay System (Promega, Shanghai, China) approved by protocol. Renilla signals were used to normalize luciferase activity.
Cell Apoptosis Detection
HGC-27 cells were seeded on a 6-well plate. After reaching 80% confluency, HGC-27 cells were transfected with SNHG15 plasmid, SNH15 siRNA, or miR-141 mimic by Lipofectamine 2000 according to the manufacturer’s protocol. After 24 h, PBMCs (MT-BIO, Shanghai) were resuscitated and activated with 100 ng/mL CD3 antibody,100 ng/mL CD28 antibody, and 10 ng/mL IL2 for 24 h. Then, activated PBMCs were incubated with HGC-27 cells at an effector-to-target ratio of 5:1 at 37 °C for another 24 h. Then, the mixed cells were rinsed with PBS for 5 times to remove PBMCs, and HCT-116 cells were collected and identified through flow cytometry by using an Annexin V-FITC/PI staining kit according to manufacturer’s protocol.
Detection of Surface PD-L1
HGC-27 cells were seeded on a 6-well plate. After reaching 80% confluency, HGC-27 cells were transfected with SNHG15 plasmid, SNH15 siRNA, or miR-141 mimic by Lipofectamine 2000 according to the manufacturer’s protocol. After 48 h, cells were collected. After wash with PBS for 3 times, cells were blocked with 0.1% BSA for 30 min, stained with FAM-labeled PD-L1 antibody (BioLegend, USA) (1:200) for another 30 min. After wash with PBS for 3 times, FAM-positive cells were then analyzed by flow cytometry.
Western Blot
Samples were lysed in RIPA with PMSF, and BCA assay was detected in protein concentrations. Next, total protein was separated on a 10% SDS–PAGE and transferred onto PVDF membrane. The membranes were incubated with primary antibodies at 4 °C overnight followed washed with TBST, and incubated with secondary antibody at room temperature for 1 h. Finally, the membranes were detected by chemiluminescence (ECL, Thermo Fisher, MA, USA). Anti-GAPDH antibody was used as a control.
Statistical Analysis
All experiments were conducted independently for 3 times. All quantitative data were presented as the mean ± SD. Statistical analyses were performed using two-tails Student’s t-tests. Values of P<0.05 were considered significant.
Results
The Expression of LncRNA SNHG15 Was Up-Regulated in Gastric Cancer Tissues and Cells
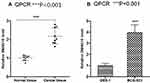
To verified the function of LncRNA SNHG15, its expression was firstly evaluated in both normal tissue and gastric tissue. Compared to normal tissue, the expression of LncRNA SNHG15 in cancer tissue was significantly increased (P < 0.001, Figure 1A). Correspondingly, the SNHG15 expression was 4-fold higher in gastric cancer cell line (HGC-27) than that in normal human gastric epithelial cells (GES-1) (P < 0.001, Figure 1B). These results indicated that SNHG15 was associated with gastric cancer.
SNHG15 Facilitated the Expression of PD-L1 and Immuno-Escape
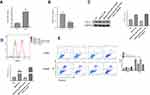
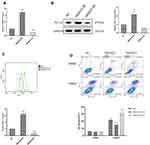
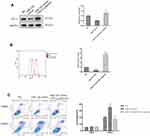
To invest the relationship between LncRNA SNHG15 and gastric cancer, LncRNA SNHG15 was overexpressed (SNHG15 OE) and knocked down (SNHG15 KD) respectively, in HGC-27 cells and the expression level of SNHG15 was then evaluated by qPCR (P < 0.001, Figure 2A). Further, the expression of whole cell PD-L1 was examined both in SNHG15 OE and SNHG15 KD cells using Western blot (P < 0.01, Figure 2B) which inferred that PD-L1 and SNHG15 were positively correlated. Consistent results were also exhibited flow cytometry (FACS) analysis of surface PD-L1 expression (P < 0.001, Figure 2C). It is reported that the expression of PD-L1 promoted immune escape of tumor cells, so we further examined whether SNHG15 promoted the immune escape of HGC-27 cells. As shown in Figure 2D, over-expression of SNHG15 significantly reduced the proportion of cell apoptosis after incubated with peripheral blood mononuclear cells (PBMCs) for 24 h, while decreased SNHG15 expression showed crosscurrent (P < 0.001, Figure 2D) when they were compared to NC. These results suggested that SNHG15 availed the expression of PD-L1 and promoted the immune escape of HGC-27 cells.
SNHG15 Mediated Immunotherapy Through Regulating miR-141
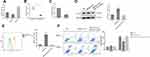
It was reported that SNHG15 could directly interact with miR-141 to promote cell proliferation, invasion and autophagy in osteosarcoma cells.21 Therefore, we also evaluated the regulation of miR-141 by SNHG15. As shown in Figure 3A, the expression of miR-141 was negatively correlated to the expression of SNHG15 (P < 0.01, Figure 3A) which suggested miR-141 was negatively regulated via SNHG15. Lower expression of miR-141 was also detected in gastric tissues compared to normal tissues or HGC-27 cells compared to normal GES-1 cells (P < 0.001, Figure 3B and C). In addition, to investigate whether SNHG15 mediated the development of gastric cancer by down-regulated miR-141, miR-141 mimic was transfected into SNHG15OE cell lines to detect the expression of PD-L1 and cell apoptosis. Using Western blot and FACS analysis, the results had shown that the sharp rise in PD-L1 expression was reversed to normal condition when SNHG15OE cell line was supplied with miR-141 mimic compared to negative controls (P < 0.001, Figure 3D and E). In addition, the cell apoptosis at the presence of PBMC was also elevated. Compared to SNHG5OE cells, the proportion of cell apoptosis was significantly increased and reached a level consistent with that of the control group when miR-141 mimic was transfected into SNHG5OE cells (P < 0.001, Figure 3D and 3E).
SNHG15 and PD-L1 Were Targets of miR-141
To study the relationship between SNHG15, miR-141 and PD-L1, the potential target binding sites were analyzed using DIANA tools – LncBase Predicted v2 (http://carolina.imis.athena-innovation.gr/diana_tools/web/index.php?r=lncbasev2/index-predicted) and targetscans - TargetScanHuman 7.2 (http://www.targetscan.org/vert_72/). As shown in Figure 4A, both SNHG15 and PD-L1 were predicted to be direct targets of miR-141. Dual-luciferase activity assays were further performed to evaluate the predicted target relationship. Both the luciferase activity of SNHG15 and PD-L1 were significantly inhibited by extra miR-141 mimic, but not the mutations (P < 0.001, Figure 4B). The expression level of PD-L1 was significantly reduced in miR-141 mimic treated group and increased in miR-141 inhibitor group compared to that in control group (P < 0.01, Figure 4C and D). These results indicated that SNHG15 and miR-141 had a corresponding binding target relationship and PD-L1 was a target of miR-141 which its expression was negatively regulated by miR-141.
miR-141 Regulated Immune Escape Through Regulating PD-L1
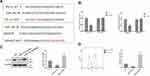
To further confirm the direct effect of miR141 regulating PD-L1 on immune escape of stomach cancer cells. PD-L1 over-expression plasmid and miR-141 mimic were used. As shown in Figure 5A and B which were detected by Western blot and FACS analysis, respectively, the decline in the level of PD-L1 expression caused by miR-141 mimic was reversed by overexpression PD-L1. Then, the immune escape of gastric cells at the presence of PBMC was also evaluated. Treating HGC-27 cells with miR-141 mimic alone significantly increased the rate of apoptosis while this increase was abandoned by extra overexpressed of PD-L1 (Figure 5C). These results suggested that miR-141 directly affected the immune escape of gastric cancer cells by regulating the expression of PD-L1.
LncRNA SNHG15 Contributes to Immuno-Escape of Gastric Cancer Through Targeting miR-141/PD-L1
We have verified in another gastric cancer cell line (MKN45) to further illustrate that LncRNA SNHG15 can promote the immune escape of gastric cancer by targeting miR-141/PD-L1. First, we examined the expression of lncRNA SNHG15 and miR-141 in MKN45 cells. Compared to normal human gastric epithelial cells (GES-1), the expression of LncRNA SNHG15 in MKN45 cells was significantly increased and miR-141 was decreased (Figure 6A and B). To further confirm the direct effect of miR141 regulating PD-L1 on the immune escape of stomach cancer cells. PD-L1 over-expression plasmid and miR-141 mimic were used. Further, the expression of whole cell PD-L1 was examined using Western blot and the results had shown that the sharp rise in PD-L1 expression was reversed to normal condition when SNHG15OE cell line was supplied with miR-141 mimic compared to negative controls (P < 0.01, Figure 6C). In the previous study, we verified the conclusion that miR-141 directly affects the immune escape of gastric cancer cells by regulating the expression of PD-L1 in HGC-27 cells. Similarly, we also further verified it in another cell line KMN45. Consistent results were also exhibited flow cytometry (FACS) analysis of surface PD-L1 expression (Figure 6D). Then, the immune escape of gastric cells at the presence of PBMC was also evaluated. Treating MKN45 cells with miR-141 mimic alone significantly increased the rate of apoptosis while this increase was abandoned by extra overexpressed of PD-L1 (P < 0.01, Figure 6E). We conducted a correlation analysis and found that SNHG15 and PD-L1 were positive correlation. In addition, immunohistochemical results also indicate that PD-L1 expression is increased in gastric cancer tissue samples. These results suggested that SNHG15 improved the expression of PD-L1 by inhibiting miR-141, which in turn promoted the resistance of stomach cancer cells to the immune responses. The above results are consistent with the previous.
LncRNA and SNHG15 PD-L1 Was Up-Regulated in Gastric Cancer Tissues
We conducted a correlation analysis and found that SNHG15 and PD-L1 were positive correlation (Figure 7A). In addition, immunohistochemical results also indicate that PD-L1 expression is increased in gastric cancer tissue samples. The above results are consistent with the previous (Figure 7B).
 |
Figure 7 LncRNA and SNHG15 PD-L1 was up-regulated in gastric cancer tissues. (A) Correlation analysis of LncRNA SNHG15 and PD-L1. (B) Immunohistochemistry of PD-L1 in gastric cancer samples. |
Discussion
The occurrence and development of cancer has a very complex regulatory mechanism and it is also a great threat to the survival of mankind. Gastric cancer is one of the most deadly cancers. However, the underlying mechanisms that regulate the progression of gastric cancer remain largely unclear. During this study, we explored the effects of up-regulated lncRNA SNHG15 on gastric cancer and found that SNHG15 promoted immune escape of gastric cell from PBMC by regulating the expression of PD-L1. Further studies suggested that SNHG15 relieved the suppression of miR-141 on PD-L1 by inhibiting the expression of miR-141 which resulted in immune escape of gastric cell. Therefore, our results revealed that SNHG15 played an important role in the development of gastric cancer.
Long noncoding RNAs (lncRNAs) constitute a group of transcripts that can change chromatin structure and regulate gene expression at both transcriptional and posttranscriptional levels.22 Emerging evidences showed that lncRNAs have an important influence on various biological processes, such as embryonic development, immune response and tumor development.23–25 Dysregulation of lncRNA is often associated with the occurrence and development of many diseases, such as inflammatory response and cancers. The human SNHG15 gene is a conserved long intergenic noncoding RNA located on chromosome 7p13. SNHG15 has been demonstrated acting as an oncogene which is closely related to the progression of cancer. It has been reported that SNHG15 is a potential prognostic biomarker for hepatocellular carcinoma26 and promotes the progression of hepatocellular carcinoma and osteosarcoma cells by sponging miR-141.17,21 In addition, SNHG15 influences human breast cancer, papillary thyroid carcinoma and glioma microvascular endothelial cells by sponging miR-211-3p, miR-200a-3p and miR-153, respectively.18,19,27 SNHG15 is also reported to stabilize Slug to promote colon cancer progression.20 In gastric cancer, SNHG15 could directly regulate MMP2/MMP9 to effect cell proliferation and invasion.16 Competitive endogenous RNA (ceRNA) is one of the main mechanisms of lncRNA regulation.22 In this study, we explored that SNHG15 sponged miR-141 (Figures 3 and 4) which served as CeRNA in gastric cancer and further broadened the role of SNHG15 in the progression of gastric cancer.
Furthermore, we also verified that PD-L1 was a functional target of miR-141 using miR-141 mimic and miR-141 inhibitor (Figures 3–5). It has been reported in the literature that the lipoprotein HP1454 of Helicobacter pylori regulates the T cell response by forming T cell receptor signals. In addition, their study also shows that HP1454 is an important bacterial factor that exerts its proinflammatory activity by directly regulating the T-cell response. Meanwhile, the results provide evidence that a significant proportion of T cells in chronic gastritis and in gastric adenocarcinoma. PD-L1 is a ligand of PD-1 which is a co-suppression receptor molecule induced on activated T and B cells and crucial in regulating peripheral tolerance.28–30 PD-L1, usually expressed on dendritic cells, macrophages and parenchymal cells, can inhibit the function of PD-1 expressing T cells which results in suppression of the immune response by inducing apoptosis, unresponsiveness and functional exhaustion of T cells.30,31 In this study, we demonstrated that SNHG15 and PD-L1 were positive correlation. In gastric cells, up-regulated SNHG15 suppressed the expression of miR-141, the inhibitor of PD-L1, resulting in an increase in PD-L1 expression which caused the resistance of gastric cancer cells to immune responses. Our study suggests a new lncRNA-mediated mechanism for gastric cancer cells to escape immune response and the SNHG15/miR-141/PD-L1 may act as a novel therapeutic target for gastric cancer.
Acknowledgments
The Six Talent Peaks Project in Jiangsu Province (No. 2016-WSN-007), the Jiangsu 333 Project Foundation (No. BRA2017560), the Zhenjiang Science and Technology Committee (No. SH 2018061, 2019061) supported this study.
Disclosure
The authors report no conflicts of interest for this work.
References
1. Van Cutsem E, Sagaert X, Topal B, Haustermans K, Prenen H. Gastric cancer. Lancet. 2016;388:2654–2664. doi:10.1016/S0140-6736(16)30354-3
2. Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–386. doi:10.1002/ijc.29210
3. McGuire S. World Cancer Report 2014. Geneva, Switzerland: world Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Adv Nutr. 2016;7:418–419. doi:10.3945/an.116.012211
4. Saracci R, Wild CP. Fifty years of the international agency for research on cancer (1965 to 2015). Int J Cancer. 2016;138:1309–1311. doi:10.1002/ijc.29929
5. Iyer MK, Niknafs YS, Malik R, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47:199–208. doi:10.1038/ng.3192
6. Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29:452–463. doi:10.1016/j.ccell.2016.03.010
7. Qi P, Zhou XY, Du X. Circulating long non-coding RNAs in cancer: current status and future perspectives. Mol Cancer. 2016;15:39. doi:10.1186/s12943-016-0524-4
8. Li T, Mo X, Fu L, Xiao B, Guo J. Molecular mechanisms of long noncoding RNAs on gastric cancer. Oncotarget. 2016;7:8601–8612. doi:10.18632/oncotarget.6926
9. Sand M, Bechara FG, Sand D, et al. Long-noncoding RNAs in basal cell carcinoma. Tumour Biol. 2016;37:10595–10608. doi:10.1007/s13277-016-4927-z
10. Xu TP, Liu XX, Xia R, et al. SP1-induced upregulation of the long noncoding RNA TINCR regulates cell proliferation and apoptosis by affecting KLF2 mRNA stability in gastric cancer. Oncogene. 2015;34:5648–5661. doi:10.1038/onc.2015.18
11. Qi P, Xu MD, Shen XH, et al. Reciprocal repression between TUSC7 and miR-23b in gastric cancer. Int J Cancer. 2015;137:1269–1278. doi:10.1002/ijc.29516
12. Yang F, Xue X, Bi J, et al. Long noncoding RNA CCAT1, which could be activated by c-Myc, promotes the progression of gastric carcinoma. J Cancer Res Clin Oncol. 2013;139:437–445. doi:10.1007/s00432-012-1324-x
13. Hu Y, Wang J, Qian J, et al. Long noncoding RNA GAPLINC regulates CD44-dependent cell invasiveness and associates with poor prognosis of gastric cancer. Cancer Res. 2014;74:6890–6902. doi:10.1158/0008-5472.CAN-14-0686
14. Liu Y, Zhao J, Zhang W, et al. lncRNA GAS5 enhances G1 cell cycle arrest via binding to YBX1 to regulate p21 expression in stomach cancer. Sci Rep. 2015;5:10159. doi:10.1038/srep10159
15. Zhang L, Yang F, Yuan JH, et al. Epigenetic activation of the MiR-200 family contributes to H19-mediated metastasis suppression in hepatocellular carcinoma. Carcinogenesis. 2013;34:577–586. doi:10.1093/carcin/bgs381
16. Chen SX, Yin JF, Lin BC, et al. Upregulated expression of long noncoding RNA SNHG15 promotes cell proliferation and invasion through regulates MMP2/MMP9 in patients with GC. Tumour Biol. 2016;37:6801–6812. doi:10.1007/s13277-015-4404-0
17. Ye J, Tan L, Fu Y, et al., LncRNA SNHG15 promotes hepatocellular carcinoma progression by sponging miR-141-3p. J Cell Biochem. 2019;12:19775–19783. doi:10.1002/jcb.29283
18. Kong Q, Qiu M. Long noncoding RNA SNHG15 promotes human breast cancer proliferation, migration and invasion by sponging miR-211-3p. Biochem Biophys Res Commun. 2018;495:1594–1600. doi:10.1016/j.bbrc.2017.12.013
19. Wu DM, Wang S, Wen X, et al. LncRNA SNHG15 acts as a ceRNA to regulate YAP1-Hippo signaling pathway by sponging miR-200a-3p in papillary thyroid carcinoma. Cell Death Dis. 2018;9:947. doi:10.1038/s41419-018-0975-1
20. Jiang H, Li T, Qu Y, et al. Long non-coding RNA SNHG15 interacts with and stabilizes transcription factor Slug and promotes colon cancer progression. Cancer Lett. 2018;425:78–87. doi:10.1016/j.canlet.2018.03.038
21. Liu K, Hou Y, Liu Y, Zheng J. LncRNA SNHG15 contributes to proliferation, invasion and autophagy in osteosarcoma cells by sponging miR-141. J Biomed Sci. 2017;24:46. doi:10.1186/s12929-017-0353-9
22. Yoon JH, Abdelmohsen K, Gorospe M. Posttranscriptional gene regulation by long noncoding RNA. J Mol Biol. 2013;425:3723–3730. doi:10.1016/j.jmb.2012.11.024
23. Xu YC, Liang CJ, Zhang DX, et al. LncSHRG promotes hepatocellular carcinoma progression by activating HES6. Oncotarget. 2017;8:70630–70641. doi:10.18632/oncotarget.19906
24. Li J, Han W, Shen X, Han S, Ye H, Huang G. DNA methylation signature of long noncoding RNA genes during human pre-implantation embryonic development. Oncotarget. 2017;8:56829–56838. doi:10.18632/oncotarget.18072
25. Heward JA, Lindsay MA. Long non-coding RNAs in the regulation of the immune response. Trends Immunol. 2014;35:408–419. doi:10.1016/j.it.2014.07.005
26. Zhang JH, Wei HW, Yang HG. Long noncoding RNA SNHG15, a potential prognostic biomarker for hepatocellular carcinoma. Eur Rev Med Pharmacol Sci. 2016;20:1720–1724.
27. Ma Y, Xue Y, Liu X, et al. SNHG15 affects the growth of glioma microvascular endothelial cells by negatively regulating miR-153. Oncol Rep. 2017;38:3265–3277. doi:10.3892/or.2017.5985
28. Agata Y, Kawasaki A, Nishimura H, et al. Expression of the PD-1 antigen on the surface of stimulated mouse T and B lymphocytes. Int Immunol. 1996;8:765–772. doi:10.1093/intimm/8.5.765
29. Keir ME, Butte MJ, Freeman GJ, Sharpe AH. PD-1 and its ligands in tolerance and immunity. Annu Rev Immunol. 2008;26:677–704. doi:10.1146/annurev.immunol.26.021607.090331
30. Chen L, Han X. Anti-PD-1/PD-L1 therapy of human cancer: past, present, and future. J Clin Invest. 2015;125:3384–3391. doi:10.1172/JCI80011
31. Sun C, Mezzadra R, Schumacher TN. Regulation and function of the PD-L1 checkpoint. Immunity. 2018;48:434–452. doi:10.1016/j.immuni.2018.03.014
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.