Back to Journals » OncoTargets and Therapy » Volume 13
LncRNA PTTG3P Sponge Absorbs microRNA-155-5P to Promote Metastasis of Colorectal Cancer
Received 5 February 2020
Accepted for publication 24 April 2020
Published 9 June 2020 Volume 2020:13 Pages 5283—5291
DOI https://doi.org/10.2147/OTT.S248457
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Leo Jen-Liang Su
Ning Liu,1 Lei Dou,1 Xinxin Zhang2
1Department of Pancreatic and Biliary Surgery, The First Affiliated Hospital of China Medical University, Shenyang, People’s Republic of China; 2Department of Pathology, China Medical University, Shenyang, People’s Republic of China
Correspondence: Ning Liu
Department of Pancreatic and Biliary Surgery, The First Affiliated Hospital of China Medical University, No. 155, Nanjing North Street, Heping District, Shenyang 110001, Liaoning, People’s Republic of China
Tel +8601599827618
Email [email protected]
Objective: To study the role of LncRNA PTTG3P in colorectal cancer (CRC) and the underlying mechanism.
Patients and Methods: The expression level of LncRNA PTTG3P was evaluated by quantitative real-time polymerase chain reaction (qRT-PCR) in CRC cell lines and tumor tissues from 43 CRC patients, and the correlations between LncRNA PTTG3P expression and the clinicopathological indicators and prognosis of CRC patients were analyzed. Then, a PTTG3P knockdown model in CRC cell lines HCT-8 and HCT-116 was constructed. Finally, the relationship between LncRNA PTTG3P and microRNA-155-5p was explored through luciferase reporter experiments and recovery experiments.
Results: qRT-PCR results showed that LncRNA PTTG3P was markedly up-regulated in CRC tumor tissues than that in adjacent tissues. Meanwhile, patients with high LncRNA PTTG3P expression had higher rates of lymph node metastasis and distant metastasis. In addition, cell functional experiments suggested that knocking down PTTG3P markedly reduced the migration abilities of CRC cells. Subsequently, bioinformatics analysis and luciferase reporter gene experiments suggested that LncRNA PTTG3P could directly bind to microRNA-155-5P. Analysis of CRC tissue samples showed that microRNA-155-5P expression was markedly reduced in CRC and was negatively correlated with LncRNA PTTG3P. Finally, the recovery experiments also suggested that there was a mutual regulation between LncRNA PTTG3P and microRNA-155-5P, and silencing microRNA-155-5P can reverse the inhibitory effect of knocking down PTTG3P on the malignant progression of CRC.
Conclusion: In summary, LncRNA PTTG3P level was markedly increased in CRC, and was highly correlated to the incidence of lymph node metastasis and distant metastasis in CRC patients. In addition, LncRNA PTTG3P might promote the ability of CRC to invade and migrate by downregulating microRNA-155-5P. Therefore, dissecting the aberrant regulation of LncRNA PTTG3P/microRNA-155-5P may be valuable for early screening, guidance treatment, and recurrence detection of CRC.
Keywords: LncRNA PTTG3P, microRNA-155-5P, CRC, metastasis
Introduction
Colorectal cancer (CRC) is a common malignant tumor of the digestive tract that is a serious threat to human health worldwide.1,2 In 2018, the number of new cases of CRC in China reached 253,000 and the number of deaths was 139,000, ranking fifth in the incidence and death spectrum of malignant tumors in China.3,4 In the coming decades, CRC will continue to threaten the health of our residents.1–3 In order to effectively prevent and control CRC, it is urgent to clarify the mechanism of CRC development.5,6 From the perspective of cellular and molecular processes, CRC is a complex genetic disease.6,7 Changes in many genes that regulate cell growth, proliferation, apoptosis, and intercellular communication may cause intestinal epithelial cells to lose their normal growth cycle, undergo malignant transformation, infinite proliferation, and even distant metastasis.7,8 Recently, various studies have revealed a series of single nucleotide polymorphisms that affect the susceptibility of CRC and further found important CRC-related genes, which further proved that related signaling pathways functioned in the development of CRC.9,10 However, many of the genetic mutations related to susceptibility to CRC found in current research are not on traditional protein-coding genes, and researchers have begun to realize the potential important role of non-coding genes in the process of tumorigenesis.11,12
In fact, less than 2% of the sequences in the human genome are involved in encoding proteins, and at least 75% of them can be transcribed into non-coding RNAs which can bind to DNA, RNA, and proteins, and regulate their functions.13,14 Some non-coding RNAs are very small, but some non-coding RNAs exceed 200bp and are called lncRNAs.15,16 Previous studies have shown that some lncRNAs can affect tumorigenesis and development by regulating proto-oncogenes or tumor suppressor genes, important metabolic enzymes, and even miRNAs, and are considered to be tumor-related lncRNAs.16–18
Here, we revealed that lncRNA PTTG3P was up-regulated in CRC cancer tissues by lncRNA databases, and could function as an indicator for the prognosis in CRC. Besides, we demonstrated that PTTG3P interference inhibited cell migration and invasion in vitro. Finally, we demonstrated that PTTG3P could target microRNA-155-5P by Bioinformatics website analysis (miRDB), thus regulating the CRC progression. Our study may provide evidence for the development of diagnostics and therapeutics of CRC.
Patients and Methods
Patients and CRC Samples
All tissue specimens from 43 CRC patients were obtained in our hospital and approved by the medical ethics committee of The First Affiliated Hospital of China Medical University. All patients had not received any radiotherapy or chemotherapy before surgery. CRC pathological classification and staging criteria were implemented in accordance with the Union for International Cancer Control (UICC) CRC staging criteria. Patients had signed informed consent. This study was conducted in accordance with the Declaration of Helsinki.
Cell Lines and Reagents
Human colon cancer cell lines HT29, HCT-8, HCT-116, and normal human intestinal epithelial cell line FHC were obtained from ATCC (Manassas, VA, USA). Human colon cancer cell lines HT29 and HCT-8 were cultured in Dulbecco’s modified eagle medium (DMEM) (Thermo Fisher Scientific, Waltham, MA, USA) high glucose medium containing 10% fetal bovine serum (FBS) (Gibco, Rockville, MD, USA), penicillin (100U/mL) and streptomycin (100ug/mL). HCT- 116 was cultured in Roswell Park Memorial Institute 1640 (RPMI 1640) medium containing 10% fetal bovine serum and antibiotics. All cells were cultured in an incubator at 37 °C and 5% CO2. When the cells grew to 80−90% confluence, they were passaged with 1 × trypsin + ethylene diamine tetraacetic acid (EDTA) digestion.
Transfection
CRC cell lines HCT-8 and HCT-116 were inoculated into 6-well plates and cultured to a cell density of about 30% to 50%. The control group (sh-NC) and PTTG3P-containing lentiviral sequence vector (sh-PTTG3P) were purchased from Shanghai Jima Company (Shanghai, China). Lentiviral transfection was performed as instructed, and cells were collected 48 hours later for function experiments.
Cell Wound Healing
Cells were digested, centrifuged, and resuspended in serum-free medium to adjust the density to 5 x 105 cells/mL. The cell density was determined according to the cell size (most cells are plated at 40,000 cells/well), which should reach a confluency of more than 90% in the following day. After scratching, cells were rinsed gently with PBS 2–3 times, and added with low-concentration serum medium (such as 1% FBS). Then cells were observed again after 24 hours. The difference in cell healing ability was evaluated based on the migration area.
Transwell Cell Migration Assay
Cell migration was measured using a Transwell cell. About 2 × 10^5 cells were seeded in the upper chamber in 100 ul of serum-free medium. Then bottom of the chamber was added with a medium containing 10% FBS. The upper and lower culture fluids were separated by a polycarbonate membrane. After culturing the cells in a standard environment for 48 hours, the cells passing through the lower layer were collected and stained with 0.5% crystal violet dye, and finally photographed with a microscope and counted.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
TRIzol (Invitrogen, Carlsbad, CA, USA) reagent was used to extract total RNA. The extracted RNA was then subjected to cDNA synthesis using the Primescript RT Reagent, and primers were designed using Primer 5.0 software. QRT-PCR reactions were performed using SYBR® Premix Ex Taq ™ and StepOne Plus Real-time PCR System. The following primers were used in the qRT-PCR reaction: LncRNA PTTG3P: F: 5ʹ-AAACGAAGAACCAGGCATCCTT-3 ‘, R: 5ʹ-GGGAGCATCGAATGTTTTGCC-3 ‘; glyceraldehyde 3-phosphate dehydrogenase (GAPDH): F: 5ʹ-GGAGCGAGATCCTCCAAAAT-3 ‘, R: 5ʹ-GGCTGTGTCATACTTCTCATGG-3 ‘; microRNA-155-5P: F: 5ʹ-UUAAUGCUAAUCGUGAUAGGGGU-3 ‘, R: 5ʹ-CUCCUACUAAUUAGCAUUAACA-3 ‘; U6: F: 5ʹ-CTCGCTTCGCAGCACA-3 ‘, R: 5ʹ-AACGCTTCACGATTTGCGT-3 ‘. GAPDH and U6 genes were used as internal reference. The gene expression was calculated by 2-ΔΔCt method.
Dual-Luciferase Reporter Gene Assay
The wild-type and mutant PTTG3P containing microRNA-155-5P binding sites on the MIR4435-2HG promoter region were ligated into the pMIR luciferase reporter vectors, and co-transfected into cells with microRNA-155-5P mimic/NC mimic using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). The dual luciferase reporter assay was performed to normalize the reporter luciferase activity to the control firefly luciferase activity.
Statistically Analysis
SPSS 22.0 statistical software (SPSS IBM, Armonk, NY USA) was used for statistical analysis. Measurement data were compared using t-test, and categorical variables were analyzed using χ2 test or Fisher ’s exact probability method. Kaplan-Meier method was used for survival analysis and survival curves were drawn. Data were represented as mean ± SD (Standard Deviation), and P <0.05 was considered statistically significant.
Results
LncRNA PTTG3P was Highly Expressed in CRC Tissues and Cell Lines
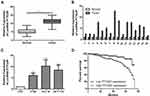
According to qRT-PCR results, compared with adjacent normal tissues, LncRNA PTTG3P was markedly increased in tumor tissues of CRC patients (Figure 1A and B). In addition, compared with FHC, LncRNA PTTG3P levels were markedly higher in CRC cell lines, especially in HCT-8 and HCT-116 cells, so these two cells were selected for subsequent experiments (Figure 1C).
LncRNA PTTG3P Expression was Correlated with Lymph Node, Distance Metastasis and Poor Prognosis in CRC Patients
According to the relative expression of LncRNA PTTG3P in tumor tissues, patients were divided into relatively high group (n=16) and relatively low groups (n=27). The relationship between LncRNA PTTG3P and clinicopathological features of CRC patients was further examined. As displayed in Table 1, high expression of LncRNA PTTG3P was positively correlated with the incidence of lymph node metastasis and distant metastases, but not with age, gender and pathological stage. In addition, Kaplan–Meier survival curve suggested that the high expression of PTTG3P was markedly associated with shorter survival of CRC patients. Among them, compared with the low PTTG3P expression group, the prognosis of CRC patients with high PTTG3P expression was relatively poor. The above results demonstrated that PTTG3P might be a new prognostic indicator for CRC (P <0.05; Figure 1D).
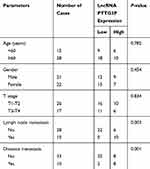 |
Table 1 Association of LncRNA PTTG3P Expression with Clinicopathologic Characteristics of Colorectal Cancer |
Knockdown of LncRNA PTTG3P Inhibited Cell Migration Ability in CRC Cell Lines
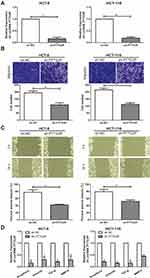
To explore the effect of LncRNA PTTG3P on the function of CRC cells, a PTTG3P knockdown expression model was successfully constructed and verified by qRT-PCR (Figure 2A). Subsequently, related experiments such as cell invasion and migration assays were performed in CRC cell lines HCT-8 and HCT-116, respectively. As a result, Transwell invasion experiments showed that the invasion ability of CRC cells was markedly reduced after knocking down PTTG3P (Figure 2B). In addition, results showed that the crawling ability of CRC cells decreased after knocking down PTTG3P, suggesting that the cell migration ability was inhibited (Figure 2C). The qRT-PCR results showed that the expression levels of key genes in the EMT signaling, including N-cadherin, Vimentin, TGF-β and MMP-9 were downregulated in CRC cells transfected with sh-PTTG3P (Figure 2D).
LncRNA PTTG3P was Bound to microRNA-155-5P
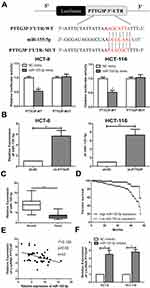
Bioinformatics studies (miRDB) was used to further identify the target genes modulated by LncRNA PTTG3P. Ultimately, the association between LncRNA PTTG3P and microRNA-155-5P was found. Subsequently, a luciferase reporter experiment showed that LncRNA PTTG3P could directly bind to microRNA-155-5P (Figure 3A). Besides, qRT-PCR experiments suggested that microRNA-155-5P expression was markedly upregulated in CRC cells after knocking down PTTG3P (Figure 3B). In addition, microRNA-155-5P expression levels were markedly reduced in tumor tissues of CRC patients (Figure 3C). Kaplan–Meier survival curve showed that low expression of microRNA-155-5P (18/43) was markedly in association with poor prognosis of CRC patients. (Figure 3D). Moreover, the expressions of LncRNA PTTG3P and microRNA-155-5P in 43 CRC tumor tissues showed a negative correlation (Figure 3E). Subsequently, microRNA-155-5P knockdown expression models were constructed and qRT-PCR assay revealed that the expression of LncRNA PTTG3P was markedly up-regulated after knockdown of microRNA-155-5P (Figure 3F). These results demonstrated that LncRNA PTTG3P could bind to microRNA-155-5P.
LncRNA PTTG3P Modulated microRNA-155-5P in Human CRC Cells
To further explore how LncRNA PTTG3P and microRNA-155-5P regulated the malignant progress of CRC, PTTG3P and microRNA-155-5P was simultaneously knocked down in HCT-8 and HCT-116. The results suggested that co-transfection of PTTG3P and microRNA-155-5P knockdown vectors markedly lowered the expression of PTTG3P (Figure 4A); meanwhile, microRNA-155-5P expression was also markedly reduced (Figure 4B). Subsequently, Transwell invasion experiments and cell scratch experiments proved that knockdown of microRNA-155-5P markedly reversed the inhibitory effects of knockdown of PTTG3P on the number of CRC cells and cell crawling ability in the transwell compartment (Figure 4C and D). Hence, LncRNA PTTG3P could modulate microRNA-155-5P in CRC.
Discussion
The etiology and pathogenesis of CRC have not yet been fully defined.1,4-6 At present, the main viewpoint is that the occurrence of CRC is a multi-factor and multi-stage process. The vast majority of CRC is the result of the interaction of genetic, environmental, psychological and lifestyle factors.5–7 Epidemiological studies have demonstrated that lifestyle-related factors such as obesity, smoking, alcohol, tea, plant fiber intake, and physical activity are associated with the risk of CRC.5,6 In terms of genetic factors, various in vivo factors such as tumor susceptibility gene polymorphism, methylation/ubiquitination, multiple gene epigenetic modifications, and non-coding RNAs have been reported with the occurrence of CRC.7,8 Among them, the association between non-coding RNA and the occurrence and development of CRC is a hot area in CRC research since the 21st century.19,20 Long-chain non-coding RNAs have attracted wide attention and may be considered to affect tumorigenesis.20
Long-chain non-coding RNA (lncRNA) is a new type of RNA molecule discovered in recent years that is closely related to the tumorigenesis of many diseases.13,14 Only about 1% of the genes in the human genome are protein-coding genes, and about 4−9% of the genes are transcribed but their transcripts cannot encode proteins, and non-coding transcripts longer than 200 nt (nucleotide units) are defined as lncRNAs.14–16 LncRNA has a variety of mechanisms of action, and it can combine with different target molecules (including DNA, mRNA, miRNA, and protein, etc.), and participates in biological processes such as epigenetic regulation and splicing regulation of genes.16,17 With the continuous research on the biological function of lncRNA, researchers have found that lncRNA is closely related to progression of many diseases, including a variety of tumors, cardiovascular diseases, respiratory system diseases, etc.16,18 This study focused on investigating the effect of LncRNA PTTG3P on the malignancy of CRC cells. In the current study, Examination of CRC tissues levels suggested that the expression levels of LncRNA PTTG3P in CRC tumor tissues were not only markedly up-regulated but also positively correlated with the lymph node metastasis and distant metastasis, which suggested that LncRNA PTTG3P may play a cancer-promoting role in CRC. Subsequently, to further explore the effect of LncRNA PTTG3P on the function of CRC cells, a lentivirus was used to construct a knockdown expression model of LncRNA PTTG3P. Cell scratch experiments and Transwell invasion experiments showed that LncRNA PTTG3P can promote the metastatic capacity of CRC cells. Bioinformatics and luciferase reporter gene analyses showed that microRNA-155-5P was involved in the development of CRC as a target gene of LncRNA PTTG3P. qRT-PCR results suggested that the expression of microRNA-155-5P in tumor tissues was markedly down-regulated. In addition, microRNA-155-5P mRNA expression in CRC cells was markedly up-regulated after knocking down PTTG3P. Subsequently, the recovery experiments verified that knockdown of microRNA-155-5P can markedly reverse the inhibition of the number of CRC cells and cell crawling ability caused by knockdown of PTTG3P. Therefore, the above experiments suggested that LncRNA PTTG3P may promote the malignant progression of CRC by regulating microRNA-155-5P. In the follow-up study, the sample size of CRC patients was need to be expand and more clinical data included to reveal the LncRNA PTTG3P in the development of CRC.
Conclusions
In summary, LncRNA PTTG3P expression was markedly increased in CRC tissues and cell lines, and was markedly correlated with the incidence of lymph node metastasis and distant metastasis in CRC patients. In addition, LncRNA PTTG3P might promote the migration ability of CRC by regulating microRNA-155-5P. Therefore, the differential expression of LncRNA PTTG3P/microRNA-155-5P can provide clinical application value for early screening, guide treatment and recurrence detection of CRC.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Brody H. Colorectal cancer. Nature. 2015;521(7551):S1. doi:10.1038/521S1a
2. Siegel RL, Miller KD, Fedewa SA, et al. Colorectal cancer statistics, 2017. CA Cancer J Clin. 2017;67(3):177–193. doi:10.3322/caac.21395
3. Zhu J, Tan Z, Hollis-Hansen K, Zhang Y, Yu C, Li Y. Epidemiological trends in colorectal cancer in China: an ecological study. Dig Dis Sci. 2017;62(1):235–243. doi:10.1007/s10620-016-4362-4
4. Pan R, Zhu M, Yu C, et al. Cancer incidence and mortality: a cohort study in China, 2008–2013. Int J Cancer. 2017;141(7):1315–1323. doi:10.1002/ijc.30825
5. Syed AR, Thakkar P, Horne ZD, et al. Old vs new: risk factors predicting early onset colorectal cancer. World J Gastrointest Oncol. 2019;11(11):1011–1020. doi:10.4251/wjgo.v11.i11.1011
6. Fayazfar S, Zali H, Arefi OA, Asadzadeh AH, Rezaei TM, Nazemalhosseini ME. Early diagnosis of colorectal cancer via plasma proteomic analysis of CRC and advanced adenomatous polyp. Gastroenterol Hepatol Bed Bench. 2019;12(4):328–339.
7. Willis JA, Vilar E. Refining prognosis in early-stage colorectal cancer: one or multiple genes at a time? Ann Oncol. 2017;28(8):1686–1688. doi:10.1093/annonc/mdx272
8. Oh HH, Joo YE. Novel biomarkers for the diagnosis and prognosis of colorectal cancer. Intest Res. 2019. doi:10.5217/ir.2019.00080
9. Thomas J, Ohtsuka M, Pichler M, Ling H. MicroRNAs: clinical relevance in colorectal cancer. Int J Mol Sci. 2015;16(12):28063–28076. doi:10.3390/ijms161226080
10. Pagani F, Randon G, Guarini V, et al. The landscape of actionable gene fusions in colorectal cancer. Int J Mol Sci. 2019;20(21). doi:10.3390/ijms20215319
11. Yiu AJ, Yiu CY. Biomarkers in colorectal cancer. Anticancer Res. 2016;36(3):1093–1102.
12. Okugawa Y, Grady WM, Goel A. Epigenetic alterations in colorectal cancer: emerging biomarkers. Gastroenterology. 2015;149(5):1204–1225. doi:10.1053/j.gastro.2015.07.011
13. Jathar S, Kumar V, Srivastava J, Tripathi V. Technological developments in lncRNA Biology. Adv Exp Med Biol. 2017;1008:283–323. doi:10.1007/978-981-10-5203-3_10
14. Ferre F, Colantoni A, Helmer-Citterich M. Revealing protein-lncRNA interaction. Brief Bioinform. 2016;17(1):106–116. doi:10.1093/bib/bbv031
15. Mishra K, Kanduri C. Understanding long noncoding RNA and chromatin interactions: what we know so far. Noncoding RNA. 2019;5(4). doi:10.3390/ncrna5040054
16. Jin SJ, Jin MZ, Xia BR, Jin WL. Long non-coding RNA DANCR as an emerging therapeutic target in human cancers. Front Oncol. 2019;9:1225. doi:10.3389/fonc.2019.01225
17. Smith KN, Miller SC, Varani G, Calabrese JM, Magnuson T. Multimodal long noncoding RNA interaction networks: control panels for cell fate specification. Genetics. 2019;213(4):1093–1110. doi:10.1534/genetics.119.302661
18. McDonel P, Guttman M. Approaches for understanding the mechanisms of long noncoding RNA regulation of gene expression. Cold Spring Harb Perspect Biol. 2019;11(12):a032151. doi:10.1101/cshperspect.a032151
19. Wang L, Cho KB, Li Y, Tao G, Xie Z, Guo B. Long noncoding RNA (lncRNA)-mediated competing endogenous RNA networks provide novel potential biomarkers and therapeutic targets for colorectal cancer. Int J Mol Sci. 2019;20(22). doi:10.3390/ijms20225758
20. Bermudez M, Aguilar-Medina M, Lizarraga-Verdugo E, et al. LncRNAs as regulators of autophagy and drug resistance in colorectal cancer. Front Oncol. 2019;9:1008. doi:10.3389/fonc.2019.01008
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.