Back to Journals » Cancer Management and Research » Volume 12
LncRNA PSMA3-AS1 Promotes Lung Cancer Growth and Invasion via Sponging MiR-4504
Received 12 March 2020
Accepted for publication 28 May 2020
Published 1 July 2020 Volume 2020:12 Pages 5277—5283
DOI https://doi.org/10.2147/CMAR.S253575
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Eileen O'Reilly
Fangfang Li,1 LianLing Yu,1 Jun Zhu2
1Department of Respiration Medicine, Qingdao Eighth People’s Hospital, Qingdao 266000, People’s Republic of China; 2Department of Inspection, The 5th People’s Hospital of Jinan, Jinan 250022, People’s Republic of China
Correspondence: Jun Zhu Email [email protected]
Background: Long noncoding RNAs (lncRNAs) have close correlation with tumorigenesis. And how lncRNAs participate in lung cancer require investigation in-depth. The aim of this study was to determine the role of lncRNA PSMA3-AS1 in lung cancer progression.
Methods: PSMA3-AS1 expression was analyzed via qRT-PCR. Kaplan–Meier method was used to analyze survival rate based on PSMA3-AS1 value. Proliferation was measured via CCK8 and colony formation assays. Transwell assay was utilized to examine migration and invasion. Luciferase reporter assay and RNA pulldown assay were utilized to analyze the interaction between PSMA3-AS1 and miR-4504.
Results: PSMA3-AS1 expression was upregulated in lung cancer tissues and cell lines. PSMA3-AS1 expression was positively correlated with clinical stage and metastasis. PSMA3-AS1 overexpression predicted a poor prognosis in lung cancer patients. PSMA3-AS1 knockdown suppressed proliferation, migration and invasion of lung cancer cells. Through bioinformatics analysis, PSMA3-AS1 was predicted to sponge miR-4504. MiR-4504 expression was inhibited by PSMA3-AS1. And inhibition of miR-4504 reversed the effects of PSMA3-AS1 depletion.
Conclusion: PSMA3-AS1 promotes the tumorigenesis of lung cancer through inhibiting miR-4504.
Keywords: PSMA3-AS1, miR-4504, lung cancer, progression
Introduction
Lung cancer is one of the most common malignancies worldwide.1 In spite of advances on treatment, the overall survival rate of lung cancer patients remains lower than 20%.2 High rates of metastasis and recurrence and drug resistance markedly restrict the curative effects of lung cancer.3 As a serious public health problem, it is urgently required to determine the pathogenesis of lung cancer and develop novel therapeutic targets.
Long noncoding RNA (lncRNA) refers to a type of noncoding RNAs with over 200 nucleotides in length and without protein-coding potential.4 Initially, lncRNA is thought to have no biological roles. Gradually, lncRNA has been reported to participate in various biological processes through transcriptional or post-transcriptional regulation and modulating proteins.5,6 Especially, lncRNAs have been demonstrated to regulate tumorigenesis via modulating proliferation, migration, differentiation and survival.7 For example, lncRNA LINC00963 enhances growth and metastasis of ovarian cancer via modulating miR-378-CHI3L1 pathway.8 LncRNA WT1-AS is downregulated in papillary thyroid cancer and inhibits tumor progression via miR-203.9 LncRNA LINC01783 contributes to proliferation, migration and invasion of cervical cancer through inhibiting miR-199b-5p and promoting GBP1 expression.10 Additionally, lncRNA SNHG11 increases growth and invasiveness of liver cancer cells by affecting miR-184/AGO2 axis.11 Therefore, illustrating the correlation between lncRNA and tumorigenesis will benefit for cancer intervention.
PSMA3-AS1 is poorly investigated. Initially, PSMA3-AS1 was reported to be involved in multiple myeloma.12 Recently, PSMA3-AS1 was demonstrated to promote esophageal cancer development.13 However, whether PSMA3-AS1 regulates lung cancer progression is unclear. Herein, we found that PSMA3-AS1 was upregulated in lung cancer tissues. PSMA3-AS1 expression was correlated with tumor advance and metastasis. Moreover, PSMA3-AS1 high expression predicted a low survival rate. PSMA3-AS1 knockdown suppressed the proliferation, migration and invasion of lung cancer cells. We further found that PSMA3-AS1 was a competing endogenous RNA (ceRNA) for miR-4504. PSMA3-AS1 inhibits miR-4504 via direct interaction. And miR-4504 suppression reversed the effects of PSMA3-AS1 silencing. In conclusion, our findings suggest an important role of PSMA3-AS1/miR-4504 axis in regulating lung cancer progression.
Materials and Methods
Lung Cancer Tissues
Forty-one lung cancer tissues and adjacent normal tissues were collected from The 5th People’s Hospital of Jinan. The patients were not treated with radiotherapy or chemotherapy prior to surgery. Tissues were frozen immediately after collection and stored in liquid nitrogen. This study was approved by the Ethics Committee of the 5th People’s Hospital of Jinan. And written informed consent was achieved from every patient. All experiments were conducted in accordance with the Declaration of Helsinki.
Cell Lines
All lung cancer cell lines and normal lung epithelial cells BEAS-2B were obtained from the Cell Bank of Type Culture Collection of the Chinese Academy of Sciences. Cells were cultured with DMEM medium (HyClone) supplemented with 10% FBS (Gibco), penicillin (100 U/mL), and streptomycin (100 mg/mL) at 37°C, 5% CO2.
Cell Transfection
The small interfering (si) RNAs targeting PSMA3-AS1 (5ʹ-CCAGCAUCAAGAUGAUUUA-3ʹ), miR-4504 mimics, miR-4504 inhibitors and negative controls were were purchased from Shanghai GenePharma Co., Ltd. Cell transfection was conducted using RNAiMAX Lipofectamine® (Invitrogen; Thermo Fisher Scientific, Inc.) according to the manufacturer’s protocol.
RT-qPCR
RNA isolation was performed using Trizol reagent and used for cDNA synthesis using High-Capacity cDNA Reverse Transcription Kits (Bio-Rad, Hercules, CA, USA) or Reverse Transcription Kit (Thermo Fisher Scientific, Waltham, MA, USA). qPCR was performed with SYBR Green Mixture (Takara, Dalian, China). Relative expression was normalized to GAPDH or U6. Primer sequences were as follows: PSMA3-AS1 Forward, 5ʹ-AACAGACCATCAGAAGAGAACA-3ʹ and reverse, 5ʹ-GAACAGAAACCAGAGCCATACA-3ʹ; GAPDH Forward, 5ʹ-AAGGTGAAGGTCGGAGTCAA-3ʹ and reverse, 5ʹ-GGAAGATGGTGATGGGATTT-3ʹ.
Cell Proliferation Assay
CCK8 assay was used for detecting proliferation. 2×103 cells were seeded into the 96-well plates and incubated for 24h, 48h, 72h or 96h. Then 10 μL CCK8 (Dojindo Molecular Technologies, Inc) was added and cultured for 2h. The OD450 values were determined using a microplate reader (BD Biosciences).
Colony Formation
500 cells were seeded into the 6-well plates and cultured for 2 weeks. Then cells were fixed and stained with crystal violet solution. Finally, colony number was counted.
Transwell Assay
Cell migration and invasion was determined via Transwell assay. In brief, cells were seeded into the upper chamber with serum-free medium. The down chamber was filled with 500 μL complete medium. After incubation for 24 h, the migrated or invaded cells were fixed with polyoxymethylene and stained with 0.1% crystal violet. Then, migrated or invaded cells were counted using a microscope. For invasion assay, a matrigel invasion chamber (BD Biosciences, San Jose, CA, USA) was used. Other steps were the same as migration assay.
Dual-Luciferase Reporter Assay
miRDB (http://mirdb.org/miRDB/index.html) was used to identify the potential target of PSMA3-AS1. PSMA3-AS1 sequence containing the wide-type or mutant binding site of miR-4504 was inserted into the pGL3 luciferase reporter vector (Promega, Madison, WI, USA). Then, the reporter vector and miR-4504 mimics or negative control were transfected into lung cancer cells using RNAiMAX Lipofectamine®. After 48 h incubation, the firefly and Renilla luciferase activities were determined via a Dual-Luciferase Reporter Assay System (Promega Corporation).
RNA Pulldown
H1299 and A549 cells were transfected with biotinylated miR-4504 (Bio-miR-4504 WT or Bio-miR-4504 MUT) or the control (Bio-miR-NC). After 48 hours, cells were lysed and cell lysates were incubated with Dynabeads M-280 Streptavidin (Thermo Fisher Scientific) for two hours. The enrichment of PSMA3-AS1 was determined by qRT-PCR.
Statistical Analysis
All experiments were conducted at least three times and results were presented as the mean ± standard deviation. Significant differences between two groups were analyzed using a Student’s t-test. Multiple group comparison was performed using one-way ANOVA followed by Tukey’s post hoc test. The Kaplan–Meier and the Log rank test were used for survival rate analysis. P<0.05 was considered to indicate a statistically significant difference. The statistical analysis was conducted using Graph Pad software (GraphPad Software 6.0, Inc.).
Results
PSMA3-AS1 Is Upregulated in Lung Cancer
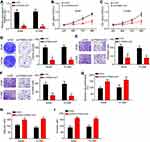
PSMA3-AS1 expression was examined by qRT-PCR in lung cancer and adjacent normal tissues. We found that PSMA3-AS1 level was upregulated in tumor tissues (Figure 1A). Notably, PSMA3-AS1 expression was increased in advanced lung cancer tissues and metastatic tissues (Figure 1B and C). Consistently, PSMA3-AS1 level was higher in lung cancer cell lines compared to BEAS-2B cells (Figure 1D). Then, 41 lung cancer tissues were divided into two groups according to PSMA3-AS1 expression value followed by survival rate analysis. Results showed that PSMA3-AS1 high expression indicated a low survival rate (Figure 1E).
PSMA3-AS1 Knockdown Suppresses Proliferation, Migration and Invasion
To analyze the potential function of PSMA3-AS1 in lung cancer, we chose A549 and H1299 cells to deplete PSMA3-AS1 (Figure 2A). Through CCK8 assay, we found that PSMA3-AS1 silencing suppressed the proliferation of A549 and H1299 cells (Figure 2B and C). Colony formation assay further validated the effect of PSMA3-AS1 on proliferation (Figure 2D). Then transwell assay was performed. Results showed that PSMA3-AS1 silencing led to decreased cell numbers of migration and invasion (Figure 2E and F). To further confirm the roles of PSMA3-AS1, we overexpressed it, followed by CCK8 and transwell assays. Results indicated that PSMA3-AS1 overexpression promoted the proliferation, migration and invasion of A549 and H1299 cells (Figure 2G–I).
PSMA3-AS1 Inhibits MiR-4504 in Lung Cancer
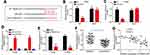
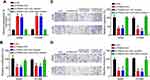
LncRNAs have been shown to sponge miRNAs in tumor.13 Thus, we performed bioinformatic analysis using miRDB and identified miR-4504 as the most potential candidate (Figure 3A) We constructed luciferase reporter vectors and found that miR-4504 mimics suppressed the luciferase activity of PSMA3-AS1-WT reporter (Figure 3B and C). Additionally, RNA pulldown assay showed that PSMA3-AS1 was precipitated by biotin-labeled miR-4504-WT (Figure 3D), suggesting that PSMA3-AS1 directly interacts with miR-4504. And PSMA3-AS1 overexpression led to decrease of miR-4504 expression (Figure 3E). Moreover, we found that miR-4504 was downregulated in lung cancer tissues (Figure 3F). And the expression of miR-4504 was reversely correlated with that of PSMA3-AS1 in lung cancer tissues (Figure 3G).
PSMA3-AS1 Promotes Lung Cancer Progression via Inhibiting MiR-4504
To determine whether PSMA3-AS1 exerts functions through miR-4504, we conducted rescue assays. We inhibited miR-4504 expression in PSMA3-AS1-depleted A549 and H1299 cells (Figure 4A). Then, CCK8 and Transwell assays were carried out. Results showed that PSMA3-AS1 silencing suppressed proliferation, migration and invasion of A549 and H1299 cells (Figure 4B–D). However, miR-4504 inhibitors reversed the effects of PSMA3-AS1 depletion on proliferation, migration and invasion (Figure 4B–D). Therefore, PSMA3-AS1 acts as an oncogenic lncRNA via inhibiting miR-4504.
Discussion
Lung cancer is a major risk factor for public health around the world. There is an urgent need to determine the molecular mechanism underlying lung cancer progression. In this study, we defined the oncogenic roles of PSMA3-AS1 in lung cancer. We found that PSMA3-AS1 was upregulated in lung cancer tissues and positively correlated with tumor severity. Additionally, PSMA3-AS1 is a potential prognostic biomarker for lung cancer patients. Moreover, we showed that PSMA3-AS1 knockdown suppressed the proliferation, migration and invasion of lung cancer cells through sponging miR-4504.
Increasing evidences have indicated that lncRNAs regulate multiple biological processes in lung cancer, such as growth and metastasis.14,15 And many lncRNAs are potential biomarkers for diagnosis and prognosis, such as NNT-AS1 and JPX.16,17 For example, lncRNA SNHG16 overexpression promotes proliferation, migration and invasion of lung cancer cells through regulating miR-520a-3p/EphA2 axis.18 LINC-PINT inhibits lung cancer cell growth, cell cycle, migration and invasion while promoting apoptosis.19 In addition, lncRNA MIAT upregulation leads to accelerated cell growth and predicts poor prognosis in lung cancer.14 There are still a lot of lncRNAs whose functions are unclear in lung cancer. PSMA3-AS1 is a rarely studied lncRNA. Only two references suggest its potential roles in multiple myeloma and esophageal cancer.12,13 In the current work, we found that PSMA3-AS1 expression was markedly upregulated in lung cancer tissues. Besides, we showed that PSMA3-AS1 dysregulation affects lung cancer progression.
Increasing studies have supported the ceRNA hypothesis that lncRNA can sponge miRNAs to form a lncRNA-miRNA regulatory network.20 LncRNAs could inhibit the activity of miRNAs to regulate gene expression. For instance, lncRNA MINCR sponges miR-223 to regulate irradiation resistance of nasopharyngeal cancer cells.21 SNHG12 targeted miR-195-5p to promote growth and metastasis in osteosarcoma.22 And lncRNA AC114812.8 initiates proliferation, migration and invasion of bladder tumor cells by modulating miR-371b-5p/FUT4 pathway.23 Besides, lncRNA AB074169 suppress thyroid cancer growth by affecting CDKN1a expression.24 Previously, PSMA3-AS1 was also reported to sponge miR-101 in esophageal cancer.13 In our study, we analyzed the possible PSMA3-AS1-related ceRNA pathway through bioinformatic method. We confirmed that PSMA3-AS1 bound to miR-4504 via luciferase reporter assay and RNA pulldown assay. We also found that miR-4504 expression was inhibited by PSMA3-AS1 overexpression in lung cancer. A study indicates that miR-4504 is involved in nacre color formation in Hyriopsis cumingii.25 Nevertheless, the role of miR-4504 in tumor is fully unknown. In our findings, we found that miR-4504 expression was downregulated in lung cancers. Moreover, miR-4504 level was negatively correlated with PSMA3-AS1 in lung cancer tissues, suggesting their regulatory relationship. Finally, we performed rescue assays. We found that effects of PSMA3-AS1 silencing on lung cancer cells were reversed by miR-4504 inhibitors, indicating miR-4504 is a tumor suppressor. However, the downstream target mRNA of miR-4504 requires further investigation. Recently, no study reveals the potential target of miR-4504. Thus, it is essential to determine the downstream mechanism of PSMA3-AS1/miR-4504 axis in the future.
In summary, this work indicates that PSMA3-AS1 contributes to proliferation, migration and invasion of lung cancer cells via sponging miR-4504. And PSMA3-AS1/miR-4504 axis may be a potential therapeutic target for lung cancer treatment. However, there is a limitation in our study. The downstream target of PSMA3-AS1/miR-4504 axis requires investigation in the future.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Youlden DR, Cramb SM, Baade PD. The international epidemiology of lung cancer: geographical distribution and secular trends. J Thorac Oncol. 2008;3(8):819–831. doi:10.1097/JTO.0b013e31818020eb
2. Reck M, Heigener DF, Mok T, Soria JC, Rabe KF. Management of non-small-cell lung cancer: recent developments. Lancet. 2013;382(9893):709–719. doi:10.1016/S0140-6736(13)61502-0
3. Gottesman MM, Lavi O, Hall MD, Gillet JP. Toward a better understanding of the complexity of cancer drug resistance. Annu Rev Pharmacol Toxicol. 2016;56:85–102. doi:10.1146/annurev-pharmtox-010715-103111
4. Chu C, Zhang YL, Yu L, Sharma S, Fei ZL, Drevet JR. Epididymal small non-coding RNA studies: progress over the past decade. Andrology. 2019;7(5):681–689. doi:10.1111/andr.12639
5. Liu B, Ye B, Yang L, et al. Long noncoding RNA lncKdm2b is required for ILC3 maintenance by initiation of Zfp292 expression. Nat Immunol. 2017;18(5):499–508. doi:10.1038/ni.3712
6. Shi X, Sun M, Liu H, Yao Y, Song Y. Long non-coding RNAs: a new frontier in the study of human diseases. Cancer Lett. 2013;339(2):159–166. doi:10.1016/j.canlet.2013.06.013
7. Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011;1(5):391–407. doi:10.1158/2159-8290.CD-11-0209
8. Liu W, Yang YJ, An Q. LINC00963 promotes ovarian cancer proliferation, migration and EMT via the miR-378g/CHI3L1 axis. Cancer Manag Res. 2020;12:463–473. doi:10.2147/CMAR.S229083
9. Le F, Luo P, Ouyang Q, Zhong X. LncRNA WT1-AS downregulates survivin by upregulating miR-203 in papillary thyroid carcinoma. Cancer Manag Res. 2020;12:443–449. doi:10.2147/CMAR.S232294
10. Chen WJ, Xiong L, Yang L, et al. Long non-coding RNA LINC01783 promotes the progression of cervical cancer by sponging miR-199b-5p to mediate GBP1 expression. Cancer Manag Res. 2020;12:363–373. doi:10.2147/CMAR.S230171
11. Huang W, Huang F, Lei Z, Luo H. LncRNA SNHG11 promotes proliferation, migration, apoptosis, and autophagy by regulating hsa-miR-184/AGO2 in HCC. Onco Targets Ther. 2020;13:413–421. doi:10.2147/OTT.S237161
12. Xu H, Han H, Song S, et al. Exosome-transmitted PSMA3 and PSMA3-AS1 promote proteasome inhibitor resistance in multiple myeloma. Clin Cancer Res. 2019;25(6):1923–1935. doi:10.1158/1078-0432.CCR-18-2363
13. Qiu BQ, Lin XH, Ye XD, et al. Long non-coding RNA PSMA3-AS1 promotes malignant phenotypes of esophageal cancer by modulating the miR-101/EZH2 axis as a ceRNA. Aging (Albany NY). 2020;12:1843–1856. doi:10.18632/aging.102716
14. Wu L, Liu C, Zhang Z. Knockdown of lncRNA MIAT inhibits proliferation and cisplatin resistance in non-small cell lung cancer cells by increasing miR-184 expression. Oncol Lett. 2020;19(1):533–541. doi:10.3892/ol.2019.11084
15. Zhao M, Xin XF, Zhang JY, Dai W, Lv TF, Song Y. LncRNA GMDS-AS1 inhibits lung adenocarcinoma development by regulating miR-96-5p/CYLD signaling. Cancer Med. 2020;9(3):1196–1208. doi:10.1002/cam4.2776
16. Shen Q, Jiang Y. LncRNA NNT-AS1 promotes the proliferation, and invasion of lung cancer cells via regulating miR-129-5p expression. Biomed Pharmacother. 2018;105:176–181. doi:10.1016/j.biopha.2018.05.123
17. Pan J, Fang S, Tian H, et al. lncRNA JPX/miR-33a-5p/Twist1 axis regulates tumorigenesis and metastasis of lung cancer by activating Wnt/beta-catenin signaling. Mol Cancer. 2020;19(1):9. doi:10.1186/s12943-020-1133-9
18. Yu L, Chen D, Song J. LncRNA SNHG16 promotes non-small cell lung cancer development through regulating EphA2 expression by sponging miR-520a-3p. Thorac Cancer. 2020;11(3):603–611. doi:10.1111/1759-7714.13304
19. Wang S, Jiang W, Zhang X, et al. LINC-PINT alleviates lung cancer progression via sponging miR-543 and inducing PTEN. Cancer Med. 2020;9(6):1999–2009.
20. Anastasiadou E, Jacob LS, Slack FJ. Non-coding RNA networks in cancer. Nat Rev Cancer. 2018;18(1):5–18. doi:10.1038/nrc.2017.99
21. Zhong Q, Chen Y, Chen Z. LncRNA MINCR regulates irradiation resistance in nasopharyngeal carcinoma cells via the microRNA-223/ZEB1 axis. Cell Cycle. 2020;19(1):53–66. doi:10.1080/15384101.2019.1692176
22. Xu N, Xu J, Zuo Z, Liu Y, Yan F, Han C. Downregulation of lncRNA SNHG12 reversed IGF1R-induced osteosarcoma metastasis and proliferation by targeting miR-195-5p. Gene. 2020;726:144145. doi:10.1016/j.gene.2019.144145
23. Li W, Li Y, Ma W, Zhou J, Sun Z, Yan X. Long noncoding RNA AC114812.8 promotes the progression of bladder cancer through miR-371b-5p/FUT4 axis. Biomed Pharmacother. 2020;121:109605. doi:10.1016/j.biopha.2019.109605
24. Gou Q, Gao L, Nie X, et al. Long noncoding RNA AB074169 inhibits cell proliferation via modulation of KHSRP-mediated CDKN1a expression in papillary thyroid carcinoma. Cancer Res. 2018;78(15):4163–4174. doi:10.1158/0008-5472.CAN-17-3766
25. Chen X, Bai Z, Li J. The mantle exosome and microRNAs of hyriopsis cumingii involved in nacre color formation. Mar Biotechnol (N Y). 2019;21(5):634–642. doi:10.1007/s10126-019-09908-8
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.