Back to Journals » Cancer Management and Research » Volume 13
LncRNA HCG18 Promotes Clear Cell Renal Cell Carcinoma Progression by Targeting miR-152-3p to Upregulate RAB14
Authors Yang Y, Gong P, Yao D, Xue D, He X
Received 22 December 2020
Accepted for publication 31 January 2021
Published 11 March 2021 Volume 2021:13 Pages 2287—2294
DOI https://doi.org/10.2147/CMAR.S298649
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Ahmet Emre Eşkazan
Yu Yang,1 Pengfeng Gong,2 Dongwei Yao,3 Dong Xue,2 Xiaozhou He2
1Department of Hepatopancreatobiliary Surgery, The Third Affiliated Hospital of Soochow University, Changzhou, Jiangsu, People’s Republic of China; 2Department of Urology, The Third Affiliated Hospital of Soochow University, Changzhou, Jiangsu, People’s Republic of China; 3Department of Urology, The Second People’s Hospital of Lianyungang, Lianyungang, Jiangsu, People’s Republic of China
Correspondence: Xiaozhou He
Department of Urology, The Third Affiliated Hospital of Soochow University, No. 185, Juqian Road, Changzhou, 213000, People’s Republic of China
Tel +86-519-68873211
Email [email protected]
Background: Long noncoding RNAs (lncRNAs) have been regarded as crucial regulators in many cancers, including clear cell renal cell carcinoma (ccRCC). This research aimed to explore the biological role and molecular mechanism of lncRNA HCG18 in ccRCC.
Materials and Methods: The expression levels of HCG18, miR-152-3p and RAB14 were examined by RT-qPCR. Cell viability, migration and invasion were examined by CCK-8 and transwell assays. Luciferase reporter and RIP assays were adopted to verify the interaction between miR-152-3p and HCG18 or RAB14.
Results: It was found that HCG18 expression was highly expressed in ccRCC tissues and cells, and patients with high expression of HCG18 had a short overall survival time. Moreover, HCG18 depletion attenuated ccRCC cell viability, migration and invasion. In addition, miR-152-3p was confirmed as a downstream target of HCG18 and was inversely regulated by HCG18, and RAB14 was a target of miR-152-3p. Functional assays demonstrated that miR-152-3p silencing or RAB14 addition abolished the inhibitory effects of HCG18 knockdown on ccRCC progression.
Conclusion: The results of the present study indicated that HCG18 accelerated the development and progression of ccRCC by upregulating RAB14 via sponging miR-152-3p, suggesting a potential therapeutic target for patients with ccRCC.
Keywords: HCG18, miR-152-3p, RAB14, clear cell renal cell carcinoma
Introduction
Renal cell carcinoma (RCC) is one of the most common urological malignancies with high mortality worldwide. Clear cell renal cell carcinoma (ccRCC) is one of the main subtypes of RCC and occupies about 80% of RCC cases.1,2 Although early ccRCC can be cured, patients with metastatic ccRCC still have a poor prognosis.3 Nowadays, increasing molecular biomarkers have been identified as diagnostic, prognostic, and therapeutic biomarkers in human cancers, which can improve the early detection of cancers and decreased the mortality rate.4–6 Therefore, it is essential to discover novel biomarkers for diagnosis, treatment, and prognosis of ccRCC.
Long non-coding RNAs (lncRNAs) are a type of non-coding RNAs with >200 nts in length.7 LncRNAs have been found to exert essential functions in many types of cancers, including ccRCC. For example, lncRNA TUG1 facilitated the proliferation of ccRCC cells by targeting miR-31-5p and regulating FLOT1 expression.8 LncRNA DARS-AS1 promoted the tumorigenesis of ccRCC through the miR-194-5p/DARS axis.9 Recently, a cancer-related lncRNA HCG18 was identified to participate in several malignant cancers, such as bladder cancer, gastric cancer, and nasopharyngeal carcinoma.10−12 However, the regulatory role of HCG18 in ccRCC remains undetermined.
MicroRNAs (miRNAs) are non-coding RNAs with 17–25 nucleotides in length.13 Emerging evidence implied that dysregulation of miRNAs was implicated in the progression of multiple tumors. For instance, miR-448 restrained the growth of pituitary adenoma cells by regulating BLC2.14 miR-335-5p overexpression suppressed the metastasis of lung adenocarcinoma cells via downregulating CCNB2.15 miR-152-3p was identified to act as a tumor suppressor in several cancers, such as prostate cancer and glioma.16,17 Nevertheless, the biological significance of miR-152-3p in ccRCC is still obscure.
The current study explored the regulatory mechanism of HCG18 in ccRCC and found that HCG18 contributed to ccRCC tumorigenesis by regulating miR-152-3p/RAB14 axis. The novel regulatory pathway might provide a potential diagnosis and treatment strategy for ccRCC.
Materials and Methods
Clinical Samples
A total of 32 paired ccRCC tissues and adjacent normal tissues were collected from patients who underwent nephrectomy at the Third Affiliated Hospital of Soochow University, and serum was also sampled from each patient before the operation. Besides, serum samples were also collected from 32 healthy individuals during the same period as a healthy control group. All the samples were rapidly frozen in liquid nitrogen and stored at ‑80°C until use. Our study was approved by the Ethics Committee of the Third Affiliated Hospital of Soochow University and conducted in accordance with the Declaration of Helsinki. Written informed consent forms were signed by all participants.
Cell Culture
Human ccRCC cell lines (Caki-1, 786-O, 769-P and ACHN) and normal human kidney cells HK-2 were obtained from the American Type Culture Collection (ATCC) and incubated in Dulbecco’s Modified Eagle Medium (DMEM; Gibco, USA) containing 10% fetal bovine serum (FBS) at 37°C with 5% CO2.
Cell Transfection
Short hairpin (sh) RNA targeting HCG18 (shHCG18; 5ʹ-UGAGCGGCUGUGCUAUACAUGUG-3ʹ), RAB14 (shRAB14; 5ʹ-GCAGUUCACAAGUAGUACUGG-3ʹ) with their negative control (shNC), miR-152-3p mimics/inhibitor with their negative controls (NC mimics/inhibitor), and pcDNA3.1/RAB14 with its negative control (pcDNA3.1) were synthesized by GenePharma (Shanghai, China). Cell transfection was performed using Lipofectamine 2000 (Invitrogen).
RT-qPCR
Total RNA was extracted using Trizol reagent (Invitrogen, CA, USA). Then, 1 μg of total RNA was reverse transcribed to cDNA using the PrimeScript RT reagent kits (Takara). RT-qPCR was performed by SYBR Premix Ex Taq II (TaKaRa) on ABI 7500 real-time PCR system (Applied Biosystems). The data were analyzed using the 2−ΔΔCq method. GAPDH or U6 were used as the internal controls for HCG18 and RAB14 or miR-152-3p, respectively.
CCK-8 Assay
Transfected cells were seeded into 96-well plates. After cultured for 0, 24, 48, 72 h, 10 μL CCK-8 solution was added into each well, and the cells were incubated for 2 h. The absorbance at 450 nm was examined using a microplate reader (Bio-Rad, USA).
Transwell Assay
Cell migration and invasion were detected using transwell chambers (Corning, NY, USA). For cell invasion, the transwell chambers were coated with Matrigel (BD Biosciences). The transfected cells (1x105 cells) were suspended in DMEM without serum and added to the upper chambers. Then, the lower chamber was filled with 600μL DMEM plus 10% FBS. After 24 incubation, the invaded cells were stained with 0.5% crystal violet. Cell migration was measured by the method same as that for cell invasion, except that the transwell chambers were not coated with Matrigel.
Luciferase Reporter Assay
HCG18-(wild type) Wt, RAB14-Wt and their mutants (HCG18-Mut and RAB14-Mut) were constructed by GenePharma (Shanghai, China). Then, the above reporters were co-transfected miR-152-3p mimics or NC mimics into HEK-293T cells for 48 h. Luciferase activities were estimated by the dual-luciferase reporter assay system (Promega, USA).
RNA Immunoprecipitation (RIP)
RIP assay was conducted using Magna RIP-Kit (Millipore, Bedford, USA). Briefly, the treated Caki-1 and 786-O cells were lysed by RIP buffer that contained magnetic beads coupled with anti-Ago2 or anti-IgG antibody. Then, the enrichment of HCG18 and miR-152-3p was detected by RT-qPCR.
In vivo Experiments
The animal study was approved by the Animal Ethics Committee of the Third Affiliated Hospital of Soochow University and conducted following the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. Briefly, cells transfected with shNC or shHCG18 were injected into BALB/C nude mice. The volume and weight of tumors were measured every 7 days. Four weeks later, mice were sacrificed by cervical dislocation after deep anesthesia with 2% isoflurane. Then, the tumors were dissected and detected.
Statistical Analysis
Data were presented as mean ± SD. Statistical analysis was conducted using SPSS 22.0 (IBM, Chicago, IL, USA). Group difference was analyzed by Student’s t-test or one-way ANOVA. Overall survival rate was assessed by Kaplan–Meier method. The correlation between miR-152-3p and HCG18 or RAB14 was estimated by Pearson’s correlation analysis. p<0.05 was thought as statistically significant.
Results
HCG18 is Overexpressed in ccRCC and Knockdown of HCG18 Represses the Tumorigenesis of ccRCC
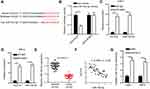
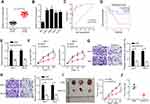
Firstly, the expression of HCG18 in ccRCC was evaluated by RT-qPCR and the results indicated that HCG18 was overexpressed in ccRCC tissues and cells (Figure 1A and B). In addition, we found that the AUC of HCG18 for diagnosing ccRCC was 0.922, indicating that it might be used as an indicator for ccRCC screening (Figure 1C). Moreover, ccRCC patients with a high level of HCG18 were associated with a low survival rate (Figure 1D). To investigate the biological function of HCG18 in ccRCC, shHCG18 was transfected into Caki-1 and 786-O cells, and the transfection efficiency was performed by RT-qPCR (Figure 1E). Moreover, CCK-8 assay showed that HCG18 depletion impaired the viability of ccRCC cells (Figure 1F). Furthermore, transwell assays revealed that HCG18 depletion repressed the migration and invasion of ccRCC cells (Figure 1G and H). In addition, an in vivo xenograft experiment showed that depletion of HCG18 reduced the tumor growth rate in mice (Figure 1I and J). Thus, these results implied that HCG18 accelerated the development of ccRCC.
HCG18 is a Molecular Sponge for miR-152-3p in ccRCC
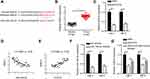
Through using StarBase website (http://starbase.sysu.edu.cn/), miR-152-3p was predicted as a potential target of HCG18 (Figure 2A). Luciferase reporter assay determined that miR-152-3p mimics remarkably decreased the luciferase activity of HCG18-Wt, but had no effect on HCG18-Mut in 293T cells (Figure 2B). Meanwhile, RIP assay manifested that miR-152-3p and HCG18 were enriched in the AGO2 group (Figure 2C and D). Then, RT-qPCR analysis revealed that miR-152-3p level was reduced in ccRCC tissues (Figure 2E). Moreover, an inverse correlation between HCG18 and miR-152-3p expression was observed in ccRCC tissues (Figure 2F). Besides, RT-qPCR results indicated that HCG18 depletion enhanced miR-152-3p expression in ccRCC cells (Figure 2G). The above data determined that miR-152-3p was a target of HCG18, and was inversely modulated by HCG18.
HCG18 Sponges miR-152-3p and Positively Modulates RAB14 Expression in ccRCC
Subsequently, starBase website predicted that RAB14 was a potential target of miR-152-3p (Figure 3A). RT-qPCR analysis exhibited that RAB14 level was increased in ccRCC tissues (Figure 3B), and RAB14 expression was decreased by silencing of HCG18 (Figure 3C). Then, we found that RAB14 expression was inversely correlated with miR-152-3p expression, and RAB14 expression was positively correlated with HCG18 expression in ccRCC tissues (Figure 3D and E). To elucidate whether HCG18 modulated RAB14 expression by targeting miR-152-3p, Caki-1 and 786-O cells were transfected with shNC, shHCG18, and shHCG18+miR-152-3p inhibitor. RT-qPCR analysis demonstrated that miR-152-3p was lowly expressed in ccRCC cells transfected with miR-152-3p inhibitor (Figure 3F). Additionally, depletion of HCG18 reduced RAB14 expression, while this effect was reversed by miR-152-3p inhibition (Figure 3G). In sum, the results elucidated that HCG18 sponged miR-152-3p and positively modulated RAB14 expression in ccRCC.
HCG18 Contributes to ccRCC Progression by Modulating the miR-152-3p/RAB14 Axis
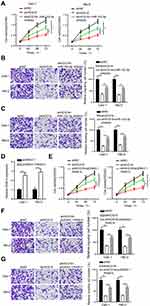
To investigate whether HCG18 participated in ccRCC progression via modulating the miR-152-3p/RAB14 axis, a string of functional experiments was conducted. Firstly, CCK-8 and transwell assays disclosed that the inhibition of miR-152-3p reversed the repressive effect of HCG18 interference on the viability, migration and invasion of Caki-1 and 786-O cells (Figure 4A–C). Next, pcDNA3.1/RAB14 and shHCG18 were co-transfected into Caki-1 and 786-O cells. RT-qPCR results confirmed that addition of RAB14 significantly upregulated RAB14 expression in ccRCC cells (Figure 4D). Functional assays elaborated that the impacts of HCG18 knockdown on cell viability, and metastasis were eliminated by RAB14 overexpression (Figure 4E–G). Taken together, HCG18 contributed to ccRCC progression via sponging miR-152-3p and upregulating RAB14.
Discussion
LncRNAs have been reported to serve as oncogenes or tumor suppressors in the tumorigenesis of ccRCC. For example, lncRNA-LET restrained the growth of ccRCC cells by modulating miR-373-3p.18 LncRNA LUCAT1 facilitated ccRCC cell viability and invasion by the AKT/GSK-3β pathway.19 LncRNA DLEU1 accelerated ccRCC cell viability and migration by sponging miRNA-194-5p.20 This research focused on the role and regulatory mechanism of HCG18 in ccRCC, and the results elucidated that HCG18 accelerated ccRCC progression through sponging miR-152-3p and upregulating RAB14.
Numerous studies have revealed that HCG18 acted as an oncogene in several tumors. For example, HCG18 expedited the progression of colorectal cancer via targeting miR-1271 and regulating MTDH/Wnt/β-catenin axis.21 HCG18 facilitated lung adenocarcinoma cells growth in vitro and promoted tumor growth in vivo via the miR-34a-5p/HMMR axis.22 Herein, we manifested that HCG18 expression was elevated in ccRCC tissues and cells. ccRCC patients with high HCG18 expression had short overall survival time. Moreover, interference of HCG18 markedly attenuated ccRCC cell viability and metastasis and suppressed tumor growth in vivo.
It is widely reported that lncRNA can serve as a ceRNA to sponge target miRNAs.23,24 MiR-152-3p was discovered to be downregulated in various cancers. For instance, miR-152 attenuated cell viability and induced apoptosis in breast cancer through regulating PIK3CA.25 miR-152-3p inhibited hepatocellular carcinoma progression by targeting CDK8.26 LINC00174 expedited carcinogenesis of glioma via modulating miR-152-3p/SLC2A1 axis.27 At present study, we confirmed that miR-152-3p expression was declined and inversely correlated with HCG18 in ccRCC tissues. Moreover, functional assays demonstrated that miR-152-3p inhibition rescued the suppressive effect of HCG18 interference on ccRCC cell viability, migration and invasion. Therefore, our data confirmed that HCG18 accelerated ccRCC progression via sponging miR-152-3p.
Additionally, miRNAs have been reported to modulate gene expressions by binding to the 3′-UTR of mRNAs.28 Previous studies have implied that RAB14 participated in the progression of human cancers, such as cervical cancer, gastric cancer and colorectal cancer.29–31 In this study, we identified that RAB14 was a downstream target of miR-152-3p, and RAB14 expression was enhanced in ccRCC tissues. Moreover, RAB14 was inversely correlated with miR-31-5p and positively correlated with HCG18 expression. Besides, RAB14 addition partially abolished the suppressive effect of HCG18 knockdown on cell viability and metastasis. Thus, above results determined that HCG18 promoted ccRCC progression through sponging miR-152-3p/RAB14 axis.
Conclusion
The present investigated the ceRNA regulatory network of HCG18/miR-152-3p/RAB14 in ccRCC for the first time, which might provide a new treatment target for ccRCC patients.
Funding
This work was supported by Young Talent Development Plan of Changzhou Health Commission (CZQM2020049).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J. Clin. 2011;61:69–90. doi:10.3322/caac.20107
2. Cheville JC, Lohse CM, Zincke H, Weaver AL, Blute ML. Comparisons of outcome and prognostic features among histologic subtypes of renal cell carcinoma. Am J Surg Pathol. 2003;27:612–624. doi:10.1097/00000478-200305000-00005
3. Motzer RJ, Molina AM. Targeting renal cell carcinoma. J Clin Oncol. 2009;27:3274–3276. doi:10.1200/JCO.2009.21.8461
4. Xie L, Li H, Zhang L, et al. Autophagy-related gene P4HB: a novel diagnosis and prognosis marker for kidney renal clear cell carcinoma. Aging. 2020;12:1828–1842. doi:10.18632/aging.102715
5. Cheng G, Li M, Ma X, et al. Systematic analysis of microRNA biomarkers for diagnosis, prognosis, and therapy in patients with clear cell renal cell carcinoma. Front Oncol. 2020;10:543817. doi:10.3389/fonc.2020.543817
6. Xie L, Dang Y, Guo J, et al. High KRT8 expression independently predicts poor prognosis for lung adenocarcinoma patients. Genes. 2019;10.
7. Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 2009;23:1494–1504. doi:10.1101/gad.1800909
8. Lv D, Xiang Y, Yang Q, Yao J, Dong Q. Long Non-Coding RNA TUG1 promotes cell proliferation and inhibits cell apoptosis, autophagy in clear cell renal cell carcinoma via MiR-31-5p/FLOT1 axis. Onco Targets Ther. 2020;13:5857–5868. doi:10.2147/OTT.S254634
9. Jiao M, Guo H, Chen Y, Li L, Zhang L. DARS-AS1 promotes clear cell renal cell carcinoma by sequestering miR-194-5p to up-regulate DARS. Biomed. Pharmacother. 2020;128:110323. doi:10.1016/j.biopha.2020.110323
10. Ma F, An K, Li Y. Silencing of long non-coding RNA-HCG18 Inhibits the tumorigenesis of gastric cancer through blocking PI3K/Akt pathway. Onco Targets Ther. 2020;13:2225–2234. doi:10.2147/OTT.S240965
11. Xu Z, Huang B, Zhang Q, He X, Wei H, Zhang D. NOTCH1 regulates the proliferation and migration of bladder cancer cells by cooperating with long non-coding RNA HCG18 and microRNA-34c-5p. J Cell Biochem. 2019;120:6596–6604. doi:10.1002/jcb.27954
12. Li L, Ma TT, Ma YH, Jiang YF. LncRNA HCG18 contributes to nasopharyngeal carcinoma development by modulating miR-140/CCND1 and Hedgehog signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2019;23:10387–10399. doi:10.26355/eurrev_201912_19678
13. Kloosterman WP, Plasterk RH. The diverse functions of microRNAs in animal development and disease. Dev Cell. 2006;11:441–450. doi:10.1016/j.devcel.2006.09.009
14. He C, Yang J, Ding J, et al. MiR-448 targets BLC2 and inhibits the growth of pituitary adenoma cells. Biochem Cell Biol. 2020;1–7.
15. Wang X, Xiao H, Wu D, Zhang D, Zhang Z. miR-335-5p regulates cell cycle and metastasis in lung adenocarcinoma by targeting CCNB2. Onco Targets Ther. 2020;13:6255–6263. doi:10.2147/OTT.S245136
16. Feng F, Liu H, Chen A, et al. miR-148-3p and miR-152-3p synergistically regulate prostate cancer progression via repressing KLF4. J Cell Biochem. 2019;120:17228–17239. doi:10.1002/jcb.28984
17. Sun J, Tian X, Zhang J, et al. Regulation of human glioma cell apoptosis and invasion by miR-152-3p through targeting DNMT1 and regulating NF2: miR-152-3p regulate glioma cell apoptosis and invasion. J Exp Clin Cancer Res. 2017;36:100. doi:10.1186/s13046-017-0567-4
18. Ye Z, Duan J, Wang L, Ji Y, Qiao B. LncRNA-LET inhibits cell growth of clear cell renal cell carcinoma by regulating miR-373-3p. Cancer Cell Int. 2019;19:311. doi:10.1186/s12935-019-1008-6
19. Zheng Z, Zhao F, Zhu D, et al. Long non-coding RNA LUCAT1 promotes proliferation and invasion in clear cell renal cell carcinoma through AKT/GSK-3beta signaling pathway. Cell Physiol Biochem. 2018;48:891–904. doi:10.1159/000491957
20. He GZ, Yu SY, Zhou QP, et al. LncRNA DLEU1 accelerates the malignant progression of clear cell renal cell carcinoma via regulating miRNA-194-5p. Eur. Rev. Med. Pharmacol. Sci. 2019;23:10691–10698. doi:10.26355/eurrev_201912_19768
21. Li S, Wu T, Zhang D, Sun X, Zhang X. The long non-coding RNA HCG18 promotes the growth and invasion of colorectal cancer cells through sponging miR-1271 and upregulating MTDH/Wnt/beta-catenin. Clin Exp Pharmacol Physiol. 2020;47:703–712. doi:10.1111/1440-1681.13230
22. Li W, Pan T, Jiang W, Zhao H. HCG18/miR-34a-5p/HMMR axis accelerates the progression of lung adenocarcinoma. Biomed. Pharmacother. 2020;129:110217. doi:10.1016/j.biopha.2020.110217
23. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146:353–358. doi:10.1016/j.cell.2011.07.014
24. Abdollahzadeh R, Daraei A, Mansoori Y, Sepahvand M, Amoli MM, Tavakkoly-Bazzaz J. Competing endogenous RNA (ceRNA) cross talk and language in ceRNA regulatory networks: a new look at hallmarks of breast cancer. J Cell Physiol. 2019;234:10080–10100. doi:10.1002/jcp.27941
25. Ge S, Wang D, Kong Q, Gao W, Sun J. Function of miR-152 as a tumor suppressor in human breast cancer by targeting PIK3CA. Oncol Res. 2017;25:1363–1371. doi:10.3727/096504017X14878536973557
26. Yin T, Liu MM, Jin RT, Kong J, Wang SH, Sun WB. miR-152-3p Modulates hepatic carcinogenesis by targeting cyclin-dependent kinase 8. Pathol Res Pract. 2019;215:152406. doi:10.1016/j.prp.2019.03.034
27. Shi J, Zhang Y, Qin B, Wang Y, Zhu X. Long non-coding RNA LINC00174 promotes glycolysis and tumor progression by regulating miR-152-3p/SLC2A1 axis in glioma. J Exp Clin Cancer Res. 2019;38:395. doi:10.1186/s13046-019-1390-x
28. Bernardo BC, Ooi JY, Lin RC, McMullen JR. miRNA therapeutics: a new class of drugs with potential therapeutic applications in the heart. Future Med. Chem. 2015;7:1771–1792. doi:10.4155/fmc.15.107
29. Guo B, Wang W, Zhao Z, et al. Rab14 act as oncogene and induce proliferation of gastric cancer cells via AKT signaling pathway. PLoS One. 2017;12:e0170620. doi:10.1371/journal.pone.0170620
30. Yang J, Liang B, Hou S. TMPO-AS1 promotes cervical cancer progression by upregulating RAB14 via sponging miR-577. J. Gene Med. 2019;21:e3125. doi:10.1002/jgm.3125
31. Li M, Bian Z, Jin G, et al. LncRNA-SNHG15 enhances cell proliferation in colorectal cancer by inhibiting miR-338-3p. Cancer Med. 2019;8:2404–2413. doi:10.1002/cam4.2105
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.