Back to Journals » Cancer Management and Research » Volume 13
LINC01224 Promotes Colorectal Cancer Progression by Sponging miR-2467
Authors Chen L, Chen W, Zhao C, Jiang Q
Received 12 September 2020
Accepted for publication 31 December 2020
Published 26 January 2021 Volume 2021:13 Pages 733—742
DOI https://doi.org/10.2147/CMAR.S281625
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Antonella D'Anneo
Lin Chen,1 Wei Chen,1 Changjie Zhao,2 Qi Jiang1
1Department of Gastroenterology, Affiliated Dongtai Hospital of Nantong University, Dongtai, Jiangsu 224200, People’s Republic of China; 2Endoscopy Center, Affiliated Dongtai Hospital of Nantong University, Dongtai, Jiangsu 224200, People’s Republic of China
Correspondence: Qi Jiang
Department of Gastroenterology, Affiliated Dongtai Hospital of Nantong University, 2 Kangfuxi Road, Dongtai, Jiangsu 224200, People’s Republic of China
Email [email protected]
Introduction: Colorectal cancer (CRC) is one of the most common human cancers and a leading cause of cancer-related death. Accumulating evidence has confirmed that long non-coding RNA (lncRNA) plays crucial roles in CRC development.
Methods: qRT-PCR was performed to examine the expressions of LINC01224 and miR-2467. CCK-8 assay, colony formation assay and transwell invasion assay were used to examine the progression of breast cancer cells. Luciferase and RNA-binding protein immunoprecipitation (RIP) assay were applied to verify the binding site. Correlation analysis of miR-2467 and LINC01224 expression in lung cancer tissues was shown. Pancreatic cancer cells growth in vivo was evaluated using xenograft tumor assay.
Results: LINC01224 expression was observed to be up-regulated in CRC tissues and cell lines. Functional studies suggested that LINC01224 silence inhibited CRC cells proliferation and invasion of CRC cells, while co-transfection with a miR-2467 inhibitor reversed these biological effects. Luciferase reporter assays illustrated that LINC01224 regulated miR-2467 directly, and RNA-binding protein immunoprecipitation (RIP) further confirmed that the suppression of LINC01224 by miR-2467 was in an RISC-dependent manner. Finally, LINC01224 silence inhibited the growth CRC cells in vivo.
Conclusion: In conclusion, our findings showed that LINC01224 promoted CRC progression through sponging miR-2467. LINC01224 may serve as a potential diagnostic biomarker and therapeutic target for CRC patients.
Keywords: LINC01224, miR-2467, colorectal cancer, proliferation, invasion
Introduction
CRC (colorectal cancer) has been recognized as one of the most common human cancers and the most dominating causes of cancer-related deaths worldwide.1,2 Although great progression has been made in CRC diagnosis and treatment in the past decades, the prognosis of CRC patients remains rather dismal due to metastasis and recurrence.3 Therefore, further exploring the potential mechanism underlying CRC development is urgently needed to identify novel diagnosis and therapy biological targets for CRC.
LncRNA (long noncoding RNA) is defined as transcripts with more than 200 nucleotides in length.4,5 Emerging studies have suggested that lncRNAs play important roles in the initiation and progression of human cancers.6,7 Numerous studies show that lncRNA is involved in CRC development.8,9 LINC01224 was a novel identified lncRNA. It was reported that LINC01224 promoted hepatocellular carcinoma progression through microRNA-330-5p-induced inhibition of CHEK1.10 Moreover, Xing et al reported that LINC01224 exhibited cancer-promoting activity in EOC (epithelial ovarian cancer) through microRNA-485-5p-mediated PAK4 upregulation.11 However, the biological function and underlying mechanism of LINC01224 in the development of human cancer, including CRC are still elusive.
In this study, we determined the expression of LINC01224 in CRC tissues and in paired adjacent normal colorectal tissues, and we further investigated the biological function of LINC01224 on CRC in vitro and in vivo. We also examined the interaction between LINC01224 and miR-2467 to reveal the potential mechanism of LINC01224 in CRC. To the best of our knowledge, our study is the first to show that LINC01224 functions as an oncogene in the development of CRC.
Materials and Methods
Clinical Samples
A total of 24 pairs (age, 40–74 years; male patients, 13; female patients, 11) of CRC tissues and matched adjacent normal tissues were sourced from patients undergoing resection surgery at the Affiliated Dongtai Hospital of Nantong University from Jan 2012 to September 2015. All patients did not receive chemotherapy or radiation before collecting specimens. All these participants signed informed consents prior to the samples collection. The samples from resection surgery were rapidly frozen and stored in liquid nitrogen until required. This study was approved by the Ethics and Research Committees of the Affiliated Dongtai Hospital of Nantong University. The research was conducted according to the principles of the World Medical Association Declaration of Helsinki.
Cell Culture
The normal colon cell line (FHC) and human CRC cell lines (LoVo, HT29, HCT116 and SW480) were bought from the Institute of Biochemistry and Cell Biology of the Chinese Academy of Sciences (Shanghai, China). The FHC cells were maintained in Ham’s F12 medium (45%); Dulbecco’s modified Eagle’s medium (45%); 25 mM HEPES; 10 ng/mL cholera toxin; 0.005 mg/mL transferrin; 0.005 mg/mL insulin; 100 ng/Ml hydrocortisone; 20 ng/mL human recombinant EGF (Thermo Fisher PHG0311); fetal bovine serum (FBS, 10%). LoVo cells were cultured in F-12K medium (Invitrogen; Thermo Fisher Scientific, Inc.). HT29, HCT116, and SW480 cells were cultured in RPMI-1640 (Gibco; Thermo Fisher Scientific, Inc.) supplemented with 10% FBS. All cells were maintained at 37°C with 5% CO2 in a humidified incubator.
Constructs, Synthesized Oligos and Transfection
siRNAs and shRNA (short hairpin RNA) targeting LINC01224 (si-LINC01224, sh-LINC01224), miR-2467 mimics, inhibitors and their corresponding negative control were bought from by GeneChem (Shanghai, China). All the DNAs were inserted into pcDNA3.1. Finally, Lipofectamine 3000 (Thermo Fisher Scientific) was utilized to transfer the oligonucleotides and constructs into the SW480 and HCT116 cells according to the manufacturer’s protocol. 36–48 hours after transfection, the cells were ready for the following experiments.
RNA Extraction and Quantitative Real-Time Polymerase Chain Reaction
Total RNA from CRC tissues and cells was extracted using Trizol reagent (Invitrogen; Thermo Fisher Scientific, Ind.) according to the manufacturer’s protocol. 2 µg RNA was reverse transcribed into cDNA using the PrimeScript RT Reagent kit (Invitrogen; Thermo Fisher Scientific, Ind.). qRT-PCR was undertaken using SYBR Green Master mix (Invitrogen; Thermo Fisher Scientific, Ind.) on ABI PRISM 7500 PCR System (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s protocol. GAPDH or U6 was used as controls and normalized the expression of mRNA and miRNA, respectively. Primer sequences are provided in Table 1.
 |
Table 1 Primer List |
Cell Proliferation Assay
Cell proliferation was examined by Cell Counting Kit-8 (CCK-8) and colony formation assays. For CCK8 assay, cells were seeded in 96-well plates at the concentration of 3000 cells/well. Then, a 10 μL of Cell Counting Kit-8 (CCK-8, Dojindo, Japan) was added after 24, 48, 72, and 96 h of incubation, respectively. After 2 h, the plates were washed using PBS (phosphate-buffered saline) and the absorbance was measured at 450 nm through a microplate reader (ELx800; BioTek Instruments, Inc, Winooski, VT, USA). For colony formation assay, 1000 cells were seeded into a 6-well plate and continuously incubated for 14 days. The colonies were fixed with 4% paraformaldehyde for 10 min and stained with 1% crystal violet for 15 min. Finally, the colonies were counted and photographed.
Transwell Invasion Assay
Transwell invasion assays were performed to determine the cell invasion potential using transwell plates (Corning, NY) that were coated with 50 µL of Matrigel (BD Biosciences, San Jose, CA, USA). Briefly, 1ⅹ105 cells were suspended in 300 μL serum-free medium and added to the upper chamber, while 800 μL complete medium was placed in the lower chamber. After 24 h incubation, cells on the upper surface of the membrane were scraped off. Cells on the lower side of the chamber were fixed with methanol and stained with 1% crystal violet. The invaded cells were counted in at least five fields under a light microscope (magnification, x200, Olympus Corp).
Luciferase Reporter Assay
Mut (mutant-type) or wt (wild-type) fragments of LINC01224 containing miR-2467 targeting site were synthesized and cloned into a dual-luciferase reporter vector (pmirGLO, GenePharma, Shanghai, China). Similarly, luciferase vectors and miR-2467 mimics or miR–2467 NC together with Renilla plasmid were cotransfected into HCT116 cells by Lipofectamine 3000. Forty-eight hours after transfection, dual-luciferase assay (Promega, Madison, WI) was adopted to examine the Renilla and firefly luciferases activity following the manufacturer’s protocol, and normalized to that of Renilla luciferase activity.
RNA-Binding Protein Immunoprecipitation Assay
Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Bedford, MA, USA) was used for RIP (RNA-binding protein immunoprecipitation) assay. Cells were harvested and lysed, and lysis buffer containing magnetic beads was incubated with human anti-Ago2 antibody (Abcam, Cambridge, MA, USA) to conjugate the antibody to the magnetic beads. Then, proteinase K was added to digest the protein and the immunoprecipitated RNAs were isolated using Trizol reagent and measured using RT-qPCR.
Tumor Xenograft Experiment
HCT116 cells (2 × 106) stably transfected with lv-sh- LINC01224 or lv-sh-NC were subcutaneously injected subcutaneously into the left flank of 6-week-old female nude mice (n = 5 mice per group). The tumor sizes were measured every week. After 4 weeks, the mice were euthanized, the tumor tissues were excised and weighted, and qRT-PCR was performed to determine LINC01224 and miR-2467 expression. This study was approved by the Ethics and Research Committees of the Affiliated Dongtai Hospital of Nantong University prior to the commencement of the study. This study was carried out in accordance with the National Institutes of Health guide for the care and use of laboratory animals (NIH Publications No.8023, revised 1978).
Statistical Analysis
All results are presented as mean ± SD from at least three independent experiments. One-way ANOVA or two-tailed Student’s t-test was performed for comparisons between groups. Pearson’s coefficient correlation was used to conduct expression correlation assays. A value of P < 0.05 was considered to be statistically significant.
Results
LINC01224 Was Up-Regulated and miR-2467 Was Down-Regulated in CRC Tissues
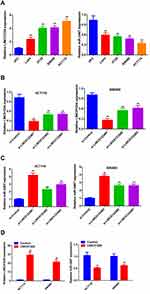
qRT-PCR analysis was conducted to determine the relative expression of LINC01224 and miR-2467 in 24 pairs of CRC and adjacent non-tumor tissues. The results showed that LINC01224 expression was significantly up-regulated (Figure 1A), and the miR-2467 level was dramatically down-regulated (Figure 1B) in CRC tissues compared with that in the corresponding normal tissues. Furthermore, we found that there was a significant negative correlation between LINC01224 expression and miR-2467 level in CRC tissues (Figure 1C). These data suggested that LINC01224 and miR-2467 may be associated with CRC development.
LINC01224 Negatively Regulated miR-2467 Expression
The result of qRT-PCR analysis showed that LINC01224 expression was up-regulated in CRC cells compared with normal cells (HFC) while the expression of miR-2467 was down-regulated in CRC cells compared with normal cells (HFC) (Figure 2A). As shown in Figure 2B, LINC01224 expression was strikingly decreased in both cells compared with the negative control group. Moreover, knockdown of LINC01224 strikingly increased the expression of miR-2467 (Figure 2C). Additionally, overexpression of LINC01224 reduced expression of miR-2467 in the two cell lines was verified by qRT-PCR (Figure 2D). Taken together, these results indicated the negative regulation existing between LINC01224 and miR-2467.
miR-2467 Was a Direct Target of LINC01224
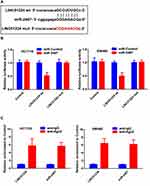
Dual-luciferase reporter assay was performed to further determine whether miR-2467 was a direct of LINC01224. The result of luciferase assay indicated that miR-2467 mimic obviously decreased luciferase reporter expression in the cells transfected with LINC01224-wt but not the cells transfected with LINC01224-mut or NC (Figure 3A and B). It is widely acknowledged that miRNA functions through regulating RISC (RNA-induced silencing complex). Ago2, a key component of RISC, exert crucial roles in RNA cleavage.12 Thus, RIP assay was performed to determine whether miR-2467 regulates LINC01224 via RISC formation. As the results show, compared to NC (IgG), LINC01224 was preferentially enriched in anti-Ago2 antibody-incubated beads (Figure 3C). Collectively, these data revealed that LINC01224 bound to miR-2467 directly.
LINC01224 and miR-2467 Effects on CRC Cell Proliferation and Invasion
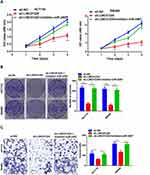
We further explored the biological effects of LINC01224 and miR-2467 on CRC cell proliferation. The results of CCK-8 analysis suggested that knockdown of LINC01224 significantly inhibited proliferation of HCT116 and SW480 cells but the inhibitory effect was reversed when the cells were co-transfected with sh-LINC01224 and the miR-2467 inhibitor (Figure 4A). Similarly, colony formation further confirmed that knockdown of LINC01224 significantly inhibited proliferation of HCT116 and SW480 cells but the inhibitory effect was reversed when the cells were co-transfected with sh-LINC01224 and the miR-2467 inhibitor (Figure 4B). We also examined whether LINC01224 and miR-2467 affected CRC cell invasion. Transwell invasion analysis suggested that LINC01224 silence could suppress the invasion abilities of CRC cells. Similar to above, this effect was also reversed when sh-LINC01224 and the miR-2467 inhibitor were cotransfected (Figure 4C).
LINC01224 Silence Inhibited CRC Tumor Growth in vivo
To further verify the in vitro results the subcutaneous xenograft tumor model was established by injecting stable sh-NC or sh-LINC01224 cells into nude mice. Consistent with the in vitro findings, the in vivo study demonstrated that the volume and weight of tumors in the sh- LINC01224 group were significantly reduced compared with the sh-NC group (Figure 5A–C). Moreover, the expression of LINC01224 was decreased in sh-LINC01224 group tumors while miR-2467 expression was increased in sh-LINC01224 group tumors compared with that of the sh-NC group (Figure 5D and E). Therefore, it was concluded that LINC01224 knockdown inhibited CRC tumor growth in vivo.
Discussion
Tumor cells proliferation and invasion are important aggressive behavior of human cancers.13,14 A large number of studies have revealed that lncRNA can function as promoter or inhibitor of human cancer proliferation, invasion and it was widely acknowledged that numerous lncRNAs were involved in the progression of CRC.2 For example, FOXCUT (a novel long noncoding RNA) promotes the colorectal cancer proliferation and invasion through activating FOXC1/PI3K/AKT pathway.15 Knockdown of long noncoding RNA linc-ITGB1 suppresses colorectal cancer metastasis by inhibiting BDNF.16 LncRNA CCAT2 facilitates chromosomal instability in colorectal cancer via BOP1-AURKB axis.17 However, the exact function and underlying mechanism of the majority of lncRNAs in CRC is still unclear.
LINC01224, a novel identified lncRNA, was reported to promote the development of hepatocellular carcinoma and epithelial ovarian cancer. In brief, LINC01224 expression was up-regulated in epithelial ovarian cancer tissues and cells, and increased expression of LINC01224 was associated with the progression of epithelial ovarian cancer patients. LINC01224 silence inhibited epithelial ovarian cancer cell proliferation, migration, and invasion in vitro, and hindered epithelial ovarian cancer tumor growth in vivo. Mechanistically, LINC01224 served as a competing endogenous RNA for microRNA-485-5p (miR-485-5p) and consequently increased PAK4 expression.11 LINC01224 could up-regulate the expression of CHEK1 by competitively binding to miR-330-5p, thus promoting hepatocellular carcinoma development.10 In the present study, similar to the previous studies, we found that the expression of LINC01224 was significantly increased in CRC tissues and cell lines. Moreover, in vitro analysis demonstrated that LINC01224 silence suppressed cell proliferation and invasion. Meanwhile, our results further confirmed that LINC01224 knockdown inhibited CRC growth in vivo.
Recently, accumulating studies suggested that lncRNA exerts its functions through binding with microRNAs as ceRNA.18 For example, LINC00963 promotes cell proliferation and migration via the miR-124-3p/FZD4 signaling pathway in CRC.19
LncRNA HOTAIR knockdown enhances radiosensitivity through regulating microRNA-93/ATG12 axis in CRC.20 In this study, an online database was used to find potential target miRNAs of LINC01224 and selected miR-2467. Previous studies have established the inhibitory function of miR-2467 in human cancers, including colorectal cancer,21 cervical cancer,22 and non-small cell lung cancer.23 In LINC01224-knockdown CRC cells, the expression of miR-2467 was significantly increased, while LINC01224overexpresion significantly up-regulated the expression of miR-2467. Additionally, the expression of miR-2467 was negatively associated with LINC01224 expression in CRC tissues. The results of luciferase reporter and RIP assay further confirmed that LINC01224 directly bound to miR-2467. Finally, the results of the functional rescue assay indicated that LINC01224 knockdown inhibited CRC cell proliferation and invasion through regulating miR-2467.
In conclusion, our findings indicated that LINC01224 expression is up-regulated in CRC tissues and cell lines. Moreover, LINC01224 promotes CRC proliferation, invasion partly through regulating miR-2467. These results demonstrated that LINC01224 is an important oncogenic player in the development of CRC and may be a promising therapeutic target for CRC patients.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Liang L, Chen Y, Yu Y, et al. SLC25A18 has prognostic value in colorectal cancer and represses Warburg effect and cell proliferation via Wnt signaling. Am J Cancer Res. 2020;10(5):1548–1567.
2. Zhao Y, Du T, Du L, et al. Long noncoding RNA LINC02418 regulates MELK expression by acting as a ceRNA and may serve as a diagnostic marker for colorectal cancer. Cell Death Dis. 2019;10(8):568. doi:10.1038/s41419-019-1804-x
3. Li Y, Liu J, Xiao Q, et al. EN2 as an oncogene promotes tumor progression via regulating CCL20 in colorectal cancer. Cell Death Dis. 2020;11(7):604. doi:10.1038/s41419-020-02804-3
4. Huang J, Pan B, Xia G, et al. LncRNA SNHG15 regulates EGFR-TKI acquired resistance in lung adenocarcinoma through sponging miR-451 to upregulate MDR-1. Cell Death Dis. 2020;11(7):525. doi:10.1038/s41419-020-2683-x
5. Han Q, Li J, Xiong J, et al. Long noncoding RNA LINC00514 accelerates pancreatic cancer progression by acting as a ceRNA of miR-28-5p to upregulate Rap1b expression. J Exp Clin Cancer Res. 2020;39(1):151. doi:10.1186/s13046-020-01660-5
6. Song Z, Zhang X, Lin Y, et al. LINC01133 inhibits breast cancer invasion and metastasis by negatively regulating SOX4 expression through EZH2. J Cell Mol Med. 2019;23(11):7554–7565. doi:10.1111/jcmm.14625
7. Sun LB, Zhao S, Zhu J, et al. Long noncoding RNA UCID sponges miR‑152‑3p to promote colorectal cancer cell migration and invasion via the Wnt/β‑catenin signaling pathway. Oncol Rep. 2020;44(3):1194–1205. doi:10.3892/or.2020.7670
8. Lai F, Deng W, Fu C, et al. Long non-coding RNA SNHG6 increases JAK2 expression by targeting the miR-181 family to promote colorectal cancer cell proliferation. J Gene Med. 2020;22(12):e3262. doi:10.1002/jgm.3262
9. Jiang Z, Li L, Hou Z, et al. LncRNA HAND2-AS1 inhibits 5-fluorouracil resistance by modulating miR-20a/PDCD4 axis in colorectal cancer. Cell Signal. 2020;66:109483. doi:10.1016/j.cellsig.2019.109483
10. Gong D, Feng P-C, Ke X-F, et al. Silencing long non-coding RNA LINC01224 inhibits hepatocellular carcinoma progression via MicroRNA-330-5p-induced inhibition of CHEK1. Mol Ther Nucleic Acids. 2020;19:482–497. doi:10.1016/j.omtn.2019.10.007
11. Xing S, Zhang Y, Zhang J. LINC01224 exhibits cancer-promoting activity in epithelial ovarian cancer through microRNA-485-5p-mediated PAK4 upregulation. Onco Targets Ther. 2020;13:5643–5655. doi:10.2147/OTT.S254662
12. Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet. 2008;9(2):102–114. doi:10.1038/nrg2290
13. An Y, Zhang S, Zhang J, et al. Overexpression of lncRNA NLIPMT inhibits colorectal cancer cell migration and invasion by downregulating TGF-β1. Cancer Manag Res. 2020;12:6045–6052. doi:10.2147/CMAR.S247764
14. Li N, Li J, Mi Q, et al. Long non-coding RNA ADAMTS9-AS1 suppresses colorectal cancer by inhibiting the Wnt/beta-catenin signalling pathway and is a potential diagnostic biomarker. J Cell Mol Med. 2020;24(19):11318–11329.
15. Zhang X, Yi S, Xing G, et al. FOXCUT promotes the proliferation and invasion by activating FOXC1/PI3K/AKT pathway in colorectal cancer. Cancer Manag Res. 2020;12:6269–6278. doi:10.2147/CMAR.S259801
16. Wan WB, Kong QL. Knockdown of long noncoding RNA linc-ITGB1 inhibits tumor metastasis in colorectal cancer through suppressing BDNF. Eur Rev Med Pharmacol Sci. 2020;24(14):7551.
17. Chen B, Dragomir MP, Fabris L, et al. The long noncoding RNA CCAT2 induces chromosomal instability through BOP1-AURKB signaling. Gastroenterology. 2020;159(6):2146–2162.e33. doi:10.1053/j.gastro.2020.08.018
18. Cesana M, Cacchiarelli D, Legnini I, et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell. 2011;147(2):358–369. doi:10.1016/j.cell.2011.09.028
19. Zheng K, Zhang TK. LncRNA LINC00963 promotes proliferation and migration through the miR-124-3p/FZD4 pathway in colorectal cancer. Eur Rev Med Pharmacol Sci. 2020;24(14):7634–7644. doi:10.26355/eurrev_202007_22264
20. Liu Y, Chen X, Chen X, et al. Long non-coding RNA HOTAIR knockdown enhances radiosensitivity through regulating microRNA-93/ATG12 axis in colorectal cancer. Cell Death Dis. 2020;11(3):175. doi:10.1038/s41419-020-2268-8
21. Xiao H, Liu M. Circular RNA hsa_circ_0053277 promotes the development of colorectal cancer by upregulating matrix metallopeptidase 14 via miR-2467-3p sequestration. J Cell Physiol. 2020;235(3):2881–2890. doi:10.1002/jcp.29193
22. Liu F, Wen C. LINC01410 knockdown suppresses cervical cancer growth and invasion via targeting miR-2467-3p/VOPP1 axis. Cancer Manag Res. 2020;12:855–861. doi:10.2147/CMAR.S236832
23. Chen H, Tan X, Ding Y. Knockdown SNHG20 suppresses nonsmall cell lung cancer development by repressing proliferation, migration and invasion, and inducing apoptosis by regulating miR-2467-3p/E2F3. Cancer Biother Radiopharm. 2020. doi:10.1089/cbr.2019.3430
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.