Back to Journals » OncoTargets and Therapy » Volume 13
LINC01224 Exhibits Cancer-Promoting Activity in Epithelial Ovarian Cancer Through microRNA-485-5p-Mediated PAK4 Upregulation
Authors Xing S, Zhang Y, Zhang J
Received 19 March 2020
Accepted for publication 12 May 2020
Published 16 June 2020 Volume 2020:13 Pages 5643—5655
DOI https://doi.org/10.2147/OTT.S254662
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr XuYu Yang
This paper has been retracted.
Shujian Xing, 1 Yaqi Zhang, 2 Jing Zhang 3
1Department of Gynaecology and Obstetrics, Zouping People’s Hospital, Zouping, Shandong 256200, People’s Republic of China; 2Department of Gynaecology and Obstetrics, Weifang Yidu Central Hospital, Weifang, Shandong 262500, People’s Republic of China; 3Department of Gynaecology and Obstetrics, The No.4 Hospital of Jinan, Jinan, Shandong 250031, People’s Republic of China
Correspondence: Jing Zhang Email [email protected]
Purpose: Long intergenic non-protein coding RNA 1224 (LINC01224) plays vital roles in the tumorigenesis and progression of hepatocellular carcinoma. Here, we determined LINC01224 expression in epithelial ovarian cancer (EOC) tissues and cells. We also assessed the effects of LINC01224 knockdown on the malignant phenotype of EOC cells both in vitro and in vivo. Furthermore, the detailed molecular mechanisms underlying the oncogenic actions of LINC01224 in EOC cells were elucidated.
Methods: Quantitative real-time polymerase chain reaction (qRT-PCR) was used to detect LINC01224 expression in EOC tissues and cells. EOC cells were transfected with small interfering RNAs, and cell proliferation, apoptosis, migration, and invasion were assessed using Cell Counting Kit-8 assay, flow cytometry, cell migration assays, and cell invasion assays, respectively. Using tumor xenografts, the effects of LINC01224 silencing on EOC tumor growth were analyzed in vivo. The mechanism underlying LINC01224 regulation of malignant processes in EOC cells was explored using bioinformatics, RNA immunoprecipitation assay, qRT-PCR, Western blotting, and rescue experiments.
Results: LINC01224 expression was upregulated in EOC tissues and cells. LINC01224 upregulation was correlated to tumor size, the International Federation of Gynecology and Obstetrics stage, and lymph node metastasis. LINC01224 depletion in EOC cells suppressed cell proliferation, migration, and invasion and facilitated cell apoptosis in vitro. LINC01224 downregulation also hindered EOC tumor growth in vivo. Mechanistically, LINC01224 served as a competing endogenous RNA for microRNA-485-5p (miR-485-5p) and consequently increased p21-activated kinase 4 (PAK4) expression in EOC cells. Furthermore, miR-485-5p inhibition or PAK4 upregulation significantly abrogated the effects of LINC01224 depletion in EOC cells.
Conclusion: LINC01224/miR-485-5p/PAK4 formed a competing endogenous RNA network regulating the aggressive behavior of EOC. Therefore, targeting this pathway may be an attractive therapeutic strategy for EOC.
Keywords: long intergenic non-protein coding RNA 1224, p21‑activated kinase 4, epithelial ovarian cancer, anticancer therapy
Introduction
Ovarian cancer is one of the most common gynecological malignancies and is the third leading cause of cancer-related mortalities among women.1 According to GLOBOCAN 2018, there are 295,414 new ovarian cancer cases and 84,799 ovarian cancer-related deaths globally each year.2 Epithelial ovarian cancer (EOC) is a major subtype of ovarian cancer and accounts for approximately 90% of all ovarian cancer cases.3 Owing to lack of obvious symptoms during the early phase of EOC and the unavailability of sensitive clinical screening techniques, patients with EOC are usually diagnosed with general dissemination metastasis in the pelvic and abdominal cavity.4 Although significant advances have been made toward the development of therapeutic and diagnostic methods, the outcomes of patients with EOC remain rather poor, and approximately 80% of patients suffer from recurrence and/or metastasis even after undergoing complex treatment.5 Taking these issues into account, there is an urgent clinical demand to thoroughly understand the key molecular mechanisms associated with EOC initiation and progression, which may lead to the identification of effective diagnostic and therapeutic targets for this malignancy.
Noncoding RNAs, including long noncoding RNAs (lncRNAs) and microRNAs (miRNAs), have been extensively studied for their roles in carcinogenesis and cancer progression.6 lncRNAs are a group of transcribed RNA molecules longer than 200 nucleotides.7 lncRNAs lack an open reading frame and do not therefore encode proteins.8 However, a substantial amount of evidence has revealed that they participate in regulating gene expression at transcriptional and/or post-transcriptional levels by interacting with DNA, proteins, or other RNAs.9–11 Abnormal expression of lncRNAs has been frequently observed in EOC, wherein they exhibit both tumor-suppressing or tumor-inhibiting activities.12,13
miRNAs are another group of noncoding RNAs with a length of 20–25 nucleotides.14 Although they do not encode proteins, miRNAs are implicated in diverse biological behaviors via direct binding to the 3′-untranslated regions (3′-UTRs) of mRNAs, resulting in mRNA degradation or translation repression. Interestingly, several studies have identified the role of an lncRNA–miRNA regulatory axis in EOC.15–17 lncRNAs can serve as competing endogenous RNAs (ceRNAs) or molecular sponges by competitively interacting with miRNAs, thereby liberating miRNA-induced actions on target mRNAs.18 Therefore, exploring the lncRNA–miRNA axis in EOC is crucial and essential for the development of potential diagnostic biomarkers and therapeutic targets.
Long intergenic non-protein coding RNA 1224 (LINC01224) plays crucial roles in the tumorigenesis and progression of hepatocellular carcinoma.19 However, its expression status, detailed roles, and possible molecular mechanisms in EOC have not yet been adequately explored. Therefore, we determined LINC01224 expression in EOC tissues and cell lines. In addition, we assessed the effects of LINC01224 knockdown on the malignant phenotype of EOC cells both in vitro and in vivo. Furthermore, the molecular mechanisms underlying the oncogenic actions of LINC01224 in EOC cells were elucidated in detail.
Materials and Methods
Ethics Statement and Clinical Tissues
This study was approved by the Ethics Committee of The No.4 Hospital of Jinan and performed in accordance with the guidelines of the Declaration of Helsinki. Written informed consent was provided by all patients prior to their participation. Paired EOC tissues and the corresponding adjacent normal tissues were obtained from 63 patients in The No.4 Hospital of Jinan. All normal tissues were obtained at least 2 cm away from EOC tissues. All normal ovarian and EOC tissues were confirmed according to histopathological evaluation. Patients who received preoperative radiotherapy, chemotherapy, or other anticancer therapies were excluded. All collected tissue specimens were instantly frozen and stored in liquid nitrogen until further use. The pathologic types of patients with EOC were summarized in Table 1.
 |
Table 1 Summary of Pathologic Types of EOC |
Cell Lines
The Cell Bank of the Chinese Academy of Science (Shanghai, China) provided four EOC cell lines: Caov-3, ES-2, OVCAR3, and SK-OV-3. Caov-3 cells were cultured in Dulbecco’s modified Eagle’s medium (Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc.) and sodium pyruvate 100 mM solution (Gibco; Thermo Fisher Scientific, Inc.). ES-2 and SK-OV-3 cells were cultured in McCoy’s 5A culture medium (Sigma-Aldrich; Merck KGaA, Darmstadt, Germany) containing 10% FBS. OVCAR3 cells were maintained in RPMI-1640 medium (Gibco; Thermo Fisher Scientific, Inc.) with 20% FBS. All culture media were supplemented with100 U/mL penicillin and 100 mg/mL streptomycin (both from Sigma-Aldrich). The normal human ovarian epithelial cell line NOEC was purchased from ScienCell Research Laboratories (Catalog #7310; Carlsbad, CA, USA) and cultured in Ovarian Epithelial Cell Medium (Catalog #7311; ScienCell Research Laboratories). All cells were grown in humidified air with 5% CO2 at 37°C.
Cell Transfection
Small interfering RNAs (siRNAs) targeting LINC01224 (si-LINC01224#1 and si-LINC01224#2) and negative control siRNA (si-NC) were obtained from Ribobio (Guangzhou, China). An miR-485-5p mimic and miR-485-5p inhibitor (GenePharma; Shanghai, China) were used to increase and decrease endogenous miR-485-5p expression, respectively, with miRNA mimic negative control (miR-NC) and NC inhibitor as controls. The p21-activated kinase 4 (PAK4) overexpression plasmid pCDNA3.1-PAK4 (pc-PAK4) was constructed by GenePharma (Shanghai, China). The empty plasmid pcDNA3.1 served as the negative control for pc-PAK4. Cells were inoculated into 6-well plates and transfected with the above molecular products using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA).
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
The RNAsimple Total RNA Kit (Tiangen Biotech, China) was used to isolate total RNA from tissue specimens or cultured cells. To quantify LINC01224 and PAK4 expression, cDNA was synthesized from total RNA using a Prime Script RT Reagent Kit (Takara, Dalian, China). The synthesized cDNA was subjected to quantitative PCR using a SYBR Premix Ex Taq kit (Takara). Expression levels of LINC01224 and PAK4 were normalized to that of GAPDH. miR-485-5p expression was detected using an All-in-One™ miRNA qRT-PCR Detection Kit (GeneCopoeia, Guangzhou, China). U6 small nuclear RNA was used to normalize miR-485-5p expression. Relative gene expression was calculated using the 2−ΔΔCt method.
Subcellular Fractionation
The separation and purification of cytoplasmic and nuclear RNA were performed using a Cytoplasmic and Nuclear RNA Purification Kit (Norgen Biotek, Thorold, Canada) following the manufacturer’s instructions. The cytoplasmic and nuclear RNA was then subjected to qRT-PCR for the determination of LINC01224 sub-cellular percentage.
Cell Counting Kit-8 (CCK-8) Assay
The CCK-8 assay was conducted to measure cell proliferation. Briefly, 24 h following transfection, cells were collected and seeded into 96-well plates at a density of 2 × 103 cells/well and then incubated at 37°C for 0, 24, 48, or 72 h. At each time point, 10 μL CCK-8 reagent (Dojindo Laboratories, Kumamoto, Japan) was added and cells were incubated at 37°C for 2 h. Optical density was detected at 450 nm using a microtiter plate reader (SpectraMax, Molecular Devices, USA).
Flow Cytometry
Cell apoptosis was determined using an Annexin V/Fluorescein Isothiocyanate (FITC) Apoptosis Detection Kit (Biolegend, San Diego, CA, USA). Transfected cells were harvested by treating with an EDTA-free trypsin reagent (Gibco; Thermo Fisher Scientific, Inc.). After two rinses with pre-cooled PBS, cells were centrifuged at 37°C for 5 min. The supernatant was then aspirated, and the cells were resuspended in 100 μL of 1× binding buffer and stained with 5 μL of Annexin V/FITC/and 5 μL of propidium iodide. Following 15-min incubation at room temperature in the dark, the apoptotic rate was quantified using flow cytometry (FACScan, BD Biosciences, Franklin Lakes, NJ, USA).
Cell Migration and Invasion Assays
Transwell polycarbonate inserts with an 8-µm pore size (BD Biosciences, San Jose, CA) were used for cell migration assays. A total of 5 × 104 cells suspended in 100 μL FBS-free culture medium were added into the upper compartment. The lower compartment was filled with culture medium containing 10% FBS. Following incubation for 24 h at 37°C, the unmigrated cells were carefully removed with a cotton bud, and the migrated cells were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. The stained cells were photographed using a light microscope, and five random fields per sample were selected to count cell numbers. Cell invasion assays were conducted as described above, except that the transwell chambers were precoated with Matrigel® (BD Biosciences).
Tumor Xenografts
To knockdown LINC01224, recombinant lentiviruses carrying LINC01224 short hairpin RNA (shRNA; sh-LINC01224) and negative control shRNA (sh-NC) were chemically synthesized by GenePharma. After transfection with the constructed lentiviruses, the stably transfected Caov-3 cells were selected with puromycin for 14 days.
All animal studies were approved by the Animal Care and Use Committee of The No.4 Hospital of Jinan. Female BALB/c nude mice (4–6-week old) were purchased from the Experimental Animal Center of Shandong University (Shandong, China), and carried out in compliance with the Animal Protection Law of the People’s Republic of China-2009 for experimental animals. The stably transfected Caov-3 cells were collected and resuspended in culture medium. Cell suspension (100 μL) containing 5 × 106 cells, uniformly mixed with 0.1 mL of extracellular matrix gel, was subcutaneously injected into the flank of mice. Each group contained three mice. After cell inoculation, the volume of tumor xenografts was measured every 4 days and analyzed using the following formula: 0.5 × width2 × length. The width and length of tumor xenografts were measured using a Vernier caliper. All mice were euthanized 4 weeks after injection, and the tumor xenografts were removed, weighed, and collected for further use.
Bioinformatics Analysis
Gene Expression Profiling Interactive Analysis (GEPIA2; http://gepia2.cancer-pku.cn/#index) was used to analyze LINC01224 expression in EOC. The interaction between LINC01224 and miRNAs was assessed using StarBase 3.0 (http://starbase.sysu.edu.cn/).
Two online databases, StarBase 3.0 and TargetScan Human 7.2 (http://www.targetscan.org/vert_72/), were used to predict the putative targets of miR-485-5p.
RNA Immunoprecipitation (RIP) Assay
The RIP assay was performed using a Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Billerica, MA, USA). EOC cells were treated with RIP lysis buffer to obtain cell extracts. The cell extracts were probed with magnetic beads conjugated with human anti-argonaute 2 (Ago2; Millipore) or anti-immunoglobulin G (IgG; Millipore) antibodies. After overnight incubation at 4°C, the magnetic beads were digested with proteinase K to remove proteins and isolate RNA. Finally, the isolated RNA was subjected to qRT-PCR to quantify LINC01224 and miR-485-5p expression.
Luciferase Reporter Assay
The partial sequences of LINC01224 carrying the predicted wild-type (wt) miR-485-5p binding site (CAGCCUC) and mutant (mut) LINC01224 sequences (GUCGGAG) were inserted into the psiCHECK™-2 luciferase reporter plasmid (Promega Corporation, Madison, WI, USA), resulting in wt-LINC01224 and mut-LINC01224 reporter plasmids. The wt-PAK4 and mut-PAK4 reporter plasmids were designed and produced in the same way.
For reporter assays, cells seeded into 24-well plates were transfected with either wt or mut luciferase reporter plasmids along with miR-485-5p mimic or miR-NC using Lipofectamine 2000. Luciferase activity was detected 48 h after transfection using a dual-luciferase assay system (Promega Corporation).
Western Blotting
Transfected cells were collected and lysed in radioimmunoprecipitation lysis buffer (Beyotime Institute of Biotechnology, Shanghai, China) containing phenylmethylsulfonyl fluoride. A Bradford Protein Quantitative Kit (Tiangen Biotech) was used to measure protein concentration. Equal amounts of proteins were isolated using 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis, transferred to polyvinylidene fluoride membranes, and blocked with 5% nonfat dried milk at room temperature for 2 h. After incubating with primary antibodies overnight at 4°C, the membranes were incubated with a horseradish peroxidase (HRP)-conjugated anti-rabbit IgG secondary antibody (ab205718; 1:5000 dilution; Abcam Cambridge, MA, USA), followed by incubation with the ECL Prime Western Blotting Detection Reagent (GE Healthcare, Parsippany, NJ, USA) for protein signal visualization. The primary antibodies against PAK4 (ab62509) or GAPDH (ab181602) were obtained from Abcam and used at a dilution of 1:1000.
Statistical Analysis
All experiments were independently repeated at least three times. The obtained data were presented as mean and standard deviation and analyzed using SPSS 21.0 (IBM Corp, Armonk, NY). The correlation of LINC01224 expression with clinicopathological characteristics of patients with EOC was examined using the chi-square test. Comparisons between two groups were performed using t-tests. One-way analysis of variance following by Tukey’s post hoc test was performed to detect differences among multiple groups. The correlation between LINC01224 and miR-485-5p expression was tested via Pearson’s correlation coefficient analysis. A value of P < 0.05 was considered statistically significant.
Results
LINC01224 Is Upregulated in EOC
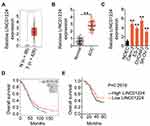
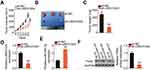
RNA sequencing data of LINC01224 expression from The Cancer Genome Atlas (TCGA) and The Genotype Tissue Expression (GTEx) projects were used to analyze LINC01224 expression in EOC using GEPIA2. The data indicated that LINC01224 expression was higher in EOC tissues (n = 426) than in normal ovarian (n = 88) tissues (Figure 1A). In agreement with this result, qRT-PCR analysis confirmed that LINC01224 expression was upregulated in EOC tissues compared with that in the corresponding adjacent normal tissues (Figure 1B). Further measurement of LINC01224 expression was performed in four EOC cell lines (Caov-3, ES-2, OVCAR3, and SK-OV-3) and the normal human ovarian epithelial cell line NOEC. All four EOC cell lines showed a much higher level of LINC01224 expression than the NOEC cell line (Figure 1C).
To examine the clinical relevance of LINC01224 in EOC, GEPIA2 was used to determine whether LINC01224 expression was associated with the overall survival of patients with EOC. The data showed that there was no significant association between LINC01224 expression and overall survival of patients with EOC (Figure 1D). Our finding was also consistent with this observation (Figure 1E; P = 0.2619). All 63 patients enrolled were classified into either low LINC01224 or high LINC01224 expression groups using the median LINC01224 expression in EOC tissues as the cutoff value. Next, the correlation of LINC01224 expression with clinicopathological characteristics of the 63 patients with EOC was analyzed using the chi-square test. As shown in Table 2, high LINC01224 expression was correlated with tumor size (P = 0.045), the International Federation of Gynecology and Obstetrics (FIGO) stage (P = 0.019), and lymph node metastasis (P = 0.011). These findings collectively suggest that LINC01224 is upregulated in EOC and plays important roles in aggressive cancer behaviors.
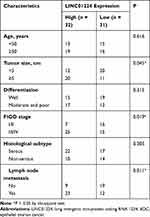 |
Table 2 Association of LINC01224 Expression with Clinicopathological Characteristics of Patients with EOC |
LINC01224 Depletion Inhibits EOC Cell Proliferation, Migration, and Invasion and Promotes Cell Apoptosis in vitro
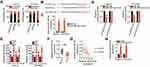
To explore the detailed functions of LINC01224 in EOC, Caov-3 and OVCAR3 cells, which showed the highest level of LINC01224 expression, were used in subsequent experiments and were transfected with si-LINC001224. qRT-PCR analysis verified that transfection with both si-LINC001224#1 and si-LINC001224#2 dramatically downregulated LINC01224 expression in Caov-3 and OVCAR3 cells (Figure 2A). The CCK-8 assay and flow cytometry were performed to determine the effects of LINC01224 downregulation on the proliferation and apoptosis of Caov-3 and OVCAR3 cells, respectively. LINC01224 downregulation suppressed Caov-3 and OVCAR3 cell proliferation (Figure 2B) but promoted cell apoptosis (Figure 2C) in vitro. In addition, the migratory (Figure 2D) and invasive (Figure 2E) capacities of Caov-3 and OVCAR3 cells were obviously reduced after LINC01224 silencing, as revealed by cell migration and invasion assays. Collectively, these results suggest that LINC01224 exhibits tumor-promoting activities in EOC progression.
LINC01224 Acts as a Molecular Sponge for miR-485-5p in EOC Cells
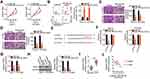
lncRNAs located in the cytoplasm serve as ceRNAs or molecular sponges for miRNAs. We first evaluated the cellular distribution of LINC01224 expression in EOC cells. Subcellular fractionation indicated that LINC01224 was mainly distributed in the cytoplasm of Caov-3 and OVCAR3 cells (Figure 3A). Bioinformatic prediction analysis was conducted to identify target miRNAs of LINC01224. The seed region of miR-485-5p could form complementary base pairing with LINC01224 (Figure 3B). In addition, miR-485-5p expression is downregulated in multiple human cancers, and it exerts antioncogenic actions during carcinogenesis and cancer progression;20–30 therefore, this miRNA was selected for further validation.
Luciferase reporter assay was conducted to verify whether miR-485-5p was able to directly bind to LINC01224 in EOC cells. To conduct this assay, the efficiency of miR-485-5p mimic was determined using qRT-PCR. The results showed that transfection with miR-485-5p mimic notably upregulated miR-485-5p expression in both Caov-3 and OVCAR3 cells (Figure 3C). The results of the luciferase reporter assay revealed that miR-485-5p upregulation evidently decreased the luciferase activity of wt-LINC01224 in Caov-3 and OVCAR3 cells, whereas mutation in the binding site within LINC01224 abrogated the inhibitory ability of miR-485-5p on luciferase activity (Figure 3D). Furthermore, the RIP assay further demonstrated that LINC01224 and miR-485-5p were clearly enriched in Ago2-containing microribonucleoprotein complexes (Figure 3E), suggesting that LINC01224 can bind to the miR-485-5p RNA-induced silencing complex in Caov-3 and OVCAR3 cells.
In addition, miR-485-5p was expression was lower in EOC tissues than in the corresponding adjacent normal tissues (Figure 3F). Furthermore, miR-485-5p expression was inversely correlated to LINC01224 expression in the 63 EOC tissues, as shown by Pearson’s correlation coefficient analysis (Figure 3G; r = −0.5315, P < 0.0001). Moreover, LINC01224 silencing clearly increased miR-485-5p expression in Caov-3 and OVCAR3 cells (Figure 3H). Collectively, these results suggest that LINC01224 functions as a molecular sponge for miR-485-5p in EOC cells.
PAK4 Is a Direct Target of miR-485-5p in EOC Cells
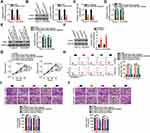
Because miR-485-5p was downregulated in EOC, we next studied the biological roles of miR-485-5p in EOC cells. The CCK-8 assay and flow cytometry indicated that exogenous miR-485-5p expression suppressed the proliferation (Figure 4A) and enhanced the apoptosis (Figure 4B) of Caov-3 and OVCAR3 cells. Additionally, ectopic miR-485-5p expression obviously impaired the migration (Figure 4C) and invasion (Figure 4D) of Caov-3 and OVCAR3 cells.
Using an online database for miRNA target predictions, PAK4 was predicted as a putative target of miR-485-5p (Figure 4E) and selected for further analysis in subsequent experiments. Luciferase reporter assay was conducted to assess whether miR-485-5p could bind to the 3′-UTR of PAK4. The luciferase activity of wt-PAK4 but not of mut-PAK4 was decreased in Caov-3 and OVCAR3 cells following miR-485-5p overexpression (Figure 4F), suggesting that miR-485-5p was could directly binding to the 3′-UTR of PAK4. PAK4 mRNA (Figure 4G) and protein (Figure 4H) expression was downregulated in miR-485 mimic-transfected Caov-3 and OVCAR3 cells, as revealed by qRT-PCR and Western blotting, respectively. Furthermore, PAK4 mRNA expression was higher in EOC tissues (Figure 4I) and was inversely correlated to miR-485-5p expression (Figure 4J; r = −0.5986, P < 0.0001). Therefore, miR-485-5p functions as an antioncogenic miRNA, and PAK4 is a direct target of miR-485-5p in EOC cells.
Increased miR-485-5p/PAK4 Output Axis Abolishes the Effects of LINC01224 Knockdown in EOC Cells
We demonstrated that LINC01224 functions as a molecular sponge for miR-485-5p, and PAK4 is a direct target of miR-485-5p. Thus, we examined whether LINC01224 regulated PAK4 expression in EOC cells via sponging miR-485-5p. qRT-PCR and Western blotting were conducted to measure PAK4 mRNA and protein levels, respectively, in LINC01224-deficient Caov-3 and OVCAR3 cells. LINC01224 depletion notably suppressed PAK4 expression in Caov-3 and OVCAR3 cells at both mRNA (Figure 5A) and protein (Figure 5B) levels. To conduct rescue experiments, qRT-PCR was used to determine the transfection efficiency of miR-485-5p inhibitor in Caov-3 and OVCAR3 cells. miR-485-5p inhibitor effectively silenced miR-485-5p expression in Caov-3 and OVCAR3 cells (Figure 5C). si-LINC01224 together with miR-485-5p inhibitor or NC inhibitor was co-transfected into Caov-3 and OVCAR3 cells. The downregulation of PAK4 mRNA (Figure 5D) and protein expression (Figure 5E) following LINC01224 knockdown was almost completely recovered in Caov-3 and OVCAR3 cells after miR-485-5p inhibition.
Furthermore, a series of rescue experiments was performed to further clarify whether the pro-oncogenic activities of LINC01224 in EOC cells were dependent on the output of the miR-485-5p/PAK4 axis. The PAK4 overexpression plasmid pc-PAK4 was used in rescue assays, and its transfection efficiency was detected using Western blotting. Treatment with pc-PAK4 led to considerable upregulation of PAK4 protein expression in Caov-3 and OVCAR3 cells (Figure 5F). Next, LINC01224-deficient Caov-3 and OVCAR3 cells were co-transfected with miR-485-5p inhibitor or pc-PAK4. The CCK-8 assay and flow cytometry revealed that reduction in LINC01224 expression attenuated Caov-3 and OVCAR3 cell proliferation (Figure 5G) and promoted their apoptosis (Figure 5H). Conversely, the LINC01224 silencing-mediated effects on cell proliferation and apoptosis were abrogated by miR-485-5p inhibitor or pc-PAK4 co-transfection. Similarly, LINC01224 downregulation impaired the migration (Figure 5I) and invasion (Figure 5J) of Caov-3 and OVCAR3 cells, but miR-485-5p inhibition or PAK4 reintroduction reversed these effects. In summary, the above results suggest that LINC01224 performs its oncogenic actions during EOC progression via regulating output of the miR-485-5p/PAK4 axis.
LINC01224 Silencing Inhibits Tumor Growth of EOC Cells in vivo
To elucidate the effect of LINC01224 silencing on EOC tumor growth in vivo, tumor xenografts were generated using recombinant lentiviruses expressing sh-LINC01224 or sh-NC. The volumes of tumor xenografts formed by sh-LINC01224-transfected Caov-3 cells were markedly smaller than those formed by sh-NC-transfected Caov-3 cells, and the growth curves of the sh-LINC01224 and sh-NC groups diverged over time (Figure 6A and B). The weights of tumor xenografts in the sh-LINC01224 group were evidently reduced compared with those in the sh-NC group (Figure 6C). In addition, qRT-PCR analysis showed that LINC01224 expression was downregulated (Figure 6D), while miR-485-5p expression was increased (Figure 6E) in the tumor xenografts derived from Caov-3 cells stably expressing sh-LINC01224. Finally, PAK4 protein expression was downregulated in the LINC01224-depleted tumor xenografts (Figure 6F), as shown by Western blotting. In short, these results suggest that LINC01224 silencing inhibited tumor growth of EOC cells in vivo via regulating the miR-485-5p/PAK4 axis.
Discussion
An increasing number of studies have recently indicated that differentially expressed lncRNAs are involved in the oncogenicity of EOC.31 Extensive evidence has demonstrated the comprehensive regulatory actions of lncRNAs in nearly all aggressive phenotypes in EOC.32–34 Hence, lncRNAs may be potential targets for the diagnosis and therapy of human EOC. Nevertheless, the expression of and important roles of lncRNAs in EOC are not completely understood. The present study attempted to explore the implications of LINC01224 in EOC pathogenesis.
LINC01224 is upregulated in hepatocellular carcinoma.19 Its high expression is correlated with TNM stage and distant metastasis.19 In a previous study, LINC01224 inhibition decreased hepatocellular carcinoma cell sphere, colony formation, proliferation, migration, and invasion in vitro; promoted cell apoptosis; induced cell cycle arrest; and impaired tumor growth in vivo.19 However, the expression profile and detailed roles of LINC01224 in EOC have not yet been explored. In this study, information obtained from TCGA and GTEx databases showed that LINC01224 is upregulated in ovarian cancer. Furthermore, qRT-PCR was used to detect LINC01224 expression in 63 pairs of EOC tissues and the corresponding adjacent normal tissues. Histological evaluation was performed to verify the EOC tissues and adjacent normal ovarian tissues. LINC01224 was found to be highly expressed in EOC tissues compared with adjacent normal tissues. The survival data from TCGA and GTEx databases showed no association between LINC01224 expression and overall survival of patients with ovarian cancer, which was in line with our results. In addition, high LINC01224 expression was found to be correlated with tumor size, FIGO stage, and lymph node metastasis in patients with EOC. Functionally, loss of LINC01224 expression decreased EOC cell proliferation, migration, and invasion in vitro and increased cell apoptosis. Additionally, LINC01224 knockdown attenuated tumor growth of EOC cells in vivo. However, in the tumor xenografts, only 3 mice each group was used, and it was a limitation of our study. We will resolve it in the near future.
We next elucidated the possible mechanisms involved in LINC01224 regulation of tumor behaviors in vitro and in vivo. lncRNAs participate in the regulation of tumorigenesis and tumor progression via different mechanisms. At present, the lncRNA/miRNA/mRNA pathway has been widely studied.35–37 LncRNAs can attenuate the expression of specific miRNAs by acting as molecular sponges, thereby increasing the expression of repressed target mRNAs.38 Thus, in this study, the localization of LINC01224 in EOC cells was first examined via subcellular fractionation, which showed that LINC01224 was mostly distributed in the cytoplasm of EOC cells. Bioinformatic analysis predicted that LINC01224 may interact with miR-485-5p, a classical tumor suppressor miRNA.20–30 The results of luciferase reporter and RIP assays showed that miR-485-5p could bind directly and interact with LINC01224 in EOC cells. qRT-PCR analysis further demonstrated that miR-485-5p was expressed at low levels in EOC tissues and showed an inverse correlation with LINC01224 expression. These findings identified LINC01224 as an upstream regulator of miR-485-5p and verified LINC01224 as a molecular sponge of miR-485-5p in EOC cells.
miR-485-5p expression is downregulated in colorectal cancer,20,21 gastric cancer,22 glioblastoma,23,24 hepatocellular carcinoma,25,26 lung cancer,27 breast cancer,28,29 and bladder cancer.30 To the best of our knowledge, the present study is the first to describe the expression, roles, and underlying mechanisms of miR-485-5p in EOC. We showed that miR-485-5p was downregulated in EOC and that miR-485-5p overexpression could prevent EOC progression. PAK4 was identified as a direct downstream target of miR-485-5p in EOC cells. Furthermore, interference of LINC01224 expression reduced PAK4 expression by sponging miR-485-5p. The present study results identified a ceRNA pathway comprising LINC01224, miR-485-5p, and PAK4.
PAK4, located on 19q13.2, is a serine/threonine protein kinase and functions as a downstream effector of Rac and Cdc42.39 A previous study reported that PAK4 was upregulated in EOC and that its expression was correlated with frequent tumor metastasis, high chemoresistance, advanced stage, high grade, and short overall and disease-free survival rates.40 Functionally, PAK4 exhibited cancer-promoting activities in EOC and was implicated in the regulation of various malignant processes both in vitro and in vivo.40,41 In this study, rescue experiments showed that increased output of the miR-485-5p/PAK4 axis, including miR-485-5p inhibition and PAK4 upregulation, significantly abrogated the effects of LINC01224 depletion in EOC cells. These observations revealed that LINC01224, miR-485-5p, and PAK4 formed a ceRNA network to regulate the aggressive behavior of EOC. Therefore, targeting the LINC01224/miR-485-5p/PAK4 pathway may be an attractive therapeutic strategy for EOC.
In this study, we found that miR-485-5p was not correlated with the overall survival of patients with EOC. Nevertheless, high expression of LINC01224 and PAK4 was notably correlated with the shorter overall survival in patients with EOC. This discrepancy may be attributed to the small sample size and short follow-up time. In later experiments, we will collect more tissues and further test the correlation between miR-485-5p expression and clinical overall survival in patients with EOC.
Conclusion
We showed for the first time that LINC01224 plays pivotal roles in the malignancy of EOC both in vitro and in vivo. Furthermore, our results revealed an explicit tumor-promoting role of LINC01224 in EOC by competing with miR-485-5p as a molecular sponge and consequently increasing PAK4 expression. These findings may facilitate the development of novel diagnostic and therapeutic techniques for EOC.
Data Sharing Statement
The datasets used and/or analyzed during the present study are available from the corresponding author on reasonable request.
Disclosure
The authors declare that they have no competing interests.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. Ca-Cancer J Clin. 2019;69(1):7–34. doi:10.3322/caac.21551
2. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
3. Suh DH, Kim JW, Kim K, Kim HJ, Lee KH. Major clinical research advances in gynecologic cancer in 2012. J Gynecol Oncol. 2013;24(1):66–82. doi:10.3802/jgo.2013.24.1.66
4. Naora H, Montell DJ. Ovarian cancer metastasis: integrating insights from disparate model organisms. Nat Rev Cancer. 2005;5(5):355–366. doi:10.1038/nrc1611
5. Jessmon P, Boulanger T, Zhou W, Patwardhan P. Epidemiology and treatment patterns of epithelial ovarian cancer. Expert Rev Anticancer Therapy. 2017;17(5):427–437. doi:10.1080/14737140.2017.1299575
6. Sun T. Long noncoding RNAs act as regulators of autophagy in cancer. Pharmacol Res. 2018;129:151–155. doi:10.1016/j.phrs.2017.11.009
7. Jarroux J, Morillon A, Pinskaya M. History, discovery, and classification of lncRNAs. Adv Exp Med Biol. 2017;1008:1–46.
8. Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Gene Dev. 2009;23(13):1494–1504. doi:10.1101/gad.1800909
9. Atianand MK, Caffrey DR, Fitzgerald KA. Immunobiology of long noncoding RNAs. Annu Rev Immunol. 2017;35(1):177–198. doi:10.1146/annurev-immunol-041015-055459
10. Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29(4):452–463. doi:10.1016/j.ccell.2016.03.010
11. Abolghasemi M, Tehrani SS, Yousefi T, et al. Critical roles of long noncoding RNAs in breast cancer. J Cell Physiol. 2020;235(6):5059–5071. doi:10.1002/jcp.29442
12. Ni H, Niu LL, Tian SC, et al. Long non-coding RNA LINC00152 is up-regulated in ovarian cancer tissues and regulates proliferation and cell cycle of SKOV3 cells. Eur Rev Med Pharmacol Sci. 2019;23(22):9803–9813.
13. Dai L, Niu JX, Feng YL. Knockdown of long non-coding RNA LINC00176 suppresses ovarian cancer progression by BCL3-mediated down-regulation of ceruloplasmin. J Cell Mol Med. 2020;24(1):202–213. doi:10.1111/jcmm.14701
14. Deb B, Uddin A, Chakraborty S. miRNAs and ovarian cancer: an overview. J Cell Physiol. 2018;233(5):3846–3854. doi:10.1002/jcp.26095
15. Li J, Zhang SL, Wu L, Pei ML. Interaction between LncRNA-ROR and miR-145 contributes to epithelial-mesenchymal transition of ovarian cancer cells. Gen Physiol Biophys. 2019;38(6):461–471. doi:10.4149/gpb_2019028
16. Zhang PF, Wu J, Luo JH, et al. SNHG22 overexpression indicates poor prognosis and induces chemotherapy resistance via the miR-2467/Gal-1 signaling pathway in epithelial ovarian carcinoma. Aging-Us. 2019;11(19):8204–8216. doi:10.18632/aging.102313
17. Sun Q, Li Q, Xie FF. LncRNA-MALAT1 regulates proliferation and apoptosis of ovarian cancer cells by targeting miR-503-5p. Onco Targets Ther. 2019;12:6297–6307. doi:10.2147/OTT.S214689
18. Schaukowitch K, Kim TK. Emerging epigenetic mechanisms of long non-coding RNAs. Neuroscience. 2014;264:25–38. doi:10.1016/j.neuroscience.2013.12.009
19. Gong D, Feng PC, Ke XF, et al. Silencing long non-coding RNA LINC01224 inhibits hepatocellular carcinoma progression via microRNA-330-5p-induced inhibition of CHEK1. Mol Ther Nucleic Acids. 2019;19:482–497. doi:10.1016/j.omtn.2019.10.007
20. Li J, Xu J, Yan X, Jin K, Li W, Zhang R. MicroRNA-485 plays tumour-suppressive roles in colorectal cancer by directly targeting GAB2. Oncol Rep. 2018;40(1):554–564. doi:10.3892/or.2018.6449
21. Hu XX, Xu XN, He BS, et al. microRNA-485-5p functions as a tumor suppressor in colorectal cancer cells by targeting CD147. J Cancer. 2018;9(15):2603–2611. doi:10.7150/jca.24918
22. Jing LL, Mo XM. Reduced miR-485-5p expression predicts poor prognosis in patients with gastric cancer. Eur Rev Med Pharmacol Sci. 2016;20(8):1516–1520.
23. Mao K, Lei D, Zhang H, You C. MicroRNA-485 inhibits malignant biological behaviour of glioblastoma cells by directly targeting PAK4. Int J Oncol. 2017;51(5):1521–1532. doi:10.3892/ijo.2017.4122
24. Wang ZQ, Zhang MY, Deng ML, Weng NQ, Wang HY, Wu SX. Low serum level of miR-485-3p predicts poor survival in patients with glioblastoma. PLoS One. 2017;12(9):e0184969. doi:10.1371/journal.pone.0184969
25. Guo GX, Li QY, Ma WL, Shi ZH, Ren XQ. MicroRNA-485-5p suppresses cell proliferation and invasion in hepatocellular carcinoma by targeting stanniocalcin 2. Int J Clin Exp Pathol. 2015;8(10):12292–12299.
26. Sun X, Liu Y, Li M, Wang M, Wang Y. Involvement of miR-485-5p in hepatocellular carcinoma progression targeting EMMPRIN. Biomed Pharmacother. 2015;72:58–65. doi:10.1016/j.biopha.2015.04.008
27. Mou X, Liu S. MiR-485 inhibits metastasis and EMT of lung adenocarcinoma by targeting Flot2. Biochem Biophys Res Commun. 2016;477(4):521–526. doi:10.1016/j.bbrc.2016.04.043
28. Lou C, Xiao M, Cheng S, et al. MiR-485-3p and miR-485-5p suppress breast cancer cell metastasis by inhibiting PGC-1α expression. Cell Death Dis. 2016;7(3):e2159. doi:10.1038/cddis.2016.27
29. Anaya-Ruiz M, Bandala C, Perez-Santos JL. miR-485 acts as a tumor suppressor by inhibiting cell growth and migration in breast carcinoma T47D cells. APJCP. 2013;14(6):3757–3760. doi:10.7314/APJCP.2013.14.6.3757
30. Chen Z, Li Q, Wang S, Zhang J. miR4855p inhibits bladder cancer metastasis by targeting HMGA2. Int J Mol Med. 2015;36(4):1136–1142. doi:10.3892/ijmm.2015.2302
31. Wang H, Fu Z, Dai C, et al. LncRNAs expression profiling in normal ovary, benign ovarian cyst and malignant epithelial ovarian cancer. Sci Rep. 2016;6:38983. doi:10.1038/srep38983
32. Gao Y, Meng H, Liu S, et al. LncRNA-HOST2 regulates cell biological behaviors in epithelial ovarian cancer through a mechanism involving microRNA let-7b. Hum Mol Genet. 2015;24(3):841–852. doi:10.1093/hmg/ddu502
33. Sheng X, Li J, Yang L, et al. Promoter hypermethylation influences the suppressive role of maternally expressed 3, a long non-coding RNA, in the development of epithelial ovarian cancer. Oncol Rep. 2014;32(1):277–285. doi:10.3892/or.2014.3208
34. Qiu JJ, Wang Y, Liu YL, Zhang Y, Ding JX, Hua KQ. The long non-coding RNA ANRIL promotes proliferation and cell cycle progression and inhibits apoptosis and senescence in epithelial ovarian cancer. Oncotarget. 2016;7(22):32478–32492. doi:10.18632/oncotarget.8744
35. Wang L, Wang C, Wu T, Sun F. Long non-coding RNA TP73-AS1 promotes TFAP2B-mediated proliferation, metastasis and invasion in retinoblastoma via decoying of miRNA-874-3p. J Cell Commun Signal. 2020. doi:10.1007/s12079-020-00550-x
36. Wang W, Hu W, Wang Y, et al. Long non-coding RNA UCA1 promotes malignant phenotypes of renal cancer cells by modulating the miR-182-5p/DLL4 axis as a ceRNA. Mol Cancer. 2020;19(1):18. doi:10.1186/s12943-020-1132-x
37. Yu Y, Gao F, He Q, Li G, Ding G. lncRNA UCA1 functions as a ceRNA to promote prostate cancer progression via sponging miR143. Mol Ther Nucleic Acids. 2019;19:751–758. doi:10.1016/j.omtn.2019.11.021
38. Ye Y, Shen A, Liu A. Long non-coding RNA H19 and cancer: A competing endogenous RNA. Bull Cancer. 2019;106(12):1152–1159. doi:10.1016/j.bulcan.2019.08.011
39. Bokoch GM. Biology of the p21-activated kinases. Annu Rev Biochem. 2003;72(1):743–781. doi:10.1146/annurev.biochem.72.121801.161742
40. Siu MK, Chan HY, Kong DS, et al. p21-activated kinase 4 regulates ovarian cancer cell proliferation, migration, and invasion and contributes to poor prognosis in patients. Proc Natl Acad Sci U S A.;107(43):18622–18627. doi:10.1073/pnas.0907481107
41. Luo P, Fei J, Zhou J, Zhang W. microRNA-126 suppresses PAK4 expression in ovarian cancer SKOV3 cells. Oncol Lett. 2015;9(5):2225–2229. doi:10.3892/ol.2015.3012
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.