Back to Journals » Infection and Drug Resistance » Volume 12
Investigation of six plasmid-mediated quinolone resistance genes among clinical isolates of pseudomonas: a genotypic study in Saudi Arabia
Authors El-Badawy MF, Alrobaian MM, Shohayeb MM, Abdelwahab SF
Received 29 January 2019
Accepted for publication 5 March 2019
Published 29 April 2019 Volume 2019:12 Pages 915—923
DOI https://doi.org/10.2147/IDR.S203288
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Mohamed F El-Badawy,1,2 Majed M Alrobaian,3 Mohamed M Shohayeb,1,4 Sayed F Abdelwahab1,5
1Division of Pharmaceutical Microbiology, Department of Pharmaceutics and Industrial Pharmacy, College of Pharmacy, Taif University, Taif 21974, Kingdom of Saudi Arabia; 2Department of Microbiology and Immunology, Faculty of Pharmacy, Misr University for Science and Technology, Al-Motamayez District 12568, Egypt; 3Department of Pharmaceutics and Industrial Pharmacy, College of Pharmacy, Taif University, Taif 21974, Kingdom of Saudi Arabia; 4Department of Microbiology and Biotechnology, Faculty of Pharmacy, Delta University for Science and Technology, Gamasa 35712, Egypt; 5Department of Microbiology and Immunology, Faculty of Medicine, Minia University, Minia, 61511, Egypt
Background: Quinolones are among the most effective antibiotics against Pseudomonas spp. Several chromosomal and/or plasmid-mediated quinolone-resistance mechanisms have been found in Pseudomonas. Plasmid-mediated quinolone-resistance (PMQR) is mediated by quinolone-resistance (QNR) proteins, modifying enzymes or efflux pumps. Only a few previous studies examined the prevalence of quinolone-resistance in the Kingdom of Saudi Arabia (KSA) and showed it is increasing. Mechanisms of quinolone-resistance among Pseudomonas spp. in the KSA; examined herein; have not been extensively studied.
Methods: Ninety-two Pseudomonas isolates were collected and their resistance to seven different types of quinolones was determined by the microbroth dilution method. PMQR mechanisms were examined using a PCR screen to identify six PMQR genes including qnrA, qnrB, qnrD, qnrS, aac(6´)-Ib-cr, and qepA. Clonal relatedness of the quinolone-resistant isolates was determined by ERIC-PCR.
Results: Of the isolates, 42.4% (39/92) were resistant to 1-7 of the tested quinolones. Gemifloxacin resistance was the lowest (28.3%) while resistance to the other six quinolones were ≥ 35%. The most common biotype among the 39 quinolone-resistant isolates was resistance to the seven tested quinolones (26/39; 66.7%). qnrD, qnrS, and aac(6´)-Ib-cr were found in 31 (79.5%), 31 (79.5%) and 28 (71.8%) of the 39 isolates, respectively, and all three genes together were found in 22 of the 39 isolates (56.4%). qnrA, qnrB, and qepA were not detected in any of the isolates and two isolates did not harbor any of the six tested genes. The isolates showed 38 different ERIC profiles and only two isolates (Pa16 and Pa17) had an identical profile.
Conclusion: This is the first description of PMQR mechanisms among clinical Pseudomonas isolates from the KSA, which appears to be mainly mediated by qnrD, qnrS, and aac(6´)-Ib-cr.
Keywords: aac(6´)-Ib-cr, flouoroquinolones, Pseudomonas, qepA, qnr, QRDR, quinolones, aac(6´)-Ib-cr, flouoroquinolones, qepA, KSA, qnr, QRDR, Taif
Introduction
Pseudomonas spp. infections are problematic due to their ability to resist most classes of antibiotics. Few studies have been published regarding the prevalence of P. aeruginosa and its resistance to quinolones in the Kingdom of Saudi Arabia (KSA). Related to this, there is a substantial problem in intensive care units (ICU) among patients infected by multi-drug resistant (MDR) Pseudomonas isolates, reviewed elsewhere.1 Infections with MDR Pseudomonas lead to increased healthcare costs, longer hospital stays, and increased mortality rates.
Fluoroquinolones are among the most effective antibiotics against Pseudomonas spp. However, Pseudomonas has acquired several resistance mechanisms against several different types of antibiotics including quinolones. The activity and spectrum of the newer quinolones are much higher than older drug generations with increased microbial resistance observed for the lower generations.2 There are four known quinolone resistance mechanisms that work separately or in combination in Pseudomonas resulting in varying degrees of resistance, ranging from reduced susceptibility to clinically relevant resistance. These mechanisms may be chromosomally- or plasmid-mediated.3
Chromosomal resistance to quinolones may occur due to target site mutation; ie, alterations in the target enzyme (DNA gyrase and/or DNA topoisomerase IV), alteration in membrane permeability and active efflux.4,5 The short mutated DNA sequence located in the gyrase and topoisomerase IV genes are responsible for fluoroquinolone resistance; and is known as the quinolone resistance-determining region (QRDR).6,7 In addition to the role of chromosomally-mediated resistance to quinolones, plasmids also play a pivotal role in resistance to quinolones. The term “resistance” in the context of plasmid-mediated quinolone resistance (PMQR) is used to refer to any increase in MIC, rather than an increase above a susceptibility breakpoint, as reviewed elsewhere.8,9 PMQR is more important in the horizontal spread of resistance between different isolates and different species further complicating the issue of microbial drug resistance. PMQR is mediated mainly by the quinolone resistance (qnr) genes. Three mechanisms are responsible for PMQR: target alteration by qnr gene products, drug modification by the aminoglycoside acetyltransferase “AAC(6′)-Ib-cr”; that is capable of reducing ciprofloxacin activity, and acquiring an efflux pump mechanism mediated by two reported quinolone efflux pumps known as olaquindox (OqxAB) and the QepA quinolone efflux plasmid.9,10
Several reports examined the prevalence of quinolone resistance in the KSA and showed that it is increasing.11–18 However, none of these studies were performed in the Taif area, which is located in the western part of the KSA. Importantly, very little is known about the mechanisms of quinolone resistance among Pseudomonas spp. in the KSA. The main objective of this study was to identify the potential genes that confer PMQR mechanisms among clinical isolates of Pseudomonas spp. and determine the clonal relatedness between quinolone-resistant isolates by ERIC-PCR. This study shows that PMQR among clinical Pseudomonas isolates from Taif, KSA, is mainly mediated by qnrD, qnrS, and aac(6´)-Ib-cr.
Subjects and methods
Bacterial isolation and identification
The Pseudomonas spp. isolates included in this study were collected from December 2016 to April 2017. They were isolated as a part of routine hospital laboratory procedures and were further identified and confirmed in the laboratory. The study protocol was approved by Taif University Research Ethics Committee (approval #38-35-0021). Ninety-two non-duplicate non-consecutive clinical isolates of Pseudomonas spp. were recovered from 92 cases. Of these cases, 56 and 36 were male (60.9%) or female (39.1%), respectively, with ages ranging between two months and 102 years, with an average age of 53.0±25.2 years. These patients were admitted to or attended different medical departments at a tertiary care hospital in Taif, KSA. The clinical isolates were recovered from different clinical specimens, namely, axillary swab (n=1), bile (n=4), blood (n=7), catheter tip (n=4), eye swab (n=3), peritoneal fluid (n=5), sputum (n=30), tracheal aspirate (n=2), urine (n=20), vaginal swab (n=2), and wound swab (n=14).
All strains were isolated primarily on MacConkey’s agar (Oxoid, UK) and blood agar and were then purified on cetrimide agar (Scharlu, Spain). Genus level identification was determined by amplification of the algD gene using the primers listed in Table 1 (Macrogen, Geumcheon-gu, Seoul, Korea) and a Master cycler® personal PCR machine (Eppendorf, California, USA). Species level identification of the isolates was determined by Vitek (Biomeriux, France).
 | Table 1 Primer sets and PCR cycling conditions used for genotyping and amplification of genes encoding quinolone resistance |
Antimicrobial susceptibility testing
All bacterial isolates were subjected to antimicrobial susceptibility testing by the broth microdilution method using cation modified Mueller-Hinton broth (Oxoid, UK) in 96 well microtiter plates. Twelve different dilutions of each quinolone were tested by the two-fold dilution method (concentrations tested ranged from 1,024 μg/ml to 0.5 μg/ml). Seven quinolone antibiotics, representing the four generations of quinolones, were tested, which included nalidixic acid (NAL), representing the first generation; ciprofloxacin (CIP), norfloxacin (NOR), and ofloxacin (OFL), representing the second generation; levofloxacin (LEV), representing the third generation, and gemifloxacin (GEM) and moxifloxacin (MOX), representing the fourth generation (all from Sigma-Aldrich, USA). P. aeruginosa ATCC27853 was used as a quality control strain. Results were interpreted according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI).19 Since the CLSI guidelines do not specify a breakpoint for gemifloxacin, its breakpoint was based on the values proposed by the British Society for Antimicrobial Chemotherapy (≤0.5 mg/L for susceptible, and ≥1 mg/L for resistant).20 Also, moxifloxacin data were interpreted based on European Committee on Antimicrobial Susceptibility Testing (EUCAST) e-coff and modal MIC, which, for Pseudomonas, are 4 μg/ml and 1 μg/ml, respectively.21
Preparation of DNA templates for PCR testing
DNA was extracted as previously described.22 PCR reactions were performed in a final reaction volume of 20µl. Reactions contained 4µl of extracted DNA, 4 µl of 5x master mix (HOT FIREPol® Blend Master Mix, Solis BioDyne, Tartu, Estonia), 0.6 µl of forward primer (10 pmol/μl), 0.6 µl of reverse primer (10 pmol/μl) and 10.8 µl distilled water.
Molecular detection of plasmid-mediated quinolone resistance
Clinical Pseudomonas isolates that were resistant to any of the tested quinolones were screened for six quinolone resistance genes: qnrA, qnrB, qnrD, and qnrS, the quinolone efflux gene qepA, and the quinolone modifying enzyme gene; acc(6ʹ)-1b-cr. PCR primers (Macrogen) and cycling conditions are listed in Table 1.
Genotyping and fingerprint analysis of clinical isolates
Clonal relatedness between quinolone-resistant clinical isolates of Pseudomonas was determined by enterobacterial repetitive intergenic consensus (ERIC)-PCR. The primer was obtained from Macrogen and PCR amplification was conducted according to the cycling conditions described in Table 1. Banding patterns generated by ERIC-PCR were analyzed using BioNumerics 7.5 software (Applied Maths, Kortrijk, Belgium). PCR fingerprint profiles were analyzed using Dice (similarity) coefficient. Cluster analysis was performed as previously described,23 based on the unweighted pair group method with arithmetic averages (UPGMA) at a position tolerance of 0.15.
Results
Isolation and identification of clinical Pseudomonas isolates
All recovered isolates were found to be related to the genus Pseudomonas based on amplification of the algD gene (data not shown). Phenotypic identification at the species level using the Vitek® 2 GN ID card revealed that 83 of the 92 isolates (90.2%) were P. aeruginosa, three isolates each were P. putida, and P. fluorescence (3.3% each), and there was one isolate (1.1%) each of P. luteola, P. stutzuri, and P. oryzihabitans.
Antimicrobial susceptibility of Pseudomonas isolates to different quinolones
The first step in examining the mechanisms of quinolone resistance was to test the susceptibility of the Pseudomonas isolates to seven selected quinolones by the microbroth dilution method. Susceptibility testing revealed that 39 isolates (42.4%) were highly resistant to at least one of the tested quinolones, while the remaining isolates were susceptible to all quinolones. The 39 resistant isolates were recovered from different clinical specimens, namely, axillary swab (n=1), bile (n=2), blood (n=4), catheter tip (n=1), sputum (n=20), urine (n=6), and wound (n=5).
Nalidixic acid was the least effective quinolone against Pseudomonas spp.; 41.3% (38/92) of the isolates were resistant. Gemifloxacin, ciprofloxacin, and levofloxacin were the most effective agents, in which 28.3% (26/92), 35.9%, and 35.9% (33/92) of the isolates were resistant, respectively (Table 2). A similar resistance rate (37.0%; 34 of the 92 isolates) was observed for norfloxacin and ofloxacin. The rate of moxifloxacin resistance was 39.1% (33/92).
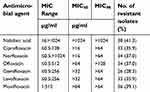 | Table 2 Percentage of resistance of clinical Pseudomonas isolates to different quinolones |
The MIC50 and MIC90 of the seven tested quinolones are shown in Table 2. Nalidixic acid had the highest MIC50 and MIC90 (>1024 μg/ml). Ciprofloxacin and norfloxacin had the lowest MIC50 and MIC90 (>16 μg/ml and > 64μg/ml, respectively).
Based on the quinolone resistance data, the 39 quinolone-resistant Pseudomonas isolates were classified into seven different quinolone-resistant biotypes, as shown in Table 3. The most common biotype was resistant to all the tested quinolones, corresponding to 66.7% of the isolates (26/39). The second most common quinolone-resistant biotype was resistant to six quinolones: NAL, CIP, NOR, OFL, LEV, and MOX, corresponding to 15.4% of the isolates (6/39). The remaining biotypes are shown in Table 3.
 | Table 3 Resistance pattern and genetic profile of quinolone resistant Pseudomonas spp. isolates |
Detection of PMQR genes among the quinolone-resistant Pseudomonas isolates
PCR was carried out to screen for the presence of the PMQR genes (qnrA, qnrB, qnrD, qnrS, qepA, and aac(6´)-Ib-cr) among the 39 quinolone-resistant Pseudomonas isolates. As shown in Table 3, qnrD, qnrS, and aac(6´)-Ib-cr were found in 31, 31 (79.5% each), and 28 (71.8%) of the 39 isolates, respectively. qnrA, qnrB and qepA were not detected and two isolates did not harbor any of the six tested genes. Additionally, qnrD, qnrS and aac(6´)-Ib-cr were collectively found in 22 of the isolates (56.4%). In addition, four of the isolates (10.2%) contained both qnrD, and qnrS. On the other hand, 5.1% (2/39) of the isolates did not contain any of the six tested PMQR genes (Table 3). The average number of PMQR genes detected among the resistant isolates was 2.3±0.9.
Fingerprint pattern analysis
Clonal relatedness of the 39 quinolone-resistant clinical isolates of Pseudomonas spp. was determined by ERIC-PCR. Representative ERIC-PCR results are shown in Figure 1. The UPGMA dendrogram at 85% similarity (Figure 2) demonstrated that the 39 Pseudomonas isolates corresponded to 38 different ERIC profiles in which only two isolates (Pa16 and Pa17) had identical profiles. The dendrogram classified the 39 isolates into two main phylogenetic groups: phylogenetic group A (PGA) and phylogenetic group B (PGB). The majority of the 39 isolates (94.8%) were categorized in PGB (Figure 2). PGB was further divided into three sub-phylogenetic groups: PGB1.1, PGB1.2, and PGB2. The major sub-phylogenetic group of PGB was PGB1.2, in which 59.5% (22/37) of the isolates clustered.
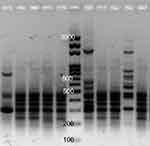 | Figure 1 A representative example of ERIC-PCR pattern for clinical Pseudomonas isolates. There are five isolates on either side of the ladder (middle lane) on the gel. |
Discussion
In this study, 92 different Pseudomonas isolates were collected from patients in a tertiary care hospital in Taif, KSA. Thirty-nine of the isolates (42.4%) were resistant to at least one quinolone. Of the seven tested quinolones, gemifloxacin was the most effective, as only 28.3% of isolates were resistant. The rate of resistance to the other six tested quinolones was ≥35.9%. The most common quinolone resistance biotype was resistant to the seven tested quinolones (66.7%; 26/39). The high fluoroquinolone resistance rate in the current study prompted us to investigate the prevalence of the PMQR genes among the quinolone-resistant isolates. qnrD, and qnrS were found in 79.5% of the isolates (31/39) and aac(6´)-Ib-cr was found in 71.8% of the resistant isolates (28/39). qnrA, qnrB, and qepA were not detected in any of the isolates. Two isolates did not harbor any of the six tested genes. Interestingly, qnrD, qnrS, and aac(6´)-Ib-cr were found together in 22 of the 39 quinolone-resistant isolates (56.4%). In addition, the isolates showed 38 different ERIC profiles and only two isolates (Pa16 and Pa17) had an identical profile. Several aspects of these data deserve further discussion.
First, antibiotic resistance represents a major health care problem due to increased mortality and longer hospital stays, which increase the cost of healthcare service provided to patients. In view of the growing number of patients with nosocomial infections worldwide, including the KSA, and the limited number of treatment options for infections caused by Pseudomonas spp., patients and their doctors are searching for effective therapies. This is the first study to address the possible role of PMQR among Pseudomonas spp. in the KSA. This study is also of interest since, to the best of our knowledge, no other studies have evaluated quinolone resistance in the Taif area, located in the western part of the KSA. The prevalence of quinolone resistance among the Pseudomonas spp. isolated in Taif during the study period showed that approximately one third or more of the isolates were resistant to at least one quinolone, which is a high proportion that limits the usage of fluoroquinolones to treat patients infected with Pseudomonas spp.
Second, 39 of the isolates (42.4%) were resistant to at least one quinolone. Gemifloxacin was the most effective quinolone (28.3% resistance rate), while the resistance rate to the other six quinolones was ≥35.9%. These data are consistent with previous findings that showed an escalation in quinolone resistance over time in the KSA.11–18 In this regard, the MIC50 and MIC90 of nalidixic acid were the highest among the tested quinolones, both measuring >1,024 μg/ml. The nalidixic acid resistance rate recorded herein is relatively low (41.3%) as compared to other reports from the KSA,18,24 suggesting that limiting the use of a certain agent can lead to a decrease in the resistance rate. Resistance to all seven tested quinolones was the most common quinolone resistance biotype, corresponding to two-thirds (26/39) of the isolates. This suggests a horizontal transfer of plasmid-mediated resistance genes between different strains.
Third, the genetic basis of PMQR among the isolated Pseudomonas spp. was determined by screening for three different resistance mechanisms using PCR, including efflux mechanisms (qepA), modifying enzymes (aac(6´)-Ib-cr), and qnr genes (qnrA, qnrB, qnrD, and qnrS). qnrD, qnrS, and aac(6´)-Ib-cr were found in 31, 31 (79.5% each) and 28 (71.8%) of the 39 isolates, respectively, while qnrA, qnrB, and qepA were not detected. A similar finding regarding qepA was recently reported in a Turkish study.25 Also, qnrD, qnrS, and aac(6´)-Ib-cr were found together in 22 (56.4%) of the 39 quinolone-resistant isolates. These data and those presented in Table 3 suggest that these clinical Pseudomonas isolates possess several resistance mechanisms that can lead to quinolone resistance. In contrast, two isolates did not harbor any of the six tested genes, suggesting that chromosomal resistance or another PMQR gene(s) not-investigated in this study may encode the mechanisms responsible for quinolone resistance in these two isolates. A recent study in Egypt reported the presence of qnrB, and qnrS in 1.8% and 2.7% of Pseudomonas isolates, respectively. The authors did not find any other PMQR genes among the isolates.26 Although the general findings of this study are similar to our findings, the frequency of PMQR gene detection is much lower than in our study. Another study in China reported the detection of qnrA in only one ciprofloxacin resistant isolate.27 Also, qnrS, and aac(6´)-Ib-cr were found in only one (2.6%) of 38 P. aeruginosa isolates in Brazil.28 On the other hand, qnrB was found in 20% of Pseudomonas isolates in a study conducted in Poland,29 while qnrS was found in 21% of the isolates in a study conducted in Iraq.30 These rates are similar to our findings. Together, these data show that PMQR genes have been found worldwide in Pseudomonas isolates at different frequencies.
Fourth, different typing methods are used in epidemiologic studies of infectious agents (including Pseudomonas), which include phage typing, serologic testing, pyocyanin typing, and DNA fingerprinting. However, no system is universally accepted.31 ERIC–PCR, which is based on amplification of highly conserved 126-bp non-coding regions, is used for molecular typing of different microorganisms.32 These conserved regions consist of imperfect palindromes that occur in multiple copies in the genomes of enteric bacteria and vibrios.33–36 Molecular typing using ERIC-PCR provides more discriminative species-specific DNA patterns) that cannot be detected by pulsed-field gel electrophoresis.37,38 In our study, the 39 quinolone-resistant isolates showed 38 different ERIC profiles while only two isolates (Pa16 and Pa17) had identical profiles. These heterogeneous ERIC profiles suggest that different strains of Pseudomonas spp. were introduced into hospital settings, indicating poor adherence to infection control measures during the study period.
Fifth, one of the strengths of this study wat that it focused on strains that were already resistant to at least one of the quinolones/fluoroquinolones. In contrast, one of the limitations of this study is that we investigated the possible role of PMQR but did not investigate the possible role of chromosomally-mediated resistance mechanisms, which may be mediated by either overexpression of active efflux pumps and/or mutations in QRDR. This is currently under investigation.
In conclusion, PMQR among clinical Pseudomonas isolates recovered from Taif in the western region of the KSA is mainly mediated by qnrD, qnrS, and aac(6´)-Ib-cr. Co-existence of more than one PMQR gene could be responsible for the high-level of fluoroquinolone resistance and horizontal spread of quinolone-resistance can lead to reduced therapeutic options for infections caused by Pseudomonas spp. This study increases our understanding of how Pseudomonas spp. resist quinolones, which could contribute to better management of patients that acquire nosocomial infections of this bacterium in the KSA.
Acknowledgments
This study was funded by the Deanship of Scientific Research, Taif University, KSA (Research Project No. 1-437-5292 to S.F. Abdelwahab). The sponsor did not have any role in the design of the study; the collection, analysis and interpretation of data; the writing of the manuscript; and in the decision to submit the article for publication.
Author contributions
All authors contributed toward data analysis, drafting and revising the paper, gave final approval of the version to be published and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Yezli S, Shibl AM, Livermore DM, Memish ZA. Prevalence and antimicrobial resistance among Gram-negative pathogens in Saudi Arabia. J Chemother. 2014;26(5):257–272. doi:10.1179/1973947814Y.0000000185
2. Oliphant CM, Green GM. Quinolones: a comprehensive review. Am Fam Physician. 2002;65(3):455–464.
3. Guan X, Xue X, Liu Y, et al. Plasmid-mediated quinolone resistance–current knowledge and future perspectives. J Int Med Res. 2013;41(1):20–30.
4. Redgrave LS, Sutton SB, Webber MA, Piddock LJ. Fluoroquinolone resistance: mechanisms, impact on bacteria, and role in evolutionary success. Trends Microbiol. 2014;22(8):438–445. doi:10.1016/j.tim.2014.04.007
5. Hooper DC, Jacoby GA. Mechanisms of drug resistance: quinolone resistance. Ann N Y Acad Sci. 2015;1354(1):12–31. doi:10.1111/nyas.12830
6. Yoshida H, Bogaki M, Nakamura M, Nakamura S. Quinolone resistance-determining region in the DNA gyrase gyrA gene of Escherichia coli. Antimicrob Agents Chemother. 1990;34(6):1271–1272.
7. Yoshida H, Bogaki M, Nakamura M, Yamanaka LM, Nakamura S. Quinolone resistance-determining region in the DNA gyrase gyrB gene of Escherichia coli. Antimicrob Agents Chemother. 1991;35(8):1647–1650.
8. Rodriguez-Martinez JM, Machuca J, Cano ME, Calvo J, Martinez-Martinez L, Pascual A. Plasmid-mediated quinolone resistance: two decades on. Drug Resist Updat. 2016;29:13–29. doi:10.1016/j.drup.2016.09.001
9. Strahilevitz J, Jacoby GA, Hooper DC, Robicsek A. Plasmid-mediated quinolone resistance: a multifaceted threat. Clin Microbiol Rev. 2009;22(4):664–689. doi:10.1128/CMR.00016-09
10. Martínez-Martínez L, Cano ME, Rodríguez-Martínez JM, Calvo J, Pascual Á. Plasmid-mediated quinolone resistance. Expert Rev Anti Infect Ther. 2014;6(5):685–711.
11. Memish ZA, Shibl AM, Kambal AM, Ohaly YA, Ishaq A, Livermore DM. Antimicrobial resistance among non-fermenting Gram-negative bacteria in Saudi Arabia. J Antimicrob Chemother. 2012;67(7):1701–1705. doi:10.1093/jac/dks091
12. Saeed NK, Kambal AM, El-Khizzi NA. Antimicrobial-resistant bacteria in a general intensive care unit in Saudi Arabia. Saudi Med J. 2010;31(12):1341–1349.
13. Babay HA. Antimicrobial resistance among clinical isolates of Pseudomonas aeruginosa from patients in a teaching hospital, Riyadh, Saudi Arabia, 2001-2005. Jpn J Infect Dis. 2007;60(2–3):123–125.
14. Al-Tawfiq JA. Occurrence and antimicrobial resistance pattern of inpatient and outpatient isolates of Pseudomonas aeruginosa in a Saudi Arabian hospital: 1998–2003. Int j infect dis. 2007;11(2):109–114. doi:10.1016/j.ijid.2005.11.004
15. Al-Harthi AA, Al-Fifi SH. Antibiotic resistance pattern and empirical therapy for urinary tract infections in children. Saudi Med J. 2008;29(6):854–858.
16. Al-Tawfiq JA, Abed MS. Prevalence and antimicrobial resistance of health care associated bloodstream infections at a general hospital in Saudi Arabia. Saudi Med J. 2009;30(9):1213–1218.
17. Akhtar N, Alqurashi AM, Abu Twibah M. In vitro ciprofloxacin resistance profiles among gram-negative bacteria isolated from clinical specimens in a teaching hospital. JPMA J Pak Med Assoc. 2010;60(8):625–627.
18. Alamri A, Hamid ME, Abid M, et al. Trend analysis of bacterial uropathogens and their susceptibility pattern: a 4-year (2013–2016) study from Aseer region, Saudi Arabia. Urology Annals. 2018;10(1):41.
19. Cockerill FR. Performance standards for antimicrobial susceptibility testing: Twenty-first informational supplement. Wayne, PA: Clinical and Laboratory Standards Institute (CLSI); 2011.
20. Wise R, Andrews JM. The in-vitro activity and tentative breakpoint of gemifloxacin, a new fluoroquinolone. J Antimicrob Chemother. 1999;44(5):679–688.
21. Grillon A, Schramm F, Kleinberg M, Jehl FJPO. Comparative activity of ciprofloxacin, levofloxacin and moxifloxacin against Klebsiella pneumoniae, Pseudomonas aeruginosa and Stenotrophomonas maltophilia assessed by minimum inhibitory concentrations and time-kill studies. PLoS One. 2016;11(6):e0156690. doi:10.1371/journal.pone.0156690
22. El-Badawy MF, Tawakol WM, El-Far SW, et al. molecular identification of aminoglycoside-modifying enzymes and plasmid-mediated quinolone resistance genes among Klebsiella pneumoniae clinical isolates recovered from Egyptian Patients. Int J Microbiol. 2017;2017:8050432. doi:10.1155/2017/8050432
23. El-Badawy MF, Tawakol WM, Maghrabi IA, Mansy MS, Shohayeb MM, Ashour MS. Iodometric and molecular detection of ESBL production among clinical isolates of E. coli fingerprinted by ERIC-PCR: the first Egyptian report declares the emergence of E. coli O25b-ST131clone Harboring blaGES. Microb Drug Resist. 2017;23(6):703–717. doi:10.1089/mdr.2016.0181
24. Asghar AH, Faidah HS. Frequency and antimicrobial susceptibility of gram-negative bacteria isolated from 2 hospitals in Makkah, Saudi Arabia. Saudi Med J. 2009;30(8):1017–1023.
25. Cayci YT, Coban AY, Gunaydin M. Investigation of plasmid-mediated quinolone resistance in Pseudomonas aeruginosa clinical isolates. Indian J Med Microbiol. 2014;32(3):285–289. doi:10.4103/0255-0857.136567
26. Saleh MA, Balboula MM. Plasmid mediated quinolone resistance determinants among nosocomial clinical Pseudomonas aeruginosa isolates. Int Curr Microbiol App Sci.2017;6(1):42–50.
27. Yang X, Xing B, Liang C, Ye Z, Zhang Y. Prevalence and fluoroquinolone resistance of pseudomonas aeruginosa in a hospital of South China. Int J Clin Exp Med. 2015;8(1):1386–1390.
28. Araujo BF, Ferreira ML, de Campos PA, et al. Clinical and Molecular Epidemiology of Multidrug-Resistant P. aeruginosa Carrying aac (6’)-Ib-cr, qnrS1 and blaSPM Genes in Brazil. 2016;11(5):e0155914.
29. Michalska AD, Sacha PT, Ojdana D, Wieczorek A, Tryniszewska E. Prevalence of resistance to aminoglycosides and fluoroquinolones among Pseudomonas aeruginosa strains in a University Hospital in Northeastern Poland. BJM 2014;45(4):1455–1458.
30. Al-Marjani MF. Presence of qnr gene in environmental and clinical Pseudomonas aeruginosa isolates in Baghdad. Int J Curr Microbiol Appl Sci 2014;3(7):853–857.
31. Baron S. Epidemiology–Medical Microbiology. Galveston, TX: University of Texas Medical Branch at Galveston; 1996.
32. Meacham KJ, Zhang L, Foxman B, Bauer RJ, Marrs CF. Evaluation of genotyping large numbers of Escherichia coli isolates by enterobacterial repetitive intergenic consensus-PCR. J Clin Microbiol. 2003;41(11):5224–5226.
33. Wilson LA, Sharp PM. Enterobacterial repetitive intergenic consensus (ERIC) sequences in Escherichia coli: evolution and implications for ERIC-PCR. Mol Biol Evol. 2006;23(6):1156–1168. doi:10.1093/molbev/msj125
34. Tripathi MK, Kumar M, Singh VK, Nath G, Asthana RK. Use of enterobacterial repetitive intergenic consensus PCR in detecting target (s) of Hapalindole-T, From a Cyanobacterium, in Escherichia Coli: in Silico Validation. Am J Bioinf. 2012;1(1):30–40.
35. Hulton CS, Higgins CF, Sharp PM. ERIC sequences: a novel family of repetitive elements in the genomes of Escherichia coli, Salmonella typhimurium and other enterobacteria. Mol Microbiol. 1991;5(4):825–834.
36. Asgarani E, Ghashghaei T, Soudi MR, Alimadadi N. Enterobacterial repetitive intergenic consensus (ERIC) PCR based genetic diversity of Xanthomonas spp. and its relation to xanthan production. Iran J Microbiol. 2015;7(1):38–44.
37. Waturangi DE, Joanito I, Yogi Y, Thomas S. Use of REP- and ERIC-PCR to reveal genetic heterogeneity of Vibrio cholerae from edible ice in Jakarta, Indonesia. Gut Pathog. 2012;4(1):2. doi:10.1186/1757-4749-4-2
38. Ishii S, Sadowsky MJ. Applications of the rep-PCR DNA fingerprinting technique to study microbial diversity, ecology and evolution. Environ Microbiol. 2009;11(4):733–740. doi:10.1111/j.1462-2920.2008.01856.x
39. Sun F, Yang S, Chen X. Detection of intestinal pathogenic klebsiella pneumoniae from fecal samples of giant panda by polymerase chain reaction. J Biotech Res. 2013;5:10.
40. Da Silva Filho LVF, Levi JE, Bento CN, Ramos SR, Rozov T. PCR identification of Pseudomonas aeruginosa and direct detection in clinical samples from cystic fibrosis patients. J Med Microbiol. 1999;48(4):357–361. doi:10.1099/00222615-48-4-357
41. Cattoir V, Poirel L, Rotimi V, Soussy C-J, Nordmann P. Multiplex PCR for detection of plasmid-mediated quinolone resistance qnr genes in ESBL-producing enterobacterial isolates. J antimicrob chemother. 2007;60(2):394–397. doi:10.1093/jac/dkm204
42. Pribul BR, Festivo ML, Souza MM, Rodrigues DD. Characterization of quinolone resistance in Salmonella spp. isolates from food products and human samples in Brazil. braz J microbiol. 2016;47(1):196–201. doi:10.1016/j.bjm.2015.04.001
43. Chen X, Zhang W, Pan W, et al. Prevalence of qnr, aac (6′)-Ib-cr, qepA, and oqxAB in Escherichia coli Isolates from Humans, Animals, and Environment. Antimicrob Agents Chemother. 2012;56(6):3423–3427.
44. Hammad E, Helal R. PMQR determinants among clinical isolates of ESBL and Amp C producing serratia marcescens in Mansoura University Hospitals: a six year study. Int Arabic J Antimicrob Agents. 2015;5:3.
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

