Back to Journals » OncoTargets and Therapy » Volume 11
Investigation of LEP and LEPR polymorphisms with the risk of hepatocellular carcinoma: a case–control study in Eastern Chinese Han population
Authors Zhang S, Jiang J, Chen Z, Wang Y, Tang W, Liu C, Liu L, Chen Y
Received 12 October 2017
Accepted for publication 16 February 2018
Published 11 April 2018 Volume 2018:11 Pages 2083—2089
DOI https://doi.org/10.2147/OTT.S153931
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Samir Farghaly
Sheng Zhang,1,* Jiakai Jiang,1,* Zhan Chen,2,* Yafeng Wang,3 Weifeng Tang,2 Chao Liu,4 Longgen Liu,5 Yu Chen6–8
1Department of General Surgery, Changzhou No. 3 People’s Hospital, Changzhou, Jiangsu Province, 2Department of Thoracic Surgery, Fujian Medical University Union Hospital, Fuzhou, Fujian Province, 3Department of Cardiology, The People’s Hospital of Xishuangbanna Dai Autonomous Prefecture, Jinghong, Yunnan Province, 4Department of Cardiothoracic Surgery, Affiliated People’s Hospital of Jiangsu University, Zhenjiang, 5Department of Liver Disease, Changzhou No. 3 People’s Hospital, Changzhou, Jiangsu Province, 6Cancer Bio-immunotherapy Center, 7Department of Medical Oncology, Fujian Cancer Hospital, Fujian Medical University Cancer Hospital, 8Fujian Provincial Key Laboratory of Translational Cancer Medicine, Fuzhou, Fujian Province, China
*These authors contributed equally to this work
Background: Leptin (LEP) and LEP receptor (LEPR) polymorphisms may be associated with the development of cancer.
Methods: In this study, we selected five functional LEP and LEPR single-nucleotide polymorphisms (SNPs) and conducted a case–control study to determine the relationship of LEP and LEPR polymorphisms with hepatocellular carcinoma (HCC) risk in Eastern Chinese Han population. There were 584 HCC cases and 923 cancer-free controls included in our study. HCC patients and controls were fully matched by age and sex. SNPscan™ genotyping method was used to analyze the genotyping of LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A SNPs.
Results: We found that LEP rs7799039 A>G and rs2167270 G>A polymorphisms were associated with the susceptibility of HCC in this population (LEP rs7799039 A>G: GG vs AA: adjusted odds ratio [OR]=2.03, 95% CI, 1.22–3.38, P=0.006 and GG vs AA/AG: adjusted OR=1.97, 95% CI, 1.20–3.22, P=0.007; rs2167270 G.A: AA vs GG: adjusted OR=2.03, 95% CI, 1.10–3.75, P=0.024 and AA vs GG/GA: adjusted OR=2.01, 95% CI, 1.10–3.68, P=0.023). However, LEPR rs6588147 G>A polymorphism decreased the risk of HCC (GA vs GG: adjusted OR=0.62, 95% CI, 0.45–0.86, P=0.005 and AA/GA vs GG: adjusted OR=0.64, 95% CI, 0.47–0.88, P=0.007).
Conclusion: This case–control study highlights that LEP rs7799039 A>G and rs2167270 G>A polymorphisms increase the susceptibility to HCC; however, LEPR rs6588147 G>A polymorphism may be a protective factor for HCC in Eastern Chinese Han population.
Keywords: LEP, LEPR, polymorphism, hepatocellular carcinoma, risk, single nucleotide polymorphism, hepatitis B virus
Introduction
Liver cancer (LC) rates are the lowest in the South-Central areas of Asia and Northern, Central, and Eastern regions of Europe and the highest in East and South-East Asia and Northern and Western Africa.1 China alone accounts for about 50% of the total number of LC patients and LC-related deaths.1,2 Also, most of the LC patients have hepatocellular carcinoma (HCC). Although chronic hepatitis B virus (HBV) infection contributes to the major etiology of HCC,3 other risk factors may also influence the development of HCC. Nowadays, obesity and overweight are becoming a prevalent problem, which may increase the susceptibility to various cancers.4 Fatty liver is found commonly in patients with chronic HBV infection and might potentiate the risk of HBV-associated HCC by 7.3-fold.5 It is thought that metabolism-related gene may influence the risk of HCC.
The excess of macronutrients stored results in overweight and obesity. Adipocytes may release many inflammatory mediators and lead to oxidative stress and proinflammatory states.6 Inflammatory and oxidative stress are related to the development of malignancy.7,8 Leptin (LEP) and LEP receptor (LEPR) may be implicated in various signal pathways, such as JAK/STAT, MAPK, PI3K, and mTOR.9 Also, these signal pathways were suggested to be correlated with carcinogenesis.10,11 Soga et al reported that fatty liver Shionogi-Lep/Lep mice developed hepatocellular adenomas and carcinomas following steatohepatitis.12 In addition, high levels of LEP were found in sera of patients with HCC.13 Thus, LEP and LEPR genes may impact on signal pathways and play an important role in the development of HCC.
Recently, some important LEP and LEPR single-nucleotide polymorphisms (SNPs) were explored for the susceptibility to a number of cancers. LEP rs7799039 A>G, a promoter SNP, was suggested this variant was particularly affecting transcriptional level and LEP expression.14 However, LEP rs2167270 G>A is located in 5′-untranslated region and correlated with the translation process of LEP mRNA. Some previous meta-analyses suggested that LEP rs7799039 A>G and rs2167270 G>A variants conferred risk to cancer.15–17 However, the association of LEP rs7799039 A>G and rs2167270 G>A polymorphisms with HCC risk is unknown.
LEPR rs1137100 G>A and rs1137101 G>A polymorphisms are two missense SNPs. Li et al reported that rs1137100 G>A and rs1137101 G>A polymorphisms in LEPR gene were correlated with susceptibility to HCC.18 In individuals with chronic HBV infection, these two LEPR SNPs may be a biomarker for the susceptibility to HCC.18 LEPR rs6588147 G>A is an intron SNP. Slattery et al found this SNP caused colon cancer susceptibility among men.19
In view of these previous studies, we found that the relationship of LEP and LEPR polymorphisms with the development of cancer was unclear, especially in Asians.20 In this study, we selected the LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A SNPs and conducted a case–control study to determine the relationship between LEP and LEPR polymorphisms and HCC risk in Eastern Chinese Han population.
Patients and methods
Subjects
This case–control study was performed on two groups. The patient group consisted of 584 HCC cases (mean age 53.17±11.76 years) who were selected from the Department of Hepatobiliary Surgery at Fuzong Clinical Medical College (Fuzhou city, China) and Union Clinical Medical College (Fuzhou city, China) of Fujian Medical University through 2002–2016. HCC diagnosis was confirmed by pathology. Two hepatobiliary surgeons independently evaluated the disease stage according to the criteria of Barcelona Clinic Liver Cancer (2010). The control group consisted of 923 cancer-free participants (mean age 53.72±9.97 years). Written informed consent was signed by each participant included in this study. Approval from the Ethics Committee of Fujian Medical University was obtained. Each participant donated a blood sample which was stored in an EDTA vacutainer tube.
DNA extraction and genotyping
The blood sample obtained from each participant was stored at −80°C immediately. Promega Genomic DNA Kit (Promega Corporation, Fitchburg, WI, USA) was used to extract and purify genomic DNA from lymphocytes. Purity and concentration of genomic DNA were determined by the NanoDrop ND-1000 spectrophotometer and 1.5% agarose gel electrophoresis. SNPscan™ genotyping method (Genesky Biotechologies Inc., Shanghai, China) was used to determine LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A genotypes. Sixty sample sizes (4%) were randomly selected for quality control. The genotyping was performed by our technicians and the results were not changed.
Statistical analysis
Data were analyzed by using SAS 9.4 software (SAS Institute, Cary, NC, USA). Continuous variables were expressed as mean±SD. Student’s t-test was used to assess the differences in age distribution. Chi-square (χ2) or Fisher’s exact test was used to assess the distribution of age, sex, chronic HBV infection status, smoking status, alcohol use, and genotypes. The deviation from Hardy–Weinberg equilibrium in the control group was determined by using an online calculator (http://ihg.gsf.de/cgi-bin/hw/hwa1.pl)21–25 to compare the observed genotype frequencies of LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A SNPs with the expected frequencies. Using additive, homozygote, dominant, and recessive models, the associations between LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A SNPs and susceptibility of HCC were assessed by crude/adjusted odds ratios (ORs) and confidence intervals (CIs). A P<0.05 (two-tailed) was considered as the threshold for statistical significance. In this study, a Bonferroni correction method was used for multiple testing.26,27
Results
Baseline characteristics
There were 584 HCC cases and 923 cancer-free controls included in our study. The frequency distributions for age, sex, chronic HBV infection status, smoking, and alcohol use among HCC patients and controls are summarized in Table 1. The characteristics of HCC cases and controls were fully matched by age and sex. The locus information of LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A polymorphisms is shown in Table 2. The genotyping success rate for all SNPs was eligible (>99%). Minor allele frequency of these SNPs was similar to the data of Chinese population. For these included SNPs, the genotype distribution in controls conformed to Hardy–Weinberg equilibrium.
Association of LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A polymorphisms with the risk of HCC
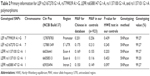
Table 3 lists the genotype distribution of LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A polymorphisms.
Frequencies of LEP rs7799039 AA, AG, and GG genotypes were 51.30%, 38.43%, and 10.26% in HCC cases and 54.83%, 39.09%, and 6.08% in controls, respectively. When compared with the frequency of LEP rs7799039 AA genotype, a significant difference was found in the frequency of LEP rs7799039 GG genotype between HCC cases and controls (crude OR=1.76, 95% CI, 1.19–2.60, P=0.005). When compared with frequency of LEP rs7799039 AA/AG genotype, there was a difference in the frequency of LEP rs7799039 GG genotype between HCC patients and controls (crude OR=1.77, 95% CI, 1.21–2.59, P=0.004). Adjustments for age, sex, chronic HBV infection status, smoking, and drinking, this association was not essentially changed (GG vs AA: adjusted OR=2.03, 95% CI, 1.22–3.38, P=0.006 and GG vs AA/AG: adjusted OR=1.97, 95% CI, 1.20–3.22, P=0.007; Table 3).
Frequencies of LEP rs2167270 GG, GA, and AA genotypes were 59.65%, 34.43%, and 5.91% in HCC patients and 61.24%, 34.85%, and 3.91% in controls, respectively. The association between LEP rs2167270 G>A polymorphism and a tendency for increased HCC risk was noted (AA vs GG: crude OR=1.52, 95% CI, 0.93–2.47, P=0.093 and AA vs GG/GA: crude OR=1.55, 95% CI, 0.96–2.50, P=0.076). Adjustments for age, sex, chronic HBV infection status, smoking, and drinking, this association was more significant (AA vs GG: adjusted OR=2.03, 95% CI, 1.10–3.75, P=0.024 and AA vs GG/GA: adjusted OR=2.01, 95% CI, 1.10–3.68, P=0.023; Table 3).
Frequencies of LEPR rs6588147 GG, GA, and AA genotypes were 77.22%, 21.39%, and 1.39% in HCC patients and 71.77%, 26.71%, and 1.52% in controls, respectively. When compared with the frequency of LEPR rs6588147 GG genotype, a significant difference was found in the frequency of LEPR rs6588147 GA genotype between HCC cases and controls (crude OR=0.73, 95% CI, 0.57–0.94, P=0.013). When compared with the frequency of LEPR rs6588147 GG genotype, there was a difference in the frequency of LEPR rs6588147 GA/AA genotype between HCC patients and controls (crude OR=0.75, 95% CI, 0.59–0.96, P=0.020). Adjustments for age, sex, chronic HBV infection status, smoking, and drinking, this association was more significant (GA vs GG: adjusted OR=0.62, 95% CI, 0.45–0.86, P=0.005 and AA/GA vs GG: adjusted OR=0.64; 95% CI, 0.47–0.88, P=0.007; Table 3).
However, we found no association between LEPR rs1137100 G>A and rs1137101 G>A polymorphisms and the risk of HCC in any genetic model (Table 3).
We used Bonferroni correction method to conduct a multiple testing. We found the genotype distribution of LEP rs7799039 A>G and LEPR rs6588147 G>A SNPs was still significantly different between HCC patients and cancer-free controls (P=0.006 and 0.007 for LEP rs7799039 A>G, and P=0.005 and 0.007 for LEPR rs6588147 G>A, respectively).
Discussion
Although chronic HBV infection status is associated with the etiology of HCC, the individual’s hereditary factor may influence the susceptibility to HCC. Recently, it is believed that the metabolic disturbance increases the susceptibility to many malignancies.4 Thus, the metabolism-related gene may affect the development of HCC. In this case–control study, we explored the relationship of LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A polymorphisms with the risk of HCC in 584 patients and 923 control subjects in Eastern Chinese Han population. We found that LEP rs7799039 A>G and rs2167270 G>A polymorphisms were associated with the susceptibility of HCC in this population. However, LEPR rs6588147 G>A polymorphism decreased the risk of HCC. To our knowledge, this is the study with the largest sample size on the relationship between LEP rs2167270 G>A, rs7799039 A>G, LEPR rs6588147 G>A, rs1137100 G>A, and rs1137101 G>A polymorphisms and HCC risk.
LEP rs7799039 A>G is a promoter SNP, which was suggested it could affect transcriptional level and LEP expression.14 In this study, we found that LEP rs7799039 A>G polymorphism was associated with the susceptibility to HCC in Chinese Han population. Recently, a meta-analysis which included 16 published studies of 6,569 cancer cases and 8,405 controls demonstrated that LEP rs7799039 A>G polymorphism may decrease the risk of cancer.20 In addition, this association between LEP rs7799039 A>G polymorphism and cancer risk was also found in other meta-analyses.16,17 However, the association between LEP rs7799039 A>G polymorphism and the risk of cancer was seldom studied in Asian populations. Clearly, these ambiguous findings suggested that the functions of LEP rs7799039 A>G variants might be different in different races. It was found that the minor allele frequency of LEP rs7799039 A>G polymorphism might alter sharply among different populations (0.201 [Chinese] vs 0.597 [Caucasians]).28 Recently, Romanowski et al reported that LEP rs2167270 G>A polymorphism increased the risk of posttransplant diabetes mellitus.29 LEP rs2167270 G>A is located in 5′-untranslated region and may influence the translation process of LEP mRNA. A functional study demonstrated that serum LEP concentrations in LEP rs2167270 GA genotype were higher than in individuals with rs2167270 GG genotype.30 In this study, the association between LEP rs2167270 G>A polymorphism and increased risk of HCC was found, which was very similar to the findings of the previous meta-analysis.15 In the future, more case–control studies should be carried out to explore the potential association of LEP rs7799039 A>G and LEP rs2167270 G>A polymorphisms with cancer risk in Asians.
Slattery et al reported that LEPR rs6588147 AA genotype had a tendency to lower susceptibility of colon cancer in a mixed population.19 We found that rs6588147 A allele may be associated with the decreased susceptibility of HCC, which was similar to previous findings. In our study, after Bonferroni correction test, this association was still significant between HCC patients and cancer-free controls. However, Nyante et al identified that the LEPR rs6588147 G>A polymorphism might confer susceptibility to breast cancer in luminal A subtype.31 Only a few investigations focused on the correlation between LEPR rs6588147 G>A polymorphism and the risk of cancer, especially in Asians. In future, the association of this SNP with cancer susceptibility should be further studied with a large sample size. Also, functional studies are needed to identify the real biologic function of LEPR rs6588147 G>A polymorphism on the development of HCC.
There are some limitations in this study. First, since the number of HCC cases and controls was moderate, the power of study might be limited. Second, the HCC cases and controls were both enrolled from local hospitals. Therefore, selection bias should have occurred. Third, for lack of some important data, such as lifestyle, dietary habits, and environmental carcinogens exposure, the interaction of gene–environment was not studied. Fourth, the information available on HCC survival has been insufficient till now; thus, we could not further analyze the potential role of LEP/LEPR functional polymorphisms in HCC progression and prognosis. Fifth, due to lack of sufficient information on height and weight, the body mass index was not considered in this study. Finally, we only selected some functional SNPs in LEP and LEPR genes, and a fine-mapping or Genome Wide Association study focusing on LEP and LEPR genes should be performed to further explore the potential relationship of LEP and LEPR SNPs with the risk of HCC.
To sum up, this study has demonstrated that LEP rs7799039 A>G and rs2167270 G>A polymorphisms are genetic risk factors for HCC with various degrees of relationship in an Eastern Chinese Han population. However, LEPR rs6588147 G>A may decrease the risk of HCC. Further epidemiologic case–control studies are needed to assess the interaction of LEP and LEPR genotypes with the exposure to environmental carcinogens, lifestyle, and dietary habits.
Acknowledgments
We appreciate all subjects who participated in this study. We wish to thank Dr Yan Liu (Genesky Biotechnologies Inc., Shanghai, China) for technical support. This study was supported in part by the Clinical Medicine Science and Technology Development Fund of Jiangsu University (JLY20140012).
Disclosure
The authors report no conflicts of interest in this work.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108. | ||
Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. | ||
Li RC, Yang JY, Gong J, et al. [Efficacy of hepatitis B vaccination on hepatitis B prevention and on hepatocellular carcinoma]. Zhonghua liu xing bing xue za zhi=Zhonghua liuxingbingxue zazhi. 2004;25(5):385–387. Chinese. | ||
Vucenik I, Stains JP. Obesity and cancer risk: evidence, mechanisms, and recommendations. Ann N Y Acad Sci. 2012;1271:37–43. | ||
Chan AW, Wong GL, Chan HY, et al. Concurrent fatty liver increases risk of hepatocellular carcinoma among patients with chronic hepatitis B. J Gastroenterol Hepatol. 2017;32(3):667–676. | ||
Ellulu MS, Patimah I, Khaza’ai H, Rahmat A, Abed Y. Obesity and inflammation: the linking mechanism and the complications. Arch Med Sci. 2017;13(4):851–863. | ||
Martinez-Useros J, Li W, Cabeza-Morales M, Garcia-Foncillas J. Oxidative stress: a new target for pPancreatic cancer prognosis and treatment. J Clin Med. 2017;6(3):pii:E29. | ||
Mahmoud AM, Zaki AR, Hassan ME, Mostafa-Hedeab G. Commiphora molmol resin attenuates diethylnitrosamine/phenobarbital-induced hepatocarcinogenesis by modulating oxidative stress, inflammation, angiogenesis and Nrf2/ARE/HO-1 signaling. Chem Biol Interact. 2017;270:41–50. | ||
Wauman J, Tavernier J. Leptin receptor signaling: pathways to leptin resistance. Front Biosci (Landmark Ed). 2011;16:2771–2793. | ||
El-Habr EA, Levidou G, Trigka EA, et al. Complex interactions between the components of the PI3K/AKT/mTOR pathway, and with components of MAPK, JAK/STAT and Notch-1 pathways, indicate their involvement in meningioma development. Virchows Arch. 2014;465(4):473–485. | ||
Pietrzyk L, Torres A, Maciejewski R, Torres K. Obesity and obese-related chronic low-grade inflammation in promotion of colorectal cancer development. Asian Pac J Cancer Prev. 2015;16(10):4161–4168. | ||
Soga M, Hashimoto S, Kishimoto Y, Hirasawa T, Makino S, Inagaki S. Insulin resistance, steatohepatitis, and hepatocellular carcinoma in a new congenic strain of Fatty Liver Shionogi (FLS) mice with the Lep(ob) gene. Exp Anim. 2010;59(4):407–419. | ||
Costantini S, Capone F, Maio P, et al. Cancer biomarker profiling in patients with chronic hepatitis C virus, liver cirrhosis and hepatocellular carcinoma. Oncol Rep. 2013;29(6):2163–2168. | ||
Hoffstedt J, Eriksson P, Mottagui-Tabar S, Arner P. A polymorphism in the leptin promoter region (-2548 G/A) influences gene expression and adipose tissue secretion of leptin. Horm Metab Res. 2002;34(7):355–359. | ||
Liu P, Shi H, Huang C, et al. Association of LEP A19G polymorphism with cancer risk: a systematic review and pooled analysis. Tumour Biol. 2014;35(8):8133–8141. | ||
Yang Y, Liu P, Guo F, et al. Genetic G2548A polymorphism of leptin gene and risk of cancer: a meta-analysis of 6860 cases and 7956 controls. J BUON. 2014;19(4):1096–1104. | ||
Liu Y, Wu H, Zhu Y, Gao Y. Genetic association between leptin-2548G/A polymorphism and risk of cancer: a meta analysis. Int J Clin Exp Med. 2015;8(1):448–455. | ||
Li Z, Yuan W, Ning S, Li J, Zhai W, Zhang S. Role of leptin receptor (LEPR) gene polymorphisms and haplotypes in susceptibility to hepatocellular carcinoma in subjects with chronic hepatitis B virus infection. Mol Diagn Ther. 2012;16(6):383–388. | ||
Slattery ML, Wolff RK, Herrick J, Caan BJ, Potter JD. Leptin and leptin receptor genotypes and colon cancer: gene-gene and gene-lifestyle interactions. Int J Cancer. 2008;122(7):1611–1617. | ||
He J, Xi B, Ruiter R, et al. Association of LEP G2548A and LEPR Q223R polymorphisms with cancer susceptibility: evidence from a meta-analysis. PLoS One. 2013;8(10):e75135. | ||
Qiu H, Cheng C, Wang Y, et al. Investigation of cyclin D1 rs9344 G>A polymorphism in colorectal cancer: a meta-analysis involving 13,642 subjects. Onco Targets Ther. 2016;9:6641–6650. | ||
Tang W, Wang Y, Jiang H, et al. Programmed death-1 (PD-1) rs2227981 C>T polymorphism is associated with cancer susceptibility: a meta-analysis. Int J Clin Exp Med. 2015;8(12):22278–22285. | ||
Tang W, Yu P, Wang Y, et al. Lack of association between cyclin D1 A870G (rs9344) polymorphism and esophageal squamous cell carcinoma risk: case-control study and meta-analysis. Int J Clin Exp Med. 2015;8(8):12685–12695. | ||
Tang W, Chen Y, Chen S, Sun B, Gu H, Kang M. Programmed death-1 (PD-1) polymorphism is associated with gastric cardia adenocarcinoma. Int J Clin Exp Med. 2015;8(5):8086–8093. | ||
Tang W, Wang Y, Chen S, et al. Investigation of Cytotoxic T-lymphocyte antigen 4 polymorphisms in gastric cardia adenocarcinoma. Scand J Immunol. 2016;83(3):212–218. | ||
Bland JM, Altman DG. Multiple significance tests: the Bonferroni method. BMJ. 1995;310(6973):170. | ||
Lesack K, Naugler C. An open-source software program for performing Bonferroni and related corrections for multiple comparisons. J Pathol Inform. 2011;2:52. | ||
Teleginski A, Welter M, Frigeri HR, et al. Leptin (rs7799039) and solute carrier family 30 zinc transporter (rs13266634) polymorphisms in Euro-Brazilian pregnant women with gestational diabetes. Genet Mol Res. 2017;16(1). | ||
Romanowski M, Dziedziejko V, Maciejewska-Karlowska A, et al. Adiponectin and leptin gene polymorphisms in patients with post-transplant diabetes mellitus. Pharmacogenomics. 2015;16(11):1243–1251. | ||
Marcello MA, Calixto AR, de Almeida JF, et al. Polymorphism in LEP and LEPR may modify leptin levels and represent risk factors for thyroid cancer. Int J Endocrinol. 2015;2015:173218. | ||
Nyante SJ, Gammon MD, Kaufman JS, et al. Common genetic variation in adiponectin, leptin, and leptin receptor and association with breast cancer subtypes. Breast Cancer Res Treat. 2011;129(2):593–606. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.