Back to Journals » Infection and Drug Resistance » Volume 13
Investigation of Antibiotic Resistance, Serotype Distribution, and Genetic Characteristics of 164 Invasive Streptococcus pneumoniae from North China Between April 2016 and October 2017
Authors Zhao C, Xie Y, Zhang F, Wang Z, Yang S, Wang Q, Wang X, Li H, Chen H, Wang H
Received 9 April 2020
Accepted for publication 9 June 2020
Published 3 July 2020 Volume 2020:13 Pages 2117—2128
DOI https://doi.org/10.2147/IDR.S256663
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Chunjiang Zhao, Yuhao Xie, Feifei Zhang, Zhanwei Wang, Shuo Yang, Qi Wang, Xiaojuan Wang, Henan Li, Hongbin Chen, Hui Wang
Department of Clinical Laboratory, Peking University People’s Hospital, Beijing, People’s Republic of China
Correspondence: Hui Wang
Department of Clinical Laboratory, Peking University People’s Hospital, Beijing, People’s Republic of China
Email [email protected]
Background: Streptococcus pneumoniae infections are the major cause of global morbidity and mortality among children and patients aged more than 65 years. This study aimed to investigate the antimicrobial resistance, bacterial serotype distribution, and genetic characteristics of invasive S. pneumoniae from different cities in North China.
Materials and Methods: A total of 164 invasive S. pneumoniae strains were collected from 8 hospitals in 5 regions of North China between April 2016 and October 2017. Minimum inhibitory concentrations (MICs) were determined using the agar dilution method. Capsular serotypes were identified using the Quellung reaction test. Molecular epidemiology was investigated using multilocus sequence typing.
Results: S. pneumoniae isolates were highly resistant to macrolides, clindamycin, and tetracycline in all age groups. The overall rate of resistance to penicillin was 56.7%. However, fluoroquinolones and vancomycin maintained excellent antimicrobial activities. The rate of resistance to β-lactam in strains isolated from children aged less than 18 years was significantly higher than that in strains from other age groups. The most prevalent serotypes were 14 (22.6%), 19F (16.5%), non-vaccine types (14.0%), 19A (9.8%), and 23F (9.1%). The coverage for PCV10 and PCV13 was 59.8% and 75.6%, respectively. The vaccine coverage rate was the highest among children aged less than 5 years. The proportion of penicillin-resistant isolates was higher among vaccine-covered strains compared with non-covered strains. S. pneumoniae showed considerable clonal dissemination, and ST876 (28, 17.1%), ST271 (22, 13.4%), ST81 (17, 10.4%) and ST320 (14, 8.5%) were the major STs.
Conclusion: All the 164 invasive S. pneumoniae isolates demonstrated high resistance to antibiotics. The coverage of S. pneumoniae vaccine was higher in children than in adults.
Keywords: antimicrobial resistance, multilocus sequence typing, serotyping, Streptococcus pneumoniae
Introduction
Streptococcus pneumoniae (S. pneumoniae) is one of the major human pathogens. In most cases, the bacterium resides asymptomatically in healthy carriers, colonizing the respiratory tract, sinuses, and nasal cavity typically. However, it may become pathogenic and spread to other locations in susceptible individuals, causing community-acquired respiratory infections and severe invasive pneumococcal diseases (IPDs), such as bacteremia and meningitis.
Invasive infection by S. pneumoniae is an important cause of morbidity and mortality, particularly in individuals aged more than 65 years of age and those with chronic health conditions. It is estimated that pneumococcal death estimates that could be as high as 515,000 deaths in 2015. Approximately 50% of all pneumococcal deaths in 2015 occurred in developing countries.1 Between 1980 and 2008, every year 12,815/100,000 cases of all-cause pneumonia were reported among children aged between 1 month and 59 months, with 526/100,000 deaths annually in China.2 This situation has become worse because the antimicrobial-resistant S. pneumoniae has spread rapidly, and the emergence of multidrug-resistant S. pneumoniae has been observed in various countries over the past decades.3–5
China is the most populous country in the world. In recent years, a huge change in the age distribution of the population has been observed. On the one hand, the average life expectancy of people in China has been extended a lot due to the economic development and improved medical conditions in the last decades, giving rise to the aging Chinese population. On the other hand, China have launched the universal two-child policy since 2015, which allowed all married couples to have two children, contributing to a significant increase in the number of children in China.6 The children and the aging people are the most susceptible individuals to S. pneumonia infection. Therefore, the prevention and control of pneumococcal infections have become an important public health challenge in China. However, the data of IPDs are difficult to obtain in China because of the low culture rate due to the inappropriate use of antibiotics and difficulties in in vitro culturing S. pneumoniae, especially in primary hospitals.
S. pneumoniae can be divided into 46 serogroups with 97 serotypes according to the antigenicity of the capsular polysaccharide.7 An obvious correlation has been found between the antimicrobial resistance and the specific serotypes.8 Many serotypes associated with multidrug resistance can spread widely across the world. The serotypes of carriage isolates may vary depending on the geographic regions, vaccine policies, and socioeconomic status.9,10
Besides antibiotic treatment, the pneumococcal conjugate vaccine (PCV) composed of capsular polysaccharide antigens has been demonstrated to be highly effective in preventing pneumococcal disease and controlling IPDs worldwide, particularly in the developed countries.11–14 Moreover, herd immunity, which is believed to be mediated by a reduction in the transmission of pneumococci to susceptible individuals, has been discovered as another benefit of universal PCV7 immunization programs.14 Three PCVs, including PCV 7, PCV 10, and PCV 13, are currently available on the market, which are immunogenic and protective among children less than 2 years old.15 In 2017, PCV13 had been introduced in China planning to take place of PCV7; however, due to the lack of epidemic data, its protective effect remains unknown in China.
Hence, the surveillance of serotype distribution of isolates obtained from patients diagnosed with IPDs in China is necessary for better managing IPDs and obtaining the information regarding the coverage of novel vaccines. Further, multilocus sequence typing (MLST) is an effective method to discover the relationship and the mode of transmission among the antimicrobial-resistant isolates. This study aimed to present the serotype distribution, genotype, antibiotic susceptibility, and epidemiology of S. pneumoniae associated with IPDs among people in China in 2016–2017 to provide guidance for the prevention and treatment of clinical diseases.
Materials and Methods
Bacterial Isolates
This study involved eight hospitals scattered in five regions (Beijing, Hebei, Shanxi, Liaoning, and Shandong) in China. Based on the number of hospital bed counts and the number of annual outpatient visits, Beijing Chaoyang Hospital (1900 beds, 3.8 million outpatients per year), Peking University People’s Hospital (1700 beds, 2.8 million outpatients per year), Hebei Children’s Hospital (1600 beds, 1.4 million outpatients per year), Shanxi Children’s Hospital (1600 beds, 1.3 million outpatients per year), China Medical University Shengjing Hospital (6750 beds, 4.7 million outpatients per year), Shandong University Qilu Children’s Hospital (1400 beds, 1.1 million outpatients per year) are considered as big hospitals; People’s Hospital of Linyi City (3562 beds, 2.4 million outpatients per year, the number is the sum of five campuses of this hospital in five different locations, in this study the strains were isolated from only one of the campuses), Tangshan City Maternal and Child Health Hospital (1200 beds, 1.0 million outpatients per year) are considered as small hospitals.
During the study period of April 2016 to October 2017, S. pneumoniae strains were isolated from adults and children patients with IPDs. All patients with IPD infection or suspected IPD infection were taken specimens from the site of infection for microbial culture. Blood was collected for blood culture in all patients with fever, and cerebrospinal fluid specimens were cultured in those with signs of intracranial infection. A total of 164 S. pneumoniae strains were isolated from patients diagnosed with IPD by laboratory testing and professional clinicians based on typical clinical manifestations and imaging examination during the study period. All the strains included in this study were cultured from sterile sites, including blood, cerebrospinal fluid (CSF) and occasionally pleural fluid. Duplicate isolates, colonization by bacteria without any clinical evidence of infection and non-IPD infections were excluded from this study. S. pneumoniae was identified based on optochin sensitivity, bile solubility, and α-hemolysis. All isolates were saved at –70°C before use.
All the patients participating in this study signed informed consent, while the guardians of children aged less than 18 years signed on behalf of them. The study protocol was approved by the Ethics Committee of the Peking University People’s Hospital (No. 2016PHB135), and all procedures were conducted according to the Declaration of Helsinki revised in 2008.
In vitro Antimicrobial Susceptibility Testing
The agar dilution method was used to determine the minimum inhibitory concentrations (MICs) of the 164 S. pneumoniae isolates against 15 antibiotics (penicillin, amoxicillin/clavulanic acid, cefaclor, cefuroxime, ceftriaxone, vancomycin, erythromycin, azithromycin, clarithromycin, tetracycline, levofloxacin, moxifloxacin, trimethoprim/sulfamethoxazole, chloramphenicol and clindamycin) in accordance with the guidelines established by the Clinical and Laboratory Standards Institute (CLSI).16 The CLSI 2017 criteria for MICs were applied to classify isolates as susceptible, intermediate, or resistant. The oral penicillin breakpoint was used to classify isolates as penicillin-susceptible (MIC ≤ 0.06 μg/mL), penicillin-intermediate (MIC between 0.12 and 1 μg/mL), or penicillin-resistant (MIC ≥ 2 μg/mL). Penicillin non-meningitis breakpoints (susceptible: MIC ≤ 2 μg/mL; resistant: MIC ≥ 8 μg/mL) and penicillin meningitis breakpoints (susceptible: MIC ≤ 0.06 μg/mL; resistant: MIC ≥ 0.12 μg/mL) were also applied to evaluation the susceptibilities. For ceftriaxone, the non-meningitis breakpoints (susceptible: MIC ≤ 1 μg/mL; resistant: MIC ≥ 4 μg/mL) and meningitis breakpoints (susceptible: MIC ≤ 0.5 μg/mL; resistant: MIC ≥ 2 μg/mL) were used to classify isolates as susceptible or resistant.17 S. pneumoniae ATCC 49,619 was used as the quality control strain and was included in each batch of tests to ensure accurate results. MICs were calculated as the MIC50 and MIC90 (MICs that inhibit 50% and 90% of the isolates, respectively).
Pneumococcal Serotyping
Pneumococcal serotypes/groups were determined for the 164 isolates using Pneumotest-Latex kit (Statens Serum Institut, Copenhagen, Denmark) and type-specific antisera (Statens Serum Institut, Copenhagen, Denmark). The Pneumotest-Latex kit consisted of the 14 latex reagent pools A–I and P–Q. By testing all 14 pools and using the chessboard identification system, the 23 vaccine serotypes were identified to type/group level.18 Traditional Quellung reaction with type-specific antisera was used for full serotyping of serogroups 6, 7, 9, 18, 19 and 23. Isolates that reacted with the Pneumotest-Latex kit but did not belong to serotypes or groups included in the PPV23 were classified as non-vaccine types (NVTs). Strains that belonged to serogroups 23, but was not serotype 23F were considered as not 23F (N23F). The coverage of PCV10 and PCV13 vaccine was calculated by the sum of the percentage of the serotypes the vaccine covered. For PCV10 it was the sum of percentage of serotypes 1, 4, 5, 6B, 7F, 9V, 14, 18C, 19F and 23F; for PCV13 it was the sum of percentage of serotypes 1, 3, 4, 5, 6A, 6B, 7F, 9V, 14, 18C, 19A, 19F and 23F.
Multilocus Sequence Typing Procedure
Multilocus sequence typing (MLST) analysis was performed according to the S. pneumoniae MLST protocol.19 The internal fragments of approximately 550 − 600 bps from the aroE, gdh, gki, recP, spi, xpt, and ddl genes were amplified by polymerase chain reaction using the primers described in a previous study.20 Alleles and sequence types (STs) were assigned using the software available at the S. pneumoniae MLST Database web page (http://pubmlst.org/spneumoniae). The analysis of STs and the assignment to clonal complexes (CCs) were performed using all STs found in the online database using the eBURST program and referred to the clonal complexes settings from data about China in another study.21 STs were grouped into clonal complexes by their similarity to a central allelic profile. The visualization of phylogenetic results based on MLST allele profile was performed using the PHYLOViZ online application (http://www.phyloviz.net/).
Statistical Analysis
Data from the antibiotic susceptibility tests were analyzed using WHONET 5.6 software, a Windows-based database software developed by the World Health Organization for managing and analyzing microbiological laboratory data with a special focus on the analysis of antimicrobial susceptibility test results. The χ2 and Fisher’s exact probability tests were performed using R version 3.5.1 to compare proportions. Data visualization was done with ggplot2 package version 3.1.1.22 A p value less than 0.05 was considered statistically significant.
Results
Profile of Pneumococcal Isolates
During the study period, a total of 164 non-duplicated isolates were collected from 8 hospitals, 4 of them were children and maternal hospitals and the other 4 hospitals were general hospitals. The minimum age of the patients from which the pneumococcal isolates were isolated was 1 month old, the maximum age was 79 years old while the median age was 2 years old. The patients less than 6 years old accounted for 78.7% of all the cases involved. More strains were isolated from male patients than from female patients (93 vs 71 strains, respectively). Of the 164 S. pneumoniae isolates, 95 were obtained from pediatric patients aged 0–36 months, 34 from pediatric patients aged 3–5 years (>3 but ≤5 years), 11 from patients aged 6–17 years (>6 but ≤17 years), 12 from adults aged 18–60 years (>18 but ≤60 years), and 12 from patients aged >60 years. Blood was the most common specimen source, accounting for 75.6% of samples (124 strains), followed by CSF (34 strains) and pleural fluid (6 strains). The patients were mainly concentrated in neurology (37 strains), pediatric ICU (33 strains), respiratory (24 strains) and hematology (19 strains). Fifty-nine strains were from Hebei province, 44 strains from Shandong province, 26 strains from Shanxi province, 23 strains from Shenyang and 12 strains from Beijing. Table 1 summarizes the detailed clinical data of the patient’s medical information. The relationships between patient age groups and IPD types are demonstrated in Table 2. Three major IPD specimen categories were identified: blood (75.6%), cerebrospinal fluid (20.7%), pleural fluid (3.7%) indicating empyema and pleuritic. Of the 164 strains, 80 were cultured form bacteremia infections, 60 strains were from central nervous system infection, 18 strains were from pneumonia and 6 were from empyema and pleuritic infections.
 |
Table 1 Characteristics of 164 Invasive Streptococcus pneumoniae Strains from 2018 in North China |
 |
Table 2 The Proportion of Different IPD Types in Different Age Groups |
Antimicrobial Susceptibility of S. pneumoniae
The in vitro activities of the tested antimicrobial agents against 164 invasive S. pneumoniae strains are shown in Table 3. Based on the MIC breakpoints of oral penicillin criteria, the overall rate of penicillin-resistant S. pneumoniae, penicillin-intermediate S. pneumoniae, and penicillin-susceptible S. pneumoniae isolates was 56.7% (93/164), 23.2% (38/164), and 20.1% (33/164), respectively. The resistant rate of S. pneumoniae to erythromycin, azithromycin, clarithromycin, clindamycin, and tetracycline was 98.2%, 97.6%, 98.2%, 96.3%, and 94.5%, respectively. Macrolides and tetracycline had very limited antibacterial activities against invasive S. pneumoniae in this study. Moreover, 161 (98.2%) strains were resistant to erythromycin, and of these, 158 were resistant to clindamycin simultaneously. S. pneumoniae is more sensitive to penicillin (non-meningitis) compared with amoxicillin/clavulanic acid, and the rate of resistance to amoxicillin/clavulanic acid was higher than to ceftriaxone (non-meningitis) and penicillin (non-meningitis). Levofloxacin and moxifloxacin have a high antibacterial effect on S. pneumoniae with a sensitivity rate of 98.8% both. The MIC90 value for levofloxacin and moxifloxacin was 1 μg/mL and 0.125 μg/mL, respectively. The resistance of S. pneumoniae to penicillin varies significantly among different age groups (Figure 1) and specimen types. The rate of resistance to penicillin in S. pneumoniae isolated from children aged less than 18 years was significantly higher than that in strains isolated from age above 18 years groups (62.9% vs 20.1%, p = 0.00046), and this trend was also observed in other β-lactams. In contrast, strains with increased resistance to levofloxacin, moxifloxacin, and chloramphenicol were found only in the adult groups, especially in the elder age groups. S. pneumoniae showed high resistance to macrolides, clindamycin, and tetracycline in all age groups, with no difference among groups. S. pneumoniae from different specimen types have different resistance rates to β-lactams. The strains isolated from the CSF and pleural fluid were more resistant to penicillin and other oral β-lactams than those isolated from blood (64.7% and 66.7% vs 54%) (Table 3).
 |
Table 3 Susceptibility of Invasive Streptococcus pneumoniae to Routine Clinical Antibiotics |
 |
Figure 1 Resistance of S. pneumoniae to routine antibiotics in different age groups. |
Serotyping and Vaccine Coverage
Of all the 164 strains, the most prevalent serotypes were 14 (22.6%), 19F (16.5%), NVT (14.0%), 19A (9.8%), and 23F (9.1%). Otherwise, the serotypes 14 (24.8%), 19F (16.3%), 19A (11.6%), 23F (10.1%), and NVT (10.1%) were the major serotypes in children aged less than 5 years. For patients aged more than 60 years, 14 (25.0%), 7F (16.6%), NVT (16.6%), 19A (8.3%), and N23F (8.3%) were the most prevalent serotypes. The average vaccine coverage for PCV10 and PCV13 in patients from all age groups was 59.8% and 75.6%, respectively. From the perspective of vaccine coverage, the protective effect of vaccines in children is stronger than that in adults. The vaccine coverage of PCV10 and PCV13 declined with an increasing age, and the vaccine coverage rate was the highest among children aged less than 5 years, suggesting a good protective effect in children. PCV10 showed a clear trend of decreasing vaccine coverage with increasing age, while PCV13 did not find a similar downward trend as PCV10 vaccine, although its vaccine coverage rate in the elder age group was lower than that in the younger age group. The coverage of PCV-10 in big hospitals and small hospitals were 64.8% and 50.8% (p = 0.1146), whereas the coverage of PCV-13 were 78.1% and 71.2% (p = 0.4241), respectively. There is no significant difference in the coverage of PCV-10 and PCV-13 among big hospitals and small hospitals. The proportion of PRSP isolates was higher among vaccine-covered strains (65.3% for PCV10 and 65.3% for PCV13) compared with non-covered strains (43.9% for PCV10 and 30.0% for PCV13), and the p-value was 0.0017 for PCV-10, 0.0003 for PCV-13. The sensitivities of S. pneumoniae serotypes to antimicrobial agents varied significantly. In serogroups 19 (including 19F and 19A), 23 (including 23F and N23F), 6B, and 14, the proportion of PRSP isolates was significantly higher compared with the other common serotypes (Figure 2).
 |
Figure 2 Resistance of S. pneumoniae to routine antibiotics against different serotypes. |
Multilocus Sequence Typing
All the 164 invasive strains were analyzed by the MLST method, and 54 unique STs were found. Among the strains, ST876 (28, 17.1%), ST271 (22, 13.4%), ST81 (17, 10.4%), and ST320 (14, 8.5%) were the most common STs. Of the 19F isolates identified with specific STs, 21 (77.8%) were ST271 and 4 (14.8%) were ST236. Of the 19A isolates, 14 (87.5%) were ST320 and 1 (6.25%) was ST271. Strains of 19F-ST271 and 19A-ST320 were found to be more resistant to several of the tested antibiotics, especially to β-lactams (Table 4). The phylogenetic tree generated using the PHYLOViZ online tool showed that all of the tested strains exhibited obvious clonal aggregation, with the vast majority of the population made up of resistant clones and serotypes, whereas susceptible clones and serotypes showed a decentralized pattern (Figure 3).
 |
Table 4 Sequence Types, Serotypes, Antibiotic Resistance Rates (%), and Age Distributions for 164 Streptococcus pneumoniae Isolates Analyzed by MLST |
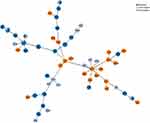 |
Figure 3 Phylogenetic tree of 164 invasive S. pneumoniae strains generated using the PHYLOViZ online application. |
Discussion
IPD is defined as an infection confirmed by isolating S. pneumoniae from a normally sterile site (eg, blood or CSF but not sputum). It was considered as a notable cause of morbidity and mortality worldwide, especially in children aged less than 5 years, adults aged more than 65 years, and individuals with risk factors (eg, immunodeficiency, splenic dysfunction, and HIV infections). The β-lactams, such as penicillin, have been recommended for treating infections caused by S. pneumoniae traditionally. The emergence of resistance to penicillin and other β-lactam antibiotics in pneumococci has led to the increased adoption of macrolides, fluoroquinolones, and other non-β-lactam antibiotics to treat pneumococcal infections. Vaccination is another way to prevent pneumococcal diseases. Childhood vaccination is recommended and is increasingly implemented across the world.23–25 The first pneumococcal conjugate vaccine was 7-valent vaccine (PCV7) which was licensed in 2008 in China and it was replaced by a 13-valent vaccine (PCV13) in 2016. Different from other developed countries, pneumococcal vaccination is not included in the national immunization program in China, and the vaccination is on an individual voluntary basis. This policy has led to a relatively low rate of vaccination against S. pneumoniae in China, especially in remote and rural regions where the economy and residence income is relatively lower compared with other major cities in China.
This study described the antibiotic resistance, serotypes, and molecular epidemiology of invasive S. pneumoniae from eight hospitals in North China of 2016–2017. Invasive pneumococcal strains can provide more reliable clinical information compared with noninvasive strains because the bias introduced by colonization could be excluded. The patients infected with non-IPD strains usually could be cured in primary community healthcare facilities by empirical therapy without a pathogenic diagnosis. The infections caused by IPD strains are usually severe, and the treatment is often based on etiological diagnosis. Therefore, IPD strains can provide more accurate clinical epidemiological information compared with non-IPD strains.
The number of IPD strains in children is much greater than that in adults and this phenomenon was more obvious in children aged less than 5 years. Even the most major proportion of patients with IPD in general hospitals comprised children aged less than 5 years. IPD cases were mainly concentrated in children aged less than 5 years, indicating that children are susceptible to IPD. In this study, children with IPD aged less than 5 years were mainly from Neurology, Pediatric Intensive Care Unit, Respiratory Medicine, and Hematology units. Of the 34 cases of neurology, only 8 cases of S. pneumoniae were isolated from the CSF. The other 26 cases were isolated from the blood, suggesting that the positive rate of CSF culture in central nervous system infection cases is low because the amount of CSF specimens is usually very limited, especially in children. Cerebrospinal fluid specimens are very precious, and other biochemical and cytological tests are usually performed together. In this way, the amount of specimens left for microbial testing is very small, which affects the positive rate of cerebrospinal fluid specimen culture. Hence, the diagnosis of meningitis in children requires supplementary blood culture.
The β-lactams were widely used to treat pneumococcal infections, but the resistant strains have emerged and become more prevalent throughout the world in recent decades.26,27 The emergence of resistance of pneumococci against penicillin and other β-lactam antibiotics has led to an increased adoption of macrolides, fluoroquinolones, and other non-β-lactam antibiotics to treat pneumococcal infections.28 The β-lactams and macrolides were considered as relatively safer antibiotics for children; they were intensively used to treat community-acquired infections among children.29 In this study, the resistance rates for β-lactams in children were significantly higher than those in adults, probably because β-lactams were the primary drugs to treat pneumococcal infection in children and more frequently prescribed. On the contrary, reduced susceptibility to levofloxacin, moxifloxacin, and chloramphenicol was observed only in adults, especially elderly people aged more than 60 years. This was because these antibiotics could be used only in adults. The resistant strains emerged with the increased use of these antibiotics, which could not be used in children. Hence, the strains isolated from children were generally very sensitive to these drugs. Macrolides, including erythromycin and azithromycin, were recommended as the primary antibiotics to treat community-acquired infections in both adults and children.28 This study found that the resistance rates of S. pneumoniae against macrolides were very high, with no difference between different age groups. The percentage of PRSP from invasive pneumococcal isolates was 56.4% in this study, which was a little higher compared with data from another study reported earlier (51.6% for PRSP).28 The high resistance rate might be related to the difference in the study participants. S. pneumoniae responsible for invasive infections among children demonstrated higher resistance compared with that from the adult noninvasive counterpart.
Besides the use of antibiotics for already occurring infections, vaccination is an alternative initiative for preventing pneumococcal infection. The prevalence of vaccine-covered serotypes faded considerably when PCV was used widely in other western countries.25,30 Many studies showed that the introduction of PCVs has not only reduced the burden of pneumococcal disease among youngsters,31 but also greatly wedged the burden of disease in adults by preventing the vaccine-related resistant strains in adults – the herd protection effect.32,33 In this study, the average vaccine coverage rate of PCV10 and PCV13 in the invasive strains was 59.8% and 75.6%, respectively, which were much higher than the previously reported coverage in noninvasive strains,34 suggesting that vaccines could play a better protective role in IPD compared with noninvasive infections. Taking into account the higher coverage in invasive strains, this study recommended that all children should be vaccinated at the appropriate age and pneumococcal vaccine should be included in the national immunization program. The coverage of PCV10 vaccine gradually declined with age, suggesting that PCV10 had a relatively lower protective effect in adults. In contrast, although PCV13 coverage in adults is lower than that in children, no decrease in coverage associated with age has been observed, suggesting that coverage of PCV13 is higher among children. It can also produce higher protection in adults. The reason is that PCV13 covered the major epidemic serotypes in adults, which were not covered by PCV10 (serotype 3, serotype 6A and serotype 19A). Based on the results of serotyping, drug resistance, and MLST classification, the present study concluded that the spread pattern of S. pneumoniae in China was clonal propagation. The prevalent clones serotype 14-ST876, serotype 19F-ST271, and serotype 19A-ST320 were responsible for the high resistance in invasive strains. The proportion of S. pneumoniae serotype 14 was very high in this study, higher than the proportion of noninvasive strains in previous studies,8,34 indicating that S. pneumoniae with serotype 14 might have a stronger invasive ability in young children. The limitation of this study is that due to the difficult nature of cerebrospinal fluid specimens for bacterial culture, there are many intracranial infections that were identified by blood culture rather than as a result of cerebrospinal fluid culture and the role of cerebrospinal fluid culture in the diagnosis of intracranial infection has been underestimated. The positive rate of diagnosing life-threatening intracranial infection can therefore be increased by using other tests, such as nucleic acid testing. Differences in the types of hospitals enrolled (four hospitals are maternal and child health centers, mainly for children) resulted in more S. pneumoniae isolated from children than from adults in this study. Moreover, the empirical use of drugs in China is very common in adults, whereas in children the use of antimicrobial drugs is more regulated, which leads to a high rate of positive microbial cultures in children and a relatively low rate of positive microbial cultures in adults, especially for S. pneumoniae.
Conclusions
High resistance to β-lactams and macrolides was observed among 164 invasive S. pneumoniae strains mainly from children aged less than 5 years. However, fluoroquinolones and vancomycin maintained excellent activity against S. pneumoniae. The serotypes 19F, 19A, 6A, 6B, 14, and 23F demonstrated higher resistance compared with other serotypes. The protective effect of vaccines in children and invasive strains was higher than that in adult and noninvasive strains. The vaccine coverage varied considerably with the age of the patients. Obvious clonal aggregation was observed by molecular biology analysis.
Abbreviations
CC, clonal complex; CSF, cerebrospinal fluid; IPDs, invasive pneumococcal diseases; MLST, multilocus sequence typing; NVTs, non-vaccine types; PCV, pneumococcal conjugate vaccine; PCV10, 10-valent vaccine; PCV13, 13-valent vaccine; S. pneumonia, Streptococcus pneumoniae; STs, sequence types.
Data Sharing Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Ethics Approval and Informed Consent
All the patients participating in this study signed informed consent, while the guardians of children aged less than 18 years signed on behalf of them. The study protocol was approved by the Ethics Committee of the Peking University People’s Hospital (No. 2016PHB135), and all procedures were conducted according to the Declaration of Helsinki revised in 2008.
Acknowledgments
We sincerely thank Wang Chunlei of Beijing Chaoyang Hospital, Li Mei of Hebei Children’s Hospital, Zhu Lei and Meng Jinhua of Shanxi Children’s Hospital, Zhang Zhijie of China Medical University Shengjing Hospital, Wang Shifu of Shandong University Qilu Children’s Hospital, Jin Fengli of People’s Hospital of Linyi City and Gao Chunyan of Tangshan City Maternal and Child Health Hospital for their outstanding contributions to the research and collection of strains.
Author Contributions
CJZ conceived and coordinated the study, designed, performed and analyzed the experiments, wrote the paper. YHX, FFZ, ZWW, SY, QW, XJW, HNL, HBC and HW carried out the data collection, data analysis, and revised the paper. All authors contributed to data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors declare that they have no conflicts of interest.
References
1. Wahl B, O’Brien KL, Greenbaum A, et al. Burden of Streptococcus pneumoniae and haemophilus influenzae type b disease in children in the era of conjugate vaccines: global, regional, and national estimates for 2000–15. Lancet Glob Health. 2018;6(7):e744–57. doi:10.1016/S2214-109X(18)30247-X
2. Chen Y, Deng W, Wang S-M, et al. Burden of pneumonia and meningitis caused by Streptococcus pneumoniae in China among children under 5 years of age: a systematic literature review. PLoS One. 2011;6(11):e27333. doi:10.1371/journal.pone.0027333
3. Purba AK, Ascobat P, Muchtar A, et al. Multidrug-resistant infections among hospitalized adults with community-acquired pneumonia in an Indonesian tertiary referral hospital. Infect Drug Resist. 2019;12:3663–3675. doi:10.2147/IDR.S217842
4. Olarte L, Kaplan SL, Barson WJ, et al. Emergence of multidrug-resistant pneumococcal serotype 35B among children in the United States. J Clin Microbiol. 2017;55(3):724–734. doi:10.1128/JCM.01778-16
5. Park M, Kim HS, Kim H-S, et al. Novel levofloxacin-resistant multidrug-resistant Streptococcus pneumoniae serotype 11A isolates, South Korea. Emerg Infect Dis. 2016;22(11):1978–1980. doi:10.3201/eid2211.151450
6. Peian W. On the two-child policy in China. Popul Res. 2016;40:3–7.
7. Geno KA, Gilbert GL, Song JY, et al. Pneumococcal capsules and their types: past, present, and future. Clin Microbiol Rev. 2015;28(3):871–899. doi:10.1128/CMR.00024-15
8. Zhao C, Zhang F, Chu Y, et al. Phenotypic and genotypic characteristic of invasive pneumococcal isolates from both children and adult patients from a multicenter surveillance in China 2005–2011. PLoS One. 2013;8(12):e82361. doi:10.1371/journal.pone.0082361
9. Block SL, Harrison CJ, Hedrick JA, et al. Penicillin-resistant Streptococcus pneumoniae in acute otitis media: risk factors, susceptibility patterns and antimicrobial management. Pediatr Infect Dis J. 1995;14(9):751–759. doi:10.1097/00006454-199509000-00005
10. Perniciaro S, Imöhl M, Fitzner C, van der Linden M, Sekaran SD. Regional variations in serotype distribution and vaccination status in children under six years of age with invasive pneumococcal disease in Germany. PLoS One. 2019;14(1):e0210278–e0210278. doi:10.1371/journal.pone.0210278
11. Fitzwater SP, Chandran A, Santosham M, Johnson HL. The worldwide impact of the seven-valent pneumococcal. Vaccine Rep. 2012;31:501–508.
12. Wang M, Wang Y, Chu C, Tu T, Shiao A, Chou P. Impact of pneumococcal conjugate vaccine on pediatric tympanostomy tube insertion in partial immunized population. Sci World J. 2015;2015:1–8. doi:10.1155/2015/248678
13. Pilishvili T, Lexau C, Farley MM, et al. Sustained reductions in invasive pneumococcal disease in the era of conjugate vaccine. J Infect Dis. 2017;201(1):32–41.
14. Miller E, Andrews NJ, Waight PA, Slack MPE, George RC. Herd immunity and serotype replacement 4 years after seven-valent pneumococcal conjugate vaccination in England and Wales: an observational cohort study. Lancet Infect Dis. 2011;11(10):760–768. doi:10.1016/S1473-3099(11)70090-1
15. Tarahomjoo S. Recent approaches in vaccine development against Streptococcus pneumoniae. J Mol Microb Biotech. 2014;24(4):215–227.
16. CLSI. Methods for Dilution Antimicrobial Susceptibility Tests f or Bacteria That Grow Aerobically; Approved St andard—Ninth Edition. CLSI Doc M07-A9. Wayne, PA: Clin Lab Stand Inst; 2012.
17. CLSI. Performance Standards for Antimicrobial Susceptibility Testing.
18. Slotved H-C, Kaltoft M, Skovsted IC, Kerrn MB, Espersen F. Simple, rapid latex agglutination test for serotyping of pneumococci (Pneumotest-Latex). J Clin Microbiol. 2004;42(6):2518–2522. doi:10.1128/JCM.42.6.2518-2522.2004
19. Enright MC, Spratt BG. A multilocus sequence typing scheme for Streptococcus pneumoniae: identification of clones associated with serious invasive disease. Microbiology. 1998;144(Pt 1):3049–3060. doi:10.1099/00221287-144-11-3049
20. Moore MR, Gertz REJ, Woodbury RL, et al. Population snapshot of emergent Streptococcus pneumoniae serotype 19A in the United States, 2005. J Infect Dis. 2008;197(7):1016–1027. doi:10.1086/528996
21. Gladstone RA, Lo SW, Lees JA, et al. International genomic definition of pneumococcal lineages, to contextualise disease, antibiotic resistance and vaccine impact. EBioMedicine. 2019;43:338–346. doi:10.1016/j.ebiom.2019.04.021
22. Wickham H. Ggplot2: Elegant Graphics for Data Analysis. Springer; 2016.
23. Galanis I, Lindstrand A, Darenberg J, et al. Effects of PCV7 and PCV13 on invasive pneumococcal disease and carriage in Stockholm, Sweden. Eur Respir J. 2016;47(4):1208–1218. doi:10.1183/13993003.01451-2015
24. Shiri T, Datta S, Madan J, et al. Indirect effects of childhood pneumococcal conjugate vaccination on invasive pneumococcal disease: a systematic review and meta-analysis. Lancet Glob Health. 2017;5(1):e51–9. doi:10.1016/S2214-109X(16)30306-0
25. Koshy E, Murray J, Bottle A, Sharland M, Saxena S. Impact of the seven-valent pneumococcal conjugate vaccination (PCV7) programme on childhood hospital admissions for bacterial pneumonia and empyema in England: national time-trends study, 1997–2008. Thorax. 2010;65(9):770–774. doi:10.1136/thx.2010.137802
26. Jacobs MR, Koornhof HJ, Robins-Browne RM, et al. Emergence of multiply resistant pneumococci. N Engl J Med. 1978;299(14):735–740. doi:10.1056/NEJM197810052991402
27. Linares J, Ardanuy C, Pallares R, Fenoll A. Changes in antimicrobial resistance, serotypes and genotypes in Streptococcus pneumoniae over a 30-year period. Clin Microbiol Infect. 2010;16(5):402–410. doi:10.1111/j.1469-0691.2010.03182.x
28. Lin H, Dyar OJ, Rosales-Klintz S, et al. Trends and patterns of antibiotic consumption in Shanghai municipality, China: a 6 year surveillance with sales records, 2009–14. J Antimicrob Chemother. 2016;71(6):1723–1729. doi:10.1093/jac/dkw013
29. Bradley JS, Byington CL, Shah SS, et al. The management of community-acquired pneumonia in infants and children older than 3 months of age: clinical practice guidelines by the Pediatric Infectious Diseases Society and the Infectious Diseases Society of America. Clin Infect Dis. 2011;53:e25–76. doi:10.1093/cid/cir531
30. Esposito S, Principi N. Impacts of the 13-valent pneumococcal conjugate vaccine in children. J Immunol Res. 2015;2015:1–6. doi:10.1155/2015/591580
31. Whitney CG, Farley MM, Hadler J, et al. Decline in invasive pneumococcal disease after the introduction of protein-polysaccharide conjugate vaccine. N Engl J Med. 2003;348(18):1737–1746. doi:10.1056/NEJMoa022823
32. Centers for Disease Control and Prevention (CDC). Direct and indirect effects of routine vaccination of children with 7-valent pneumococcal conjugate vaccine on incidence of invasive pneumococcal disease–United States, 1998–2003. MMWR Morb Mortal Wkly Rep. 2005;54(36):893–897.
33. Whitney CG, Klugman KP. Vaccines as tools against resistance: the example of pneumococcal conjugate vaccine. Semin Pediatr Infect Dis. 2004;15(2):86–93. doi:10.1053/j.spid.2004.01.011
34. Zhao C, Li Z, Zhang F, et al. Serotype distribution and antibiotic resistance of Streptococcus pneumoniae isolates from 17 Chinese cities from 2011 to 2016. BMC Infect Dis. 2017;17(1):804. doi:10.1186/s12879-017-2880-0
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
