Back to Journals » OncoTargets and Therapy » Volume 13
Incidence and PD-L1 Expression of MET 14 Skipping in Chinese Population: A Non-Selective NSCLC Cohort Study Using RNA-Based Sequencing
Authors Xu Z, Li H, Dong Y , Cheng P, Luo F, Fu S, Gao M, Kong L, Che N
Received 5 December 2019
Accepted for publication 17 May 2020
Published 30 June 2020 Volume 2020:13 Pages 6245—6253
DOI https://doi.org/10.2147/OTT.S241231
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Takuya Aoki
Ziguang Xu,1,* Hongxia Li,2,* Yujie Dong,3,* Peng Cheng,4 Fang Luo,4 Shijun Fu,5 Min Gao,5 Lingfei Kong,1 Nanying Che3
1Department of Pathology, Henan Provincial People’s Hospital, People’s Hospital of Zhengzhou University, Zhengzhou, Henan, People’s Republic of China; 2Department of Pathology, Beijing Chest Hospital, Capital Medical University, Beijing, People’s Republic of China; 3Department of Pathology, Beijing Tuberculosis & Thoracic Tumor Research Institute, Beijing Chest Hospital, Capital Medical University, Beijing, People’s Republic of China; 4Novogene Co., Ltd, Beijing, People’s Republic of China; 5Medical Affairs Department, Pfizer Oncology, Shanghai, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Nanying Che
Department of Pathology, Beijing Tuberculosis & Thoracic Tumor Research Institute, Beijing Chest Hospital, Capital Medical University, No. 97 Ma Chang, Tongzhou District, Beijing 101149, People’s Republic of China
Email [email protected]
Lingfei Kong
Department of Pathology, Henan Provincial People’s Hospital, People’s Hospital of Zhengzhou University, No. 7, Weiwu Road, Zhengzhou, Henan 450003, People’s Republic of China
Email [email protected]
Background: Mesenchymal–epithelial transition (MET) exon14 skipping mutations represent a clinically unique molecular subtype of NSCLC. The prevalence rates of MET exon 14 skipping in lung adenocarcinoma (ADC) range from 0.9% to 4.0% in Asian populations. Since some somatic variants that do not encompass the MET exon 14 splice sites might also induce MET exon 14 skipping, the RNA-based sequencing is speculated as the most accurate method for detecting exon 14 skipping.
Patients and Methods: A total of 951 NSCLC patients from two hospitals were enrolled in this study. MET exon14 skipping was detected using RNA-based next-generation sequencing (NGS). Also, immunohistochemistry (IHC) was performed in 405 samples simultaneously.
Results: The overall estimated prevalence of MET exon 14 skipping was approximately 1.8% in ADCs and 1.7% in NSCLCs. The detection rate of MET exon 14 skipping from surgical resection specimen was 2.3% in NSCLCs and 2.0% in ADCs. The MET exon 14 skipping was identified in 6.6% of EGFR/KRAS/ALK/ROS1/RET-negative ADCs. Additionally, PD-L1 was found to be highly expressed in NSCLC patients harboring MET exon 14 skipping (P< 0.01).
Conclusion: The prevalence of MET exon14 skipping in lung ADCs in the East Asian population was similar to that of the Western population as assessed by RNA-based NGS. The NSCLC patients with MET exon 14 skipping were older than those with other oncogenic driver mutations, such as EGFR, ALK, and ROS1. In addition, PD-L1 was highly expressed in NSCLC patients with MET exon 14 skipping.
Keywords: non-small cell lung cancer, MET exon14 skipping, next-generation sequencing, PD-L1
Introduction
Currently, lung cancer is the leading cause of cancer-related deaths worldwide.1 Non-small cell lung cancer (NSCLC) accounts for about 85% of all lung cancers; of these, ADC and squamous cell carcinoma (SCC) are the most frequent histological subtypes, accounting for 50% and 30%, respectively. In the past decade, remarkable progress has been made in the treatment of NSCLC in patients whose tumors harbor targetable somatic mutations.2 Recently, the activating mutations in the MET gene have been recognized as potential therapeutic targets in NSCLC.3,4
MET is a receptor tyrosine kinase that is activated upon binding to the hepatocyte growth factor (HGF) ligand, forming homodimers and subsequently activating the kinase domain. This process triggers the downstream signaling cascade that promotes proliferation, cell cycle progression, cell migration, and invasion.5 The aberrant activation of MET triggers a constitutive activation of downstream signaling pathways, PI3K/AKT/mTOR and RAS/ERK/MAPK, involved in tumorigenesis and tumor development.6–8
MET exon14 skipping as a therapeutically targetable mutation has been reported in lung cancer and comprises of 4.3% of lung ADCs in The Cancer Genome Atlas dataset.9 MET exon 14 skipping results in the deletion of the juxtamembrane domain that is essential for the efficient binding of E3 ubiquitin protein ligase (CBL). These alterations lead to increased MET stability and oncogenic potential and confer sensitivity to MET tyrosine kinase inhibitors (TKIs), such as crizotinib and cabozantinib.3,4,10
The prevalence rates of MET exon 14 skipping in Asian populations with lung ADC are reported to be approximately 4.0% in Taiwan, 2.6% in Hong Kong, 0.9% in China, and 2.13% in Korea11–14 as compared to the rates reported in the USA, 3% (131/4403), 2.9% (205/7140), and 2.1% (18/873).10,15,16 The diverse detection rate of MET exon 14 skipping might be attributed to the methodology, specimen source, and ethnic differences of the patients. In this study, we detected MET exon 14 alterations in 951 Chinese NSCLC patients using targeted RNA-based next-generation sequencing (NGS). In addition, the molecular alterations of the main oncogenic drivers and PD-L1 expression was elucidated in these patients.
Patients and Methods
Study Cohort
Formalin-fixed paraffin-embedded (FFPE) tumor tissues were obtained from patients with NSCLC, who had undergone surgical resection, percutaneous transthoracic needle biopsy (PTNB), and transbronchial lung biopsy (TBLB) from January to December 2018 at the Beijing Chest Hospital, Capital Medical University and Henan Provincial People’s Hospital, Zhengzhou University. The patients were reviewed according to the 2015 World Health Organization classification.17 The demographic data and clinicopathological parameters were collected from the electronic medical records. The staging of patients was assessed according to the 8th edition of the tumor, node, and metastasis (TNM) classification for lung cancer.18 A total of 951 patients with NSCLC, who fulfilled the selection criteria were included in the present study that was approved by the Ethics Committee of the Beijing Chest Hospital. This study was conducted in accordance with the Declaration of Helsinki and all patients signed the informed consent to allow the use of the tumor samples in future studies. All clinical data and samples were received anonymously.
DNA and RNA Preparations
According to the manufacturer’s instructions, DNA was isolated using TIANamp Genomic DNA Kit (Tiangen, Beijing, China) and the qualitative check was performed on 1% agarose gel electrophoresis. RNA was isolated using the RecoverAll™ Total Nucleic Acid Isolation Kit (Thermo Fisher Scientific, Waltham, MA, USA), and RNA integrity was evaluated by qPCR and agarose gel electrophoresis.
DNA/RNA-Based NGS
An equivalent of 10 ng DNA served as a template to construct the amplicon libraries using an Ion AmpliSeqTM Library Kit 2.0 (Thermo Fisher Scientific), followed by targeted NGS as reported previously.19 The custom-designed panel encompassed 21 cancer-related genes, including AKT1, ALK, BRAF, CTNNB1, DDR2, EGFR, ERBB2, ERBB4, FGFR1, FGFR2, FGFR3, KRAS, MAP2K1, MET, NOTCH1, NRAS, PIK3CA, PTEN, SMAD4, STK11, and TP53 (Supplementary Table 1).
RNA-based NGS was conducted using a custom-designed panel which included probes spanning the MET exon 13–15 junction. Also, fusions of ALK, RET, ROS1, and NTRK1 with other genes were detected. The libraries were prepared using the one-step PCR amplification method. Then, the amplicon libraries were sequenced on an Ion Torrent Systems Proton system and a PI chip with barcoding performed using an Ion Xpress Barcode Adapter 1–96 Kit (Thermo Fisher Scientific).
NGS Data Analysis
Torrent Suite Software (version 5.0) was used for signal processing, base calling, quality score assignment, and adapter trimming after sequencing reaction. The mutant allele frequency ≥ 1%, fusion mutants with ≥ 1000× coverage, and Normalized Detection Fractions (NDF) ≥ −2.8 were accepted.
RT-PCR and Sanger Sequencing
In order to confirm MET exon 14 skipping mutation in the specimens of patients, FFPE tumor samples were assessed by quantitative RT-PCR using the following primers: forward, 5ʹ-AGATCAGAATTTCACAGGATTGATTGC-3ʹ; reverse, 5ʹ-CTGTCAG AGGATACTGCACTTGTC-3ʹ. The forward primer probed MET exon 13, and the reverse primer targeted MET exon 15. Finally, the PCR products were analyzed by agarose gel electrophoresis and sequenced by standard procedures.
MET and PD-L1 Immunohistochemistry (IHC) Assay
The specimens were subjected to IHC staining to evaluate the expression of PD-L1 and MET using a mouse monoclonal PD-L1 antibody (22-c3, Dako, California, USA) and a rabbit monoclonal c-Met antibody (SP44, 1:50 dilution, Ventana Medical Systems, AZ, London, UK). The PD-L1 protein expression was scored using the tumor proportion score (TPS), which is the percentage of viable tumor cells showing partial or complete membrane staining.20 The staining intensity was classified into TPS ≥ 50% (high expression), 1–49% (low expression), and 0 (negative). The IHC staining intensity of MET was scored semi-quantitatively on a 4-titer scale as follows: 0 (none), 1+ (weak membrane staining in > 10% of the tumor cells), 2+ (moderate staining of the entire membrane in > 10% of the tumor cells), or 3+ (strong staining of the entire membrane in > 10% of the tumor cells).21 The MET-positive specimens were defined by a score of 2+ or 3+. Representative IHC images of c-MET and PD-L1 are shown in Supplementary Figure 1.
Statistical Analysis
The correlation between the clinicopathological variables and MET status was analyzed by χ2 test, Fisher’s exact test, and Student’s t-test using the Statistical Package for the Social Sciences 19 (SPSS, Chicago, IL, USA) software. All tests were two-sided. P-value < 0.05 was considered as statistically significant.
Results
Patient Characteristics
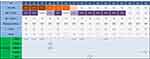
A total of 951 NSCLC patients were enrolled in this study, and the clinical characteristics are summarized in Table 1. The median age of the patient cohort was 63 (range, 25–91) years. The cohort comprised of 38 patients (4.0%) who were < 45-years-old. Moreover, 55.3% of the patients (526/951) were male, and 44.7% (425/951) were female. Regarding smoking history, 51.1% (454/888) were never-smokers and 48.9% (434/888) were ever-smokers (63 cases had missing data). The histological subtypes of the cancer were primarily ADC (81.2%, 772/951) and SCC (15.9%, 151/951). Other histological subtypes included adenosquamous carcinoma (ASC, 1.2%, 11/951), large cell carcinoma (LCC, 0.7%, 7/951), sarcomatoid carcinomas (SC, 0.4%, 4/951), and mucoepidermoid carcinoma (MEC, 0.1%, 1/951). In addition, the histological subtype of 5 (0.5% 5/951) patients was not specified (NOS). Among 608 patients with complete pathological stage information, 17.3% were stage I (105/608), 4.4% were stage II (27/608), 16.4% were stage III (100/608), and 61.9% were stage IV (376/608).
 |
Table 1 Patient Characteristics and Correlation Between Clinical Features and MET Exon 14 Skipping in NSCLC Patients |
Prevalence of MET Exon 14 Skipping in Patients with NSCLC
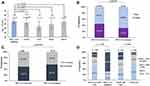
MET exon 14 skipping was identified in 16 cases, which accounted for 1.7% (16/951) of all the NSCLC patients (Figure 1A). Of these, 14 cases were ADC (14/16; 87.5%), 1 case was ASC (1/16, 6.3%), and 1 case was SC (1/16; 6.3%). MET exon 14 skipping was not detected in squamous cell carcinoma or other histological types in this study. The incidence was 1.8% (14/772, Figure 1B) in ADCs and 2.0% (16/800) in non-squamous NSCLCs. In the present study, tumor samples were acquired in three different ways: surgical resection, PTNB, and TBLB. The detection rate of MET exon 14 skipping from surgical resection specimen was 2.3% (8/344) in NSCLCs and 2.0% (6/307) in ADCs (Figure 1A). The prevalence in the surgical resected sample was a representative of the overall population.
Other known tumor genetic alterations have also been evaluated using a DNA- and RNA-based NGS panel. The oncogenic driver mutation status of 951 NSCLC patients is shown in Figure 1A: EGFR mutation (n = 444, 46.7%), KRAS mutation (n = 99, 10.4%), ALK rearrangement (n = 52, 5.5%), ERBB2 mutation (n = 31, 3.3%), BRAF mutation (n = 18, 1.9%), ROS1 rearrangement (n = 13, 1.4%), ERBB4 mutation (n = 12, 1.3%), and RET rearrangement (n = 10, 1.1%). The proportion of genetic alterations was also evaluated in the ADC population (Figure 1, left panel). Consequently, the proportion of EGFR mutation and ALK rearrangement in ADCs was slightly higher as compared to that in the NSCLC population.
Clinical Characteristics of Patients with MET Exon 14 Skipping
The clinicopathological characteristics of 16 patients harboring MET exon 14 skipping are demonstrated in Table 1. Except 1 patient (59-year-old), all were > 60-years-old. Significant differences were observed in the median age between NSCLC patients with MET exon 14 skipping and other oncogenic driver mutations, such as ALK (66 vs 54, P = 0.001), and ROS1 (66 vs 59, P = 0.010; Figure 2A), indicating that patients with MET exon 14 skipping were older than those with other oncogenic driver mutations.
Further statistical analyses assessed the gender and smoking history. Between MET exon 14 skipping and wild-type population, no significant difference was observed in the proportion of gender and never-smokers. Furthermore, no significant association was detected between MET exon 14 skipping and tumor location and stage (Figure 2B and C).
c-MET Expression in NSCLC Patients with MET Exon 14 Skipping
A total of 364 patients underwent c-MET IHC assay and were enrolled in the c-MET cohort. According to the staining intensity (negative, weak, moderate, or strong) and prevalence in tumor cells, patients in the c-MET cohort were divided into four subgroups: 3+ (n = 49, 13.5%), 2+ (n = 160, 44.0%), 1+ (n = 122, 33.5%), and 0 (n = 33, 9.0%). Furthermore, MET exon 14 skipping was identified in 7 cases (Figure 3), 3 cases (42.9%) scored 3+, 3 cases (42.9%) scored 2+, and 1 case (14.2%) scored 1+. According to the definition of c-MET-positive described before, MET overexpression occurred in 85.7% (6/7) MET exon 14 skipping NSCLCs and in 56.9% (203/357) MET exon 14 wild-type NSCLCs. As shown in Supplementary Table 2, the χ2 test did not show any significant difference (P = 0.253) between the two groups.
PD-L1 Expression in NSCLC Patients with MET Exon 14 Skipping
Simultaneously, the PD-L1 expression status was estimated by IHC when sufficient tissue is available. The tumor proportion score (TPS) was applied to evaluate the PD-L1 protein expression as introduced before. Among 401 patients who were subjected to PD-L1 IHC assay, 196 cases (48.9%) were negative (TPS < 1%), 132 cases (32.9%) showed low expression (1% ≤ TPS ≤ 49%), and 73 cases (18.2%) showed high expression (TPS ≥ 50%; Table 1). The PD-L1 expression status of 13 patients harboring the MET exon 14 skipping is shown in Figure 3. Of these, 9 (69.2%) patients showed high PD-L1 expression, and 4 (30.8%) patients showed low or negative PD-L1 expression (Table 1). The prevalence of PD-L1 expression status in patients with MET exon 14 skipping was significantly different from those with MET exon 14 wild-type. As shown in Figure 2D, PD-L1 tended to be highly expressed in NSCLC patients harboring MET exon 14 skipping (69.2% vs 17.3%, P < 0.01). Conversely, compared to the EGFR wild-type cohort, the proportion of PD-L1 high expression was significantly lower in the EGFR mutant cohort (9.3% vs 26.9%, P < 0.01; Figure 2D).
Co-Occurrence of Genetic Alterations in NSCLC Patients with MET Exon 14 Skipping
DNA- and RNA-based NGS was performed to identify the lung cancer-associated genetic alterations including fusions, truncations, and in-frame and missense mutations. Among the 16 patients harboring MET exon 14 skipping, EGFR exon 19 deletion was detected with low mutation abundance in 1 case (Figure 3, EGFR c.2240_2257del, 1.41%). The other common oncogenic driver genes, such as KRAS, ALK, ROS1, and RET were not found to be co-mutated with MET exon 14 skipping. As shown in Table 2, the incidence of MET exon 14 skipping was 4.7% (16/340) in EGFR/KRAS/ALK/ROS1/RET-negative NSCLCs.
 |
Table 2 Co-Mutation Between MET Ex14 Skipping and EGFR/KRAS/ALK/ROS1/RET in NSCLC and ADC Populations |
As a major tumor suppressor, TP53 co-mutations were widespread in the driver gene mutation-positive population. In this study, TP53 mutations were detected in 48% (444/935) NSCLC patients, 46.2% (205/444) showed EGFR mutants, 49.5% (49/99) showed KRAS mutants, 21.2% (11/52) showed ALK-rearrangements, 38.5% (5/13) patients showed ROS1 rearrangement, and 20% (2/10) presented RET rearrangements (Figure 4). The proportion of TP53 co-mutation was 37.5% (6/16) and 47.5% (444/935) in MET exon 14 skipping and wild-type patients, respectively (Supplementary Figure 2). However, no significant difference was observed in both groups.
Discussion
Lung ADC patients harboring MET exon14 skipping mutations have been reported to represent a clinically unique molecular subtype of NSCLC. In the present study, RNA-based NGS was used for the detection of MET exon 14 skipping in 951 NSCLC patients. Since some somatic variants not encompassing the MET exon14 splice sites might induce MET exon 14 skipping, we speculated that RNA could serve as a favorable source of material for analyzing the phenomenon.
The prevalence rates of MET exon 14 skipping in lung ADC were 0.9–4.0% in Asian populations and 2.1–3% in the Western population. In the current study, the overall estimated prevalence of MET exon 14 skipping was 1.9% in ADCs and 1.7% in NSCLCs (Figure 1). Furthermore, we compared the detection rate of MET exon 14 skipping from different specimen sources, including surgical resection, PTNB, and TBLB. As shown in Figure 1A, the detection rate of MET exon 14 skipping from surgical resection specimen was 2.3% in NSCLC and 2.0% in ADC, respectively. Liu et al reported that the prevalence rate of MET exon 14 skipping in Chinese population with lung ADC was 0.9% as assessed by biopsy specimens in 2016. The low detection rate of MET exon 14 skipping in the study by Liu et al might be attributed to diverse specimen sources rather than ethnic differences.
Notably, MET exon 14 skipping is mutually exclusive of other known driver gene alterations.10,12 In this study, MET exon 14 skipping was identified in 6.6% of EGFR/KRAS/ALK/ROS1/RET-negative ADCs (13/196). This incidence was lower than that reported previously. Furthermore, the prevalence rate of MET exon 14 skipping was 19% (10/54 NSCLC cases) in Western patients without the mutation of other driver genes, such as EGFR, KRAS, BRAF, ERBB2, ALK and ROS1, while it is 37.8% (17/45) in East Asian patients.14,22 Thus, it is beneficial to screen MET exon14 skipping mutations in patients without other oncogenic driver mutations, as described previously23.
TP53 mutations were detected in 37.5% of the samples with MET exon 14 skipping, 46.2% of the samples with EGFR mutations, and 21.2% of the samples with ALK fusions. A previous study demonstrated that TP53 mutations reduced the responsiveness to crizotinib and worsened the prognosis in NSCLC patients with ALK rearrangements.23 Thus, additional studies are essential to determine the impact of TP53 mutations in response to crizotinib in NSCLC patients with MET exon 14 skipping. One patient with ADC harbored the MET exon 14 skipping and EGFR exon19 deletions (Figure 3), which could be attributed to tumor heterogeneity.24 Presently, the treatment of patients with multiple mutations lacks consensus on the appropriate TKI combination for optimal efficacy, thereby necessitating further investigation.
Several recent studies25,26 reported that PD-L1 expression in NSCLCs is inversely correlated with EGFR mutation and positively correlated with TP53 mutation and MET amplification. The current results also showed that PD-L1 is highly expressed in NSCLC patients with MET exon 14 skipping (69.2% vs 17.3%, P < 0.01), and significantly lower in patients with EGFR mutant cohort (9.3% vs 26.9%, P < 0.01). These data suggested that in the EGFR wild-type patients, MET mutation status and PD-L1 expression might be critical factors for personalized targeted therapy that downregulates the RAS/MAPK pathway.
The present study has several inherent limitations due to the retrospective design and insufficient follow-up period. The prevalence of MET exon 14 skipping in lung ADCs in the East Asian population was similar to that in the Western population as assessed using RNA-based NGS. The NSCLC patients with MET exon 14 skipping were older than those with other oncogenic driver mutations, such as EGFR, ALK, and ROS1. However, no significant differences were observed in the proportion of gender, smoking history, tumor location, and tumor stage between MET exon 14 skipping and wild-type NSCLCs. Moreover, PD-L1 was highly expressed in NSCLC patients with MET exon 14 skipping.
Conclusion
The prevalence of MET exon14 skipping in lung ADCs in the East Asian population was similar to that in the Western population as assessed by RNA-based NGS. The NSCLC patients with MET exon 14 skipping were older than those with other oncogenic driver mutations, such as EGFR, ALK, and ROS1. In addition, PD-L1 was highly expressed in NSCLC patients with MET exon 14 skipping.
Ethics Approval and Consent to Participate
This study was approved by the Ethics Committee of Beijing Chest Hospital (2017CHB06), and written informed consent was obtained from all patients.
Acknowledgments
We acknowledge all the patients who have participated in this study. We also appreciate the doctors from the Departments of Oncology and Pathology for technical help and the staff of Novogene Bioinformatics.
Author Contributions
All authors contributed to data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The abstract of this paper was presented at the 2019 ASCO Annual Meeting as a poster presentation with interim findings. The poster’s abstract was published in “Identification of MET exon 14 skipping by targeted RNA-based next-generation sequencing in Chinese Patients with NSCLC” in Journal of Clinical Oncology, DOI: 10.1200/JCO.2019.37.15_suppl.e20018. Drs Ziguang Xu and Lingfei Kong report grants from Major Science and Technology Projects in Henan Province (No.161100311400), during the conduct of the study. Peng Cheng and Fang Luo are employees of Novogene Co., Ltd., Beijing, China. Shijun Fu and Min Gao are employees of Pfizer Inc. The authors declare no other potential conflicts of interest.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68(1):7–30. doi:10.3322/caac.21442
2. Hirsch FR, Scagliotti GV, Mulshine JL, et al. Lung cancer: current therapies and new targeted treatments. Lancet. 2017;389(10066):299–311. doi:10.1016/S0140-6736(16)30958-8
3. Jenkins RW, Oxnard GR, Elkin S, Sullivan EK, Carter JL, Barbie DA. Response to crizotinib in a patient with lung adenocarcinoma harboring a MET splice site mutation. Clin Lung Cancer. 2015;16(5):e101–e104. doi:10.1016/j.cllc.2015.01.009
4. Paik PK, Drilon A, Fan PD, et al. Response to MET inhibitors in patients with stage IV lung adenocarcinomas harboring MET mutations causing exon 14 skipping. Cancer Discov. 2015;5(8):842–849. doi:10.1158/2159-8290.CD-14-1467
5. Birchmeier C, Birchmeier W, Gherardi E, Vande Woude GF. Met, metastasis, motility and more. Nat Rev Mol Cell Biol. 2003;4(12):915–925. doi:10.1038/nrm1261
6. Ma PC, Kijima T, Maulik G, et al. c-MET mutational analysis in small cell lung cancer: novel juxtamembrane domain mutations regulating cytoskeletal functions. Cancer Res. 2003;63(19):6272–6281.
7. Ma PC, Jagadeeswaran R, Jagadeesh S, et al. Functional expression and mutations of c-Met and its therapeutic inhibition with SU11274 and small interfering RNA in non-small cell lung cancer. Cancer Res. 2005;65(4):1479–1488. doi:10.1158/0008-5472.CAN-04-2650
8. Sadiq AA, Salgia R. MET as a possible target for non-small-cell lung cancer. J Clin Oncol. 2013;31(8):1089–1096. doi:10.1200/JCO.2012.43.9422
9. Cancer Genome Atlas Research N. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511(7511):543–550. doi:10.1038/nature13385
10. Frampton GM, Ali SM, Rosenzweig M, et al. Activation of MET via diverse exon 14 splicing alterations occurs in multiple tumor types and confers clinical sensitivity to MET inhibitors. Cancer Discov. 2015;5(8):850–859. doi:10.1158/2159-8290.CD-15-0285
11. Gow CH, Hsieh MS, Wu SG, Shih JY. A comprehensive analysis of clinical outcomes in lung cancer patients harboring a MET exon 14 skipping mutation compared to other driver mutations in an East Asian population. Lung Cancer. 2017;103:82–89.
12. Tong JH, Yeung SF, Chan AW, et al. MET amplification and exon 14 splice site mutation define unique molecular subgroups of non-small cell lung carcinoma with poor prognosis. Clin Cancer Res. 2016;22(12):3048–3056. doi:10.1158/1078-0432.CCR-15-2061
13. Liu SY, Gou LY, Li AN, et al. The unique characteristics of MET Exon 14 mutation in chinese patients with NSCLC. J Thorac Oncol. 2016;11(9):1503–1510. doi:10.1016/j.jtho.2016.05.016
14. Lee GD, Lee SE, Oh DY, et al. MET Exon 14 skipping mutations in lung adenocarcinoma: clinicopathologic implications and prognostic values. J Thorac Oncol. 2017;12(8):1233–1246. doi:10.1016/j.jtho.2017.04.031
15. Awad MM, Oxnard GR, Jackman DM, et al. MET Exon 14 mutations in non-small-cell lung cancer are associated with advanced age and stage-dependent met genomic amplification and c-met overexpression. J Clin Oncol. 2016;34(7):721–730. doi:10.1200/JCO.2015.63.4600
16. Schrock AB, Frampton GM, Suh J, et al. Characterization of 298 patients with lung cancer harboring MET Exon 14 skipping alterations. J Thorac Oncol. 2016;11(9):1493–1502. doi:10.1016/j.jtho.2016.06.004
17. Travis WD, Brambilla E, Burke AP, Marx A, Nicholson AG. WHO classification of tumors of the lung, pleura, thymus, and heart. Lyon. 2015;1:9–152.
18. Detterbeck FC, Boffa DJ, Kim AW, Tanoue LT. The eighth edition lung cancer stage classification. Chest. 2017;151(1):193–203. doi:10.1016/j.chest.2016.10.010
19. Li W, Qiu T, Guo L, Ying J. Major challenges related to tumor biological characteristics in accurate mutation detection of colorectal cancer by next-generation sequencing. Cancer Lett. 2017;410:92–99. doi:10.1016/j.canlet.2017.09.014
20. Roach C, Zhang N, Corigliano E, et al. Development of a companion diagnostic PD-L1 Immunohistochemistry assay for pembrolizumab therapy in non-small-cell lung cancer. Appl Immunohistochem Mol Morphol. 2016;24(6):392–397. doi:10.1097/PAI.0000000000000408
21. Spigel DR, Ervin TJ, Ramlau RA, et al. Randomized Phase II trial of Onartuzumab in combination with erlotinib in patients with advanced non-small-cell lung cancer. J Clin Oncol. 2013;31(32):4105–4114. doi:10.1200/JCO.2012.47.4189
22. Heist RS, Shim HS, Gingipally S, et al. MET Exon 14 skipping in non-small cell lung cancer. Oncologist. 2016;21(4):481–486. doi:10.1634/theoncologist.2015-0510
23. Forde PM, Chaft JE, Smith KN, et al. Neoadjuvant PD-1 blockade in resectable lung cancer. N Engl J Med. 2018;378(21):1976–1986. doi:10.1056/NEJMoa1716078
24. Fan J, Dai X, Nie X. Concomitant epidermal growth factor receptor mutation and EML4-ALK fusion in a patient with multifocal lung adenocarcinomas. J Thorac Oncol. 2018;13(3):e45–e48. doi:10.1016/j.jtho.2017.11.130
25. Gainor JF, Shaw AT, Sequist LV, et al. EGFR mutations and ALK rearrangements are associated with low response rates to PD-1 pathway blockade in non-small cell lung cancer: a retrospective analysis. Clin Cancer Res. 2016;22(18):4585–4593. doi:10.1158/1078-0432.CCR-15-3101
26. Albitar M, Sudarsanam S, Ma W, et al. Correlation of MET gene amplification and TP53 mutation with PD-L1 expression in non-small cell lung cancer. Oncotarget. 2018;9(17):13682–13693. doi:10.18632/oncotarget.24455
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.