Back to Journals » Journal of Hepatocellular Carcinoma » Volume 6
Genetic Biomarkers For Hepatocellular Carcinoma In The Era Of Precision Medicine
Authors Duan J, Wu Y, Liu J , Zhang J, Fu Z, Feng T, Liu M, Han J, Li Z, Chen S
Received 26 July 2019
Accepted for publication 18 September 2019
Published 18 October 2019 Volume 2019:6 Pages 151—166
DOI https://doi.org/10.2147/JHC.S224849
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Ahmed Kaseb
Jingxian Duan,1 Yuling Wu,2 Jikui Liu,3 Jiajia Zhang,2 Zhichao Fu,2 Tieshan Feng,2 Ming Liu,2 Jie Han,2 Zhicheng Li,1 Shifu Chen1,2
1Department of Biomedical and Health Engineering, Shenzhen Institutes of Advanced Technology, Chinese Academy of Sciences, Shenzhen, People’s Republic of China; 2Department of Oncology, HaploX Biotechnology Co. Ltd, Shenzhen 518000, People’s Republic of China; 3Department of Hepatobiliary and Pancreatic Surgery, Peking University Shenzhen Hospital, Shenzhen 518036, People’s Republic of China
Correspondence: Shifu Chen
Department of Biomedical and Health Engineering, Shenzhen Institutes of Advanced Technology, Chinese Academy of Sciences, Shenzhen, People’s Republic of China
Tel +86 135 1042 7830
Email [email protected]
Abstract: Being one of the most lethal cancers that exhibit high levels of heterogeneity, hepatocellular carcinoma (HCC) is associated with diverse oncogenic pathways underpinned by varied driver genes. HCC can be induced by different etiological factors including virus infection, toxin exposure or metabolic disorders. Consequently, patients may display varied genetic profiles, and may respond differently to the treatments involving inhibition of target pathways. These DNA/RNA mutations, copy number variations, chromatin structural changes, aberrant expression of non-coding RNAs and epigenetic modifications were considered as biomarkers in the application of precision medication. To explore how genetic testing could contribute to early diagnosis, prognosis, treatment and postoperative monitoring of HCC, we conducted a systematic review of genetic markers associated with different pathologies. Moreover, we summarized on-going clinical trials for HCC treatment, including the trials for multiple kinase inhibitors and immune checkpoint blockade (ICB). The efficacy of ICB treatment in HCC is not as good as what was observed in lung cancer and melanoma, which might be due to the heterogeneity of the microenvironment of the liver.
Keywords: genetic biomarkers, hepatocellular carcinoma, genomic sequencing, clinical trials
Introduction
Liver cancer is considered to be the fourth most lethal cancer globally, and hepatocellular carcinoma (HCC) accounts for 75–85% of liver cancer cases.1 In addition to the high mortality rate, the prognosis and treatment of HCC are suboptimal, most of the patients reach malignancy within a year of initial diagnosis.2 The survival statistics of the American cancer society from 2008 to 2014 showed that the the overall 5-year survival rate was 18% for liver cancer patients, but the 5-year survival rate for patients with distant metastasis was only 2%. In great contrast, among early-stage HCC patients who were diagnosed and treated before extrahepatic metastasis, the 5-year survival rate would be increased to 31%. To improve HCC early diagnosis rate, HCC biomarkers with higher sensitivity and specificity are required. Postoperative monitoring, which aims to evaluate disease progression and predict cancer recurrence, also heavily relies on the exploration of HCC biomarkers. Recently, targeted therapy, immune checkpoint blockade therapy, and combinational therapy showed promising efficacy in clinical trials. Biomarkers also play an important role in the design of personalized treatment plans. In the new era of genomic oncology, genetic biomarkers are becoming the core of cancer biomarkers. To bring a panoramic view of HCC genetic markers to academic and clinical experts, we conducted a systemic review of these genetic biomarkers for HCC early diagnosis, prognosis, treatment and postoperative monitoring.
Etiology And Pathogenesis
The primary risk factors of HCC are chronic hepatitis B and hepatitis C virus infection, alcohol consumption, non-alcoholic fatty liver disease, exposure to dietary aflatoxin, genetic hemochromatosis, and metabolic disorders.3,4 The resulting chronic liver inflammation may develop to severe liver fibrosis and cirrhosis, which were predispositions of HCC. It was reported that up to 90% of HCC cases occurred on the background of liver cirrhosis or fibrosis.5 Increased production of ROS was predicted to cause the accumulation of oxidative stress and DNA instability, which were accompanied by hepatocytes proliferation, telomeres shortening, and chromosomal alterations. These processes were associated with tumor development in fibrosis according to early studies.6,7 Interestingly, each HCC risk factor is involved in differed signaling pathways during carcinogenesis as Figure 1 shows, and the resulting HCC patients often exhibit distinct genomic profiles.
Hepatitis B Virus Infection
In HBV endemic regions such as Asia-Pacific and sub-Saharan Africa, HBV infection accounts for 75–90% of HCC incidences.8 Once entered the host cell, the HBV DNA transcribes to 4 viral mRNA for 7 HBV proteins,9,10 one of which is the 17kDa polypeptide HBV X (HBx) that regulates cell proliferation and apoptosis by modulating Wnt/β-catenin expression.11 As Figure 1 shows, overexpression of HBx could also activate NF-κB to block tumor necrosis factor-α(TNFα)- and FAS-mediated apoptosis. In addition, both HBV and HCV can cause mitochondrial stress and increase reactive oxygen species (ROS) levels, which triggers endoplasmic reticulum (ER) stress and the unfolded protein response (UPR), leading to autophagy promoted cell survival and virus persistence.12
Given that different risk factors induce HCC through varied mechanisms, the genomic profiles of patients affected by those risk factors may differ. An early study screened biomarkers for HBV induced HCC and identified 7 up-regulated genes in those patients including RPS5, KRT8, CFLAR, ATP5F1, IGFBP2, MAP3K5, and MMP9. The genes regulate diverse cellular processes ranging from protein synthesis to cytoskeleton organization.13 More recent research showed that CCND1,14 BCL2, Mcl-1,15 NFKB116 and SOCE17 were also up-regulated in HBV induced HCC. Additionally, one whole-exome sequencing study reported that TP53, CTNNB1, RB1, AXIN1, SELPLG, and FGF19 appear to be the candidates of driver mutation genes for HBV induced HCC.18 The unique genomic profile of HBV induced HCC can be applied to the early diagnosis of HCC, which attributes to the implement of precise curative treatment that may improve the survival of patients.
Hepatitis C Virus Infection
Being the second most common cause of HCC worldwide, HCV infection is responsible for at least 10% of HCC incidences.19 HCV infection is a major cause of HCC in Western countries, Africa and Japan.8,20 The core proteins of HCV such as E2, NS5A, and NS5B were shown to interfere with the E2F1 pathway and RAF/MAPK/ERK kinase pathways (Figure 1), by which they may modulate cell proliferation and tumor development.21 The other HCV produced protein, NV5a, was reported to inhibit the p53 pathway, which consequently modulates cell cycle.22
Apart from carcinogenesis, data suggest that HCV induced HCC can be distinguished from HBV induced HCC based on the genomic profile of the patients. An early study compared the gene expression pattern of HCV and HBV induced HCC, they concluded that different genes were up-regulated in the two scenarios.13 Unlike the HBV infected population, VIM, ACTB, GAPD, and CD58 were up-regulated in HCV induced HCC cases. A later study identified 40 up-regulated genes in HCV triggered HCC cases compared to the controls, including RYBP, ATP1B3, TMC, ZNF567, GPR108, CD19.23 These studies identified potential biomarkers for HCV induced HCC, which are crucial to the design of treatment strategy for precision medicine.
Alcohol Consumption
Alcohol-related cirrhosis is the third most common trigger of HCC worldwide, which appears to correlate with alcohol consumption behavior.24 Alcohol intake may increase the production of iron-induced reactive oxygen species (ROS), which would interfere with DNA repair mechanisms (Figure 1). Moreover, acetaldehyde is formed during ethanol metabolization, the accumulation of acetaldehyde has negative effects on DNA and proteins.25
Patients carrying alcohol-related HCC also exhibit unique genetic profiles. It was reported that mutations in TRET promoter, CTNNB1, ARID1A are more common in alcohol-related HCC incidences.26 A recent study identified 5 up-regulated genes (CSMD1, MAGEA3, MAGEA6, CSAG1, and CSAG3) and 4 down-regulated genes (CD5L,UROC1, IGF2, and SLC22A10) that were associated with alcohol-related HCC.27
Exposure To Aflatoxin
Exposure to aflatoxin B1 (AFB1) is identified as a risk factor for HCC, which is propagated via food contaminations. AFB1 and HBV were proposed to have synergistic interaction.28 Although the mechanism remains unclear, HBV infection seems to sensitize hepatocytes to the carcinogenic effects of AFB1.29 AFB1 may trigger carcinogenesis via the transformation to aflatoxin-8, 9-exo epoxide, which interacts with the p53 tumor suppressor gene (Figure 1), and facilitates the mutation at R249S.29,30
Compared to other risk factors, the biomarker for AFB associated HCC was not extensively studied. In addition to TP53, One recent study claimed that frequent mutation in the adhesion G protein coupled receptor B1 (ADGRB1), AXINI and TERT were observed in AFB associated HCC cases.31
Non-Alcoholic Fatty Liver Disease And Non-Alcoholic Steatohepatitis
Non-alcoholic fatty liver disease (NAFLD) is one of the most common risk factors of chronic liver disease in the US, it is frequently associated with cirrhosis, which might lead to HCC.32 NAFLD represents a variety of chronic liver diseases ranging from hepatic steatosis to the progressed and inflammatory form non-alcoholic steatohepatitis (NASH). In particular, NASH-related HCC incidence increased by 63% from 2002 to 2012 in the US.33 A study reported that NAFLD and alcohol consumption contributed to steatohepatitis.34 Together with HCV-NS5A stimulated TLR4-NANOG and the leptin-phosphorylate STAT3 signaling pathways, NAFLD results in HCC by up regulating TWIST1 in tumor-initiating stem-like cells. NAFLD may alter the expression of KCTD17, PHLPP2, and CPT2, which promotes hepatic steatosis, NASH, and hepatocarcinogenesis.35,36 Similarly, The missense mutation of the PNPLA3 gene was reported to promote NAFLD/NASH-related HCC.37 Moreover, one study identified 41 candidate genetic markers for NAFLD associated HCC including Sav1, Son, Slc25a17, Fbxo11, Myo10, and Pten, the mutation seems to promote apoptosis and fibrogenesis.38
Staging Systems
The severity of cancer is classified by cancer staging systems. A number of systems have been proposed to predict the prognosis of HCC patients, they considered different variables and were tested in varied populations. Therefore, none of the staging schemes has been universally applied. The commonly adopted staging and scoring system for HCC prognosis are the tumor, node, metastasis (TNM) staging, the Okuda stage, the Barcelona Clinic Liver Cancer (BCLC) systems, the Cancer of the Liver Italian Program (CLIP) score, the Japan Integrated Staging (JIS) scores, the Chinese University Prognostic Index (CUPI) scores, the French scores, and the albumin-bilirubin (ALBI) grading system (Table 1).
 |
Table 1 Staging And Scoring System Of HCC |
Different staging and scoring system were selected based on the clinical and scientific requirement. For instance, BCLC scoring was applied in clinical trials evaluating sorafenib, and sorafenib is recommended to be the treatment option for BCLC grade C.39,40
It is generally accepted that diagnosis at the early stage could significantly improve the survival of patients by allowing the implement of curative treatment. Furthermore, the design of treatment strategy is based on the understanding of the cause and stage of each individual. In that sense, diagnosis methods for precision medication should be developed for the identification of HCC at very early stage.
Early Diagnosis Of HCC
Diagnosis at an early stage is the key to HCC patient survival. Currently, the most widely used biomarker for HCC diagnosis is serum alfa-fetoprotein (AFP), but its sensitivity and specificity are both around 50%.41 It is worth noting that liquid biopsy has been extensively developed and put into clinical practice over the past decades. It mainly detects circulating tumor DNA (ctDNA), circulating tumor cell (CTC), exosomes, and circulating tumor RNA (ctRNA) in body liquid including plasma (Figure 2), urine, and cerebrospinal fluid. Among them, ctDNA is the most widely applied genetic biomarker. It is derived from tumor tissue, and carries somatic mutations, CNVs, DNA methylations, viral sequences, and physical characteristics associated with carcinogenesis.
Somatic Mutations
The genomic landscape of HCC has been revealed by multiple genome sequencing studies. The three most frequently mutated genes are TERT (40–60%), TP53 (31%), and CTNNB1 (27%). One systematic analysis summarized frequently mutated genes reported, including the tumor suppressor genes AXIN1 (8%) and RB1 (4%), the chromatin remodeling genes ARID1A (7%), ARID2 (5%) and BAP1 (5%), the cellular anti-oxidant defense genes NFE2L2 (3%) and its interactor KEAP1 (5%), Albumin (13%) and 10% APOB mutations.42 These mutational features were applied in the design of sequencing panels for HCC early screening.
One research group developed a liquid biopsy assay named hepatocellular carcinoma screen (HCCscreen), which could detect HCC in asymptomatic HBsAg-seropositive individuals. The assay simultaneously assesses the ctDNA gene status of TERT, TP53, CTNNB1, and AXIN1, and the level of serum AFP and DCP, as well as the HBV integration profile.In the training cohort, the assay effectively distinguished HCC patients from non-HCC individuals, and the sensitivity and specificity of the assay were 85% and 93% correspondingly. In the validation cohort, the assay showed 100% sensitivity and 94% specificity. Notably, while the training cohort recruited individuals who had liver nodules and/or elevated serum AFP levels, the validation cohort only enrolled individuals with normal serum AFP levels and liver ultrasonography results.43 Another study reported that the TP53 mutation at codon 249 could be used as a biomarker to identify HCC caused by aflatoxin exposure or HBV infection. Its sensitivity and specificity reached 40% and 88% respectively.44 These results were encouraging in the sense that it provided a method for HCC screening at a very early, or even asymptomatic stage.
DNA Methylations
The alteration of DNA methylation status is an early event in carcinogenesis, so it is also considered as a potential biomarker for HCC early detection. One study developed an HCC-specific methylation marker panel and built a diagnostic prediction model with ten markers. The sensitivity and specificity of the model were 85.7% and 94.3% in the training cohort (715 HCC patients and 560 non-HCC individuals), and 83.3% and 90.5% in the validation cohort (383 HCC patients and 275 non-HCC individuals). Furthermore, the model could effectively distinguish HCC induced by varied risk factors.45 Another study performed by Abderrahim Oussalah et al evaluated the accuracy of a PCR-based cfDNA assay for SEPT9 promoter methylation. The area under the receiver operating characteristic curve was 0.944, showing it could be served as a potential biomarker for HCC diagnosis.46
Physical Characteristics
The physical characteristics of ctDNA are different from non-tumor-derived cfDNA in the aspects of size profiles and preferred end coordinates. Dennis Lo et al performed a detailed analysis of the size profiles of plasma DNA in patients with HBV induced HCC and healthy controls. The results showed that aberrantly short or long DNA molecules existed in the plasma of HCC patients. The short DNAs preferentially carried the tumor-associated copy number aberrations.47 Moreover, they revealed specific end coordinates of tumor-associated plasma DNA.48 By calculating the ratio of sequence with tumor-associated DNA ends versus sequence without the characteristic, the possibility of carrying HCC can be estimated, and the area under the receiver operating curve was 0.88.
CTCs, Exosomes And ncRNAs
CTCs are derived from advanced tumors, they may promote tumor metastasis. In a meta-analysis study, the pooled sensitivity and specificity of CTC for the detection of tumor were 67% and 98% respectively.49 Similarly, tumor-derived exosomes were shown to deliver oncogenes, pathogens and microRNAs. It was reported that a number of exosomal proteins, RNAs and miRNAs may serve as biomarkers for HCC including miR-122, miR-21, and miR192.50 Consistently, Huang et al performed a systematic review and meta-analysis of published studies to evaluate the utility of microRNAs (miRNAs) in the early diagnosis of HCC. They analyzed 50 studies that included 3423 cases of HCC, 2403 chronic hepatic disease (CH) patients, and 1887 healthy controls. It showed that miRNA could be used as a marker to discriminate HCC patients from healthy individuals, and the sensitivity and specificity were 75.8% and 75.0% correspondingly.51 LncRNAs (long non-coding RNAs) are non-coding mRNA-like trasncripts that are longer than 200 nucleotides. Many novel lncRNAs were proposed to be predictive biomarkers for cancer diagnosis or prognosis, including HULC, HOTAIR, MALAT1, and H19.52 In the case of HCC, lnc-Myd88 was suggested to be a new diagnostic and therapeutic target for HCC because that it could increase Myd88 expression, which in turn activates NF-κB and PI3K/AKT signaling pathways.53
Prognosis And Post-Operational Monitoring Of HCC
The primary treatment strategy for HCC is to remove the tumor by surgery, but HCC may relapse after the operation. It was reported that the 5-year recurrence rate of HCC after surgery was 74.2% in Japan and Korea.54 Therefore, post-operational monitoring and precise prognosis are necessary for cancer treatment. Genetic sequencing might be an alternative tool for cancer post-operational monitoring. For example, ALB1 mutation was observed in recurrent liver cancer, the time that the mutation can be detected in ctDNA is associated with the time when the HCC relapses.55 A unique mutation of HCK p.V174M was also observed in recurrent HCC and metastatic HCC, which was diminished after surgery but would increase rapidly if HCC relapses.56 Similarly, the IL-28B (rs8099917) TT genotype was interrelated with HCC recurrence.57 Taken those facts together, genetic sequencing may serve as an important tool for the post-operational surveillance of cancer.
Genetic testing also serves as a useful tool for HCC prognosis evaluation. A number of studies have identified biomarkers associated with the prognosis of HCC, such as miR-203,58 ATXN7,59 and Alpha-1-fucosidase.60 A study demonstrated that the concentration of HMGB1 in blood was linked to prognosis of patients with HCC after receiving sorafenib treatment for 4 weeks.61 In addition, it was reported that the profile of circulating tumor DNA in blood samples of HCC patients could reveal the heterogeneity of tumors and monitor the process of disease in real time.56
Targeted Therapy Of HCC
U.S Food and Drug Administration has approved 7 drugs for HCC treatment (Table 2), which were applied for targeted therapy or immunotherapy. Sorafenib, lenvatinib, regorafenib, ramucirumab and cabozantinib are inhibitors of receptor tyrosine kinase (TK). These drugs can suppress the activities of major TKs including vascular endothelial growth factor receptors (VEGFR1-3), platelet-derived growth factor receptor β (PDGFR-β) and fibroblast growth factor receptors 1–4 (FGFR1-4), thereby inhibiting the growth of tumor cell by antiangiognic effects.62–65 HCC is majorly supplied by hepatic arteries whereas liver parenchymas are supported primarily by portal vein. In fact, radio-graphically visible tumors rely entirely on the oxygen and nutrients supply via hepatic vasculature.66 Therefore, one of the strategy applied in HCC treatment is to target the angiogenesis pathways.
 |
Table 2 FDA Approved HCC Drugs And Their Molecular Targets |
As the first drug that has been shown to increase the overall survival of patients with liver cancer, sorafenib inhibits the serine/threonine kinases that are the crucial components of the Raf/MEK/ERK pathway (Figure 3). Moreover, the drug could suppress the activity of vascular endothelial growth factor receptors (VEGFR1-3) and platelet-derived growth factor receptor β (PDGFR-β), thereby inhibiting the growth of tumor cell.67 Consistently, a phase III clinical trial named SHARP demonstrated that sorafenib significantly improved the median survival time (mOS) of the patients compared with the placebo (10.7 months vs 7.9 months).68 A trial in the Asia-Pacific region also showed that sorafenib extended the mOS of patients to 6.5 months, whereas the mOS of the placebo group was 4.2 months.69
Lenvatinib was discovered by the Tsukuba Research Laboratory in Japan as an angiogenesis inhibitor,70 it is a multiple receptor tyrosine kinase inhibitor that targets vascular endothelial growth factor receptors (VEGFR1-3), fibroblast growth factor receptors (FGFR1-4), PDGRα, KIT and RET.62,63,70 Lenvatinib blocks the VEGF pathway and inhibits angiogenesis, exerting anti-tumor activity (Figure 3). The drug has been tested in Phase II and Phase III trials for the treatment of advanced HCC. The phase III trial showed that lenvatinib is comparable to sorafenib in terms of OS. In addition, the incidence of fatal adverse events associated with lenvatinib treatment (including liver failure, cerebral hemorrhage, and respiratory failure) was 2%, which is higher compared to sorafenib (1%).71,72 A series of studies showed that lenvatinib could be an alternative first- line treatment for patients with advanced- stage HCC.71 In August 2018, the US FDA approved lenvatinib as a first-line therapy for advanced liver cancer.
Regorafenib is a tyrosine kinase inhibitor that shares structural similarity with sorafenib, its pharmacological targets include VEGFR1-3, c-TKITIE-2, PDGFR-β, FGFR-1, RET, c-RAF, BRAF and p38MAP kinase.73 Regorafenib inhibits multiple protein kinases that are involved in tumor angiogenesis and oncogenesis, proliferation, and metastasis.74 A Phase II clinical trial conducted by Bruix et al showed that regorafenib was safe as a second-line treatment for advanced liver cancer.73 The progression free survival (PFS) and mOS of the 36 patients were 4.3 months and 13.8 months, respectively. Another multicenter, phase III clinical trial demonstrated that regorafenib increased survival of advanced liver cancer patients with disease progression after sorafenib treatment. Compared with the placebo group (mOS 7.8 months, mortality rate 20%), regorafenib can extend the median survival to 10.6 months and reduce the mortality rate to 13%.75
Cabozantinib inhibits tyrosine kinases (Figure 3), including vascular endothelial growth factor receptors 1, 2, and 3, MET, and AXL.65,76 It is a more potent inhibitor of MET, AXL, RET, FLT3, and TIE-2 compared to regorafenib.77 VEGF, MET, and AXL are involved in tumor proliferation and angiogenesis, thereby cabozantinib can inhibit tumor growth. Abou-Alfa et al showed that among patients who have been treated for HCC, the overall survival was 10.2 months in cabozantinib treated group, and the overall survival of the placebo group was 8.0 months.78 Moreover, the median progression-free survival was 5.2 months in the cabozantinib treated group whereas that of the placebo group was 1.9 months. These results showed that cabozantinib could improve the overall survival of patients who were intolerant to or had progressive disease after sorafenib treatment.
Biomarkers For Targeted Therapy
Therapeutic effects of drugs might be estimated based on the genetic profiles of patients. Yeon-Su Lee et al identified a total of 1813 genomic variations associated with sorafenib responsiveness, 708 of which located within regions transcribing genes associated with sorafenib responsiveness. These genes are involved in drug absorption, distribution, and drug metabolism pathway.79 Another study that focused on precision medicine reported that the mutation in RSK2 led to the lasting activation of RAS, which was associated with HCC resistance to sorafenib. Moreover, Teufel et al reported 9 plasma miRNAs and 49 variants in 27 oncogenes or tumor suppressor genes, these miRNA and mutations might be used to identify HCC patients that are most likely to respond to regorafenib.77 In terms of targeted therapy by antibodies, data suggested that the growth of HCC cells harboring FGF or CCND1 amplification were selectively inhibited by the anti-FGF19 antibody 1A6.80
Immunotherapy Of HCC
Immune Microenvironment Of The Liver
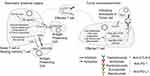
Various types of immune cells reside in the liver and produce different cytokines and growth factors in response to local stimulation. As the result, the immune cell repertoire built up an immune microenvironment that maintains the balance between immune tolerance and immune activation in the liver. The key components of the immune repertoire are macrophages, dendritic cells, myeloid-derived suppressor cells, natural killer (NK) cells, NK T cells, T cells and B cells. By single cell RNA sequencing, studies have identified subsets of immune cells that may conduct distinct immune functions. Two populations of intrahepatic CD68+ macrophages were observed in liver, one was characterized as inflammatory macrophages while the other was suggested to be tolerogenic.81 Similarly, the T cell and B cell population can be further divided to subsets that expressing various gene markers, including CD3+ γδ T cells and phosphoantigen-reactive γδ T cells, antigen inexperienced B cells as well as plasma B cells. The heterogeneity of intrahepatic immune cell subsets was confirmed by another single cell RNA sequencing study, which discovered CD163 + VSIG4 + Kupffer cells, MS4A1 + CD37 + subset of B cells, and CD56- and CD56 + CD8A + NKT cells.82 Based on the characterization of the immune microenvironment, HCC patients can be subdivided to high, medium or low immunity groups for prognosis prediction. It was reported that patients with high immunity exhibited poorly cytokeratin 19 (CK19)+, and/or Sal‐like protein 4 (SALL4)+ high‐grade HCC, which was associated with significantly better prognosis.83 The medium and low immunity groups were suggested to adapt distinct treatment strategies for better outcomes. Therefore, the efficacy of immunotherapy was partially determined by the immune microenvironment of individuals.
Immunotherapy
Immunotherapy has been extensively studied in the past decades, and has made significant progress in the field of cancer therapy.84 By targeting the immune checkpoint receptors or ligands including cytotoxic T lymphocyte antigen 4 (CTLA-4), programmed cell death protein 1 (PD-1), and programmed cell death 1 ligand 1 (PD-L1), immunotherapy breaks immune tolerance and delays tumor progression.85
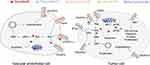
CTLA-4 is mainly expressed on the surface of activated T cells and regulates the activation of T lymphocytes at the early stages of the tumor immune cycle (Figure 4). By binding to ligand B7-1/B7-2, and CTLA-4 suppresses the activity of T cells and facilitates tumor immune escape.86 CTLA-4 is mainly expressed on the surface of activated T cells and regulates the activation of T lymphocytes at the early stages of the tumor immune cycle (Figure 4). By binding to ligand B7-1/B7-2, and CTLA-4 suppresses the activity of T cells and facilitates tumor immune escape (137). CTLA-4 antibodies can block the interaction between CTLA-4 and B7-1/B7-2, promote the binding of B7 to the stimulatory receptor CD28 and restore the activity of T cells.
PD-1 is mainly expressed on the surface of immune cells in peripheral tissues, especially in the tumor microenvironment. It was an unexpected discover when studying T cell apoptosis. PD-1 was later identified as a receptor that negatively regulates the immune response. The PD-1 ligands PD-L1 and PD-L2 were discovered in 2000.87 Studies have shown that PD-1 and PD-L2 can block T cell activation, proliferation and the production of cytokines, such as interferon-γ (IFN-γ), thereby exerting a negative immune-modulatory effect which results in immunosuppression.88 Receptor PD-1 on the surface of T lymphocytes binds to PD-L1 on the surface of tumor cells to attenuate the immune responses and inhibits tumor cell elimination by T lymphocytes in the tumor bed.89 Therefore, blocking the PD-1/PD-L1 interaction can alleviate tumor-induced immunosuppression and restore the body’s immune activity.
Based on a series of clinical trials (Supplementary Table 1) including CheckMate 057, CheckMate 040,90 KEYNOTE-224,91 and KEYNOTE-042, both nivolumab and pembrolizumab were approved by the FDA for treatment of patients with HCC who have been previously treated with sorafenib.
Biomarkers For Immunotherapy
The efficacy of immunotherapy can be predicted by genetic markers. A recent study has shown that low PD-L1 expression is associated with poor clinical outcomes in respond to immunotherapy. The objective clinical responses were observed in 26% (9/34) of patients with PD-L1 expressed in at least 1% of their tumor cells. In contrast, among patients carrying tumor cells with less than 1% of PD-L1 expression, only 19% (26/140) of them responded to ICB treatment.90 On a larger scale, high tumor mutation burden (TMB) is an emerging biomarker that reflects the sensitivity of patients to immune checkpoint inhibitors. TMB measured by hybrid capture-based NGS interrogating 1.2 Mb of the genome can predict clinical outcomes of anti-PD-1/PD-L1 immunotherapy in many tumor types.92 One study has shown that HCC patients with Wnt-β-catenin pathway mutations responded poorly to immune checkpoint blockers. All patients experienced disease progression after treatment, and their median survival was significantly shorter than the patients without the aforementioned mutations (9.1 months vs 15.2 months).93 Despite the fact that the incidence of high microsatellite instability (MSI-H) in HCC is estimated to be low,94 the FDA has approved pembrolizumab for the treatment of advanced-stage cancers with MSI-H regardless of the origin of the cancer.95 Collectively, these results implicate that specific DNA mutations may serve as biomarkers for ICB response prediction.
Clinical Trials For HCC Treatment
Clinical trials have been conducted to assess the performance of targeted therapy and immune therapy for HCC treatment. Several large-scale on-going trials focus on the FDA approved drugs for targeted therapy. Sorafenib and lenvatinib have been approved by the FDA as the first-line treatment for HCC patients whose disease cannot be removed by surgery. Notably, recent phase III trial confirmed that lenvatinib was non-inferior to sorafenib in overall survival in untreated HCC (NCT01761266), the progression free survival (7.4 vs 3.7 months, median) and time to progression (8.9 vs 3.7 months, median) of the lenvatinib group were both better than the sorafenib group.71 The trial (NCT01908426) that enrolled 707 participants has reported that cabozantinib could lead to longer overall survival and progression-free survival than placebo in patients with previously treated advanced HCC.78 The median overall survival and progression-free survival were extended to 10.3 months and 5.2 months respectively, with the objective response rate of 4%. Similarly, a trial involving 1000 participants for regorafenib is also underway (NCT03289273). Interestingly, a monoclonal antibody named ramucirumab was applied in targeted therapy as a vascular endothelial growth factor receptor 2 (VEGFR2) inhibitor. The results showed that ramucirumab significantly improved the median overall survival and progression free survival compared to the placebo group.96,97 At the median follow-up of 7.6 months, the median overall survival and pregression free survival reached 8.5 months and 2.8 months correspondingly.
Numerous data were generated from clinical trials with drugs that have not been approved by FDA. Apatinib, a reversible dual tyrosine kinase inhibitor that selectively targets and inhibits HER2 and EGFR,98 is tested against placebo in a phase III clinical trial (NCT02329860). The preliminary results showed that 50% patients survived longer that 11.4 months following apatinib administration.99 The stable disease rate (40.9%) was considerably higher than the progressive disease rate (18.2%). Furthermore, the highly selective FGFR4 inhibitor BLU-554 (NCT02508467), the ATP-competitive cyclin A/CDK2 inhibitor milciclib (NCT03109886) and the dual TORC1/TORC2 inhibitor ATG-008 (NCT03591965) were also tested against HCC. The primary results suggest that BLU-554 might be selective for FGF19 IHC-positive HCC.100 No conclusions have been drawn for milciclib and ATG-008.
Nivolumab (Opdivo) and pembrolizumab (Keytruda) were widely applied in clinical trials of immunotherapy. CheckMate-040 (NCT01658878) had confirmed that nivolumab induced active responses in patients with advanced HCC.90 The follow-up of CheckMate 459 was announced on EMSO 2019 congress, which reported that clinical benefits following nivolumab treatment were observed in predefined subgroups including hepatitis infection status, regions (Asia vs non-Asia), and vascular invation/matastasis status.101 However, the Bristol-Myers Squib announced in June 2019 that the randomized Phase 3 study evaluating Opdivo (nivolumab) versus sorafenib as a first-line treatment in patients with unresectable HCC failed to achieve statistical significance for its primary endpoint of overall survival (OS) per the pre-specified analysis. Similarly, the KEYNOTE-224 trial (NCT02702414) reported that pembrolizumab is effective in HCC patients previously treated with sorafenib.91 The primary result of KEYNOTE-240 was released on 2019 ASCO, which showed that pembrolizumab improved progression free survival over placebo. However, the comparisons were not statistically significant according to the prespecified statistical criteria.102
Ipilimumab (Yervoy) is an inhibitor targeting CTLA-4. Concurrent clinical trials primarily focused on the combined effects of ipilimumab and nivolumab. The follow-up of CheckMate-040 reported that the combination obtained meaningful clinical responses, the result of arm A showed that the median overall survival was extended to 23 months.103
Combination Of Treatments
The combined application of targeted therapy, immunotherapy or locoregional therapy may give rise to great clinical outcomes. A trial in the US assessed the combined effect of sorafenib and carotuximab (TRC105), an antibody against an important angiogenic target named endoglin. The primary results indicated that TRC205+ sorafenib may give rise to additional activity regarding the treatment of advanced/metastatic HCC.104 A follow-up trial with the same combination is on-going, the combined effect of target therapy and immunotherapy is very promising and the outcome is expected in 2020 (NCT02560779). Another on-going multi-center study is testing the combined efficacy of nivolumab and a novel transforming growth factor beta receptor I kinase inhibitor named galunisertib. The study aimed to include 100 participants and finish in December 2019 (NCT02423343). Similarly, the effect of lenvatinib in combination with pembrolizumab as first-line treatment for HCC is tested in 750 participants, and the multi-centered, phase III trial would be completed in 2022 (NCT03713593). The efficacy of such combinations were confirmed by the primary result of one clinical trial (NCT03289533), which exhibited that the combined use of avelumab and axitiib reduced the sizes of tumors in more than 68% of patients.105
Regarding the combination of immunotherapies, AstraZeneca sponsored a global phase III trial that aimed to enroll 1310 patients for the assessment of durvalumab plus tremelimumab as first-line HCC treatment (NCT03298451). The trial was designed based on the promising results of a previous study, which reported that the combined treatment of durvalumab and tremelimumab showed positive clinical activity in 70% of uninfected HCC patients at ≥16 weeks follow-up.106
Targeted therapy is also combined with locoregional treatment. One clinical trial (NCT03838796) in China involved 482 HCC patients for the observation of the effect of lenvatinib combined with transcatheter arterial chemoembolization (TACE), and the primary study might be completed in 2021 (NCT03838796). Another clinical trial revealed that TACE plus radiofrequency ablation achieved superior efficacy compared to TACE alone, the median overall survival was improved to 29 months whereas that of the TACE group was 18 months.107
Geographic Distribution Of Clinical Trials In Relation To HCC Incidence Rates
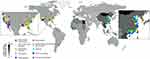
To overview the geographic distribution of clinical trials, 96 on-going clinical trials for HCC treatment were summarized in Supplementary Table 2 and tagged onto the HCC incidence world map, in which the HCC incidence rate was reflected by color intensity (Figure 5). It appears that the high HCC incidence rates were mostly observed in East Asian and African countries according to the latest global cancer statistics,1 however, the clinical trials for HCC treatments were conducted majorly in developed regions including America and European countries. In addition, China also has an increasing number of trials initiated in recent years. It is not unexpected that most clinical trials were planned in regions with good medical care and research facilities. Nevertheless, regions with high incidence rates would have more potential participants for clinical trials, and it would be beneficial for the patients if they can sign up for clinical trials at accessible locations.
Treatment Strategies Received Varied Clinical Responses In Different Cancer
Being the effective PD-1 inhibitor antibodies, nivolumab and pembrolizumab are widely applied in the treatment of various malignant tumors including lung cancer and melanoma. The results of CheckMate 017 and 057 showed that nivolumab significantly extended the overall survival of squamous and nonsquamous non-small-cell lung cancer (NSCLC) patients while exhibiting lower symptom burden,44,108 which accelerated the approval of nivolumab for the treatment of NSCLC. A 5-year follow-up study reported that the median OS of NSCLC patients received nivolumab second-line treatment was 9.9 months (95% CI, 7.8 to 12.4), with the 5-year survival rate of 16%. Comparing to the low survival rate of patients with metastatic lung cancer, the clinical outcome of nivolumab treatment was considered as a milestone in the advancement of lung cancer treatment.109 FDA approved nivolumab for advanced melanoma treatment in 2014. The recently published result of CheckMate 172 revealed that the median overall survival was 25.3–25.8 months for acral or non-acral cutaneous melanoma after second-line nivolumab treatment, and the survival rates were 57.5% to 59%.110 There is no doubt that nivolumab treatment received positive clinical outcomes that significantly prolonged the overall survival of patients suffering from lung cancer or melanoma. However, the recent press release regarding CheckMate 459, a randomized Phase 3 study evaluating nivolumab versus sorafenib as a first-line treatment in patients with unresectable hepatocellular carcinoma, showed that the primary endpoint of overall survival showed no statistical difference between groups per the pre-specified analysis. The unsatisfying clinical outcomes against HCC were also observed with another PD-1 antibody pembrolizumab. The primary result of KEYNOTE-240 showed that pembrolizumab improved progression free survival over placebo, However, the comparisons between groups were not statistically significant according to the prespecified statistical criteria.102 In great contrast, pembrolizumab achieved meaningful clinical outcomes as second-line treatment in advanced melanoma,111 it also significantly improved the overall survival of lung cancer patients in comparison to chemotherapy.112 It was proposed that the main barrier to successful immunotherapy of HCC is the inherent immunosuppressive function of the liver. The resident immune cells subsets exhibit varied and complex immune functions, which were not fully understood.113 Moreover, the inter- and intra-tumor heterogeneity of HCC immune microenvironment was also believed to be the reason that the efficacy of immunotherapy appeared to be unsatisfactory.114 Apart from immunotherapy, chemotherapies are also limitedly applied in HCC treatment because of the adverse events and toxicities.115 A clinical trial applied doxorubicin and PIAF for HCC chemotherapy, PIAF showed better responsiveness (10.5% vs 20%), however, 82% patients experienced neutropenia and 57% patients suffered from thrombocytopenia. More importantly, the prognosis of the patients remained poor.116 The choice of treatment strategy for HCC has been limited, clinical targets that used in other liver disease was also not effective in HCC clinical trials. For example, the Ligand-activated nuclear receptor peroxisome proliferator-activated receptor alpha (PPARα) was a good clinical target for NASH/NAFLD, but it was not clear yet whether it may benefit HCC treatment.117 Given the facts above, it seems that finding a safe and efficient treatment for HCC still remains a global challenge nowadays.
Conclusion
HCC patients often exhibit varied genetic profiles, and these differences can be applied in the early screening of HCC and prognosis prediction by genetic markers. In relation to diagnosis, differed treatment strategy can be designed for precision medicine, and potential biomarkers may be utilized to predict responses to drugs. Moreover, genetic markers were involved in the post-operational surveillance of HCC, which provides evidence of tumor reoccurrence at asymptomatic state. Apart from established treatment strategies, clinical trials for the investigation of new treatment plans were conducted globally. Although geographic disparity was observed, these studies enlightened new paths for HCC treatment, which would significantly improve patient survival. The results of the clinical trials suggest that the efficacy of immunotherapy in HCC is not desirable compared to lung cancer or melanoma. The underlying barrier would be the heterogeneity of the immune microenvironment of the liver. Given the above, studies on the immune microenvironment of the liver and the understanding of its heterogeneity are required for the improvement of HCC treatment efficacy.
Acknowledgment
The work received financial support from the Bureau of Industry and Information Technology of Shenzhen, programme grant 20170922151538732.
Author Contributions
All authors contributed to data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
Yuling Wu, Jiajia Zhang, Zhichao Fu, Tieshan Feng, Ming Liu, Jie Han, and Shifu Chen are affiliated with HaploX Biotechnology Co. Ltd. The authors report no other conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Boyle DA. Hepatocellular carcinoma: implications for Asia-Pacific oncology nurses. Asia Pac J Oncol Nurs. 2017;4(2):98–103. doi:10.4103/2347-5625.204497
3. Yang JD, Roberts LR. Hepatocellular carcinoma: a global view. Nat Rev Gastroenterol Hepatol. 2010;7(8):448–458. doi:10.1038/nrgastro.2010.100
4. Balogh J, Victor D
5. Fattovich G, Stroffolini T, Zagni I, Donato F. Hepatocellular carcinoma in cirrhosis: incidence and risk factors. Gastroenterology. 2004;127(5 Suppl 1):S35–S50. doi:10.1053/j.gastro.2004.09.014
6. Frank A, Seitz HK, Bartsch H, Frank N, Nair J. Immunohistochemical detection of 1N6-ethenodeoxyadenosine in nuclei of human liver affected by diseases predisposing to hepato-carcinogenesis. Carcinogenesis. 2004;25(6):1027–1031. doi:10.1093/carcin/bgh089
7. O’Rourke JM, Sagar VM, Shah T, Shetty S. Carcinogenesis on the background of liver fibrosis: implications for the management of hepatocellular cancer. World J Gastroenterol. 2018;24(39):4436–4447. doi:10.3748/wjg.v24.i39.4436
8. Zhu RX, Seto WK, Lai CL, Yuen MF. Epidemiology of hepatocellular carcinoma in the Asia-Pacific region. Gut Liver. 2016;10(3):332–339. doi:10.5009/gnl15257
9. Summers J, Mason WS. Replication of the genome of a hepatitis B–like virus by reverse transcription of an RNA intermediate. Cell. 1982;29(2):403–415. doi:10.1016/0092-8674(82)90157-x
10. Zeisel MB, Lucifora J, Mason WS, et al. Towards an HBV cure: state-of-the-art and unresolved questions–report of the ANRS workshop on HBV cure. Gut. 2015;64(8):1314–1326. doi:10.1136/gutjnl-2014-308943
11. Kuo CY, Wang JC, Wu CC, Hsu SL, Hwang GY. Effects of hepatitis B virus X protein (HBx) on cell-growth inhibition in a CCL13-HBx stable cell line. Intervirology. 2008;51(1):26–32. doi:10.1159/000118793
12. Arzumanyan A, Reis HM, Feitelson MA. Pathogenic mechanisms in HBV- and HCV-associated hepatocellular carcinoma. Nat Rev Cancer. 2013;13(2):123–135. doi:10.1038/nrc3449
13. Lee CF, Ling ZQ, Zhao T, Lee KR. Distinct expression patterns in hepatitis B virus- and hepatitis C virus-infected hepatocellular carcinoma. World J Gastroenterol. 2008;14(39):6072–6077. doi:10.3748/wjg.14.6072
14. Patil MA, Lee SA, Macias E, et al. Role of cyclin D1 as a mediator of c-Met- and beta-catenin-induced hepatocarcinogenesis. Cancer Res. 2009;69(1):253–261. doi:10.1158/0008-5472.CAN-08-2514
15. Hung JH, Teng YN, Wang LH, et al. Induction of Bcl-2 expression by hepatitis B virus pre-S2 mutant large surface protein resistance to 5-fluorouracil treatment in Huh-7 cells. PLoS One. 2011;6(12):e28977. doi:10.1371/journal.pone.0028977
16. He Y, Zhang H, Yin J, et al. IkappaBalpha gene promoter polymorphisms are associated with hepatocarcinogenesis in patients infected with hepatitis B virus genotype C. Carcinogenesis. 2009;30(11):1916–1922. doi:10.1093/carcin/bgp226
17. Lu JW, Hsia Y, Yang WY, et al. Identification of the common regulators for hepatocellular carcinoma induced by hepatitis B virus X antigen in a mouse model. Carcinogenesis. 2012;33(1):209–219. doi:10.1093/carcin/bgr224
18. Kan Z, Zheng H, Liu X, et al. Whole-genome sequencing identifies recurrent mutations in hepatocellular carcinoma. Genome Res. 2013;23(9):1422–1433. doi:10.1101/gr.154492.113
19. Ghouri YA, Mian I, Rowe JH. Review of hepatocellular carcinoma: epidemiology, etiology, and carcinogenesis. J Carcinog. 2017;16:1. doi:10.4103/jcar.JCar_9_16
20. Parkin DM. The global health burden of infection-associated cancers in the year 2002. Int J Cancer. 2006;118(12):3030–3044. doi:10.1002/ijc.21731
21. Goossens N, Hoshida Y. Hepatitis C virus-induced hepatocellular carcinoma. Clin Mol Hepatol. 2015;21(2):105–114. doi:10.3350/cmh.2015.21.2.105
22. Majumder M, Ghosh AK, Steele R, Ray R, Ray RB. Hepatitis C virus NS5A physically associates with p53 and regulates p21/waf1 gene expression in a p53-dependent manner. J Virol. 2001;75(3):1401–1407. doi:10.1128/JVI.75.3.1401-1407.2001
23. De Giorgi V, Monaco A, Worchech A, et al. Gene profiling, biomarkers and pathways characterizing HCV-related hepatocellular carcinoma. J Transl Med. 2009;7:85. doi:10.1186/1479-5876-7-85
24. Ganne-Carrie N, Nahon P. Hepatocellular carcinoma in the setting of alcohol-related liver disease. J Hepatol. 2019;70(2):284–293. doi:10.1016/j.jhep.2018.10.008
25. Seitz HK, Stickel F. Molecular mechanisms of alcohol-mediated carcinogenesis. Nat Rev Cancer. 2007;7(8):599–612. doi:10.1038/nrc2191
26. Schulze K, Nault JC, Villanueva A. Genetic profiling of hepatocellular carcinoma using next-generation sequencing. J Hepatol. 2016;65(5):1031–1042. doi:10.1016/j.jhep.2016.05.035
27. Zhang X, Kang C, Li N, et al. Identification of special key genes for alcohol-related hepatocellular carcinoma through bioinformatic analysis. PeerJ. 2019;7:e6375. doi:10.7717/peerj.6375
28. Wang JS, Qian GS, Zarba A, et al. Temporal patterns of aflatoxin-albumin adducts in hepatitis B surface antigen-positive and antigen-negative residents of Daxin, Qidong County, People’s Republic of China. Cancer Epidemiol Biomarkers Prev. 1996;5(4):253–261.
29. Kew MC. Aflatoxins as a cause of hepatocellular carcinoma. J Gastrointestin Liver Dis. 2013;22(3):305–310.
30. Qi LN, Bai T, Chen ZS, et al. The p53 mutation spectrum in hepatocellular carcinoma from Guangxi, China: role of chronic hepatitis B virus infection and aflatoxin B1 exposure. Liver Int. 2015;35(3):999–1009. doi:10.1111/liv.12460
31. Zhang W, He H, Zang M, et al. Genetic Features of Aflatoxin-Associated Hepatocellular Carcinoma. Gastroenterology. 2017;153(1):249–262 e242. doi:10.1053/j.gastro.2017.03.024
32. Degasperi E, Colombo M. Distinctive features of hepatocellular carcinoma in non-alcoholic fatty liver disease. Lancet Gastroenterol Hepatol. 2016;1(2):156–164. doi:10.1016/S2468-1253(16)30018-8
33. Wong RJ, Cheung R, Ahmed A. Nonalcoholic steatohepatitis is the most rapidly growing indication for liver transplantation in patients with hepatocellular carcinoma in the U.S. Hepatology. 2014;59(6):2188–2195. doi:10.1002/hep.26986
34. Uthaya Kumar DB, Chen CL, Liu JC, et al. TLR4 signaling via NANOG cooperates with STAT3 to activate twist1 and promote formation of tumor-initiating stem-like cells in livers of mice. Gastroenterology. 2016;150(3):707–719. doi:10.1053/j.gastro.2015.11.002
35. Fujiwara N, Nakagawa H, Enooku K, et al. CPT2 downregulation adapts HCC to lipid-rich environment and promotes carcinogenesis via acylcarnitine accumulation in obesity. Gut. 2018;67(8):1493–1504. doi:10.1136/gutjnl-2017-315193
36. Kim K, Ryu D, Dongiovanni P, et al. Degradation of PHLPP2 by KCTD17, via a glucagon-dependent pathway, promotes hepatic steatosis. Gastroenterology. 2017;153(6):1568–1580 e1510. doi:10.1053/j.gastro.2017.08.039
37. Romeo S, Kozlitina J, Xing C, et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat Genet. 2008;40(12):1461–1465. doi:10.1038/ng.257
38. Kodama T, Yi J, Newberg JY, et al. Molecular profiling of nonalcoholic fatty liver disease-associated hepatocellular carcinoma using SB transposon mutagenesis. Proc Natl Acad Sci U S A. 2018;115(44):E10417–E10426. doi:10.1073/pnas.1808968115
39. Llovet JM, Bru C, Bruix J. Prognosis of hepatocellular carcinoma: the BCLC staging classification. Semin Liver Dis. 1999;19(3):329–338. doi:10.1055/s-2007-1007122
40. Jun CH, Yoon JH, Cho E, et al. Barcelona clinic liver cancer-stage C hepatocellular carcinoma: a novel approach to subclassification and treatment. Medicine (Baltimore). 2017;96(17):e6745. doi:10.1097/MD.0000000000006745
41. Bruix J, Sherman M. American Association for the Study of Liver D. Management of hepatocellular carcinoma: an update. Hepatology. 2011;53(3):1020–1022. doi:10.1002/hep.24199
42. Zucman-Rossi J, Villanueva A, Nault JC, Llovet JM. Genetic landscape and biomarkers of hepatocellular carcinoma. Gastroenterology. 2015;149(5):1226–1239 e1224. doi:10.1053/j.gastro.2015.05.061
43. Qu C, Wang Y, Wang P, et al. Detection of early-stage hepatocellular carcinoma in asymptomatic HBsAg-seropositive individuals by liquid biopsy. Proc Natl Acad Sci U S A. 2019;116(13):6308–6312. doi:10.1073/pnas.1819799116
44. Brahmer J, Reckamp KL, Baas P, et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med. 2015;373(2):123–135. doi:10.1056/NEJMoa1504627
45. Xu RH, Wei W, Krawczyk M, et al. Circulating tumour DNA methylation markers for diagnosis and prognosis of hepatocellular carcinoma. Nat Mater. 2017;16(11):1155–1161. doi:10.1038/nmat4997
46. Oussalah A, Rischer S, Bensenane M, et al. Plasma mSEPT9: a novel circulating cell-free DNA-based epigenetic biomarker to diagnose hepatocellular carcinoma. EBioMedicine. 2018;30:138–147. doi:10.1016/j.ebiom.2018.03.029
47. Jiang P, Chan CW, Chan KC, et al. Lengthening and shortening of plasma DNA in hepatocellular carcinoma patients. Proc Natl Acad Sci U S A. 2015;112(11):E1317–E1325. doi:10.1073/pnas.1500076112
48. Jiang P, Sun K, Tong YK, et al. Preferred end coordinates and somatic variants as signatures of circulating tumor DNA associated with hepatocellular carcinoma. Proc Natl Acad Sci U S A. 2018;115(46):E10925–E10933. doi:10.1073/pnas.1814616115
49. Sun C, Liao W, Deng Z, et al. The diagnostic value of assays for circulating tumor cells in hepatocellular carcinoma: a meta-analysis. Medicine (Baltimore). 2017;96(29):e7513. doi:10.1097/MD.0000000000007513
50. Sato K, Meng F, Glaser S, Alpini G. Exosomes in liver pathology. J Hepatol. 2016;65(1):213–221. doi:10.1016/j.jhep.2016.03.004
51. Huang JT, Liu SM, Ma H, et al. Systematic review and meta-analysis: circulating miRNAs for diagnosis of hepatocellular carcinoma. J Cell Physiol. 2016;231(2):328–335. doi:10.1002/jcp.25135
52. Zhou H, Xu Q, Ni C, et al. Prospects of noncoding RNAs in hepatocellular carcinoma. Biomed Res Int. 2018;2018:6579436. doi:10.1155/2018/6579436
53. Xu X, Yin Y, Tang J, et al. Long non-coding RNA Myd88 promotes growth and metastasis in hepatocellular carcinoma via regulating Myd88 expression through H3K27 modification. Cell Death Dis. 2017;8(10):e3124. doi:10.1038/cddis.2017.518
54. Kim DS, Kim BW, Hatano E, et al. Surgical outcomes of hepatocellular carcinoma with bile duct tumor thrombus: a Korea-Japan multicenter study. Ann Surg. 2018. doi:10.1097/SLA.0000000000003014
55. Ye SL, Yang J, Bie P, et al. Safety assessment of sorafenib in Chinese patients with unresectable hepatocellular carcinoma: subgroup analysis of the GIDEON study. BMC Cancer. 2018;18(1):247. doi:10.1186/s12885-018-4242-8
56. Cai ZX, Chen G, Zeng YY, et al. Circulating tumor DNA profiling reveals clonal evolution and real-time disease progression in advanced hepatocellular carcinoma. Int J Cancer. 2017;141(5):977–985. doi:10.1002/ijc.30798
57. Hodo Y, Honda M, Tanaka A, et al. Association of interleukin-28B genotype and hepatocellular carcinoma recurrence in patients with chronic hepatitis C. Clin Cancer Res. 2013;19(7):1827–1837. doi:10.1158/1078-0432.CCR-12-1641
58. Hur K, Toiyama Y, Okugawa Y, et al. Circulating microRNA-203 predicts prognosis and metastasis in human colorectal cancer. Gut. 2017;66(4):654–665. doi:10.1136/gutjnl-2014-308737
59. Han C, Yu L, Liu X, et al. ATXN7 gene variants and expression predict post-operative clinical outcomes in hepatitis B virus-related hepatocellular carcinoma. Cell Physiol Biochem. 2016;39(6):2427–2438. doi:10.1159/000452511
60. Wang K, Guo W, Li N, et al. Alpha-1-fucosidase as a prognostic indicator for hepatocellular carcinoma following hepatectomy: a large-scale, long-term study. Br J Cancer. 2014;110(7):1811–1819. doi:10.1038/bjc.2014.102
61. Masuda K, Ono A, Aikata H, et al. Serum HMGB1 concentrations at 4 weeks is a useful predictor of extreme poor prognosis for advanced hepatocellular carcinoma treated with sorafenib and hepatic arterial infusion chemotherapy. J Gastroenterol. 2018;53(1):107–118. doi:10.1007/s00535-017-1348-8
62. Llovet JM, Ricci S, Mazzaferro V, et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359(4):378–390. doi:10.1056/NEJMoa0708857
63. Matsuki M, Hoshi T, Yamamoto Y, et al. Lenvatinib inhibits angiogenesis and tumor fibroblast growth factor signaling pathways in human hepatocellular carcinoma models. Cancer Med. 2018;7(6):2641–2653. doi:10.1002/cam4.1517
64. Sartore-Bianchi A, Siena S, Tonini G, Bardelli A, Santini D. Overcoming dynamic molecular heterogeneity in metastatic colorectal cancer: multikinase inhibition with regorafenib and the case of rechallenge with anti-EGFR. Cancer Treat Rev. 2016;51:54–62. doi:10.1016/j.ctrv.2016.10.006
65. Xiang Q, Chen W, Ren M, et al. Cabozantinib suppresses tumor growth and metastasis in hepatocellular carcinoma by a dual blockade of VEGFR2 and MET. Clin Cancer Res. 2014;20(11):2959–2970. doi:10.1158/1078-0432.CCR-13-2620
66. Carmeliet P. VEGF as a key mediator of angiogenesis in cancer. Oncology. 2005;69(Suppl 3):4–10. doi:10.1159/000088478
67. Wilhelm SM, Carter C, Tang L, et al. BAY 43-9006 exhibits broad spectrum oral antitumor activity and targets the RAF/MEK/ERK pathway and receptor tyrosine kinases involved in tumor progression and angiogenesis. Cancer Res. 2004;64(19):7099–7109. doi:10.1158/0008-5472.CAN-04-1443
68. Rimassa L, Santoro A. Sorafenib therapy in advanced hepatocellular carcinoma: the SHARP trial. Expert Rev Anticancer Ther. 2009;9(6):739–745. doi:10.1586/era.09.41
69. Keating GM. Sorafenib: a review in hepatocellular carcinoma. Target Oncol. 2017;12(2):243–253. doi:10.1007/s11523-017-0484-7
70. Kudo M. Extremely high objective response rate of lenvatinib: its clinical relevance and changing the treatment paradigm in hepatocellular carcinoma. Liver Cancer. 2018;7(3):215–224. doi:10.1159/000492533
71. Kudo M, Finn RS, Qin S, et al. Lenvatinib versus sorafenib in first-line treatment of patients with unresectable hepatocellular carcinoma: a randomised phase 3 non-inferiority trial. Lancet. 2018;391(10126):1163–1173. doi:10.1016/S0140-6736(18)30207-1
72. Ikeda K, Kudo M, Kawazoe S, et al. Phase 2 study of lenvatinib in patients with advanced hepatocellular carcinoma. J Gastroenterol. 2017;52(4):512–519. doi:10.1007/s00535-016-1263-4
73. Bruix J, Tak WY, Gasbarrini A, et al. Regorafenib as second-line therapy for intermediate or advanced hepatocellular carcinoma: multicentre, open-label, phase II safety study. Eur J Cancer. 2013;49(16):3412–3419. doi:10.1016/j.ejca.2013.05.028
74. Wilhelm SM, Dumas J, Adnane L, et al. Regorafenib (BAY 73-4506): a new oral multikinase inhibitor of angiogenic, stromal and oncogenic receptor tyrosine kinases with potent preclinical antitumor activity. Int J Cancer. 2011;129(1):245–255. doi:10.1002/ijc.25864
75. Bruix J, Qin S, Merle P, et al. Regorafenib for patients with hepatocellular carcinoma who progressed on sorafenib treatment (RESORCE): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2017;389(10064):56–66. doi:10.1016/S0140-6736(16)32453-9
76. Yakes FM, Chen J, Tan J, et al. Cabozantinib (XL184), a novel MET and VEGFR2 inhibitor, simultaneously suppresses metastasis, angiogenesis, and tumor growth. Mol Cancer Ther. 2011;10(12):2298–2308. doi:10.1158/1535-7163.MCT-11-0264
77. Teufel M, Seidel H, Kochert K, et al. Biomarkers associated with response to regorafenib in patients with hepatocellular carcinoma. Gastroenterology. 2019;156(6):1731–1741. doi:10.1053/j.gastro.2019.01.261
78. Abou-Alfa GK, Meyer T, Cheng AL, et al. Cabozantinib in patients with advanced and progressing hepatocellular carcinoma. N Engl J Med. 2018;379(1):54–63. doi:10.1056/NEJMoa1717002
79. Lee YS, Kim BH, Kim BC, et al. SLC15A2 genomic variation is associated with the extraordinary response of sorafenib treatment: whole-genome analysis in patients with hepatocellular carcinoma. Oncotarget. 2015;6(18):16449–16460. doi:10.18632/oncotarget.3758
80. Sawey ET, Chanrion M, Cai C, et al. Identification of a therapeutic strategy targeting amplified FGF19 in liver cancer by Oncogenomic screening. Cancer Cell. 2011;19(3):347–358. doi:10.1016/j.ccr.2011.01.040
81. MacParland SA, Liu JC, Ma XZ, et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat Commun. 2018;9(1):4383. doi:10.1038/s41467-018-06318-7
82. Aizarani N, Saviano A, Sagar, et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature. 2019;572(7768):199–204. doi:10.1038/s41586-019-1373-2
83. Kurebayashi Y, Ojima H, Tsujikawa H, et al. Landscape of immune microenvironment in hepatocellular carcinoma and its additional impact on histological and molecular classification. Hepatology. 2018;68(3):1025–1041. doi:10.1002/hep.29904
84. Sprinzl MF, Galle PR. Current progress in immunotherapy of hepatocellular carcinoma. J Hepatol. 2017;66(3):482–484. doi:10.1016/j.jhep.2016.12.009
85. Tunger A, Kiessler M, Wehner R, et al. immune monitoring of cancer patients prior to and during CTLA-4 or PD-1/PD-L1 inhibitor treatment. Biomedicines. 2018;6(1). doi:10.3390/biomedicines6010026
86. Vogel I, Kasran A, Cremer J, et al. CD28/CTLA-4/B7 costimulatory pathway blockade affects regulatory T-cell function in autoimmunity. Eur J Immunol. 2015;45(6):1832–1841. doi:10.1002/eji.201445190
87. Okazaki T, Honjo T. PD-1 and PD-1 ligands: from discovery to clinical application. Int Immunol. 2007;19(7):813–824. doi:10.1093/intimm/dxm057
88. Butte MJ, Keir ME, Phamduy TB, Sharpe AH, Freeman GJ. Programmed death-1 ligand 1 interacts specifically with the B7-1 costimulatory molecule to inhibit T cell responses. Immunity. 2007;27(1):111–122. doi:10.1016/j.immuni.2007.05.016
89. Postow MA, Callahan MK, Wolchok JD. Immune checkpoint blockade in cancer therapy. J Clin Oncol. 2015;33(17):1974–1982. doi:10.1200/JCO.2014.59.4358
90. El-Khoueiry AB, Sangro B, Yau T, et al. Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): an open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet. 2017;389(10088):2492–2502. doi:10.1016/S0140-6736(17)31046-2
91. Zhu AX, Finn RS, Edeline J, et al. Pembrolizumab in patients with advanced hepatocellular carcinoma previously treated with sorafenib (KEYNOTE-224): a non-randomised, open-label phase 2 trial. Lancet Oncol. 2018;19(7):940–952. doi:10.1016/S1470-2045(18)30351-6
92. Goodman AM, Kato S, Bazhenova L, et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol Cancer Ther. 2017;16(11):2598–2608. doi:10.1158/1535-7163.MCT-17-0386
93. Pinyol R, Sia D, Llovet JM. Immune exclusion-Wnt/CTNNB1 class predicts resistance to immunotherapies in HCC. Clin Cancer Res. 2019;25(7):2021–2023. doi:10.1158/1078-0432.CCR-18-3778
94. Llovet JM, Montal R, Sia D, Finn RS. Molecular therapies and precision medicine for hepatocellular carcinoma. Nat Rev Clin Oncol. 2018;15(10):599–616. doi:10.1038/s41571-018-0073-4
95. Marcus L, Lemery SJ, Keegan P, Pazdur R. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors. Clin Cancer Res. 2019;25(13):3753–3758. doi:10.1158/1078-0432.CCR-18-4070
96. Llovet JM. Ramucirumab (RAM) for sorafenib intolerant patients with hepatocellular carcinoma (HCC) and elevated baseline alpha fetoprotein (AFP): Outcomes from two randomized phase 3 studies (REACH, REACH2). J Clin Oncol. 37;2019(suppl; abstr 4073).
97. Zhu AX, Finn RS, Galle PR, Llovet JM, Kudo M. Ramucirumab in advanced hepatocellular carcinoma in REACH-2: the true value of alpha-fetoprotein. Lancet Oncol. 2019;20(4):e191. doi:10.1016/S1470-2045(19)30310-9
98. Segovia-Mendoza M, Gonzalez-Gonzalez ME, Barrera D, Diaz L, Garcia-Becerra R. Efficacy and mechanism of action of the tyrosine kinase inhibitors gefitinib, lapatinib and neratinib in the treatment of HER2-positive breast cancer: preclinical and clinical evidence. Am J Cancer Res. 2015;5(9):2531–2561.
99. Kong Y, Sun L, Hou Z, et al. Apatinib is effective for treatment of advanced hepatocellular carcinoma. Oncotarget. 2017;8(62):105596–105605. doi:10.18632/oncotarget.22337
100. Kim R, Sharma S, Meyer T, et al. First-In-Human Study of BLU-554, a Potent, Highly-Selective FGFR4 Inhibitor Designed for Hepatocellular Carcinoma (HCC) with FGFR4 Pathway Activation. Oxford: Elsevier; 2016.
101. Yau T, Park JW, Finn RS, et al. CheckMate 459: A randomized, multi-center phase III study of nivolumab (NIVO) vs sorafenib (SOR) as first-line (1L) treatment in patients (pts) with advanced hepatocellular carcinoma (aHCC). Ann Oncol. 2019;30(suppl_5):v851–v934. doi:10.1093/annonc/mdz394.029
102. Finn RS. Results of KEYNOTE-240: phase3 studyofpembrolizumab(Pembro)vsbest supportive care (BSC) for second line therapy in advanced hepatocellular carcinoma (HCC). J Clin Oncol. 37;2019(suppl; abstr 4004).
103. Yau T. Nivolumab (NIVO) + ipilimumab (IPI) combination therapy in patients (pts) withadvancedhepatocellularcarcinoma (aHCC): Results from CheckMate040. J Clin Oncol. 37;2019(suppl; abstr 4012).
104. Raghav KPS, Mody K, Greten TF, et al. An open label phase 1b/2 trial of TRC105 and sorafenib in patient with advanced/metastatic hepatocellular carcinoma (HCC) (NCT01806064). J Clin Oncol. 2018;36(4_suppl):301. doi:10.1200/JCO.2018.36.4_suppl.301
105. Kudo M. First-line avelumab + axitinib in patients with advanced hepatocellular carcinoma: Results from a phase 1b trial (VEGF Liver 100). J Clin Oncol. 37;2019(suppl; abstr 4072).
106. Kelley RK, Abou-Alfa GK, Bendell JC, et al. Phase I/II study of durvalumab and tremelimumab in patients with unresectable hepatocellular carcinoma (HCC): phase I safety and efficacy analyses. J Clin Oncol. 2017;35(15_suppl):4073. doi:10.1200/JCO.2017.35.15_suppl.4073
107. Yin X. Randomized clinical trial of transcatheter arterial chemoembolization plus radiofrequencyablationversustranscatheterarterialchemoembolizationfor hepatocellular carcinoma with intermediate stage (BCLC stage B) hepatocellular carcinoma beyondMilan criteria. J Clin Oncol. 37;2019(suppl; abstr 4077).
108. Borghaei H, Paz-Ares L, Horn L, et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med. 2015;373(17):1627–1639. doi:10.1056/NEJMoa1507643
109. Gettinger S, Horn L, Jackman D, et al. Five-year follow-up of nivolumab in previously treated advanced non-small-cell lung cancer: results from the CA209-003 study. J Clin Oncol. 2018;36(17):1675–1684. doi:10.1200/JCO.2017.77.0412
110. Nathan P, Ascierto PA, Haanen J, et al. Safety and efficacy of nivolumab in patients with rare melanoma subtypes who progressed on or after ipilimumab treatment: a single-arm, open-label, phase II study (CheckMate 172). Eur J Cancer. 2019;119:168–178. doi:10.1016/j.ejca.2019.07.010
111. Si L, Zhang X, Shu Y, et al. a phase ib study of pembrolizumab as second-line therapy for chinese patients with advanced or metastatic melanoma (KEYNOTE-151). Transl Oncol. 2019;12(6):828–835. doi:10.1016/j.tranon.2019.02.007
112. Nosaki K, Saka H, Hosomi Y, et al. Safety and efficacy of pembrolizumab monotherapy in elderly patients with PD-L1-positive advanced non-small-cell lung cancer: pooled analysis from the KEYNOTE-010, KEYNOTE-024, and KEYNOTE-042 studies. Lung Cancer. 2019;135:188–195. doi:10.1016/j.lungcan.2019.07.004
113. Obeid JM, Kunk PR, Zaydfudim VM, Bullock TN, Slingluff CL
114. Zhang Q, Lou Y, Yang J, et al. Integrated multiomic analysis reveals comprehensive tumour heterogeneity and novel immunophenotypic classification in hepatocellular carcinomas. Gut. 2019. doi:10.1136/gutjnl-2019-318912
115. Deng GL, Zeng S, Shen H. Chemotherapy and target therapy for hepatocellular carcinoma: new advances and challenges. World J Hepatol. 2015;7(5):787–798. doi:10.4254/wjh.v7.i5.787
116. Yeo W, Mok TS, Zee B, et al. A randomized phase III study of doxorubicin versus cisplatin/interferon alpha-2b/doxorubicin/fluorouracil (PIAF) combination chemotherapy for unresectable hepatocellular carcinoma. J Natl Cancer Inst. 2005;97(20):1532–1538. doi:10.1093/jnci/dji315
117. Tanaka N, Aoyama T, Kimura S, Gonzalez FJ. Targeting nuclear receptors for the treatment of fatty liver disease. Pharmacol Ther. 2017;179:142–157. doi:10.1016/j.pharmthera.2017.05.011
118. Denoix P, Lacour J, Wolff JP, Weiler J, Michel G. [TNM classification of cancer of the uterine cervix. Apropos of contents and treatment of T2]. J Gynecol Obstet Biol Reprod (Paris). 1972;1(2):161–166.
119. Okuda K, Ohtsuki T, Obata H, et al. Natural history of hepatocellular carcinoma and prognosis in relation to treatment. Study of 850 patients. Cancer. 1985;56(4):918–928. doi:10.1002/1097-0142(19850815)56:4<918::aid-cncr2820560437>3.0.co;2-e
120. The CLIP investigator group. Prospective validation of the CLIP score: a new prognostic system for patients with cirrhosis and hepatocellular carcinoma. The Cancer of the Liver Italian Program (CLIP) Investigators. Hepatology. 2000;31(4):840–845. doi:10.1053/he.2000.5628
121. Kubo S, Tanaka H, Shuto T, et al. Prognostic effects of causative virus in hepatocellular carcinoma according to the Japan integrated staging (JIS) score. J Gastroenterol. 2005;40(10):972–979. doi:10.1007/s00535-005-1681-1
122. Leung TW, Tang AM, Zee B, et al. Construction of the Chinese University Prognostic Index for hepatocellular carcinoma and comparison with the TNM staging system, the Okuda staging system, and the Cancer of the Liver Italian Program staging system: a study based on 926 patients. Cancer. 2002;94(6):1760–1769. doi:10.1002/cncr.10384
123. Johnson PJ, Berhane S, Kagebayashi C, et al. Assessment of liver function in patients with hepatocellular carcinoma: a new evidence-based approach-the ALBI grade. J Clin Oncol. 2015;33(6):550–558. doi:10.1200/JCO.2014.57.9151
124. Finkelmeier F, Waidmann O, Trojan J. Nivolumab for the treatment of hepatocellular carcinoma. Expert Rev Anticancer Ther. 2018;18(12):1169–1175. doi:10.1080/14737140.2018.1535315
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.