Back to Journals » OncoTargets and Therapy » Volume 13
Genetic and Clinical Characterization of HOXB2 in Glioma
Authors Pan X, Liu W, Chai Y, Wang J, Zhang Y
Received 23 June 2020
Accepted for publication 5 October 2020
Published 14 October 2020 Volume 2020:13 Pages 10465—10473
DOI https://doi.org/10.2147/OTT.S268635
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Prof. Dr. Takuya Aoki
Xin Pan,1,2,* Wei Liu,1,2,* Yi Chai,1,2,* Junhua Wang,1,2 Yuqi Zhang1,2
1Department of Neurosurgery, Yuquan Hospital, School of Clinical Medicine, Tsinghua University, Beijing 100040, People’s Republic of China; 2School of Clinical Medicine, Tsinghua University, Beijing 10084, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Yuqi Zhang
School of Clinical Medicine, Tsinghua University, No. 1, Tsinghua Yuan, Handian District, Beijing 10084, People’s Republic of China
Email [email protected]
Background: Glioma is a highly aggressive and heterogeneous cancer with poor survival rates. Homeobox (HOX) genes are transcription factors that play pivotal roles in many aspects of cellular physiology, embryonic development, and tissue homeostasis. Mutations in HOX genes can lead to increased cancer predisposition. Abnormal expression of HOXB2 may result in the development and progression of tumors. However, its prognostic value and mechanism of dysregulation remain unclear.
Methods: The present study included 1001 glioma patients. The correlations between the expression of HOXB2 and subgroups of glioma were investigated by t-test analyses. The prognostic value of HOXB2 was explored by Kaplan–Meier analysis as well as univariate and multivariate Cox analyses. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment were employed to detect the biological function of HOXB2 in glioma. CCK-8 and transwell assays were performed to determine the role of HOXB2 in cell proliferation and invasion.
Results: HOXB2 was positively correlated with tumor grade and enriched in patients with isocitrate dehydrogenase 1 wild-type and age > 41 years. HOXB2 was identified as an independent prognostic biomarker in glioma patients. HOXB2 was associated with cell invasion and promoted the proliferation of glioma cells in vitro.
Conclusion: HOXB2 is an independent prognostic factor and contributes to tumor invasion in glioma patients.
Keywords: glioma, gene expression, neoplasm invasiveness, prognosis, HOXB2
Introduction
Glioma is a malignant intracranial tumor that accounts for nearly 50% of all central nervous system malignant neoplastic lesions.1 According to the World Health Organization (WHO) classification, gliomas can be divided into four tumor grades for prognosis prediction, with survival ranging from 1 to 15 years.2 Although various treatment options, including neurosurgical resection and adjuvant chemoradiotherapy, have been explored to prolong the lives of glioma patients, most gliomas exhibit recurrence, thereby reducing life expectancy. Notably, most glioblastoma patients do not survive beyond 1 year, and only about 5% survive beyond 5 years under the best existing medical interventions.3,4 Despite many years of painstaking research, the status quo for the prognosis of glioma patients remains dismal. Thus, there is a substantial need to identify more effective and specific targets for treatment.
In the past few years, studies in the glioma field focusing on molecular characterization have resulted in the generation of datasets such as The Cancer Genome Atlas (TCGA) and Chinese Glioma Genome Atlas (CGGA).5–7 For example, mutations in isocitrate dehydrogenase (IDH) were identified as key markers for not only glioblastoma but also low-grade gliomas in adults.8,9 Mutations are found in >70% of WHO class II and III gliomas and can predict the prognosis more accurately.1 However, even in grade II or III tumors, IDH wild-type is usually accompanied by a poor prognosis.
The homeobox (HOX) genes are transcription factors encoded by a subset of homeodomain superfamily genes that are involved in cellular physiology, embryonic development, and tissue homeostasis.10,11 Aberrant expression of HOX genes can result in the development and progression of tumors. For example, overexpression of HOXC8 in lung cancer was positively associated with tumor metastasis and poor prognosis.12,13 Recent studies indicated that HOXB2 may participate in tumor progression. Pan et al14 demonstrated that LncRNA RGMB-ASI promoted glioma progression by targeting HOXB2. Nevertheless, the expression profiles of HOXB2 in glioma subtypes and their complex carcinogenic mechanisms remain unclear. In this study, we analyzed the expression of HOXB2 in different glioma subtypes and investigated their possible regulatory mechanisms.
Methods
Data Collection
A total of 1001 glioma patients in two independent data sets were enrolled in this study. The RNAseq data, including 691 gliomas, from The Cancer Genome Atlas (TCGA), were downloaded from public databases (https://portal.gdc.cancer.gov/). Three hundred and ten (310) glioma cases were obtained from the Chinese Glioma Genome Atlas (CGGA) dataset (http://cgga.org.cn/).
Cell Lines and Cultures
The human glioma cell lines including U87, LN229, U251, U343 and U373 were provided by the American Type Culture Collection. The HEB human glial cell line was supplied by the Cell Culture Center (Dongge Biotech, Beijing, China). All cell lines were incubated in DMEM (Invitrogen, USA) with 10% FBS (HyClone, USA) in a 37°C incubator containing 5% CO2.
Cell Transfection
Specific siRNAs for HOXB2 were purchased from RiboBio (Guangzhou, China). The transfection was performed using Lipofectamine 3000 (Invitrogen, Carlsbad, California).
Si-HOXB2-1,5ʹ-GCCUUUAGCCGUUCGCUUATT-3ʹ;
Si-HOXB2-2,5ʹ-GGCAGGUCAAAGUCUGGUUTT-3ʹ.
Cell Viability Assay
A total of 2000 cells were seeded in 96-well plates and cultured for 24 h, 48 h,72 h and 96 h, respectively. Cell Counting Kit-8 (NCM Biotech, Suzhou, China) was employed to determine the cell viability according to the manufacturer’s instructions. The absorbance was detected at an optical density (OD) 450 nm.
Western Blot Analysis
The expression of proteins was detected using Western blot as previously described.15 The primary antibodies used in the study were HOXB2 (ab136856) and MMP-2 (ab2462)(abcam, USA). β-actin (A8418) (Sigma, USA) was used as an internal control.
Transwell Assay
The invasion assay was performed using the transwell system (24wells, 8μm pore size with polycarbonate membrane, Corning Costar) as previously described.15
Statistical Analysis
Statistical analysis was performed using R software version 3.6.3 and Prism 7 (GraphPad, San Diego, CA, USA). The Student’s t-test was used to compare the difference between binary groups. One-way ANOVA, followed by the Holm-Sidak test, was employed to compare differences among multiple groups. The predictive value of HOXB2 expression for overall survival was estimated using Kaplan-Meier (K-M) survival analysis with the long-rank test. The prognostic value of HOXB2 was determined by univariate and multivariate cox proportional hazard model. Data are presented as mean ± SD from three independent experiments. P<0.05 was considered significant.
Results
Relationships Between HOXB2 Expression and Glioma Subtypes
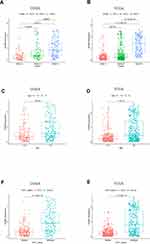
We obtained 310 glioma samples with full clinical information from the CGGA RNA-seq dataset. Expression of HOXB2 was positively correlated with tumor grade (Figure 1A and B). Moreover, high expression of HOXB2 was observed in patients aged >41 years (Figure 1C and D) and with IDH1 wild-type (Figure 1E and F). These findings were verified in 691 glioma samples from the TCGA dataset. The baseline information is presented in Tables 1 and 2.
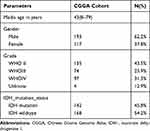 |
Table 1 Clinicopathological Parameters Baseline in CGGA Cohort |
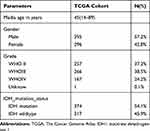 |
Table 2 Clinicopathological Parameters Baseline in TCGA Cohort |
HOXB2 is an Independent Predictive Marker in Glioma Patients
The association between HOXB2 mRNA expression and survival time was investigated by Kaplan–Meier survival curve analysis. The results showed that patients with high HOXB2 expression had a worse prognosis in both the CGGA and TCGA datasets (Figure 2A and B). Univariate and multivariate Cox regression analyses were performed to identify independent risk factors related to the prognosis of glioma patients. In the CGGA dataset, the results revealed that high HOXB2 expression (HR = 1.237; 95% CI: 1.094–1.398; P < 0.001) and tumor grade (WHO IV vs WHO II/III) (HR = 3.447; 95% CI: 2.456–4.837; P < 0.001) were independent risk factors for evaluating the prognosis of glioma patients (Table 3). In the TCGA dataset, high HOXB2 expression (HR = 1.580; 95% CI: 1.320–1.892; P < 0.001), age (HR = 1.037; 95% CI: 1.027–1.048; P < 0.001), tumor grade (WHO IV vs WHO II/III) (HR = 2.500; 95% CI: 1.782–3.505; P < 0.001), and IDH1 mutant (HR = 0.509; 95% CI: 0.360–0.719; P < 0.001) were identified as independent factors affecting the prognosis of glioma patients (Table 4). From these results, high HOXB2 expression signaled a poor prognosis for patients. Therefore, HOXB2 can be regarded as an indicator for predicting the prognosis of glioma patients.
 |
Table 3 Univariate and Multivariate Analyses of Overall Survival in the Chinese Glioma Genome Atlas Microarray Database |
 |
Table 4 Univariate and Multivariate Analyses of Overall Survival in the Cancer Genome Atlas Database |
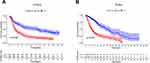 |
Figure 2 The negative association between the expression of HOXB2 and the patient outcome in CGGA (A) and TCGA (B). |
HOXB2 is Associated with Cell Invasion and Cell Proliferation
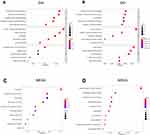
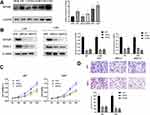
To investigate the biological function of HOXB2, the relationships between HOXB2 and other genes were detected by Pearson correlation analysis (|r| > 0.4; P < 0.05). A total of 1930 and 4223 genes were screened in the CGGA and TCGA datasets, respectively, and subjected to GO and KEGG analyses to determine the biological processes. HOXB2 was found to be closely related to the extracellular matrix (ECM), immune response, inflammatory response, and other pathways (Figure 3A–D). ECM can inhibit the invasion of tumor cells. However, tumor cells can degrade the ECM by secreting components such as matrix metalloproteinases (MMPs). When the correlations of HOXB2 with MMP expressions were examined in the TCGA and CGGA datasets, significant positive correlations were observed between HOXB2 and MMP-2, MMP-9, MMP-11, and MMP-14 (Figure 4A–H). These results suggested that HOXB2 can promote the degradation of tumor ECM components, thereby promoting tumor invasion.
HOXB2 Promotes the Proliferation and Invasion of Glioma Cells in vitro
We detected the expression of HOXB2 in glioma cell lines adjacent to the HEB. Western blot analyses revealed that U87 and U251 cells contained the highest levels of HOXB2 among the cell lines examined (Figure 5A). We explored the function of HOXB2 in glioma by silencing endogenous HOXB2 expression in U87 and U251 cells using a siRNA (Figure 5B). CCK-8 assays revealed that the proliferation ability of U87 and U251 cells was significantly decreased following the transfection of the HOXB2 siRNA (Figure 5C). Transwell assays indicated that the depletion of HOXB2 suppressed the invasion capacity of U87 and U251 cells (Figure 5D). Western blot analyses further showed that the expression of MMP-2 was significantly reduced in siRNA-transfected U87 and U251 cells (Figure 5B). Taken together, these findings demonstrate that HOXB2 may play a vital role in cell proliferation and invasion.
Discussion
Glioma is characterized by rapid proliferation and high invasiveness that pose difficulties for treatment.16,17 Previous studies demonstrated that different subtypes of glioma exhibited significant differences in treatment and prognosis.2,18–20 Identification of effective markers that can predict the prognosis of glioma patients is essential for the development of successful treatment plans. HOXB2 is a transcription factor located on chromosome 17.21–23 Numerous studies have demonstrated that abnormal expression of HOXB2 was carcinogenic for various tumors.14,21,22 In breast cancer, overexpression of HOXB2 inhibited tumor proliferation,24 while in lung cancer, pancreatic cancer, and other tumors,25–27 HOXB2 promoted tumor progression and metastasis. However, the correlations between HOXB2 expression and clinical characteristics of glioma patients have not been fully elucidated.
In the present study, we analyzed the transcriptomes and clinical data of 1001 glioma patients in two independent tumor datasets. Expression of HOXB2 was positively correlated with tumor grade and was enriched in patients with IDH1 wild-type and age >41 years. By performing Kaplan–Meier survival analysis and Cox regression analyses, we further confirmed that HOXB2 could be used as an independent factor to predict the prognosis of glioma patients. Based on these findings, we speculate that HOXB2 has a role in promoting the progression of glioma.
Invasive growth has been recognized as a critical feature of glioma and plays a vital role in promoting tumorigenesis and development.15,28 It is a complex biological process with multifactor and multicell involvement, including ECM degradation, epithelial–mesenchymal transition, and neovascularization.29,30 ECM components are large molecules that are synthesized by cells, secreted to the outside of the cells, and become distributed on the surface of the cells or between cells. The components form a complex grid structure that supports and connects the tissue structure and regulates the tissue. Compared with normal cells, tumor cells secrete large amounts of enzymes such as MMPs that degrade the ECM and allow easy spreading. MMPs can degrade various components of the ECM, leading to the destruction of the tissue barrier, thereby accelerating tumor progression. Zhou et al31 found that glioma patients with high expression of MMP-2 and MMP-9 had a poor prognosis and were insensitive to radiotherapy. MMP-11 can promote the progression of gastric cancer, while high expression of MMP-14 was positively correlated with breast cancer.32,33
To further identify the molecular mechanism of HOXB2 in glioma, we performed GO and KEGG analyses. The results indicated that the biological function of HOXB2 was enriched in the ECM, immune response, and inflammatory response pathways. Thus, we analyzed the correlations between HOXB2 expression and MMPs in the CGGA and TCGA datasets. HOXB2 expression was positively correlated with the expression of MMP-2, MMP-9, MMP-11, and MMP-14, suggesting that HOXB2 is involved in the invasion process of glioma. In vitro experiments demonstrated that knockdown of HOXB2 suppressed the proliferation, decreased the invasion, and inhibited the expression of MMP-2 in glioma cells. Besides, functional enrichment analysis indicated that HOXB2 was involved in a variety of immune processes. However, owing to the limitations of the experimental conditions, we did not conduct further analyses on the function of HOXB2 in the glioma immune responses.
In summary, our results showed that the expression of HOXB2 is upregulated in high-grade glioma and can be regarded as an independent predictive factor for the prognosis of glioma patients. Therefore, HOXB2 may be a new target for the treatment of glioma, although its specific mechanisms remain to be elucidated in further experimental studies.
Funding
This work was supported by funding from the National Natural Science Foundation of China (NO:81071885). Xin Pan, Wei Liu and Yi Chai contributed equally to this work.
Disclosure
The authors declare that there are no conflict of interests.
References
1. Lapointe S, Perry A, Butowski NA. Primary brain tumours in adults. Lancet. 2018;392(10145):432–446. doi:10.1016/S0140-6736(18)30990-5
2. Ostrom QT, Gittleman H, Liao P, et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2010–2014. Neuro Oncol. 2017;19(suppl_5):v1–v88. doi:10.1093/neuonc/nox158
3. Stupp R, Mason WP, van den Bent MJ, et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 2005;352(10):987–996. doi:10.1056/NEJMoa043330
4. Sampson JH, Gunn MD, Fecci PE, Ashley DM. Brain immunology and immunotherapy in brain tumours. Nat Rev Cancer. 2020;20(1):12–25.
5. Molinaro AM, Taylor JW, Wiencke JK, Wrensch MR. Genetic and molecular epidemiology of adult diffuse glioma. Nat Rev Neurol. 2019;15(7):405–417.
6. Chai Y, Liu W, Wang C, Rao M, Zhang Y. Prognostic role of chicken ovalbumin upstream promoter transcription factor II in isocitrate dehydrogenase-mutant glioma with 1p19q co-deletion. J Mol Neurosci. 2019;68(2):234–242.
7. Pan X, Liu W, Chai Y, Hu L, Wang J, Zhang Y. Identification of hub genes in atypical teratoid/rhabdoid tumor by bioinformatics analyses. J Mol Neurosci. 2020.
8. Sanai N, Berger MS. Surgical oncology for gliomas: the state of the art. Nat Rev Clin Oncol. 2018;15(2):112–125.
9. Hong JB, Roh TH, Ahn SS, et al. Predicting survival using the 2016 World Health Organization classification for anaplastic glioma. Clin Neuropathol. 2020.
10. Hamada J, Omatsu T, Okada F, et al. Overexpression of homeobox gene HOXD3 induces coordinate expression of metastasis‐related genes in human lung cancer cells. Int J Cancer. 2001;93(4):516–525.
11. Miyazaki YJ, Hamada J, Tada M, et al. HOXD3 enhances motility and invasiveness through the TGF-beta-dependent and -independent pathways in A549 cells. Oncogene. 2002;21(5):798–808. doi:10.1038/sj.onc.1205126
12. Zhang J, Yang M, Li D, et al. Homeobox C8 is a transcriptional repressor of E-cadherin gene expression in non-small cell lung cancer. Int J Biochem Cell Biol. 2019;114:105557. doi:10.1016/j.biocel.2019.06.005
13. Liu H, Zhang M, Xu S, et al. HOXC8 promotes proliferation and migration through transcriptional up-regulation of TGFβ1 in non-small cell lung cancer. Oncogenesis. 2018;7(2):1. doi:10.1038/s41389-017-0016-4
14. Pan B, Zhao M, Wang N, Xu L, Wu T, Li Z. LncRNA RGMB-AS1 promotes glioma growth and invasion through miR-1200/HOXB2 axis. Onco Targets Ther. 2019;12:10107–10114.
15. Liu W, Chai Y, Hu L, et al. Polyphyllin VI induces apoptosis and autophagy via reactive oxygen species mediated JNK and P38 activation in glioma. Onco Targets Ther. 2020;13:2275–2288. doi:10.2147/OTT.S243953
16. Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100(1):57–70. doi:10.1016/S0092-8674(00)81683-9
17. Yuan YL, Jiang N, Li ZY, et al. Polyphyllin VI induces apoptosis and autophagy in human osteosarcoma cells by modulation of ROS/JNK activation. Drug Des Devel Ther. 2019;13:3091–3103. doi:10.2147/DDDT.S194961
18. Sonoda Y. Clinical impact of revisions to the WHO classification of diffuse gliomas and associated future problems. Int J Clin Oncol. 2020.
19. Ohba S, Kuwahara K, Yamada S, Abe M, Hirose Y. Correlation between IDH, ATRX, and TERT promoter mutations in glioma. Brain Tumor Pathol. 2020. doi:10.1007/s10014-020-00360-4
20. Le Rhun E, Preusser M, Roth P, et al. Molecular targeted therapy of glioblastoma. Cancer Treat Rev. 2019;80:101896. doi:10.1016/j.ctrv.2019.101896
21. Jing P, Zou J, Zhang L, et al. HOXB2 and FOXC1 synergistically drive the progression of Wilms tumor. Exp Mol Pathol. 2020;115:104469.
22. Miguez A, Ducret S, Di Meglio T, et al. Opposing roles for Hoxa2 and Hoxb2 in hindbrain oligodendrocyte patterning. J Neurosci. 2012;32(48):17172–17185. doi:10.1523/JNEUROSCI.0885-12.2012
23. Liu HC, Shih LY, May Chen MJ, et al. Expression of HOXB genes is significantly different in acute myeloid leukemia with a partial tandem duplication of MLL vs. a MLL translocation: a cross-laboratory study. Cancer Genet. 2011;204(5):252–259. doi:10.1016/j.cancergen.2011.02.003
24. Boimel PJ, Cruz C, Segall JE. A functional in vivo screen for regulators of tumor progression identifies HOXB2 as a regulator of tumor growth in breast cancer. Genomics. 2011;98(3):164–172. doi:10.1016/j.ygeno.2011.05.011
25. Cheng Y, Jutooru I, Chadalapaka G, Corton JC, Safe S. The long non-coding RNA HOTTIP enhances pancreatic cancer cell proliferation, survival and migration. Oncotarget. 2015;6(13):10840–10852.
26. Liang L, Shen YY, Pan XW, et al. Adaptive evolution of the Hox gene family for development in bats and dolphins. PLoS One. 2013;8(6):e65944. doi:10.1371/journal.pone.0065944
27. Segara D, Biankin AV, Kench JG, et al. Expression of HOXB2, a retinoic acid signaling target in pancreatic cancer and pancreatic intraepithelial neoplasia. Clin Cancer Res. 2005;11(9):3587–3596. doi:10.1158/1078-0432.CCR-04-1813
28. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674. doi:10.1016/j.cell.2011.02.013
29. Vollmann-Zwerenz A, Leidgens V, Feliciello G, Klein CA, Hau P. Tumor cell invasion in glioblastoma. Int J Mol Sci. 2020;21(6):1932. doi:10.3390/ijms21061932
30. Dochiţ CM, Stepan AE, Mărgăritescu C, Florescu MM, Simionescu CE. Immunoexpression of E-, P- and N-cadherins in ovarian serous malignant tumors. Rom J Morphol Embryol. 2019;60(4):1215–1219.
31. Zhou F, Sun Y, Chi Z, Gao Q, Wang H. Long noncoding RNA SNHG12 promotes the proliferation, migration, and invasion of trophoblast cells by regulating the epithelial-mesenchymal transition and cell cycle. J Int Med Res. 2020;48(6):300060520922339. doi:10.1177/0300060520922339
32. Su C, Wang W, Wang C. IGF-1-induced MMP-11 expression promotes the proliferation and invasion of gastric cancer cells through the JAK1/STAT3 signaling pathway. Oncol Lett. 2018;15(5):7000–7006.
33. Di D, Chen L, Guo Y, Wang L, Wang H, Ju J. Association of BCSC-1 and MMP-14 with human breast cancer. Oncol Lett. 2018;15(4):5020–5026.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.