Back to Journals » Infection and Drug Resistance » Volume 13
Gastrointestinal Tract Colonization Rate of Extended-Spectrum Beta-Lactamase- and Carbapenemase-Producing Enterobacteriaceae and Associated Factors Among Hospitalized Patients in Arba Minch General Hospital, Arba Minch, Ethiopia
Authors Aklilu A , Manilal A, Ameya G, Woldemariam M , Siraj M
Received 25 November 2019
Accepted for publication 7 May 2020
Published 22 May 2020 Volume 2020:13 Pages 1517—1526
DOI https://doi.org/10.2147/IDR.S239092
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Joachim Wink
Addis Aklilu,1 Aseer Manilal,1 Gemechu Ameya,2 Melat Woldemariam,1 Munira Siraj1
1Arba Minch University, College of Medicine and Health Sciences, Department of Medical Laboratory Science, Arba Minch, Ethiopia; 2Kotebe Metropolitan University, College of Health Sciences, Department of Medical Laboratory Science, Addis Ababa, Ethiopia
Correspondence: Addis Aklilu
Arba Minch University, P.O. Box No. 21
Tel +251912718569
Email [email protected]
Background: The incidence of hospital-acquired enterobacteria that produce extended-spectrum beta-lactamases (ESBLs) is on the rise worldwide. Colonization of gastrointestinal tract by extended-spectrum beta-lactamase Enterobacteriaceae, a prominent causative agent, results in life-threatening infections.
Objective: To determine the rate of gastrointestinal colonization by extended-spectrum beta-lactamase- and carbapenemase-producing Enterobacteriaceae and also to elucidate the antibiotic susceptibility profile and associated risk factors among hospitalized patients in Arba Minch General Hospital, Ethiopia.
Methodology: A facility-based cross-sectional study was conducted in Arba Minch General Hospital from May 2018 to July 2019. Sociodemographic data and associated factors were collected using a pre-tested-structured questionnaire. Stool specimens were collected using sterile stool cups. Each sample was then inoculated onto MacConkey agar. Bacterial isolates were identified using various biochemical tests. Screening and confirmatory tests for extended-spectrum beta-lactamase- and carbapenemase-producing Enterobacteriaceae were performed using the modified Kirby–Bauer disc diffusion technique. Statistical package for Social Science was used to analyze the data. The P-value ≤ 0.05 was considered as statistically significant.
Results: A total of 421 hospitalized patients were enrolled in this study of which there were 240 (57%) females. The mean age of the study participants was 28.8 with SD of 15.7. Majority of participants were in the age range of 25– 40 years 179 (42.5%). About 146 (34.7%) participants were found to be colonized by extended-spectrum beta-lactamase-producing Enterobacteriaceae. The predominant ESBL-producing isolates were Escherichia coli 62 (42.46%) followed by Klebsiella pneumoniae 60 (41.09%). Six (1.43%) carbapenemase-producing K. pneumoniae were isolated. ESBL-producing Enterobacteriaceae showed higher resistance against tetracycline (91.1%) and cotrimoxazole (93.84%). Colonization of the gastrointestinal tract by ESBL showed statistically significant association with regard to chronic diseases (p< 0.001) and the administration of oral antibiotics after admission (p=0.020).
Conclusion: The overall colonization rate of the gastrointestinal tract by extended-spectrum beta-lactamase-producing Enterobacteriaceae was prominent. The extended-spectrum beta-lactamase-producing isolates exhibited a higher level of resistance against the commonly used antibiotics which further needs greater attention.
Keywords: colonization, gastrointestinal tract, ESBL, enterobacteriaceae, susceptibility
Introduction
Drug resistance is an increasing public health problem worldwide. Antibiotic resistance among Gram-negative bacilli is a rapidly increasing conundrum due to the organisms’ ability to mutate and to acquire and transmit plasmids and other mobile genetic elements encoded with resistance genes. Beta-lactam antibiotics are among the most commonly used antibiotics which include penicillins, cephalosporins and carbapenems and the emergence of resistance to these agents had resulted in a major clinical crisis. Infections caused by drug-resistant organisms pose important challenges during the treatment of both common and life-threatening diseases.1,2
β-lactamase production is one of the most important mechanisms of resistance displayed by the Gram-negative bacilli to penicillins and cephalosporins. Extended-spectrum beta-lactamases (ESBLs) are enzymes capable of hydrolyzing many beta-lactam antibiotics including penicillins and cephalosporins, thereby protecting ESBL-producing bacteria from the action of these groups of drugs. Carbapenems are the drug of choice for infections caused by these bacteria.2–4 Cephamycins and carbapenems are the only beta-lactam families that remained fully effective against extended-spectrum beta-lactamase-producing Enterobacteriaceae (ESBL-PE). Combinations with beta-lactam inhibitors like clavulanic acid can restore the activity of several beta-lactams. However, nowadays these ESBL-PE are becoming increasingly resistant to carbapenems due to the production of carbapenemase. Klebsiella pneumoniae is the major carbapenemase producer worldwide. Besides, species of Acinetobacter and Pseudomonas are also becoming resistant to carbapenems due to mono-therapy.5,6,7
ESBL-producing bacteria can be transmitted through fecal-oral route and/or via direct person to person contact. In a hospital setting, the transmission can be among patients, from patients to health-care workers, or/and to visitors. Especially when health-care workers contract the bacterial infections, there exists a chance of disseminations to a larger population. Therefore, hospitalized patients are considered as the main source of the causative agent, ESBL-PE.8
The incidence of MDR is increasing globally and even there are reports of pan drug-resistant (PDR) isolates. The World Health Organization (WHO) has declared infections caused by multidrug-resistant bacteria as an emerging global health problem.3,9
MDR Gram-negative bacteria are a major public health threat. For more than two decades, the incidence of hospital-acquired enterobacteria that produce ESBLs has increased worldwide. The widespread occurrence of ESBL-producing bacteria not only affects the choice of antibiotics but may also cause excessive morbidity, mortality and economic crisis.10,11
The rates of gastrointestinal tract colonization caused by ESBL-PE vary considerably in different geographical areas and also change over time. For instance, in various European countries reported carriage rates fall between 8% and 28%. And in recent years, the higher prevalence rate had been reported, viz., 50.5%, 63.3% and 69.3% in China, Egypt, and Thailand, respectively. The rates are found to be increasing over time especially in low-income countries including Ethiopia.12,13
In hospitalized patients, ESBL-PE has become the most common type and is highly prevalent in developing regions of the subtropics. According to some reports, the international/local travel may contribute substantially and certain proportion of visitors reach the ESBL burdened destinations resulting in the contraction of ESBL-PE. They may spread the bacteria to their close contacts and local hospitals and contribute further to the dissemination of MDR bacteria worldwide.7,14
Most commonly, patients infected with ESBL-producing organisms are more at risk. This is because frequently they are treated with antibiotics to which the organisms exhibit a high level of resistance. The mortality rates in infections caused by ESBL-producing bacteria are higher and some studies even show that the rate may range from 42% to 100%.15 On the other hand, infections caused by carbapenems resistant organisms are associated with mortality rates up to 70%. The spread of carbapenems resistant Enterobacteriaceae (CRE) in health-care settings is another important medical problem. Most of the nations are at risk of falling victim to the emergence of CRE. Therefore, infection prevention and control system should be strengthened to combat the spread of such resistant bacteria.16
ESBLs also add to the burden on health-care systems causing prolonged hospital stay. Report from Europe revealed that there could be more than 2700 deaths and excess burden of 18.1 million euro (in cost-wise), and also more than 120,000 days of hospital stay. ESBL-PE often displays MDR phenotypic characteristics, further limiting the therapeutic options.17,18
Survey of the literature indicated that little is known about the gastrointestinal carriage of ESBLs producers and antibiotic susceptibility patterns in Ethiopia, with only one study, which was conducted in Addis Ababa, Ethiopia. Therefore, this study was conducted to investigate the colonization rate of the gastrointestinal tract with ESBL-PE, antibiotics susceptibility pattern and also to identify associated risk factors of carriage rate among hospitalized patients in Arba Minch General Hospital, Arba Minch, Ethiopia.
Materials and Methods
Study Design, Area and Period
A facility-based prospective cross-sectional study was conducted from May 2018 to July 2019 at Arba Minch General Hospital which is located in Arba Minch town the south of Addis Ababa. It was established in 1961 during the period of Emperor Haile Selassie and was the only hospital in the town at that time, and even today continues to serve a large number of patients annually, around 1000,000 individuals visit the hospital. The hospital has different wards including medical inpatient, surgical, emergency, gyne-obstetrics, and neonatal/pediatric. There are more than 300 beds that are actively serving. The hospital serves for both outpatients and inpatients. Annually there are more than 10,000 admissions including those admitted for less than 48 hrs and for more than 48 hrs.
Population
Source population
All the patients who were admitted into Arba Minch General Hospital.
Study Population
Entire patients who were admitted to medical, surgical, gynecology and pediatrics wards for more than 48 hrs in Arba Minch General Hospital were our study population.
Sample Size Determination and Sampling Technique
Sample Size Determination
The sample size was determined by using a single population proportion formula. P-value of 0.52 from a previous study done in Ethiopia was opted.19 After considering 95% of confidence interval (Zα/2= 1.96) and 5% of marginal error (d=0.05), the initial sample size of 383 was computed using the following expression:
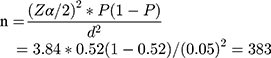
Finally, by considering the 10% of contingency, 38 subjects were added and the final sample size was consolidated as 421.
Sampling Technique
A total of 421 patients were included in this study. A systematic random sampling technique was used to recruit the study participants. The first participant was selected by lottery method and by using the formula (K= 880/421=2.09). Therefore, every two individuals were taken. Individual participants were selected randomly in every Kth interval during the study period. We have included participants from four wards (Medical inpatient, surgical, pediatrics and gynecology) proportionally.
Eligibility Criteria
The inclusion criteria for the study subjects was the patients who were admitted to medical, surgical, gynecology and pediatric wards of Arba Minch General Hospital for more than 48 hrs. The exclusion criteria for the study subjects were: (1) Patients who took antibiotics on the day of sample collection, (2) Patients who are admitted for less than 48 hrs.
Data Collection and Laboratory Investigation
Data Collection
On the starting day of data collection, patients who were admitted to the hospital 48 hrs ago were recruited by taking written consent from each one individually. Details pertaining to the socio-demographic data (age, residence and occupation), clinical history (existing antibiotic treatment, previous antibiotic therapy if any, previous history of hospitalization for longer period and also the incidence of earlier colonization by any multidrug-resistant bacteria) were collected by using a pre-tested-structured questionnaire.
Sample Collection
Two stool specimens were collected from each patient using sterile stool cups after 48 hrs of admission and just before discharging him/her from the hospital. The collected stool specimens were then transported to the Microbiology and Parasitology laboratory of College of Medicine and Health Sciences, Arba Minch University (within 1 hr of collection).
Laboratory Investigation
Bacterial Isolation
Stool samples were inoculated onto MacConkey agar and then incubated for 24 hrs at 37°C. After incubation, bacterial growth was observed and pure cultures of bacterial isolates were subsequently subjected to species identification and confirmation. Biochemical (indole, oxidase, TSI, citrate, urease, LDC, motility, and H2S gas production), morphological, and physiological characteristics of isolated bacteria were determined by adopting standard laboratory methods as described elsewhere. Corresponding American Type Culture Collection strains were utilized as reference standards to validate the biochemical identification.20
Screening of ESBL- and Carbapenemase-Producing Enterobacteriaceae
Phenotypic characterization of ESBL-PE was performed for the screening of patients. Screening for the presence of ESBL or carbapenemase production was done according to CLSI guidelines.21
Confirmatory Test for ESBL Production
ESBL phenotypic confirmatory test was performed by standard disc diffusion technique using indicator drugs. Briefly, organism suspected for the production of ESBL was spread onto a Mueller-Hinton agar by preparing a suspension equivalent to 0.5 CFU McFarland standards. Two indicator antibiotic discs were then placed 15 mm apart (center to center). One of the discs contains amoxicillin/clavulanic acid and the other contains an expanded-spectrum cephalosporin (ceftriaxone, cefotaxime or ceftazidime). After 24 hrs of incubation, if the zone of inhibition in between the discs were enhanced, the test was considered positive. The carbapenemases production of K. pneumoniae was confirmed by meropenem/cloxacillin combination discs test technique. The test was considered positive if an increase in the inhibition zone by 5 mm was observed compared to the inhibition zone of carbapenems alone. K. pneumoniae CCUG 56233 was used as a positive control for carbapenemase production and K. pneumoniae ATCC 25955 obtained from EPHI was used as a negative control in all phenotype tests.21
Antibiotic Susceptibility Testing
Antimicrobial susceptibility testing was performed by Kirby–Bauer disk diffusion technique on Mueller-Hinton agar according to CLSI guidelines (CLSI, 2017). Antibiotics (Oxoid Ltd Basingstoke, Hampshire, UK) which are recommended by CLSI for susceptibility test such as aztreonam (15 µg), ciprofloxacin (5 μg), chloramphenicol (30μg), trimethoprim-sulfamethoxazole (co-trimoxazole) (1.25/23.75 µg), amikacin (10μg), tetracycline (30μg), gentamycin (10μg), tobramycin (10 µg), meropenem (10 µg), imipenem (10 µg), and cefoxitin (30 µg) were included. The bacteria that were resistant for three or more classes of antibiotics were considered as multidrug-resistant (MDR).22
Quality Assurance
Data quality was ensured from collection up to the final laboratory identification by following a standard operating procedure. Culture media was prepared according to manufacturer’s instruction, and the sterility was checked by incubating 5% of the prepared media at 35–37°C overnight and also by observing the bacterial growth. Those batches of the media that show the growth was discarded and prepared again. The performance of the prepared media was also checked by inoculating control strain E. coli ATCC 25922 and K. pneumoniae ATCC 700603, which were obtained from Ethiopian Public Health Institute (EPHI). The performance of antibiotic discs were checked by using Enterococcus faecalis ATCC 29122 and co-trimoxazole disc. Five percentage of the questionnaire was pretested prior to the data collection and process was supervised on a daily basis. Incompletely filled questionnaires were discarded.
Statistical Analysis
Data were entered, edited, cleaned and analyzed by using Statistical Package for Social Sciences (SPSS) software Version 21.0. Descriptive statistics were calculated according to the objectives of study such as frequency, mean and standard deviations. Binary logistic regression analysis was used to determine the association between colonization rate and associated factors. All variables with P<0.25 in the bivariate analysis were included in the final model/multivariate analysis. P ≤0.05 was considered as a statistically significant association. Finally, the magnitude of the association among different variables in relation to the outcome variable was measured by the odds ratio with a 95% confidence interval. Compiled results were presented in the form of text, tables or graphs.
Ethical Considerations
The study protocol was ethically approved by the institutional review board of Arba Minch University, College of Medicine and Health Sciences (Ref. No. CMHS/10635/21). This study was conducted in accordance with the declaration of Helsinki. Besides, separate permission was procured from Arba Minch General Hospital administrators. Formal written consent was obtained from each study participant. Confidentiality was strictly maintained from sample collection up to the final report writing.
Operational Definitions
Previous antibiotics treatment – History of antibiotic treatments for the last one year if any.
Prolonged hospitalization – Hospitalization for more than two weeks
Presence of chronic disease – Diseases like HIV, diabetes mellitus and hypertension
Habit of handwashing – (Less frequent – participants who never wash their hands at all and who wash their hands once a day, Frequent – participants who wash their hands two to three times a day, More frequent – participants who have the habit of washing their hands more than three times a day).
Results
Socio-Demographic Characteristics
A total of 421 hospitalized patients were included in this study of which there were 240 (57%) females. The mean age of the study participant was 28.8 years with SD of ±15.7. Majority of participants were in the age range of 25–40 years 179 (42.5%). Half of the study participants were rural dwellers 211 (50.1%), whereas, most of them had the habit of eating raw foods, 389 (92.4%). Number of participants who had the habit of eating more than one type of raw foods were 167 (39.7%). Also, chronic diseases were found in 126 (38.5%) patients (Table 1).
 |
Table 1 Socio-Demographic Characteristics of Hospitalized Patients at Arba Minch General Hospital, 2019 |
Fecal Carriage Rate of ESBL-PE
Among the total 421 admitted patients, 140 (33.25%) were found to be colonized by ESBL-PE. Majority of the carriers about 92 (69.02%) were rural dwellers. Among the carriers, 74 (50.68%) were males. Patients with a family income of below 1000 birr per month and participants who have had the habit of eating raw foods showed higher carriage rates of ESBL-PE (Table 2). It was found that the predominant ESBL-producing organism was E. coli 62 (44.28%) followed by K. pneumoniae 60 (42.85%) (Figure 1). On the other hand, only six (1.43%) isolates of carbapenemase-producing K. pneumoniae were identified.
 |
Table 2 Prevalence and Association of ESBL-PE Colonization Rate with Demographic and Clinical Factors Among Hospitalized Patients in Arba Minch General Hospital, 2019 |
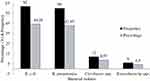 |
Figure 1 Frequency of ESBL-producing bacteria. |
ESBL-PE Colonization Rate and Associated Factors
Multivariate analysis revealed that colonization of the gastrointestinal tract by ESBL showed statistically significant association with the presence of chronic diseases (p<0.001, AOR=0.136, 95% CI=0.048–0.380), prolonged previous history of hospitalization (p=0.004, AOR=0.312, 95% CI=0.141–0.69), administration of oral antibiotics after admission (p=0.020, AOR= 2.882, 95% CI=1.179–7.043), residence in rural area (p<0.001, AOR=2.23, 95% CI= 1.47–3.37) and habit of handwashing (p=0.008, AOR= 0.13, 95% CI= 0.03–0.59) (Table 2).
Antimicrobial Susceptibility Pattern
Totally, nine antibiotics were tested against ESBL-PE by using the Kirby–Bauer disc diffusion technique. ESBL-producing Enterobacteriaceae showed a higher resistance against tetracycline (91.1%), cotrimoxazole (93.84%), chloramphenicol (69.18%) and less resistance against aztreonam (4.11%) and amikacin (4.79%). The isolates of K. pneumoniae showed higher resistance against tetracycline (91.70%), cotrimoxazole (96.7%), and chloramphenicol (73.3%). In the case of E. coli, resistance was produced against tetracycline, cotrimoxazole (93.5%), and ciprofloxacin (72.6%) (Table 3).
 |
Table 3 Antimicrobial Resistance Pattern of ESBL-PE Isolates Among Hospitalized Patients in Arba Minch General Hospital, 2019 |
Multidrug Resistance Pattern of ESBL-PE
In this study, the overall prevalence of MDR bacteria (resistant to three or more classes of antibiotics tested) was 99 (70.71%). Most of the Gram-negative enteric bacteria were resistant to at least one of the antibiotics tested (Table 4).
 |
Table 4 Multidrug Resistance Pattern of ESBL-PE Isolates Among Hospitalized Patients in Arba Minch General Hospital, 2019 |
Discussion
The incidence of MDR is increasing globally and even there are reports of PDR isolates. Recently, WHO has declared infections caused by multidrug-resistant bacteria as an emerging global health problem.3,9 The incidence of hospital-acquired enterobacteria that produce ESBLs had increased worldwide. The widespread presence of ESBL-producing bacteria not only affects the choice of antibiotics but also may cause excessive morbidity, mortality and economic crisis on a global scale.10,11
In this study, the overall colonization rate of the gastrointestinal tract by ESBL-PE was high 140 (33.25%) 95% CI (29.36%–42.67%). This data is comparable with results of a study reported in India 43%.23 Our results showed higher carriage rate compared to studies conducted in Turkey 24%,24 Israel 8%,25 India 9.3%26 and USA 2%,27 but lower than the rates found in other studies conducted in Ethiopia 52%,19 Ecuador 56%,28 Morocco 58%,29 Egypt 68%,30 and Cambodia 61.9%.31 The possible reason for the variability of colonization rate found among different studies could be attributed to variations in the socioeconomic status of the population included and the study design.
The predominant ESBL-PE was E. coli (44.28%), followed by K. pneumoniae (42.85%), Citrobacter species (8.57%) and Enterobacter species (4.3%). This is in line with previous studies conducted in Ethiopia,19 Turkey,24 Ecuador,28 and Egypt.30 However, in studies conducted in Morocco,29,32 India23,26 and the USA,25 the isolates of K. pneumoniae were found to be the predominant ESBL producer.
In the present study, the colonization rate of carbapenemase-producing K. pneumoniae was 6(1.43%) 95% CI (0.83 −3.54%). This is in accordance with the results of two similar studies reported in Ethiopia (2%)19 and in Morocco (1.8%).32 However, the study conducted in Egypt and Cambodia reported higher rates of colonization with carbapenemase such as 5% and 7.5%, respectively.30,31
In the present study, the gastrointestinal colonization rate with ESBL-PE has statistical correlation with various factors such as prolonged hospitalization, poor handwashing habits, presence of chronic diseases and habit of eating raw foods. Similar studies conducted in the USA, Israel, and Ecuador had shown that prolonged hospital stays are significantly associated with ESBL carriage rate.25,27,28 These factors could contribute to the transmission and/or dissemination of resistant bacteria among patients in the hospital setup. Probably they may also contribute to the onset of outbreaks both in hospital and in the community. On the other hand, the study conducted in Turkey showed that catheterization and surgical procedures were associated with the colonization of ESBL-PE in the gastrointestinal tract.24 Furthermore, the study done in Ethiopia also showed a greater extent of ESBL colonization within lower age group, specifically neonates.19
Administration of oral antibiotics after admission also showed a statistically significant association with gastrointestinal colonization rate by ESBL-PE. This is in concordance with a study conducted related to antibiotic treatment in Israel.25 Conversely, studies done in the USA revealed previous exposure to antibiotics and it showed a statistically significant association with colonization rate.27,33
In the present study, ESBL-PE showed higher resistance to commonly used antibiotics. This was in consonant with the findings obtained from studies done in India23 and Ethiopia.19 Besides, the results of our study revealed lower resistance against cefoxitin (14.38%), aztreonam (5.16%) and amikacin (4.79%). Another study performed in Ethiopia showed higher resistance rates against aztreonam, and cefoxitin such as 97% and 28%, respectively. Moreover, ESBL-producing E. coli and K. pneumoniae showed a higher level of resistance against tetracycline, cotrimoxazole, chloramphenicol, and ciprofloxacin. These observations are well-nigh similar to several studies conducted in Ethiopia19 and other locales of the world.23,24,27,29,30 The higher level of resistance showed by the isolates may be due to the overuse or irrational use of antibiotics. This finally makes the treatment of infection caused by ESBL-producing bacteria a very challenging task.
The overall prevalence of MDR ESBL-producing isolates that colonized the gastrointestinal tract was 99 (70.71%). This result is comparable to the findings of studies done in Ethiopia19 and Sweden 68%.6 However, this value is higher compared to the data obtained from a study reported in Egypt 47.83%.30 However, it was much lower than the overall prevalence reported from in Tanzania 94%.3 Most of the ESBL-PE can display co-resistance to non-beta-lactam antibiotics. Such variations in the drug resistance pattern may be attributed to the difference in rational use of antibiotics in clinical setups.
Conclusion
The overall colonization rate of the gastrointestinal tract by ESBL-PE among hospitalized patients at Arba Minch General Hospital was higher (34.7%). The predominant isolates of ESBL-producing bacteria were E. coli and K. pneumoniae. ESBL-producing isolates showed higher degree of resistance against the commonly used antibiotics such as tetracycline, cotrimoxazole, ciprofloxacin, and chloramphenicol which need a great attention from the concerned bodies at different levels. The carriage rate of ESBL-PE among hospitalized patients is increasing, so that it needs a nationwide survey with a large population size. Besides, awareness should be created in hospitals on how such drug-resistant bacteria develop the mechanism of resistance, spread and also the hospital should work on infection prevention strategies, especially on hospital-acquired drug-resistant bacteria. Finally, further in-depth studies pertaining to the molecular characterization of ESBL strains are warranted.
Acknowledgment
We would like to thank Arba Minch University for financial support.
Author Contributions
All authors have contributed in designing, data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors declared that they have no conflicts of interest in this work.
References
1. Siddiqui N, Bhakre J, Damle A, Bajaj J. Prevalence of Extended Spectrum Beta Lactamase (ESBL) producing gram negative bacilli from various clinical isolates. IOSR J Dent Med Sci. 2014;13(9):08–11. doi:10.9790/0853-13980811
2. Kumar D, Singh AK, Ali MR, Chander Y. Antimicrobial Susceptibility Profile of Extended Spectrum β-Lactamase (ESBL) producing Escherichia coli from various clinical samples. Infect Dis. 2014;7:1–8.
3. Tellevik MG, Blomberg B, Kommedal Ø, et al. High prevalence of faecal carriage of ESBL producing enterobacteriaceae among children in Dares Salaam, Tanzania. PLoS One. 2016;11(12):1–13. doi:10.1371/journal.pone.0168024
4. Juma BW, Kariuki S, Waiyaki PG, Mutugi MM, Bulimo WD. The prevalence of TEM and SHV genes among extended-spectrum beta-lactamase-producing klebsiella pneumoniae and escherichia coli. Afr J Pharmacol Ther. 2016;5(1):1–7.
5. Ibrahim AH, Abdelhalim KA. Detection of extended spectrum beta-lactamase in Klebsiella pneumoniae isolated from sputum in Khartoum, Sudan. World J Pharm Res. 2015;4(3):131–140.
6. Balkhed AO. Extended-Spectrum ß-Lactamase-Producing Enterobacteriaceae Antibiotic Consumption, Detection and Resistance Epidemiology. Sweden: Linköping University Medical Dissertations, LiU tryk, Linkoping; 2014. Available from: http://liu.diva-portal.org/smash/get/diva2:695475/FULLTEXT01.pdf.
7. Woerther P, Andremont A, Kantele A. Travel-acquired ESBL-producing Enterobacteriaceae: impact of colonization at individual and community level. J Travel Med. 2017;24(1):S29–S34. doi:10.1093/jtm/taw101
8. Haverket MR Carriage and transmission dynamics of multidrug-resistant Enterobacteriaceae PhD thesis. Netherlands: university of Utrecht; 2015. Available from: file:///C:/Users/user/Downloads/Haverkate.pdf.
9. World Health Organization. Antimicrobial resistance—global report on surveillance; 2014. Available from: http://apps.who.int/iris/bitstream/10665/112642/1/9789241564748_eng.pdf?ua=1.
10. Villar HE, Baserni MN, Jugo MB. Faecal carriage of ESBL-producing Enterobacteriaceae and carbapenems resistant Gram negative bacilli in community settings. J Infect Dev Ctries. 2013;7(8):630–634. doi:10.3855/jidc.2900
11. Luvsansharav U, Hirai I, Niki M, et al. Analysis of risk factors for a high prevalence of extended-spectrum b-lactamase-producing Enterobacteriaceae in asymptomatic individuals in rural Thailand. J Med Microbiol. 2011;60(5):619–624. doi:10.1099/jmm.0.026955-0
12. Ebrahimi F, Kardos G. Epidemiology of Faecal Carriage of Extended Spectrum Beta Lactamase Producing Enterobacteriaceae in Healthy Community and in Different Patient Population. Debrecen, Hungary; 2016. Available from: https://dea.lib.unideb.hu/dea/.
13. Erdogan DC, Comert F, Aktaş E, Koktürk F, Kulah C. Fecal carriage of extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella spp. in a Turkish community. Turk J Med Sci. 2017;47:172–179. doi:10.3906/sag-1512-9
14. Morosini M, Valverde A, Garcıa-Castillo M, Nordmann P, Canton R. Persistent isolation of Salmonella Concord harbouring CTX-M-15, SHV-12 and QnrA1 in an asymptomatic adopted Ethiopian child in Spain also colonized with CTX-M-14- and QnrB-producing enterobacteriaceae. J Antimicrob Chemother. 2010;65(7):1545–1546. doi:10.1093/jac/dkq168
15. Rupp ME, Fey PD. Extended Spectrum Beta-Lactamase (ESBL)-producing enterobacteriaceae considerations for diagnosis, prevention and drug treatment. Drugs. 2003;63(4):353–365. doi:10.2165/00003495-200363040-00002
16. Friedman ND, Carmeli Y, Walton AL, Schwaber MJ. Carbapenem-resistant Enterobacteriaceae: a strategic roadmap for infection control. Infect Control Hosp Epidemiol. 2017;38(5):580–594. doi:10.1017/ice.2017.42
17. Brolund A. Overview of ESBL-producing Enterobacteriaceae from a Nordic perspective. Infect Ecol Epidemiol. 2014;4:10.
18. Schechner V, Temkin E, Harbarth S, Carmeli Y, Schwaber MJ. Epidemiological interpretation of studies examining the effect of antibiotic usage on resistance. Clin Microbiol Rev. 2013;26(2):289–307. doi:10.1128/CMR.00001-13
19. Desta K, Woldeamanuel Y, Azazh A, et al. High gastrointestinal colonization rate with extended-spectrum β-lactamase-producing enterobacteriaceae in hospitalized patients: emergence of carbapenemase-producing K. pneumoniae in Ethiopia. PLoS One. 2016;11(8):e0161685. doi:10.1371/journal.pone.0161685
20. Cheesbrough M. District Laboratory Practice in Tropical Countries, Second Edition. 2nd Editio. District Laboratory Practice in Tropical Countries.
21. CLSI. Performance Standards for Antimicrobial Susceptibility Testing.
22. Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. doi:10.1111/j.1469-0691.2011.03570.x
23. Patel SM, Sood N, Patel P. A study of colonization with ESBL and Ampc E. coli in gut of patients of tertiary care hospital, Ahmedabad. Natl J Community Med. 2016;7(4):278–280.
24. Kiremitci A, Dinleyici EC, Yargıc ZA, et al. Prevalence and Risk factors of fecal carriage of extended-spectrum β-lactamase (ESBL)-Producing enterobacteriaceae in hospitalized and ambulatory children. J Pediatr Inf. 2011;5(2):54–58. doi:10.5152/ced.2011.22
25. Friedmann R, Raveh D, Zartzer E, et al. Prospective evaluation of colonization with extended-spectrum beta-lactamase (ESBL)-producing enterobacteriaceae among patients at hospital admission and of subsequent colonization with ESBL-producing enterobacteriaceae among patients during hospitalization. Infect Control Hosp Epidemiol. 2009;30(6):534–542. doi:10.1086/597505
26. Rashid M, Modi S, Sarwat T, Chander Y, Rastogi V, Manocha H. Carriage of ESBL and AmpC-positive enterobacteriaceae in gastrointestinal tract of healthy community subjects and hospitalized patients and detection of blaCTX-M gene in ESBL positive isolates. Ind Med Gaz. 2015;1:212–217.
27. Han JH, Nachamkin I, Theoklis D, et al. Risk factors for gastrointestinal tract colonization with extended-spectrum β lactamase (ESBL)–producing Escherichia coli and Klebsiella species in hospitalized patients. Infect Control Hosp Epidemiol. 2012;33(12):1242–1245. doi:10.1086/668443
28. Nordberg V, Peralta AQ, Galindo T, et al. High proportion of intestinal colonization with successful epidemic clones of ESBL-producing enterobacteriaceae in a neonatal intensive care unit in ecuador. PLoS One. 2013;8(10):e76597. doi:10.1371/journal.pone.0076597
29. Cheikh A, Belefquih B, Chajai Y, Cheikhaoui Y, Hassani AE, Benouda A. Enterobacteriaceae producing extended-spectrum β-lactamases (ESBLs) colonization as a risk factor for developing ESBL infections in pediatric cardiac surgery patients: “retrospective cohort study”. BMC Infect Dis. 2017;17(1):237. doi:10.1186/s12879-017-2346-4
30. Abdallah HM, Alnaiemi N, Reuland EA, et al. Fecal carriage of extended-spectrum β-lactamase- and carbapenemase-producing Enterobacteriaceae in Egyptian patients with community-onset gastrointestinal complaints: a hospital-based cross-sectional study. Antimicrob Resist Infect Control. 2017;6(1):62. doi:10.1186/s13756-017-0219-7
31. Turner P, Pol S, Soeng S, et al. High prevalence of antimicrobial resistant gram negative colonization in hospitalized Cambodian infants. Pediatr Infect Dis J. 2016;35(8):856–861. doi:10.1097/INF.0000000000001187
32. Arhoune B, Oumokhtar B, Hmami F, et al. Rectal carriage of extended-spectrum b-lactamase- and carbapenemase-producing Enterobacteriaceae among hospitalized neonates in a neonatal intensive care unit in Fez, Morocco. J Global Antimicrob Resist. 2017;8:90–96. doi:10.1016/j.jgar.2016.11.004
33. Harris AD, McGregor JC, Johnson JA, et al. Risk factors for colonization with extended-spectrum beta-lactamase-producing bacteria and intensive care unit admission. Emerg Infect Dis. 2007;13(8):1144–1149. doi:10.3201/eid1308.070071
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
