Back to Journals » Drug Design, Development and Therapy » Volume 9
Prevalence of the rs7903146C>T polymorphism in TCF7L2 gene for prediction of type 2 diabetes risk among Iranians of different ethnicities
Authors Allahdini M, Kamalidehghan B, Akbari L, Azadfar P, Rahmani A, Ahmadipour F, Yong Meng G, Masserrat A, Houshmand M
Received 9 February 2015
Accepted for publication 30 March 2015
Published 28 October 2015 Volume 2015:9 Pages 5835—5841
DOI https://doi.org/10.2147/DDDT.S82485
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Shu-Feng Zhou
Mojgan Allahdini,1 Behnam Kamalidehghan,2 Leila Akbari,3 Parisa Azadfar,3 Ali Rahmani,1 Fatemeh Ahmadipour,2 Goh Yong Meng,4 Abbas Masserrat,5 Massoud Houshmand6
1Department of Molecular Biology, Ahar Branch Islamic Azad University, Ahar, Iran; 2Pharmacy Department, Faculty of Medicine, University of Malaya, Kuala Lumpur, Malaysia; 3Department of Biology, Sciences and Research Branch, Azad University, Tehran, Iran; 4Department of Veterinary Preclinical Sciences, Faculty of Veterinary Medicine, Universiti Putra Malaysia, Serdang, Malaysia; 5Department of Medical Genetics, Taban Clinic, 6Department of Medical Genetics, National Institute of Genetic Engineering and Biotechnology (NIGEB), Tehran, Iran
Background: Pharmacogenetics is the study of genetic polymorphisms affecting responses to drug therapy. The common rs7903146 (C>T) polymorphism of the TCF7L2 gene has recently been associated with type 2 diabetes (T2D). In this study, prevalence of the rs7903146 (C>T) polymorphism in the TCF7L2 gene for prediction of T2D risk was examined in an Iranian population of different ethnicities.
Methods: The prevalence of rs7903146 (C>T) and the predicted phenotypes, including extensive metabolizers, intermediate metabolizers, and poor metabolizers were investigated in blood samples of 300 unrelated healthy individuals in an Iranian population, including Fars, Turk, Lure, and Kurd, using polymerase chain reaction restriction fragment length polymorphism and direct genomic DNA sequencing.
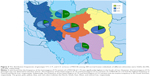
Results: The homozygous wild-type (C/C), heterozygous (C/T), and homozygous (T/T) allelic frequencies of rs7903146 (C>T) in the TCF7L2 gene were 29% (extensive metabolizers), 66.34% (intermediate metabolizers), and 4.66% (poor metabolizers), respectively. The C/C, C/T, and T/T genotypic frequencies of the rs7903146 (C>T) allele were significantly different (P<0.01) among Iranians of different ethnicities. The frequency of the homozygous T/T variant of the rs7903146 (C>T) allele was significantly low in the Lure (P<0.01) and high in the Fars (P<0.001) ethnicities. Additionally, the frequency of the T/T variant of the rs7903146 (C>T) allele in the South of Iran was the highest (P<0.04), while the East of Iran had the lowest frequency (P<0.01).
Conclusion: The prediction of rs7903146 (C>T) is required in drug research and routine treatment, where the information would be helpful for clinicians to optimize therapy and adverse drug reactions and predict drug response in individuals at risk of T2D.
Keywords: pharmacogenetics of type 2 diabetes (T2D), transcription factor 7-like 2 gene, TCF7L2, rs7903146 (C>T) SNP, Fars Turk Lure and Kurd, Iranian populations of different ethnicities
Introduction
Pharmacokinetics is the basis of personalized medicine to determine the activity of drug-metabolizing enzymes, where the occurrence of mutations in genes that are associated with drug metabolism alters drug efficacy.1 Type 2 diabetes (T2D) is a common complex disorder resulting from the interplay of genetic and environmental factors.2,3
A strong association was identified between T2D and a single nucleotide rs7903146C>T polymorphism, located within intron 4 of the TCF7L2 gene.4,5 TCF7L2 gene consists of 17 exons and is located on chromosome 10q25.3, where the rs7903146C>T polymorphism is the most important of these improved variants with negative effects on metabolism. Certain variants of TCF7L2 gene increase the risk of T2D and reduce insulin secretion.6,7 Prevalence of T2D is increasing in an epidemic fashion worldwide, attributed to a collision between genes and increasing globalization of the Western lifestyle, and a few genetic variants consistently associated with T2D.8,9 Using gene-locus genotyping, this variant has been consistently associated with T2D in European, Asian, and African populations.10 In this study, the prevalence of the rs7903146C>T polymorphism in the TCF7L2 gene for the prediction of T2D risk was investigated among Iranians of different ethnicities, and in comparison with populations of other countries.
Materials and methods
In this study, 300 blood samples of unrelated and healthy donors from Iranian populations of different ethnicities, including 178 Fars, 75 Turk, 29 Lure, and 18 Kurd, were obtained from the Special Medical Centre, Tehran, Iran. Blood samples (2 mL) were taken and collected in tubes with ethylenediaminetetraacetic acid. Written informed consent, including consent to participate in the study for genetic analysis and consent to publish, was obtained from individuals, and the Medical Ethics Committee of the Special Medical Center specifically approved this study (approval number AA-25-2008). The exclusion criterion for the healthy control group was any history of cancer, metabolic diseases and nuclear and mitochondrial DNA-related diseases that may affect the DNA.
DNA extraction and primer sequences
Genomic DNA was extracted using an Molecular Biological System Transfer kit (salting-out method) from 2 mL of whole blood. Primer sequences for the rs7903146 (C>T) were ordered as described by Palizban et al.11,12 The oligonucleotide forward and reverse primers that were used to amplify the rs7903146C>T are as follows: F: 5′-TTA GAG AGC TAA GCA CTT TTT AGGTA-3′, R: 5′-AGA GAT GAA ATG TAG CAG TGA AGTG-3′.
Restriction fragment length polymorphism (RFLP)
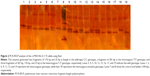
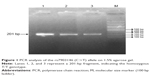
Genotyping of the rs7903146 (C>T) alleles was performed by polymerase chain reaction RFLP (PCR-RFLP). The PCR amplification was carried out with 20–60 ng of genomic DNA, 0.3 U of Taq DNA polymerase, 20 mM of each primers, 10× of PCR buffer, 2.5 mM of MgCl2, and 10 mM of deoxynucleotide triphosphate (dNTP). The reaction mixture was initially denatured at 95°C for 3 minutes, followed by 35 cycles of 95°C for 1 minute, 58.3°C for 50 seconds, 72°C for 50 seconds, and a final extension at 72°C for 10 minutes. The 201-bp amplified fragment was analyzed on 1.5% agarose gel stained with ethidium bromide before restriction digestion. One unit of RsaI restriction enzyme was added to each PCR product (5 μL) and incubated at 37°C for 16 hours. Digested products were analyzed on 8% polyacrylamide gel electrophoresis. The PCR product with a length of 201 bp was subsequently digested with the restriction endonuclease RsaI. The digestion of the homozygous wild type (C/C) leads to 176-bp and 25-bp fragments, a fragment of 201 bp for the homozygous mutant (T/T) and three fragments of 201 bp, 176 bp, and 25 bp for the heterozygous (C/T) genotype (Figures 1 and 2).
Sequencing analysis
The PCR products were examined for specificity using 1.5% agarose gel electrophoresis. Double-stranded DNA automated sequencing was performed using an ABI 3100 sequencing machine (Applied Biosystems, Kavosh Fanavaran Kawsar Company, Iran). All fragments were sequenced in both the forward and reverse directions for confirmation of any nucleotide variation. Sequence variants were analyzed using FinchTV software (http://www.geospiza.com/finchtv/).
Calculation of genotypic and allelic frequencies
Genotypic and allelic frequencies were calculated among Iranians of different ethnicities using the equation which is available online.13
Statistical analysis
The statistical analyses were conducted using SPSS 17.0 to perform the chi-square and confidence interval (95%) tests to calculate the rs7903146C>T frequencies in different ethnicities and different geographical regions of Iran. A P-value of <0.01 is considered statistically significant.
Results
Allele and genotype frequency distributions of the rs7903146 (C>T) variants were analyzed from the blood samples of 300 unrelated healthy individuals in an Iranian population of different ethnicities. The allele frequencies of the homozygous (T/T), heterozygous (C/T), and wild type (C/C) variants of the rs7903146 (C>T) were observed in 14 (4.66%), 199 (66.34%), and 87 (29%) individuals, respectively (Table 1 and Figure 3). The frequency of the homozygous (T/T) variant of rs7903146 (C>T) was significantly high in the Fars ethnicity and low in the Lure (χ2=13, P<0.01). However, the frequency of the homozygous (T/T) variant of rs7903146 (C>T) in Kurd ethnicity was similar to that of Turk (Table 1). The frequency of the heterozygous (C/T) variant of rs7903146 (C>T) was significantly high in Fars and low in Kurd (χ2=198, P<0.01). Additionally, the frequency of the wild type (C/C) variant of rs7903146 (C>T) was significantly highest in Fars, yet lowest in Lure (χ2=86, P<0.01) (Table 1).
The results indicated that the frequency of the homozygous (T/T) variant of rs7903146 (C>T) was significantly highest in the South of Iran (χ2=2.571, P<0.01) with poor metabolizers, while the homozygous (T/T) genotype was not detected in the East of Iran. The frequency of the heterozygous (C/T) variant of rs7903146 (C>T) had the highest frequency in central Iran with intermediate metabolizers, while the South had the lowest frequency (χ2=121.327, P<0.01). Moreover, the frequency of the homozygous (C/C) variant of rs7903146 (C>T) had the highest frequency in the North with extensive metabolizers, while the East (χ2=41.908, P<0.001) had the lowest frequency (Table 1; Figure 4). The sequencing and PCR-RFLP analysis of the rs7903146 (C>T) allele using the RsaI enzyme demonstrates wild-type C/C, the heterozygous C/T and homozygous T/T genotypes (Figures 2 and 3).
Discussion
The TCF7L2 gene is strongly associated with T2D, impaired insulin secretion,4,14–16 and cardiovascular risk,17 encoding for a transcription factor involved in Wnt (wingless and INT-1) signaling.18 Heterodimerization of TCF7L2 with β-catenin induces transcription of a number of genes, including intestinal GCG.19 Here, the prevalence of the rs7903146C>T polymorphism in the TCF7L2 gene for prediction of T2D was investigated in Iranian populations of different ethnicities, with comparisons to populations in other countries. Previous reports indicated that the TCF7L2 gene is associate with T2D risk in different ethnic groups.11,20,21
The meta-analysis by Dou et al22 indicated that among different single nucleotide polymorphisms of the TCF7L2 gene, only the rs7903146 polymorphism has a significant association with T2D risk. However, another meta-analysis study indicated that the rs11196218 polymorphism in the TCF7L2 gene was associated with risk of T2D mellitus among the Asian population.23 Additionally, a study demonstrated that the rs7903146, rs12255372, and rs290487 polymorphisms of TCF7L2 were associated with T2D in a Kurdish ethnic group of Iran.24
Several studies demonstrated that the impact of a single rs7903146(T) polymorphism of TCF7L2 gene in therapeutic response to sulfonylureas and metformin is lower in mutant alleles.25,26 In contrast to previous studies,27–29 the rs7903146 polymorphism in individuals leads to reduced enzyme activity and decrease of appropriate response to metformin and sulfonylurea drug treatments.30–33 The rs7903146 polymorphism in different ethnicities, including Lure, Turk, Fars, and Kurd, is considered as a possibly important determinant for the prevention of the harmful effects of metformin and sulfonylurea drug interactions, resulting in better survival and reduced mortality risk. However, in the present study, T2D patients and enzyme activity responses to metformin and sulfonylurea were not examined due to the fact that this study included only healthy individuals among Iranians of different ethnicities.
The frequencies of the rs7903146 polymorphism in our Iranian population with different ethnicities, including Fars, Turk, Lure, and Kurd, were compared with the frequencies seen in other ethnic populations (Table 2). Of 31 provinces of Iran, studies on only two provinces, Isfahan and Rafsanjan, showed that the frequency of the rs7903146 polymorphism was 0.4111 and 0.33,26 respectively. However, in the present study, the samples were selected from all 31 provinces of Iran, where the frequency of the homozygous (T/T) variant of rs7903146 (C>T) was high in the Fars ethnicity, but low in the Lure, similar to that of ethnic Turk and Kurd. The rs7903146 polymorphism was found with a frequency of 0.37 in the Iranian population, which is higher than the Asian Indians,25 Mexicans,12 Italians,34 Polish,35 and one province of Iran, Rafsanjan,26 but not as high as what was reported in other populations such as North India,36 and one province of Iran, Isfahan.11
In conclusion, this study demonstrates the presence of the rs7903146 polymorphism in the Iranian population, suggesting susceptibility to T2D and adverse or poor drug responses, which may lead to increase in mortality risk. It is clinically important to identify those individuals who probably have altered pharmacokinetics for the rs7903146 polymorphism in order to avoid adverse drug reactions. The rs7903146 polymorphism analysis in different ethnicities is a very useful forensic tool toward personalized medicine. These data also allow comparisons of pharmacogenetic variation between populations with different ethnicities. Therefore, the determination of the presence of this allele in individuals is necessary to prevent the harmful effects of drugs.
Acknowledgments
We would like to thank all the participants for their blood donation to the Medical Genetics Department at the Special Medical Centre, Tehran, Iran.
Disclosure
The authors declare no conflicts of interest in this work.
References
Klotz U. The role of pharmacogenetics in the metabolism of antiepileptic drugs. Clin Pharmacokinet. 2007;46(4):271–279. | ||
Gerich JE. The genetic basis of type 2 diabetes mellitus: impaired insulin secretion versus impaired insulin sensitivity. Endocr Rev. 1998;19(4):491–503. | ||
Ahlqvist E, Ahluwalia TS, Groop L. Genetics of type 2 diabetes. Clin Chem. 2011;57(2):241–254. | ||
Grant SF, Thorleifsson G, Reynisdottir I, et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat Genet. 2006;38(3):320–323. | ||
Nicod N, Pradas-Juni M, Gomis R. Role of the single nucleotide polymorphism rs7903146 of TCF7L2 in inducing nonsense-mediated decay. Springerplus. 2014;3(1):41. | ||
Bo S, Gambino R, Ciccone G, et al. Effects of TCF7L2 polymorphisms on glucose values after a lifestyle intervention. Am J Clin Nutr. 2009;90(6):1502–1508. | ||
Guja C, Gagniuc P, Ionescu-Tîrgovişte C. Genetic factors involved in the pathogenesis of type 2 diabetes. Proc Rom Acad, Series B1. 2012;1:44–61. | ||
Florez JC, Burtt N, De Bakker PI, et al. Haplotype structure and genotype-phenotype correlations of the sulfonylurea receptor and the islet ATP-sensitive potassium channel gene region. Diabetes. 2004;53(5):1360–1368. | ||
Gloyn AL, Weedon MN, Owen KR, et al. Large-scale association studies of variants in genes encoding the pancreatic β-cell KATP channel subunits Kir6. 2 (KCNJ11) and SUR1 (ABCC8) confirm that the KCNJ11 E23K variant is associated with type 2 diabetes. Diabetes. 2003;52(2):568–572. | ||
Cauchi S, El Achhab Y, Choquet H, et al. TCF7L2 is reproducibly associated with type 2 diabetes in various ethnic groups: a global meta-analysis. J Mol Med (Berl). 2007;85(7):777–782. | ||
Palizban A, Nikpour M, Salehi R, Maracy MR. Association of a common variant in TCF7L2 gene with type 2 diabetes mellitus in a Persian population. Clin Exp Med. 2012;12(2):115–119. | ||
Parra EJ, Cameron E, Simmonds L, et al. Association of TCF7L2 polymorphisms with type 2 diabetes in Mexico City. Clin Genet. 2007;71(4):359–366. | ||
Deriving genotypic and allelic frequencies [webpage on the Internet]. Population and Evolutionary Genetics; 1998. Available from: https://www.ndsu.edu/pubweb/~mcclean/plsc431/popgen/popgen2.htm. Accessed August 15, 2015. | ||
Groves CJ, Zeggini E, Minton J, et al. Association analysis of 6,736 UK subjects provides replication and confirms TCF7L2 as a type 2 diabetes susceptibility gene with a substantial effect on individual risk. Diabetes. 2006;55(9):2640–2644. | ||
Florez JC, Jablonski KA, Bayley N, et al. TCF7L2 polymorphisms and progression to diabetes in the Diabetes Prevention Program. N Engl J Med. 2006;355(3):241–250. | ||
Scott LJ, Bonnycastle LL, Willer CJ, et al. Association of transcription factor 7-like 2 (TCF7L2) variants with type 2 diabetes in a Finnish sample. Diabetes. 2006;55(9):2649–2653. | ||
Perez-Martinez P, Perez-Caballero AI, Garcia-Rios A, et al. Effects of rs7903146 variation in the Tcf7l2 gene in the lipid metabolism of three different populations. PloS One. 2012;7(8):e43390. | ||
Prunier C, Hocevar BA, Howe PH. Wnt signaling: physiology and pathology. Growth Factors. 2004;22(3):141–150. | ||
Yi F, Brubaker PL, Jin T. TCF-4 mediates cell type-specific regulation of proglucagon gene expression by β-catenin and glycogen synthase kinase-3β. J Biol Chem. 2005;280(2):1457–1464. | ||
Cauchi S, Meyre D, Dina C, et al. Transcription Factor TCF7L2 Genetic Study in the French Population Expression in Human β-Cells and Adipose Tissue and Strong Association With Type 2 Diabetes. Diabetes. 2006;55(10):2903–2908. | ||
Miyake K, Horikawa Y, Hara K, et al. Association of TCF7L2 polymorphisms with susceptibility to type 2 diabetes in 4,087 Japanese subjects. J Hum Genet. 2008;53(2):174–180. | ||
Dou H, Ma E, Yin L, Jin Y, Wang H. The association between gene polymorphism of TCF7L2 and type 2 diabetes in Chinese Han population: a meta-analysis. PloS One. 2013;8(3):e59495. | ||
Zhai Y, Zhao J, You H, et al. Association of the rs11196218 polymorphism in TCF7L2 with type 2 diabetes mellitus in Asian population. Meta Gene. 2014;2:332–341. | ||
Shokouhi S, Delpisheh A, Haghani K, Mahdizadeh M, Bakhtiyari S. Association of rs7903146, rs12255372, and rs290487 polymorphisms in TCF7L2 gene with type 2 diabetes in an Iranian Kurdish ethnic group. Clin Lab. 2013;60(8):1269–1276. | ||
Uma Jyothi K, Jayaraj M, Subburaj KS, et al. Association of TCF7L2 gene polymorphisms with T2DM in the population of Hyderabad, India. PloS One. 2013;8(4):e60212. | ||
Amoli MM, Amiri P, Tavakkoly-Bazzaz J, et al. Replication of TCF7L2 rs7903146 association with type 2 diabetes in an Iranian population. Genet Mol Biol. 2010;33(3):449–451. | ||
Holstein A, Hahn M, Körner A, Stumvoll M, Kovacs P. TCF7L2 and therapeutic response to sulfonylureas in patients with type 2 diabetes. BMC Med Genet. 2011;12:30. | ||
Pearson ER, Donnelly LA, Kimber C, et al. Variation in TCF7L2 Influences Therapeutic Response to Sulfonylureas A GoDARTs Study. Diabetes. 2007;56(8):2178–2182. | ||
Schroner Z, Javorsky M, Tkacova R, et al. Effect of sulphonylurea treatment on glycaemic control is related to TCF7L2 genotype in patients with type 2 diabetes. Diabetes Obes Metab. 2011;13(1):89–91. | ||
Gardiner SJ, Begg EJ. Pharmacogenetics, drug-metabolizing enzymes, and clinical practice. Pharmacol Rev. 2006;58(3):521–590. | ||
Ramon y Cajal T, Altes A, Pare L, et al. Impact of CYP2D6 polymorphisms in tamoxifen adjuvant breast cancer treatment. Breast Cancer Res Treat. 2010;119(1):33–38. | ||
Punglia RS, Burstein HJ, Winer EP, Weeks JC. Pharmacogenomic variation of CYP2D6 and the choice of optimal adjuvant endocrine therapy for postmenopausal breast cancer: a modeling analysis. J Natl Cancer Inst. 2008;100(9):642–648. | ||
Rodriguez-Antona C, Ingelman-Sundberg M. Cytochrome P450 pharmacogenetics and cancer. Oncogene. 2006;25(11):1679–1691. | ||
Melzer D, Murray A, Hurst AJ, et al. Effects of the diabetes linked TCF7L2 polymorphism in a representative older population. BMC Med. 2006;4:34. | ||
Moczulski D, Gawlik B, August R, Strojek K, Grzeszczak W. TCF7L2 gene is associated with type 2 diabetes in Polish population. Diabetologia Doświadczalna i Kliniczna. 2007;7(2):109–111. | ||
Gupta V, Khadgawat R, Ng HK, et al. A Validation Study of Type 2 Diabetes-related Variants of the TCF7L2, HHEX, KCNJ11, and ADIPOQ Genes in one Endogamous Ethnic Group of North India. Ann Hum Genet. 2010;74(4):361–368. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.