Back to Journals » Drug Design, Development and Therapy » Volume 9
PACE4 regulates apoptosis in human prostate cancer cells via endoplasmic reticulum stress and mitochondrial signaling pathways
Authors Yao Z, Sun B, Hong Q, Yan J, Mu D, Li J, Sheng H, Guo H
Received 18 April 2015
Accepted for publication 22 May 2015
Published 5 November 2015 Volume 2015:9 Pages 5911—5923
DOI https://doi.org/10.2147/DDDT.S86881
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Shu-Feng Zhou
Zhiyong Yao, Bin Sun, Quan Hong, Jingmin Yan, Dawei Mu, Jianye Li, Haibo Sheng, Heqing Guo
Department of Urology, Air Force General Hospital of People’s Liberation Army, Beijing, People’s Republic of China
Backgrounds: PACE4 is a proprotein convertase capable of processing numerous substrates involved in tumor growth, invasion, and metastasis. However, the precise role of PACE4 during prostate cancer cell apoptosis has not been reported.
Methods: In the present study, human prostate cancer cell lines-DU145, LNCaP, and PC3 were transfected with PACE4 small interfering (si)RNA to investigate the underlying mechanisms of apoptosis.
Results: We revealed that PACE4 siRNA exhibited antitumor activity by inducing apoptosis, as determined by Cell Counting Kit-8 (CCK-8), 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay, cell cycle analysis, Hoechst staining, caspase-3/7 activity, and western blot analysis. In addition, PACE4 siRNA significantly increased the ratio of Bax/Bcl-2, which led to the release of cytochrome c. Moreover, PACE4 siRNA also induced endoplasmic reticulum stress by increasing the expression of GRP78, GRP94, p-PERK, and p-eIF2α. The ratio of Bax/Bcl-2 and GRP78 were also increased in PACE4 gene knockdown prostate cancer cells compared with the control cells.
Conclusion: These data demonstrate that PACE4 siRNA may exert its antitumor activity through mitochondrial and endoplasmic reticulum stress signaling pathways, indicating it may be a novel therapeutic target for prostate cancer.
Keywords: tumor growth, small interfering RNA, GRP78, Bax/Bcl-2, gene knockdown
Introduction
Prostate cancer (PCa) is now the most commonly diagnosed cancer in men over 60 years of age, with high incidence rates also found in younger age groups, and is the second leading cause of cancer deaths behind lung cancer.1,2 Treatment for PCa is still unsatisfactory, with an almost inevitable development of hormone resistance.3 Even new-generation androgen ablation drugs fail to deliver a life extension beyond several months.4 In addition, PCa has shown poor response to chemotherapy, alongside unpleasant side effects and reduced quality of life.5 Some studies have shown that PCa development involves various molecular mechanisms, but the specific molecular regulatory pathways or markers for PCa remain poorly understood so far. Therefore, exploring the molecular mechanisms involved in the pathogenesis of PCa and identifying novel therapeutic targets or effective agents is very important. Repeated documents have reported that cell death signaling may be involved in the occurrence and development of PCa, suggesting cell death or proliferation may be the key pathway to treat PCa.6–8
The proprotein convertases (PCs) are a family of enzymes that is responsible for the activation of numerous protein precursors. So far, nine different PCs have been identified, namely, furin, PACE4, PC1/3, PC2, PC4, PC5/6, PC7, PCSK9, and SKI-1/S1P.9,10 In recent years, the PACE4 has been thought to play important roles in cancer development and progression. It activates several biologically relevant substrates, some of which have been shown to play significant roles in tissue homeostasis and cancer growth.11–13 Among these are numerous metalloproteinases, growth factors, growth factor receptors, and adhesion molecules directly associated with tumor development.14–16 PACE4 is expressed at low levels in many mammalian tissues and has been demonstrated to be upregulated in some tumor cell lines, including murine squamous cell carcinomas.17 In addition, mice overexpressing PACE4 have been shown to exhibit tumors of increased growth rate.18 Longuespée et al19 revealed that PACE4 could promote cell proliferation in ovarian cancer, providing further evidence for PACE4 as a potential therapeutic target.
Recently, two independent studies showed overexpression of PACE4 mRNA in PCa tissues.20,21 This overexpression was correlated with higher circulating protein levels in some patients.21 There are no data regarding the possible role of PACE4 in the prostate cancer cell apoptosis and the potential molecular mechanisms. In the current study, we used molecular silencing, with small interfering (si)RNA to knock down endogenously expressed PACE4 in three human PCa cell lines, DU145, LNCaP, and PC3 cells, and then, to test for cell proliferation and an apoptosis response. We found that PACE4 siRNA significantly increased apoptosis of these cells. The sum of our data confirms that PACE4 has an important role in PCa cell proliferation and further suggests that PACE4 is a potential therapeutic target.
Materials and methods
Reagents and antibodies
Rabbit anti-human cleaved caspase-3, Bcl-2, Bax, Akt, GRP78, GRP94, PERK, COXIV, XIAP, survivin antibodies, and mouse anti-GAPDH antibody were purchased from Proteintech (Wuhan, People’s Republic of China). Rabbit anti-human PACE4 and cytochrome c (cyto c) antibodies were purchased from Abcam (Cambridge, UK). Rabbit anti-human phosphor-Akt (p-Akt), phosphor-PERK (p-PERK), eIF2α, and phosphor-eIF2α (p-eIF2α) antibodies were purchased from Cell Signaling Technology, Inc (Danvers, MA, USA). Anti-rabbit or mouse immunoglobulin G-horseradish peroxidase (IgG-HRP) secondary antibodies were purchased from Proteintech. A Cell Counting Kit-8 (CCK-8) and Hoechst 33258 stain were purchased from Beyotime (Haimen, People’s Republic of China). Other reagents were of analytical grade.
Cell culture and RNA interference
Three human PCa cell lines, DU145, LNCaP, and PC3 cells, were obtained from American Type Culture Collection (Manassas, VA, USA). Cells were routinely grown in Roswell Park Memorial Institute (RPMI) 1640 medium (Hyclone®; Thermo Fisher Scientific Inc, Waltham, MA, USA) containing 5% fetal bovine serum (FBS) (Hyclone; Thermo Fisher Scientific Inc), 100 U/mL penicillin (Sigma-Aldrich Corp, St Louis, MO, USA), and 100 μg/mL streptomycin (Sigma-Aldrich Corp) in a humidified atmosphere of 5% CO2 at 37°C.
Cells were transfected with 100 nM PACE4 siRNA (Genbank ID for PACE4: NM_001291309) (sc-43990; Santa Cruz Biotechnology Inc, Dallas, TX, USA) or control siRNA (scrambled siRNA, a universal negative control), using GeneSilencer siRNA Transfection Reagent (Genlantis, San Diego, CA, USA), according to the manufacturer’s instructions. At 48 hours after transfection, the efficiency of siRNA-mediated PACE4 knockdown was determined by western blot.
Cell proliferation assay
Cells were seeded into 96-well plates (4×103 cells per well). After 24 hours of incubation, cells were transfected with PACE4 siRNA or control siRNA for 12, 24, 36, and 48 hours as described above, followed by the addition of 10 μL CCK-8 solution. The cells were then incubated for 4 hours at 37°C. The optical density for each well was measured at 450 nm with a microculture plate reader (Bio-Rad Laboratories, Hercules, CA, USA). The experiments were performed in triplicate.
MTT assay
Cells were cultured in 96-well culture plates and transiently transfected PACE4 siRNA and control siRNA. After transfection, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) was added to each well to a final concentration of 5 mg/mL in culture medium and incubated at 37°C for 4 hours. The reaction was terminated by removal of the supernatant and addition of 150 μL dimethyl sulfoxide (DMSO) to dissolve the formazan product. The plates were read at 405 nm on a microELISA plate reader (Thermo MK3; Thermo Fisher Scientific Inc). Each assay was performed at least three times.
Cell cycle analysis
Cells were seeded overnight on 60 mm-diameter plates with a complete medium, placed in a serum-free medium for 48 hours to synchronize the cells, and then kept again in the complete medium. At 24 hours, cells were recovered. After washing with ice-cold phosphate-buffered saline (PBS), cells were suspended in about 0.5 mL of 70% alcohol and kept at 4°C for 30 minutes. The suspension was filtered through a 50 mm nylon mesh, and the DNA content of stained nuclei was analyzed by a flow cytometer (EPICS XL; Coulter, Miami, FL, USA). Cell cycle was analyzed using Multicycle-DNA Cell Cycle Analyzed Software (FACScan; BD Biosciences, Franklin Lakes, NJ, USA).
Morphological analysis after Hoechst 33258 staining
Cells were seeded in 24-well plates (5×104 cells per well). After 24 hours of incubation, cells were transfected with PACE4 siRNA or control siRNA for 48 hours. Then the cells were fixed and stained with Hoechst 33258. The apoptotic cells were visualized with fluorescence microscope (Leica Microsystems, Wetzlar, Germany).
Detection of caspase-3/7 protein activity
Caspase-3/7 activity was measured using a colorimetric method following the manufacturer’s instructions (Caspase-Glo® 3/7 Assay kit; Promega Corp, Fitchburg, WI, USA). Briefly, 2×104 cells were seeded in 96-well plates and left for 24 hours and then, transfected with PACE4 siRNA or control siRNA for another 48 hours. Next, lysate of cells was mixed with equilibrated Caspase-glo 3/7 reagents for 1 hour at room temperature. Luminescence was measured using a GloMax 96 luminometer (Promega Corp).
Preparation of mitochondria and cytosol
A mitochondria/cytosol kit (Beyotime) was used to isolate mitochondria and cytosol, according to the manufacture’s protocol. After transfection as above, cells (2×107 cells) were collected by centrifugation at 800× g for 5 minutes at 4°C, washed twice with ice-cold PBS, and then resuspended in 500 μL of isolation buffer containing protease inhibitors for 10 minutes, on ice. The cells were mechanically homogenized using a Dunce grinder. The unbroken cells, debris, and nuclei were discarded by centrifugation at 800× g for 10 minutes at 4°C. The supernatants were centrifuged at 12,000× g for 20 minutes at 4°C. The supernatant cytosol was collected, and the pellet fraction of the mitochondria was dissolved in 50 μL of lysis buffer.
Western blotting
Cells were transfected as described above. Cells were lysed in RIPA buffer supplemented with protease inhibitors (Complete Mini; F. Hoffman-La Roche Ltd, Basel, Switzerland). Protein concentrations were measured using a BCA Protein Assay Kit (Dingguo, Beijing, People’s Republic of China), and 50 μg of protein samples were separated on a 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS–PAGE) and transferred to a polyvinylidene difluoride (PVDF) membrane (Merck Millipore, Billerica, MA, USA). Before immunodetection, membranes were blocked with 5% (wt/vol) bovine serum albumin (BSA) in a 0.1% Tween-PBS solution. Membranes were then incubated with indicated antibodies overnight at 4°C. After washing, blots were incubated for 1 hour with HRP-conjugated anti-rabbit or anti-mouse secondary antibodies. The blots were revealed using the ECL Plus detection system (Thermo Fisher Scientific Inc) under conditions recommended by the manufacturer. Images were captured directly using the Gel 3100 chemiluminescent and fluorescent imaging system (Sagecreation, Beijing, People’s Republic of China). Relative protein expression levels were calculated using the Quantity One software (Bio-Rad Laboratories), with normalization to the GAPDH signal.
Establishment of PACE4 gene knockdown human PCa cell lines
The human DU145, LNCaP, and PC3 cell lines were purchased from American Type Culture Collection. The PACE4 gene was knocked out from these three types of cells, respectively. The PACE4 gene knockdown cells were synthesized and purchased from Sangon Biotechnology, Int. (Shanghai, People’s Republic of China). The PACE4 gene knockdown cells were routinely grown in RPMI 1640 medium containing 5% FBS, 100 U/mL penicillin, and 100 μg/mL streptomycin in a humidified atmosphere of 5% CO2 at 37°C.
Statistical analysis
All data were presented as mean ± standard deviation and analyzed using Student’s t-test and one-way analysis of variance (ANOVA) analysis to determine the levels of significance. A P-value less than 0.05 or 0.01 was considered statistically significant. Statistical analysis was done with SPSS/Win11.0 software (SPSS Inc., Chicago, IL, USA).
Results
Effect of PACE4 on DU145, LNCaP, and PC3 cells proliferation
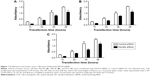
Cells proliferation was examined using CCK-8 following transfection with PACE4 siRNA or control siRNA. As shown in Figure 1, PACE4 siRNA decreased proliferation of DU145, LNCaP, and PC3 cells as compared with the control siRNA group. Furthermore, the cell proliferation at 48 hours after transfection with PACE4 siRNA was reduced significantly as compared with control siRNA.
Furthermore, the cell death was also examined using the MTT assay. The result indicated that PACE4 siRNA (at 36 and 48 hours) significantly increased the cell death rate of DU145, LNCaP, and PC3 cells compared with the control siRNA group (P<0.05) (Figure S1).
Thus, these data imply that PACE4 siRNA may influence cellular proliferation in human PCa cells.
NDRG2 induced the cell cycle arrest of PCa cells
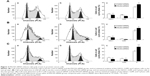
To further investigate the mechanisms by which PACE4 siRNA inhibits PCa cell growth, we studied the effects of PACE4 siRNA on the cell cycle using fluorescence-activated cell sorting analysis. The results of the cell cycle analysis showed that many more PACE4 siRNA-transfected cells were in S-phase compared with the control siRNA-transfected cells, whereas fewer PACE4 siRNA-transfected cells were in G1-phase compared with the control siRNA-transfected cells (for all of the DU145, LNCaP, and PC3 PCa cells) (P<0.05) (Figure 2).
PACE4 siRNA induces human PCa cells apoptosis
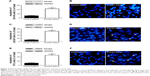
In order to assess whether the proliferation inhibition induced by PACE4 siRNA in these cells was associated with apoptosis, we examined the morphologic changes with Hoechst 33258 staining (Figure 3). DU145, LNCaP, and PC3 cells were transfected with PACE4 siRNA for 48 hours, and the apoptotic morphologic changes were observed as compared with the control group. In the control siRNA group, the nuclei of cells were round and homogeneously stained. However, PACE4 siRNA-transfected cells exhibited evident apoptosis characteristics, including cell shrinkage and membrane integrity loss or deformation, nuclear fragmentation, and chromatin compaction of late apoptotic appearance (Figure 3B, D, and F). The percentage of apoptotic cells was significantly increased in the PACE4 siRNA-transfected cells, as compared with the group transfected with the negative control siRNA (Figure 3A, C, and E). Together, these data indicate that PACE4 siRNA induces apoptosis in human PCa cells.
PACE4 siRNA induces apoptosis via the caspase-dependent pathway
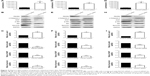
Caspase-3 is a critical executioner of apoptosis, and its activation is essential for DNA fragmentation and some of the typical biochemical and morphological changes of cells undergoing apoptosis. So, to evaluate whether or not PACE4 siRNA-induced apoptosis is involved with activation of caspase-3/7, we investigated the caspase-3/7 activity by measuring the bioluminescent intensities. The activity of caspase-3/7 was significantly activated after PACE4 siRNA transfection (Figure 4A, D, and G).
To further assess the role of PACE4 in apoptosis, we next evaluated expression of apoptosis-related proteins (Figure 4B, E and H). These include proapoptotic cleaved caspase-3, antiapoptotic XIAP, p-Akt, and survivin. Western blot results indicated that PACE4 siRNA increased the levels of cleaved caspase-3 by approximately 2.2-fold (P<0.05) in DU145 cells, 2.3-fold (P<0.05) in LNCaP cells, and by 3.8-fold (P<0.01) in PC3 cells. On the contrary, the levels of XIAP, p-Akt, and survivin in DU145, LNCaP, and PC3 cells were decreased appropriate 75% (P<0.01, XIAP in DU145), 60% (P<0.01, XIAP in LNCaP), 70% (P<0.05, XIAP in PC3), 50% (P<0.01, p-Akt in DU145), 65% (P<0.01, p-Akt in LNCaP), 75% (P<0.01, p-Akt in PC3), and 40% (P<0.01, survivin in DU145), 55% (P<0.01, survivin in LNCaP), 60% (P<0.05, survivin in PC3), respectively, after PACE4 siRNA transfection (Figure 4C, F, and I).
PACE4 siRNA induces apoptosis via the mitochondrial apoptotic pathway
In order to better understand the molecular mechanisms by which PACE4 siRNA exerts proapoptosis effects, we followed the protein expression of mediators in the mitochondrial signaling pathway. First, we determined whether PACE4 siRNA stimulated the release of cyto c into the cytosolic fraction in DU145, LNCaP, and PC3 cells. As expected, cyto c was redistributed after PACE4 siRNA transfection. The level of cyto c in the mitochondria of the DU145, LNCaP, and PC3 cells was significantly decreased, by 50% (P<0.05), 60% (P<0.01), and 58% (P<0.01), respectively. Correspondingly, the level of cyto c in cytosol was increased by 2.9-fold (P<0.05), 2.5-fold (P<0.01), and 2.4-fold (P<0.05), respectively (Figure 5).
Since the Bcl-2 family proteins play a critical role in regulating the release of cyto c, we then investigated the possible involvement of Bax and Bcl-2 in the process of PACE4 siRNA-mediated apoptosis in the cells. As shown in Figure 5, the level of Bax was significantly increased, and Bcl-2 was obviously decreased in PACE4 siRNA-transfected cells. Statistical analysis showed that PACE4 siRNA increased the ratio of Bax/Bcl-2 by approximately 5.2-fold (P<0.05), 1.2-fold (P<0.01), and 4.1-fold (P<0.01), respectively, in the DU145, LNCaP, and PC3 cells.
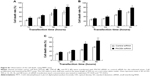
PACE4 siRNA induces activation of the endoplasmic reticulum (ER) stress pathway
There is little information on the effect of PACE4 on ER stress in cultured PCa cells. In order to determine whether ER stress was induced after PACE4 siRNA transfection, we assessed the protein levels of p-PERK and p-eIF2α, which are considered signature ER stress markers. As shown in Figure 6, compared with control siRNA, the expression of p-PERK and p-eIF2α was significantly increased in PACE4 siRNA-transfected cells, while total PERK and eIF2α were not changed. We next examined the expression of GRP78 andGRP94, which serve as gatekeeper to the activation of ER stress transducers. The data in Figure 6 demonstrates that the expression of GRP78 and GRP94 by PACE4 siRNA transfection dramatically increased, respectively, by 2.1-fold (P<0.05) and 3.8-fold (P<0.05) in DU145 cells, by 3.0-fold (P<0.05) and 2.4-fold (P<0.05) in LNCaP cells, and by 3.1-fold (P<0.01) and 1.9-fold (P<0.05) in PC3 cells. These results demonstrate that ER stress is partially involved in PACE4 siRNA-induced apoptosis.
Apoptosis occurs in PACE4 gene knockdown cells via mitochondrial apoptotic and ER stress pathway
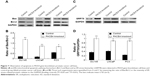
In order to investigate the effects of the PACE4 gene on the apoptosis of PCa cells, we investigated biomarkers of the mitochondrial apoptotic pathway (Bcal-2 and Bax) and a biomarker of the ER stress pathway (GRP78). The results indicated that the Bax/Bcl-2 ratio in PACE4 knockdown cells (DU145, LNCaP, and PC3 cells) was significantly increased compared with the control group (P<0.05) (Figure 7A and B). Furthermore, the GRP78 level was also significantly increased in PACE4 knockdown cells compared with the control group (P<0.05) (Figure 7C and D).
Discussion
At present, several surgical therapies and effective radiation can be offered for the clinical treatment of PCa; however, the therapies for PCa are far from satisfactory.22 Therefore, the search for new methods of cancer therapy and prognosis is a key task for many researchers. PACE4 has already been highlighted for its potential role in numerous neoplasias, such as oral and tongue carcinoma,23 hepatocellular carcinoma,24 glioma,25 skin cancer,26 and PCa.20 Whereas these studies mostly examined PACE4 overexpression, our present study focused on gene silencing as a predictive approach to define potential therapeutic benefits. Here, our results indicate that PACE4 siRNA inhibits the proliferation of DU145, LNCaP, and PC3 cells. Based on the results of Hoechst 33258 staining, caspase-3/7 activity, and western blotting, we conclude that PACE4 siRNA induces apoptosis in PCa cells. Thus, the present study constitutes the first evidence that PACE4 has an antiapoptotic effect in PCa cells.
As a primary executioner caspase in most systems, the activation of caspase-3 often results in the irreversible commitment of a cell to apoptosis. Therefore, the activation of caspase-3 is considered a reliable marker for cells undergoing apoptosis.27 We found that the activity of caspase-3/7 was significantly activated after PACE4 siRNA transfection (Figure 4A, D, and G). It is documented that a good strategy for killing cancer cells is to induce cell apoptosis. Members of the IAP family, survivin, and XIAP contribute to apoptosis resistance in cancer cells.28 Akt is a promoter of cell proliferation and survival and is found to be overexpressed in the tumor formation.29 Our investigation confirmed the role of PACE4 in the apoptosis of these cells, based on the following lines of evidence: PACE4 siRNA increased the apoptosis of cells by regulating the apoptosis-related factors cleaved caspase-3, XIAP, p-Akt, and survivin (Figure 4). The inactivation of XIAP, p-Akt, or survivin by PACE4 siRNA may prevent the development and progression of cancers.
It has been well-documented that the Bcl-2 family proteins function through different pathways in the regulation of cell apoptosis. The Bcl-2 family primarily affects the mitochondrial pathways.8 Bcl-2 and its homologs prevent mitochondrial membrane disruption and the release of cyto c, while Bax promotes these events. The ratio of Bax/Bcl-2 is usually regarded as a criterion for apoptosis.30 The result from this study demonstrated that the level of cyto c in mitochondria was significantly decreased and in cytosol, was increased (Figure 5). Meanwhile, PACE4 siRNA increased the levels of Bax and decreased the level of Bcl-2, leading to the changes of the ratio of Bax/Bcl-2 (Figure 5). The results indicate that PACE4 siRNA is able to influence mitochondrial membrane stability. This was evidenced by increased Bax/Bcl-2 ratio and the release of cyto c into the cytoplasm. Taken together, these data demonstrate that PACE4 siRNA may exert its antitumor activity through the mitochondrial signaling pathway in PCa cells.
ER stress is another pathway mediating apoptosis.31 Repeated documents have recorded that ER stress triggered apoptosis in some types of cells through the PERK-eIF2α signaling pathway.32,33 For example, dissociation of GRP78 from PERK, initiates transphosphorylation and subsequent activation of PERK during ER stress,34 and activated PERK phosphorylates eIF2α, which is essential for ER stress-induced apoptosis.35,36 The results from western blotting showed that PACE4 siRNA promotes the levels of GRP94, GRP78, p-PERK, and p-eIF2α, which are vital features for an unfolded protein response (UPR) and mean that PACE4 siRNA could induce apoptosis through the ER stress signaling pathway (Figure 6).
In recent years, gene therapy has become a hot topic in the study of a different diseases.37,38 Previous study has also indicated that silencing of the PACE4 gene could trigger the apoptosis of other cancer cells, such as, PCa, ovarian cancer, breast cancer, etc.39–41 However, the specific mechanism of the PACE4 regulation has not been elucidated. For PACE4 inhibition, pharmacological inhibition has also been investigated. Levesque et al42 used the analog AC-[dlEU]LLLRVK-Amba to inhibit PCa progression in vitro and in vivo; however, the side effect of the pharmacological treatment is that it may also damage the normal cells. Therefore, our study provided a novel and safe therapeutic strategy against PCa progression.
In conclusion, our results suggest that PACE4 siRNA possesses the activity of antiproliferation and apoptosis induction in human PCa cell lines-DU145, LNCaP, and PC3. PACE4 siRNA-induced apoptosis of cells might be mediated through the ER stress and mitochondrial pathway. Therefore, the PACE4 inhibitor might be used as a potential agent for treatment of PCa.
Disclosure
The authors report no conflicts of interest in this work.
References
Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64(1):9–29. | ||
Williamson SC, Hartley AE, Heer R. A review of tasquinimod in the treatment of advanced prostate cancer. Drug Des Devel Ther. 2013;7:167–174. | ||
Ansari J, Hussain SA, Zarkar A, Tanguay JS, Bliss J, Glaholm J. Docetaxel chemotherapy for metastatic hormone refractory prostate cancer as first-line palliative chemotherapy and subsequent re-treatment: Birmingham experience. Oncol Rep. 2008;20(4):891–896. | ||
de Bono JS, Logothetis CJ, Molina A, et al. Abiraterone and increased survival in metastatic prostate cancer. N Engl J Med. 2011;364(21):1995–2005. | ||
Southwest Oncology Group; Berry DL, Moinpour CM, Jiang CS, et al. Quality of life and pain in advanced stage prostate cancer: results of a Southwest Oncology Group randomized trial comparing docetaxel and estramustine to mitoxantrone and prednisone. J Clin Oncol. 2006;24(18):2828–2835. | ||
Johnstone RW, Ruefli AA, Lowe SW. Apoptosis: a link between cancer genetics and chemotherapy. Cell. 2002;108(2):153–164. | ||
Lowe SW, Cepero E, Evan G. Intrinsic tumour suppression. Nature. 2004;432(7015):307–315. | ||
Adams JM, Cory S. The Bcl-2 apoptotic switch in cancer development and therapy. Oncogene. 2007;26(9):1324–1337. | ||
Couture F, D’Anjou F, Day R. On the cutting edge of proprotein convertase pharmacology: from molecular concepts to clinical applications. Biomol Concepts. 2011;2(5):421–438. | ||
Seidah NG, Prat A. The biology and therapeutic targeting of the proprotein convertases. Nat Rev Drug Discov. 2012;11(5):367–383. | ||
Tsuji A, Hashimoto E, Ikoma T, et al. Inactivation of proprotein convertase, PACE4, by alpha1-antitrypsin Portland (alpha1-PDX), a blocker of proteolytic activation of bone morphogenetic protein during embryogenesis: evidence that PACE4 is able to form an SDS-stable acyl intermediate with alpha1-PDX. J Biochem. 1999;126(3):591–603. | ||
Yuasa K, Masuda T, Yoshikawa C, Nagahama M, Matsuda Y, Tsuji A. Subtilisin-like proprotein convertase PACE4 is required for skeletal muscle differentiation. J Biochem. 2009;146(3):407–415. | ||
Tsuji A, Sakurai K, Kiyokage E, et al. Secretory proprotein convertases PACE4 and PC6A are heparin-binding proteins which are localized in the extracellular matrix. Potential role of PACE4 in the activation of proproteins in the extracellular matrix. Biochim Biophys Acta. 2003;1645(1):95–104. | ||
Yana I, Weiss SJ. Regulation of membrane type-1 matrix metalloproteinase activation by proprotein convertases. Mol Biol Cell. 2000;11(7):2387–2401. | ||
Dubois CM, Blanchette F, Laprise MH, Leduc R, Grondin F, Seidah NG. Evidence that furin is an authentic transforming growth factor-beta1-converting enzyme. Am J Pathol. 2001;158(1):305–316. | ||
Sun GG, Lu YF, Zhang J, Hu WN. Filamin A regulates MMP-9 expression and suppresses prostate cancer cell migration and invasion. Tumour Biol. 2014;35(4):3819–3826. | ||
Hubbard FC, Goodrow TL, Liu SC, et al. Expression of PACE4 in chemically induced carcinomas is associated with spindle cell tumor conversion and increased invasive ability. Cancer Res. 1997;57(23):5226–5231. | ||
Kang S, Zhao Y, Hu K, et al. miR-124 exhibits antiproliferative and antiaggressive effects on prostate cancer cells through PACE4 pathway. Prostate. 2014;74(11):1095–1106. | ||
Longuespée R, Couture F, Levesque C, et al. Implications of proprotein convertases in ovarian cancer cell proliferation and tumor progression: Insights for PACE4 as a therapeutic target. Transl Oncol. Epub 2014 May 9. | ||
D’Anjou F, Routhier S, Perreault JP, et al. Molecular validation of PACE4 as a target in prostate cancer. Transl Oncol. 2011;4(3):157–172. | ||
Klee EW, Bondar OP, Goodmanson MK, et al. Candidate serum biomarkers for prostate adenocarcinoma identified by mRNA differences in prostate tissue and verified with protein measurements in tissue and blood. Clin Chem. 2012;58(3):599–609. | ||
Kuefer R, Hofer MD, Altug V, et al. Sodium butyrate and tributyrin induce in vivo growth inhibition and apoptosis in human prostate cancer. Br J Cancer. 2004;90(2):535–541. | ||
Estilo CL, O-charoenrat P, Talbot S, et al. Oral tongue cancer gene expression profiling: Identification of novel potential prognosticators by oligonucleotide microarray analysis. BMC Cancer. 2009;9:11. | ||
Kurokawa Y, Matoba R, Nakamori S, et al. PCR-array gene expression profiling of hepatocellular carcinoma. J Exp Clin Cancer Res. 2004;23(1):135–141. | ||
Delic S, Lottmann N, Jetschke K, Reifenberger G, Riemenschneider MJ. Identification and functional validation of CDH11, PCSK6 and SH3GL3 as novel glioma invasion-associated candidate genes. Neuropathol Appl Neurobiol. 2012;38(2):201–212. | ||
Mahloogi H, Bassi DE, Klein-Szanto AJ. Malignant conversion of non-tumorigenic murine skin keratinocytes overexpressing PACE4. Carcinogenesis. 2002;23(4):565–572. | ||
Tait SW, Green DR. Mitochondria and cell death: outer membrane permeabilization and beyond. Nat Rev Mol Cell Biol. 2010;11(9):621–632. | ||
LaCasse EC, Mahoney DJ, Cheung HH, Plenchette S, Baird S, Korneluk RG. IAP-targeted therapies for cancer. Oncogene. 2008;27(48):6252–6275. | ||
Serrano ML, Sánchez-Gómez M, Bravo MM, Yakar S, LeRoith D. Differential expression of IGF-I and insulin receptor isoforms in HPV positive and negative human cervical cancer cell lines. Horm Metab Res. 2008;40(10):661–667. | ||
Pettersson F, Dalgleish AG, Bissonnette RP, Colston KW. Retinoids cause apoptosis in pancreatic cancer cells via activation of RAR-gamma and altered expression of Bcl-2/Bax. Br J Cancer. 2002;87(5):555–561. | ||
Dufey E, Sepúlveda D, Rojas-Rivera D, Hetz C. Cellular mechanisms of endoplasmic reticulum stress signaling in health and disease. 1. An overview. Am J Physiol Cell Physiol. 2014;307(7):C582–C594. | ||
Duan Z, Zhao J, Fan X, et al. The PERK-eIF2α signaling pathway is involved in TCDD-induced ER stress in PC12 cells. Neurotoxicology. 2014;44:149–159. | ||
Cui WY, Liu Y, Zhu YQ, Song T, Wang QS. Propofol induces endoplasmic reticulum (ER) stress and apoptosis in lung cancer cell H460. Tumour Biol. 2014;35(6):5213–5217. | ||
Liu CY, Schröder M, Kaufman RJ. Ligand-independent dimerization activates the stress response kinases IRE1 and PERK in the lumen of the endoplasmic reticulum. J Biol Chem. 2000;275(32):24881–24885. | ||
Szegezdi E, Logue SE, Gorman AM, Samali A. Mediators of endoplasmic reticulum stress-induced apoptosis. EMBO Rep. 2006;7(9):880–885. | ||
Nordin N, Majid NA, Hashim NM, Rahman MA, Hassan Z, Ali HM. Liriodenine, an aporphine alkaloid from Enicosanthellum pulchrum, inhibits proliferation of human ovarian cancer cells through induction of apoptosis via the mitochondrial signaling pathway and blocking cell cycle progression. Drug Des Devel Ther. 2015;9:1437–1448. | ||
Rodríguez L, Villalobos X, Dakhel S, et al. Polypurine reverse Hoogsteen hairpins as a gene therapy tool against survivin in human prostate cancer PC3 cells in vitro and in vivo. Biochem Pharmacol. 2013;86(11):1541–1554. | ||
Swift SL, Rivera GC, Dussupt V, et al. Evaluating baculovirus as a vector for human prostate cancer gene therapy. PLoS One. 2013;8(6):e65557. | ||
Kang S, Zhao Y, Hu K, et al. miR-124 exhibits antiproliferative and antiaggressive effects on prostate cancer cells through PACE4 pathway. Prostate. 2014;74(11):1095–1106. | ||
Yuasa K, Suzue K, Nagahama M, Matsuda Y, Tsuji A. Transcriptional regulation of subtilisin-like proprotein convertase PACE4 by E2F: possible role of E2F-mediated upregulation of PACE4 in tumor progression. Gene. 2007;402(1–2):103–110. | ||
Lapierre M, Siegfried G, Scamuffa N, et al. Opposing function of the proprotein convertases furin and PACE4 on breast cancer cells’ malignant phenotypes: role of tissue inhibitors of metalloproteinase-1. Cancer Res. 2007;67(19):9030–9034. | ||
Levesque C, Couture F, Kwiatkowska A, et al. PACE4 inhibitors and their peptidomimetic analogs block prostate cancer tumor progression through quiescence induction, increased apoptosis and impaired neovascularisation. Oncotarget. 2015;6(6):3680–3693. |
Supplementary material
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.