Back to Journals » Cancer Management and Research » Volume 11
F factor plasmid-mediated Epstein-Barr virus genome introduction establishes an EBV positive NPC cell model
Authors Duan J, Yang Y, Wu Z, Lin S, Zhou C, Sheng G, Yang F, Bian L, Zhang X , Xiao S
Received 6 April 2019
Accepted for publication 23 July 2019
Published 5 August 2019 Volume 2019:11 Pages 7377—7389
DOI https://doi.org/10.2147/CMAR.S211372
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Yong Teng
Jingling Duan,1,2 Yang Yang,1,2 Zhen Wu,3 Shiang Lin,4 Chen Zhou,1,2 Guowen Sheng,1,2 Fan Yang,1,2 Lihui Bian,5 Xiaoling Zhang,6 Shengjun Xiao1
1Department of Pathology, The Second Affiliated Hospital, Guilin Medical University, Guilin 541199, People’s Republic of China; 2Graduate College, Guilin Medical University, Guilin 541199, People’s Republic of China; 3Xiangya School of Medicine, Central South University, Changsha 410083, People’s Republic of China; 4Department of Otorhinolaryngology, The Affiliated Hospital, Guangdong Medical University, Zhanjiang 524001, People’s Republic of China; 5Department of Pathology, The Affiliated Hospital, Hebei University, Baoding 071000, People’s Republic of China; 6Department of Physiology, Faculty of Basic Medical Science, Guilin Medical University, Guilin 541199, People’s Republic of China
Background: Most Epstein-Barr virus (EBV)-positive cells lose the EBV episomes upon prolonged propagation.
Purpose: The purposes of this study were to establish a simple cell model for nasopharyngeal carcinoma (NPC) research by introducing a plasmid with the EBV genome into NPC cells and then to investigate the resulting changes in malignant biological behaviour and NPC-associated signalling pathways.
Methods: HONE1 NPC cells were transfected with F-factor plasmids including the EBV genome (HONE1-EBV cells). Then cell proliferation, migration, cell cycle distribution and apoptosis were evaluated in vitro by using CCK8, transwell and flow cytometry assays respectively. EBV-encoded proteins and cell signal tranducting proteins were detected by western blot assays. EBV-encoded RNAs were detected by in situ hybridization. EBV particles were assayed by transmission electron microscope (TEM). The morphology of cells were detected by immunofluorescence assays for alpha-tubulin. 
Results: Latent membrane protein 1 (LMP1), latent membrane protein 2A (LMP2A), Epstein-Barr nuclear antigen 1 (EBNA1) and EBV-encoded small RNAs (EBERs) were successfully expressed in HONE1-EBV cells. No EBV particles were founded by TEM. Introduction of the EBV genome significantly promoted proliferation, cell cycle progression and migration and inhibited apoptosis in HONE1 cells. Immunofluorescence assays showed that the morphology of HONE1-EBV cells changed into spindle. Furthermore, EBV genome introduction significantly inhibited the JAK/STAT signalling pathway, while it activated the PI3K-AKT and NF-κB signalling pathways in HONE1 cells.
Conclusion: These findings suggest that F-factor plasmid-mediated EBV genome introduction was successful in constructing an EBV positive cell model, which showed deteriorated biological behavior and activated NPC-associated signalling pathways. This model can serve as a good tool for studying EBV in NPC, but the subtle differences in cancer-associated pathways must be considered.
Keywords: Epstein-Barr virus, cell cycle, signal transduction, oncogenesis
Background
Nasopharyngeal carcinoma (NPC) is an endemic cancer of the head and neck with a high incidence in Southern China, reaching a peak incidence of 20–30 cases per 100,000 individuals.1 NPC is caused by a combination of factors, including Epstein-Barr virus (EBV), environmental influences, and heredity.2,3 The EBV genome can be detected in practically all cancer cells in undifferentiated NPC.4
EBV is a human gamma herpes virus that primarily infects B lymphocytes and certain epithelial cells.5,6 Stable EBV infection and expression of latent EBV genes are postulated to drive the transformation of pre-invasive nasopharyngeal epithelial cells into cancer cells, which are then involved in NPC progression.7,8 In NPC cells, EBV often exhibits latency II infection, during which the viral genes EBNA1, LMP1, LMP2A, LMP2B, EBERs, BamHI-A rightward transcripts (BART) and BART miRNAs are expressed.9 The expressed products of the EBV genome, including oncoproteins and EBERs, can activate multiple cancer-related signalling pathways.10,11 In particular, LMP1 activates multiple cancer-related pathways (JAK/STAT, PI3K/AKT and NF-κB),12–14 which promote the growth and survival of NPC cells, ultimately facilitating the development and progression of NPC.15–17 The EBV genome exists as episomes in NPC cells. Upon infection of new host cells, the linearized EBV genome undergoes circularization to form episomes, thereby maintaining latency state.18 However, EBV episomes are readily lost in NPC cells propagated in culture using conventional media. At present, only one EBV positive NPC cell line (C666-1, established in Hong Kong) is currently available for NPC investigations.19,20 Biological and pharmacological investigations of NPC have long been hampered by the lack of genuinely representative cell lines.21
In view of this deficiency, EBV infection strategy were used to construct EBV-infected nasopharyngeal epithelial cell model, and stable infection of EBV was achieved in a telomerase-immortalized nasopharyngeal epithelial cell line.22 Here, we sought to establish an EBV positive cell model by transfecting NPC cells with F-factor plasmids carrying the EBV genome, which can epxress some of the latent products. After introduction of the EBV genome, the HONE1-EBV cells successfully expressed LMP1, LMP2A, EBNA1 and EBERs and showed progressive malignant biological behaviours with activated cancer-related signalling pathways.
Materials and methods
Cell lines and tissue culture
The human HONE1 NPC cell line was obtained from the Cancer Center of Southern Medical University (Guangzhou, China), cultured in RPMI-1640 supplemented with 10% (v/v) fetal bovine serum (GIBCO, 10270, South America), 1% (v/v) 10 kU/mL penicillin and 10 mg/mL streptomycin (Hete biological, Xi’an, China) in a humidified atmosphere containing 5% (v/v) CO2 at 37 °C. All experiments were performed using cells in the logarithmic phase.
Plasmid construction and transient transfection
The plasmid p2089 (Maxi-EBV) containing the complete wild-type EBV genome (EBV strain: B95-8) cloned into a F-factor plasmid was kindly provided by Professor Dr. W. Hammerschmidt, and it also carries the green fluorescence protein (GFP) and the hygromycin resistance gene (hyg) for selection in cell culture.23 The plasmid p2089 of F-factor was transfected in parallel as a control following the instructions using Lipofectamine™ 3000 (ThermoFisher, Carlsbad, USA). After transfection, HONE1-Vector (transfected with plasmid Vector) and HONE1-EBV (transfected with plasmid p2089 carrying EBV genome), cells were selected by hygromycin (Calbiochem, 100 μg/mL) for one week and then collected for subsequent assays. After hygromycin selection, most of all cells were successfully introduced with F plasmids (Figure 1A) and then submitted to subsequent assays. HONE1-vector cells and HONE1-EBV cells were authenticated via short Tandem Repeat (STR) profiling by the Shanghai Biowing Applied Biotechnology Company (Figure S1).
Electron microscopy detection
After transfection 48 h, the cells of HONE1-vector and HONE-EBV were fixed in 2.5% glutaraldehyde for 2 h, washed three times by Phosphate buffer saline (PBS) and then treated with 1% osmium tetraoxide in the same buffer at 4 °C for 2 h. The samples were dehydrated through an ethanol series and embedded in epoxy resin. Ultrathin sections were stained with uranyl actetate followed by lead citrate, and examined in a Tecnai G220 TWIN transmission electron microscope at an accelerating voltage of 200 kV(FEI company, Hillsboro, USA).
Cell proliferation assay
NPC cells in the logarithmic phase were seeded into 96-well plates at a density of 2,000 cells per well in 100 µL of RPMI-1640 medium and cultured at 37 °C overnight. Plasmids were transfected by Lipofectamine™ 3000. After transfection at 24, 48, 72 and 96 h, 10 µL of CCK8 (Dojindo, Shanghai, China) was added and incubated respectively for 4 h at 37 °C before detecting optical densities (ODs). ODs of the resultant purple solutions were measured at 450 nm.
Wound-healing assay
After transfection, the same number of cells (4×106/well) in two groups were seeded into in a 6-well plate. After being attached overnight, the cells were scratched with a 1 mL pipette tip, washed twice with PBS, and further incubated in serum-free RPMI-1640. Images were taken at 0, 6, 18 and 24 h with a Leica light microscope (DM4B, Leica Corporation, Germany). The Image-Pro plus image analysis system was used to measure the migration area.
Transwell migration assay
In vitro cell migration assays were examined by transwell system (Corning, 3422, USA). After transfection, the same number of cells (2×106/well) in two groups were seeded on a polycarbonate membrane insert and incubated. RPMI-1640 without anything was supplemented in the upper chamber in the traswell assay. RPMI-1640 with 10% (v/v) fetal bovine serum were supplemented in the lower chamber in the transwell assay. After 24 h, cells migrated into the lower surface were fixed with 4% paraformaldehyde for 15 min, dried out at a room temperature, and then stained with 0.1% crystal violet, seen under a microscope (Olympus, IXTIFL, Japan).
Cell cycle assay
Cells (4×106/well) were seeded into T25 cell culture bottle. After transfection, the cells were incubated in medium with 10% FBS. The cells were cultured for 24 h, 36 h and 48 h respectively. Next, they were collected and fixed in 1 mL of cold 70% alcohol for 2 h at 4 °C. These fixed cells were collected by centrifuging at 1000×g for 5 min, washed with 1 mL of cold PBS and then resuspended in 500 µL, and incubated in propidium iodide (PI) dyeing buffer (mixing 500 µL dyeing buffer, 10 µL RNase A and 12.5 µL PI of solution) for 30 min at 37 °C in the dark. Finally, cells were added to each tube. The samples were immediately examined with red fluorescence at a wavelength of 488 nm. Light scattering was detected at the same time. The cellular DNA content and light scattering were analysed using the BD FACSCalibur flow cytometer (BD Biosciences, Franklin Lakes, NJ, USA).
Cell apoptosis assay
Cell apoptosis was assessed by measuring the membrane redistribution of phosphatidylserine using PE Annexin V Apoptosis Detection Kit l (BD Biosciences, Franklin Lakes, NJ, USA), according to the manufacturer’s instructions. Briefly, cells (4×106/well) were seeded into a T25 cell culture bottle. After transfection, the cells were cultured for 36 h. Subsequently, the cells were in the logarithmic phase, washed twice with cold PBS, resuspended in 1×Binding Buffer at a concentration of 1×106 cells/mL, and then 100 µL of the solution (1×105 cells) was transferred to a 2 mL culture tube. Finally, cells were stained with 5 µL of PE and 5 µL of 7-AAD, gently vortexed and incubated for 15 min at RT in the dark. Then, the cells were filtered with a filter screen and analysed using the BD FACSCalibur flow cytometer (BD Biosciences, Franklin Lakes, NJ, USA). The cells in Q2 (late stage apoptosis) and Q4 (early stage apoptosis) area were all calculated as apoptotic cells.
Immunofluorescence assay
After transfection for 36 h in 24-well plates (2×105 cells), cells were harvested and washed three times with PBS for 3 min; slides were fixed with 4% paraformaldehyde for 15 min, cells were washed three times with PBST for 3 min, and cells were covered with 0.2% Triton X-100 for 10 min. cells were washed three times with PBST for 3 min. After 1 h of blocking in PBS +20% fetal bovine serum, cells were incubated with primary antibodies of α -tubulin (Santa Cruz, sc-23948, USA; dilution, 1:100) for 2 h at room temperature. Then, cells were washed three times with PBST for 3 min with fluorescein-conjugated goat anti-mouse IgG(H+L) (ZSGB-BIO, ZF-0312, China; dilution, 1:100) for 1 h at room temperature. Then, cells were washed three times with PBST for 3 min. After counterstaining with 1 μg/mL DAPI (Solarbio, coo65, China) for 5 min, cells were washed three times with PBS for 3 min and ten times using distilled water. Cells were then air-dried. Slides were mounted with FluoromountTM Aqueous Mounting Medium (Sigma, Darmstadt, Germany). Then observed and photographed using a fluorescence microscope (Leica DM3000, Leica Microsystems Inc, Wetzlar, Germany).
In situ hybridization (ISH) assay
The oligonucleotide probe of EBERs was digoxigenin-labelled at the 3ʹ terminus. Fresh cell specimens were seeded onto coverslips, washed three times gently and fixed for 10 min in 10% PBS-buffered formaldehyde. EBV detection was performed by using an enhanced sensitive in situ hybridization (ISH) detection kit (ZSGB-BIO, ISH-6022, China) according to the manufacturer’s instructions. Briefly, after dehydtated by 100% alcohol and air-dried, the cells were digested by 20 µg/mL proteinase K for 10 min. Then incubated with the Digoxin-congugated probe for 3 h and followed with HRP-conjugated anti-Digoxin antibody (ready-to-use) for 30 min at 37 °C. Finnally, the cells were colorated with DAB for 5 min and hydrated and mounted with neutral resin. Slides were observed and photographed using a general optical microscope (Leica DM2500, Leica Microsystems Inc, Wetzlar, Germany).
Western blot analysis
Cell lysates were collected by scraping the cells with RIPA lysis buffer (Solarbio, Beijing, China) from the culture bottle, and the protein concentrations were determined using the DC Protein Assay Kit (Beyotime Biotechnology, Shanghai, China) according to the manufacturer’s protocol. An equal amount of protein lysate per sample was resolved on 8% or 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) (using 80 V constant voltage electrophoresis for 30 min, and then 120 V constant voltage electrophoresis for 90 min) and transferred onto a polyvinylidene fluoride (PVDF) membrane (Solarbio, Beijing, China) (wet transfer 260 mA for 1.5 h). Incubated with primary antibodies overnight at 4 °C in Primary Antibodies Dilution Buffer (Beyotime, P0023A, Shanghai, China) (Table S1), membranes were probed with desired primary antibodies followed by the detection of chemiluminescent signals of the peroxidase-conjugated secondary antibody using eECL Western Blot Kit detection system (Cwbiotech, cm0049m, Beiijing, China). Secondary antibodies were incubated for 1 h at room temperature. β-actin was used as an internal control to verify basal expression levels and equal protein loading. The ratio of the specific proteins to β-actin was calculated.
Statistical analysis
Statistical analysis of the data was carried out using the one-way ANOVA test (among multiple groups) and LSD and Bonferroni tests (between groups) by software SPSS 20.0 and GraphPad Prism (version 5; GraphPad Software, Inc., La Jolla, CA, USA). The data are presented as the mean ± SEM. Values are significant at *P<0.05, **P<0.01 and #P<0.001.
Results
EBV genome transfection resulted in successful expression of EBV-encoded products in HONE1 cells, while no virus particles produced
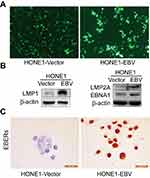
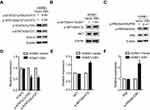
To validate whether the EBV genome was successfully introduced into HONE1 cells, the presence of LMP1, LMP2A and EBNA1 proteins were confirmed using WB and EBERs was ISH assay after transfection. HONE1-vector and HONE1-EBV cells were observed green fluorescent protein (GFP) under fluorescence microscope (Figure 1A). LMP1, LMP2A,EBNA1 and EBERs were all highly expressed in HONE1-EBV cells after transfection (Figure 1B and C). Similar to the phenotype observed in NPC cells, HONE1-EBVcells expressed two essential type II EBV latency products. Meanwhile, transmission electron microscopy showed no virus particles in HONE1-vector and HONE1-EBV cells (Figure S2). These data implied that introdution of EBV genome by F plasmid successfully simulated an latency of EBV in HONE1 cells, which partially expressed products of type II latent infection with no virus particle produced.
EBV genome introduction promoted significant proliferation and accelerated cell cycle progression in HONE1 cells
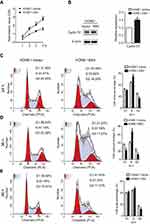
To observe the phenotypes of EBV infected NPC cells, the CCK8 method and flow cytometry were used to measure cell proliferation. The OD values of HONE1-EBV cells were clearly increased compared to those of HONE1-vector cells (Figure 2A). This result demonstrated that EBV infection enhanced the proliferation of NPC cells. Furthermore, EBV genes are involved in the regulation of the cell cycle-related protein cyclin D1. Introduction of the EBV genome increased the protein levels of cyclin D1 in NPC cells (Figure 2B). As shown in Figure 2C–E, flow cytometric analysis showed that the G1 to S phase transitions were significantly accelerated in HONE1-EBV cells compared with those in HONE1-vector cells at 24, 36 and 48 h. Taken together, these data indicated that the introduction of the EBV genome in NPC cells promotes cell proliferation by accelerating the transition from G1 phase to S phase.
EBV genome introduction promotes migration in HONE1 cells
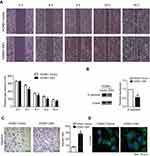
To evaluate the effect of EBV genome introduction on cell migration, wound-healing and transwell assays were employed to detect the mobility of HONE1 cells. The results showed that HONE1-EBV cells had significantly higher motility (Figure 3A and C). Meanwhile, E-cadherin was also down-regulated in HONE1-EBV cells (Figure 3B). In addition, an immunofluorescence assay revealed that the cellular morphology of HONE1-EBV cells changed from round or polygonal to spindle shaped (Figure 3D). The data above suggest that the introduction of the EBV genome enhanced cell migration in HONE1 cells.
EBV genome introduction significantly inhibited apoptosis in HONE1 cells
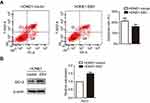
Next, in order to further explore the effect of EBV on apoptosis. The results of the 7-AAD and PI double staining assay showed that the apoptosis rate of the HONE1-EBV cells was decreased, and this effect was significant (Figure 4A). Furthermore, the expression of the anti-apoptotic protein Bcl-2 was upregulated in HONE1-EBV cells (Figure 4B). These results indicate that EBV genome introduction inhibited apoptosis in NPC cells.
EBV genome introduction significantly regulates the JAK/STAT, PI3K/AKT, and NF-κB signalling pathways in HONE1 cells
In this study, the expression levels of proteins associated with signalling pathway proteins were assessed. The JAK/STAT signalling pathway was impaired in HONE1-EBV cells. There was no statistically significant change in the expression level of the STAT3 protein, but the expression levels of the phosphorylated forms, p-STAT3(Tyr705) and p-STAT3(Ser727), were downregulated after EBV genome introduction in the NPC cell line (Figure 5A and D). This effect is different from NPC cells with EBV infection. This result requires additional attention in future studies. Both the PI3K/AKT and NF-κB signalling pathways were activated following EBV genome introduction into the NPC cells. The expression level of the p-AKT(Ser423) protein was upregulated in HONE1-EBV cells, and the expression level of AKT protein showed no statistically significant change (Figure 5B and E). Furthermore, while the level of the p-P65(Ser536) protein, part of the NF-κB signalling pathway, was upregulated in the HONE1-EBV cells, the expression level of total p65 showed no difference between HONE1-vector and HONE1-EBV cells (Figure 5C and F). Thus, these findings demonstrated that the introduction of the EBV genome negatively regulated the JAK/STAT pathways and positively regulated the NF-κB and PI3K/AKT pathway in NPC cells. Research on the major factors affecting the biological behaviour of EBV and how it affects disease progression may support the development of targeted interventions that could improve prognosis.
Discussion
EBV infection has been postulated to play an important role in NPC development and progression by inducing multiple signal transducers and activators that regulate transcription.24 However, the mechanism underlying EBV’s role in NPC development and progression is still elusive. EBV infected NPC cells are not ideal for in vitro studies of the relationship between EBV and NPC, as most NPC cells (except for the C666-1)19 frequently lose the EBV episomes upon prolonged propagation. In this study, we established a simple EBV positive NPC cell model using F-factor plasmid-mediated EBV genome introduction. F-factor plasmids exist in cells as episomes, which mimic EBV latency in NPC cells. After transfection with the EBV-containing plasmids, NPC HONE1 cells successfully expressed LMP1, LMP2A, EBNA1 and EBERs, which is characteristic of type II latent EBV infection in NPC.2 However, no virus particles produced in this model, which implied it’s defective to package lytic infectious virus.
Although C666-1 is a perfect cell model for NPC research, C666 cell represents only an undifferentiated NPC subtype.25 So more representive EBV-positive cell lines with different histological subtypes still need to be constructed. When compared to HK1-EBV cell, a recombinant EBV-infected of HK1 cell,26 F-plasmid transfection can provide a simple model for NPC research. In view of that C666-1 and HK1-EBV expresses a type II latency profile characterized by the expression of EBNA1, EBER1/2, LMP1, LMP2A, LMP2B, and BART transcripcts, in this study the expression of BART transcripts and other EBV genome coding products still need to be investigated. These results revealed that this method was effective for introducing the EBV genome into the NPC cells and that some of the latent virus-encoded genes were expressed. Using this approach, additional EBV positive NPC cell models can be successfully established by EBV genome introducing.
EBV latent infection constitutively activates several cancer associated signal pathway and then involved in aggressive biological behavior.27,28 To invesitgate whether these effects could be induced by EBV genome introducing in the NPC cells, several fundamental biological behavior and multiple signalling pathways associated proteins were assessed.
The results showed that the exogenous EBV genome promoted cell proliferation, accelerated the transition from G1 to S phase and enhanced the expression of cyclin D1, a key cell cycle regulator. Wound healing and transwell assays detection showed that the introduction of the EBV genome into NPC cells promoted cells movement. The cell migration capacity was clearly enhanced in HONE1-EBV cells. Meanwhile, expression of E-cadherin, an epithelial adhesion molecule, which frequently down-regulated in multiple malignancies, decreased.29 Furthermore, apoptosis was inhibited, and the anti apoptotic protein Bcl-2 was up-regulated in the HONE1-EBV cells. These datas implied that the expression of exogenous EBV genes, such as LMP1 and EBERs, which function are well known for their roles in NPC progression,30,31 promotes cell survival and migration. The biological characters changes confirmed this cell model is an appropriated model for NPC research. The close association between EBV and NPC had been well elucidated, as the EBV genome is present in virtually all NPC cells.2,28 Several cancers associated signalling pathways (JAK/STAT, NF-κB, PI3K/AKT, AP-1, and ERK-MAPK) were constitutively activated by latent associated proteins LMP1 and also EBERs.12–14,32 However, unlike the activated JAK/STAT3 signalling in C666-126 and HK1-EBV cells,27 exogenous EBV genes suppressed the JAK/STAT3 signalling pathway in this cell model, which was activated in EBV positive NPC cells.33 Constitutive STAT3 activation attributes to LMP1 expression in NPC cells and STAT3 is a candidate therapeutic target in NPC.34,35 Activated JAK/STAT3 signaling showed anti-apoptotic, promoting proliferative and migrative effects.17 The causative factors lead to JAK/STAT3 inhibition still need to be identified in this cell model.
While the (PI3K)-AKT and NF-κB signalling pathways, which are all constitutively activated in EBV-positive NPC cells and clinical tissue samples,36,37 were activated in HONE1 cells with exogenous EBV genome. Activation of the PI3K/AKT and NF-κB signalling pathways were fully accepted to affect many cell biological processes, including enhancement of protein synthesis, cell growth and proliferation.12,36 Taken together, the results suggested that the artificial introducing EBV genome into NPC cells may serve as a good tool for studying EBV in NPC, but the subtle differences in cancer-associated pathways must be considered. Despite of the EBV associated the HONE1-EBV cell model, which mimicking of latent EBV infection, may also have potential applicating value in NPC studies.
Conclusion
The research provided a new simple cellular model for the study of the relationship between EBV and NPC. EBV genes expression in NPC cells introduced with EBV genome by F plasmid promoted proliferation, promoted cell cycle progression and migration, and inhibited apoptosis. NPC associated signaling pathway activated except for JAK/STAT3 pathway, which differ in EBV positive NPC. This cell model provided an alternative model for NPV research.
Acknowledgments
The authors thank Prof. Dong Xiao (School of Life Sciences, Southern Medical University, Guangzhou) for donating the HONE1 cell, Pei Zhang and An-na Du (the Core Facility and Technical Support, Wuhan Institute of Virology, Wuhan) for their help with producing TEM micrographs. This work is supported by grants from the National Natural Science Foundation of China (Grant No. 81560441 and 81760491, to SJ Xiao) and the Natural Science Foundation of Guangxi Province of China (Grant No. 2015GXNSFAA139131, to SJ Xiao).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Seto E, Ooka T, Middeldorp J, Takada K. Reconstitution of nasopharyngeal carcinoma-type EBV infection induces tumorigenicity. Cancer Res. 2008;68:1030–1036. doi:10.1158/0008-5472.CAN-07-5252
2. Tsao SW, Tsang CM, Lo KW. Epstein-Barr virus infection and nasopharyngeal carcinoma. Philos Trans R Soc Lond B Biol Sci. 2017;372:20160270. doi:10.1098/rstb.2016.0270
3. Lo KW, Chung GT, To KF. Deciphering the molecular genetic basis of NPC through molecular, cytogenetic, and epigenetic approaches. Semin Cancer Biol. 2012;22:79–86. doi:10.1016/j.semcancer.2011.12.011
4. Nakanishi Y, Wakisaka N, Kondo S, et al. Progression of understanding for the role of Epstein-Barr virus and management of nasopharyngeal carcinoma. Cancer Metastasis Rev. 2017;36:435–447. doi:10.1007/s10555-017-9693-x
5. Shair KH, Schnegg CI, Raab-Traub N. Epstein-Barr virus latent membrane protein-1 effects on junctional plakoglobin and induction of a cadherin switch. Cancer Res. 2009;69:5734–5742. doi:10.1158/0008-5472.CAN-09-0468
6. Petersson F. Nasopharyngeal carcinoma: a review. Semin Diagn Pathol. 2015;32:54–73. doi:10.1053/j.semdp.2015.02.021
7. Husaini R, Ahmad M, Soo-Beng Khoo A. Epstein-Barr virus Latent Membrane Protein LMP1 reduces p53 protein levels independent of the PI3K-Akt pathway. BMC Res Notes. 2011;4:551–661. doi:10.1186/1756-0500-4-551
8. Correia S, Palser A, Elgueta Karstegl C, et al. Natural variation of Epstein-Barr virus genes, proteins, and primary MicroRNA. J Virol. 2017;91:e00375–e00317. doi:10.1128/JVI.00955-17
9. Choi SJ, Jung SW, Huh S, et al. Phylogenetic comparison of Epstein-Barr virus genomes. J Microbiol. 2018;56:525–533. doi:10.1007/s12275-018-8039-x
10. Turunen Aaro1 RJ, Reidar G. Epstein-Barr virus (EBV)-encoded small RNAs (EBERs) associated with poor prognosis of head and neck carcinomas. oncotarget. 2017;8:27328–27338. doi:10.18632/oncotarget.16033
11. Zhang L, Hong K, Zhang J, et al. Multiple signal transducers and activators of transcription are induced by EBV LMP-1. Virology. 2004;323:141–152. doi:10.1016/j.virol.2004.03.007
12. Liu H, Zheng H, Duan Z, et al. LMP1-augmented kappa intron enhancer activity contributes to upregulation expression of Ig kappa light chain via NF-kappaB and AP-1 pathways in nasopharyngeal carcinoma cells. Mol Cancer. 2009;8:92–100. doi:10.1186/1476-4598-8-92
13. Ting CM, Wong CK, Wong RN, et al. Role of STAT3/5 and Bcl-2/xL in 2-methoxyestradiol-induced endoreduplication of nasopharyngeal carcinoma cells. Mol Carcinog. 2012;51:963–972. doi:10.1002/mc.20867
14. Li SS, Yang S, Wang S, et al. Latent membrane protein 1 mediates the resistance of nasopharyngeal carcinoma cells to TRAIL-induced apoptosis by activation of the PI3K/Akt signaling pathway. Oncol Rep. 2011;26:1573–1579. doi:10.3892/or.2011.1423
15. Chung GT, Lou WP, Chow C, et al. Constitutive activation of distinct NF-kappaB signals in EBV-associated nasopharyngeal carcinoma. J Pathol. 2013;231:311–322. doi:10.1002/path.4239
16. Hu Q, Lin X, Ding L, et al. ARHGAP42 promotes cell migration and invasion involving PI3K/Akt signaling pathway in nasopharyngeal carcinoma. Cancer Med. 2018;7:3862–3874. doi:10.1002/cam4.1552
17. Wang Z, Luo F, Li L, et al. STAT3 activation induced by Epstein-Barr virus latent membrane protein1 causes vascular endothelial growth factor expression and cellular invasiveness via JAK3 And ERK signaling. Eur J Cancer. 2010;46:2996–3006. doi:10.1016/j.ejca.2010.07.008
18. Lu JH, Tang YL, Yu HB, et al. Epstein-Barr virus facilitates the malignant potential of immortalized epithelial cells: from latent genome to viral production and maintenance. Lab Invest. 2010;90:196–209. doi:10.1038/labinvest.2009.130
19. Yip YL, Lin W, Deng W, et al. Establishment of a nasopharyngeal carcinoma cell line capable of undergoing lytic Epstein-Barr virus reactivation. Lab Invest. 2018;98:1093–1104. doi:10.1038/s41374-018-0034-7
20. Xiao K, Yu Z, Li X, et al. Genome-wide Analysis of Epstein-Barr Virus (EBV) integration and strain in C666-1 and Raji cells. J Cancer. 2016;7:214–224. doi:10.7150/jca.13150
21. Ngan HL, Wang L, Lo KW, et al. Genomic landscapes of EBV-associated nasopharyngeal carcinoma vs. HPV-associated head and neck cancer. Cancers. 2018;10:210–230. doi:10.3390/cancers10070210
22. Tsang CM, Zhang G, Seto E, et al. Epstein-Barr virus infection in immortalized nasopharyngeal epithelial cells: regulation of infection and phenotypic characterization. Int J Cancer. 2010;127:1570–1583. doi:10.1002/ijc.25173
23. Mo AJ, Kurzeder C, Mautner J, et al. Infectious Epstein-Barr virus lacking major glycoprotein BLLF1 (gp350/220) demonstrates the existence of additional viral ligands. J Virol. 2000;74:10142–10152. doi:10.1128/jvi.74.21.10142-10152.2000
24. Elgui de Oliveira D, Muller-Coan BG, Pagano JS. Viral carcinogenesis beyond malignant transformation: EBV in the progression of human cancers. Trends Microbiol. 2016;24:649–664. doi:10.1016/j.tim.2016.03.008
25. Cheung ST, Huang DP, Hui AB, et al. Nasopharyngeal carcinoma cell line (C666-1) consistently harbouring Epstein-Barr virus. Int J Cancer. 1999;83(1):121–126. doi:10.1002/(SICI)1097-0215(19990924)83:1<121::AID-IJC21>3.0.CO;2-F
26. Lo AK, Lo KW, Tsao SW, et al. Epstein-Barr virus infection alters cellular signal cascadesin human nasopharyngeal epithelial cells. Neoplasia. 2006;8(3):173–180. doi:10.1593/neo.05625
27. Lo KW, To KF, Huang DP. Focus on nasopharyngeal carcinoma. Cancer Cell. 2004;5:423–428.
28. Tsang CM, Tsao SW. The role of Epstein-Barr virus infection in the pathogenesis of nasopharyngeal carcinoma. Virol Sin. 2015;30(2):107–121. doi:10.1007/s12250-015-3592-5
29. Wu C, Peng S, Sun W, et al. Association of E-cadherin methylation with risk of nasopharyngeal cancer: a meta-analysis. Head Neck. 2018;40:2538–2545. doi:10.1002/hed.25319
30. Tsao SW, Tramoutanis G, Dawson CW, et al. The significance of LMP1 expression in nasopharyngeal carcinoma. Semin Cancer Biol. 2002;12(6):473–487.
31. Pathmanathan R, Prasad U, Sadler R, et al. Clonal proliferations of cells infected with Epstein–barr virus in preinvasive lesions related to nasopharyngeal carcinoma. N Engl J Med. 1995;333:693–698. doi:10.1056/NEJM199509143331103
32. Morris MA, Laverick L, Wei W, et al. The EBV-encoded oncoprotein, LMP1, induces an Epithelial-to-Mesenchymal Transition (EMT) via its CTAR1 domain through integrin-mediated ERK-MAPK Signalling. Cancers. 2018;10:130–156. doi:10.3390/cancers10050130
33. Chen H, Hutt-Fletcher L, Cao L, et al. A positive autoregulatory loop of LMP1 expression and STAT activation in epithelial cells latently infected with Epstein-Barr virus. J Virol. 2003;77:4139–4148. doi:10.1128/jvi.77.7.4139-4148.2003
34. Leeman RJ, Lui VW, Grandis JR. STAT3 as a therapeutic target in head and neck cancer. Expert Opin Biol Ther. 2006;6:1471–2598. doi:10.1517/14712598.6.3.231
35. Tan YN, Tao YG, Song X, et al. Expression of JAK3 in nasopharyngeal carcinoma cell line associated with STAT activation regulated by EB virus encoded protein LMP1. Prog Biochem Biophys. 2003;30:560–565.
36. Dawson CW, Tramountanis G, Eliopoulos AG, et al. Epstein-Barr virus latent membrane protein 1 (LMP1) activates the phosphatidylinositol 3-kinase/Akt pathway to promote cell survival and induce actin filament remodeling. J Biol Chem. 2003;278:3694–3704. doi:10.1074/jbc.M209840200
37. Hammarskjold ML, Simurda MC. Epstein–Barr virus latent membrane protein transactivates the human immunodeficiency virus type 1 long terminal repeat through induction of NF-kappa B activity. J Virol. 1992;66:6496–6501.
Supplementary materials
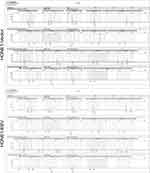 |
Figure S1 STR loci of HONE1-Vector and HONE1-EBV have basically matched with HONE1. |
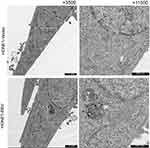 |
Figure S2 Transmission electron microscopy of HONE1-Vector and HONE1-EBV cells observed no virus particles. |
 |
Table S1 Antibodies used in Western blot |
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.