Back to Journals » International Journal of General Medicine » Volume 14
Exploring the Potential Targets and Mechanisms of Huang Lian Jie Du Decoction in the Treatment of Coronavirus Disease 2019 Based on Network Pharmacology
Authors Liu W, Zeng Y , Li Y, Li N, Peng M, Cheng J, Tian B, Chen M
Received 6 September 2021
Accepted for publication 29 November 2021
Published 16 December 2021 Volume 2021:14 Pages 9873—9885
DOI https://doi.org/10.2147/IJGM.S337025
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Wang Liu,1,* Yu Zeng,1,* Yanda Li,2,* Nanhong Li,3 Min Peng,1 Junfen Cheng,1 Binbin Tian,4 Mingdi Chen5
1Department of Respiration, The Second Affiliated Hospital of Guangdong Medical University, Zhanjiang, Guangdong, People’s Republic of China; 2Department of Internal Medicine, Central People’s Hospital of Zhanjiang, Zhanjiang, Guangdong, People’s Republic of China; 3Department of Pathology and Pathophysiology, Southern Medical University, Guangzhou, Guangdong, People’s Republic of China; 4Department of Critical Care Medicine, Central People’s Hospital of Zhanjiang, Zhanjiang, Guangdong, People’s Republic of China; 5Department of Critical Care Medicine, The Second Affiliated Hospital of Guangdong Medical University, Zhanjiang, Guangdong, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Mingdi Chen; Binbin Tian Email [email protected]; [email protected]
Background: In December 2019, coronavirus disease 2019 (COVID-19) caused by a novel coronavirus (severe acute respiratory syndrome coronavirus 2, SARS-CoV-2; previously known as 2019-nCoV) emerged in Wuhan, China, and caused many infections and deaths. At present, there are no specific drugs for the etiology and treatment of COVID-19. A combination of traditional Chinese and western medicine is proposed to treat COVID-19, in which Huang Lian Jie Du decoction (HLJDD) is recommended for the treatment of COVID-19 in many provinces in China and has been widely used in the clinic. This study explored the potential targets of HLJDD in the treatment of COVID-19 based on network pharmacology.
Methods: First, the chemical composition and targets of HLJDD and COVID-19-related targets were obtained through the TCMSP, UniProt, GeneCards and OMIM databases. Second, HLJDD target and HLJDD-COVID-19 target networks were constructed via the STRING database and Cytoscape software. Finally, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of the HLJDD-COVID-19 targets was applied via the DAVID database.
Results: Our study identified a total of 67 active ingredients of HLJDD and 204 targets of HLJDD. A total of 502 COVID-19-related targets were obtained, of which 47 were intersecting targets of HLJDD and COVID-19. A total of 179 GO terms and 77 KEGG terms, including the TNF signaling pathway, NF-κB signaling pathway and HIF-1 signaling pathway, were identified.
Conclusion: The present study explored the potential targets and signaling pathways of HLJDD during the treatment of COVID-19, which may provide a basis for the research and development of drugs for the treatment of COVID-19.
Keywords: Huang Lian Jie Du decoction, coronavirus disease 2019, COVID-19, SARS-CoV-2, network pharmacology, traditional Chinese medicine
Introduction
In December 2019, a pneumonia caused by a special pathogen emerged in Wuhan, Hubei Province, China, and quickly speeded around the world. The disease is transmitted mainly by droplets or contact and has the characteristics of a long incubation period, strong infectivity, fast transmission speed and so on.1 The special pathogen was later isolated and identified as novel coronavirus (severe acute respiratory syndrome coronavirus 2, SARS-CoV-2; previously known as 2019-nCoV) by scientists.2 As of Nov. 17, 2021, more than 255,000,000 people have been diagnosed with Coronavirus disease 2019 (COVID-19), and more than 5,000,000 people have died of COVID-19 worldwide. Both the number of patients diagnosed with COVID-19 and COVID-19-related deaths surpassed those of the 2003 outbreak of severe acute respiratory syndrome (SARS).3 The common susceptibility of people to the novel coronavirus is one of the major causes of the large outbreak of COVID-19.4 Currently, COVID-19 has become a serious social problem affecting human health.5
SARS-CoV-2 and SARS-CoV (severe acute respiratory syndrome coronavirus) and MERS-CoV (middle east respiratory syndrome coronavirus) belong to β-coronavirus genus.6,7 The genomic sequence homology of SARS-CoV-2 and SARS-CoV is 79.5%.8 The homology of SARS-COV-2 and BAT-SL-COVZC45 is more than 85%, the bat may be the natural host of SARS-CoV-2.8 At present, the mechanism of infection and invasion of COVID-19 is not completely clear. The possible mechanism is that novel coronavirus infects human body by binding with angiotensin-converting enzyme 2 (ACE2) on cells,9,10 causing body damage which may be related to inducing over immune response, leading to “self-attack” and causing multi-organ damage.11
Transmission by respiratory droplets and close contact is the main transmission route of SARS-CoV-2.8 The clinical manifestations of COVID-19 were fever, fatigue, and dry cough, and chest radiographs showed bilateral pulmonary infiltration.12 Severely ill patients may develop complications such as acute respiratory distress syndrome (ARDS), septic shock, refractory metabolic acidosis, coagulation dysfunction and multiple organ dysfunction syndrome (MODS).13 Clinical treatment is mainly symptomatic, but critically ill patients need mechanical ventilation treatment. Most critically ill patients are over 60 years old and have one or more complications.13,14 At present, there is no specific antiviral treatment; in addition, the current main treatments are all supportive. The National Health Commission of China has released a series of diagnosis and treatment plans on COVID-19 and recommended traditional Chinese medicine (TCM) to treat COVID-19. Chinese medicine stresses syndrome differentiation and treatment. The COVID-19 lesion is in the lung and has the characteristics of “wet”, “heat”, “ toxins “ and “stasis”. Chinese provinces have also released TCM treatment plans on COVID-19 that are suitable for local areas in combination with the local climate and disease characteristics. Among them is a classic Chinese medicine prescription called Huang Lian Jie Du decoction (HLJDD), which is recommended to treat COVID-19 in TCM treatment plans in many provinces.
HLJDD, a famous prescription of TCM, is derived from “Zhou Hou Bei Ji Fang” by Ge Hong and first recorded in “Wai Tai Mi Yao” in the Tang dynasty. HLJDD is composed of Phellodendri Chinensis Cortex, Scutellariae Radix, Gardeniae Fructus and Coptidis Rhizoma. HLJDD has the function of clearing heat and detoxification, and it has been reported that HLJDD may be a suitable treatment for patients with a fever of unknown origin.15,16 Furthermore, HLJDD plays an important role in anti-inflammatory, neuroprotection, anti-atherosclerosis, and blood sugar-lowering activities.17–19 As mentioned above, “heat” and “toxins” have come to be considered characteristics of COVID-19. Thus, these features seem to suggest that HLJDD can be used in the treatment of COVID-19.
Network pharmacology is a new discipline based on the fusion of big data regarding pharmacology, molecular biology, computer networks and so on. Through biological pedigree analysis, we can understand the potential mechanism between active components, target proteins and pathways related to the occurrence and development of diseases.20 This study aimed to explore the potential targets and signaling pathways of HLJDD involved in the treatment of COVID-19 based on network pharmacology and to provide a reference for clinical drug therapy in COVID-19. The workflow of the study is shown in Figure 1.
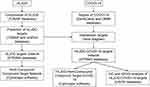 |
Figure 1 Workflow of this systematic analysis based on network pharmacology. Abbreviations: HLJDD, Huang Lian Jie Du decoction; COVID-19, coronavirus disease 2019. |
Materials and Methods
Compounds and Targets of HLJDD
The Traditional Chinese Medicine Systems Pharmacology (TCMSP, http://lsp.nwu.edu.cn/tcmsp.php) database is a unique pharmacology database of the Chinese herbal medicine system, which includes information on Chinese herbal medicine, chemical composition, targets, and drug target networks.21,22 Chinese medicine prescriptions are often composed of a variety of Chinese herbs and contain thousands of compounds, but only some of them have good pharmacokinetics and potential biological effects. In brief, the oral Chinese medicine must overcome the barriers to absorption, distribution, metabolism and excretion (ADME) process to be effective. In ADME process, oral bioavailability (OB) is defined as “the rate and extent at which the active ingredient or active part is absorbed from the drug product and available at the site of action”, which is one of the most significant pharmacokinetic parameters.23 The compounds with OB ≥ 30% were considered to have high OB. Another index, drug-likeness (DL), as a qualitative concept, is used in drug design to evaluate the usability of a molecule and can be used to quickly screen active substances. In the DrugBank database, the average DL index is 0.18 and compounds with DL ≥ 0.18 were regarded to have high druggability.24 Furthermore, drug half-life (HL) has important implications for dosing regimen and peak-to-trough ratio at the steady state, which is regarded as an important index for screening effective molecules and HL ≥ 4 is the recommended threshold in the TCMSP database.25 Therefore, the compounds in HLJDD with OB ≥ 30%, DL ≥ 0.18 and HL ≥ 4 were selected as active ingredients in this study. Next, the Related Targets function in the TCMSP database was used to identify the targets of the active ingredients of each herb in HLJDD. Moreover, the obtained targets were further screened by using UniProt database (https://www.uniprot.org/)26 to compare target information and standardize gene names.
Prediction of Drug Targets for HLJDD in the Treatment of COVID-19
GeneCards database (https://www.genecards.org/) is a comprehensive and authoritative compendium of annotative information about human genes, which provides genomic, transcriptomic, proteomic, genetic, clinical and functional information.27 In addition, the Online Mendelian Inheritance in Man (OMIM, http://www.omim.org/) database is a freely accessed database that can be used to screen disease-related targets.28 In this study, the GeneCards and OMIM databases were applied to identify the COVID-19-related targets with “2019 novel coronavirus infection”, “2019-nCoV infection”, “coronavirus disease-19”, “2019-nCoV disease”, “2019 novel coronavirus disease”, “coronavirus disease 2019” or “coronavirus disease” as keywords, and duplicate targets were eliminated. The drug targets of HLJDD in the treatment of COVID-19 were obtained by intersecting the HLJDD-related targets and COVID-19 related targets.
Target Network Analysis
The Search Tool for the Retrieval of Interacting Genes (STRING, http://string-db.org/) database is a freely accessible database that can be used to identify possible protein-protein interaction (PPI).29 The STRING database was applied to determine the HLJDD target network and the drug targets network. In STRING database, the species was set as “Homo sapiens” and PPIs with a confidence score of > 0.9 were selected. Cytoscape (https://cytoscape.org/) is software for graphically displaying, analyzing, and editing networks.30 Here, Cytoscape (version 3.7.2) was used to construct the herb-compound-target network and the herb-compound-drug target network.
Functional Enrichment Analysis
The Database for Annotation, Visualization and Integrated Discovery (DAVID) (https://david.ncifcrf.gov/) is an online bioinformatics database that can extract biological functional annotation information for large-scale genes.31 The DAVID database was employed to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of predicted drug targets of HLJDD, with the criteria of P-value < 0.01. The GO enrichment analysis consisted of three terms, namely, cellular component (CC), biological process (BP), and molecular function (MF).
Results
Screening of Active Compounds and Target Genes in HLJDD
All compounds of each Chinese herb in HLJDD were identified by using the TCMSP database with OB ≥ 30%, DL ≥ 0.18, and HL ≥ 4 as criteria. Ultimately, 27 compounds in Huangbai, 32 compounds in Huangqin, 11 compounds in Zhizi, and 10 compounds in Huanglian were screened (Table S1). After removing the repeated compounds, a total of 67 compounds were obtained from the four kinds of Chinese herbs in HLJDD. Next, the targets of compounds were predicted, and 204 target genes were found in four kinds of Chinese herbs in HLJDD via TCMSP and the UniProt databases. Specifically, there were 345 targets in Huangbai, 333 targets in Huangqin, 274 targets in Zhizi, and 220 targets in Huanglian (Table S2).
Prediction of Potential Targets of HLJDD in Anti-COVID-19
After searching the GeneCards and OMIM databases, a total of 502 targets related to COVID-19 were obtained (Table S3). Then, these targets were compared with the 204 target genes of HLJDD by using R software (version 3.6.2), and finally 47 overlapping genes were identified to be potential drug targets in the treatment of COVID-19 (Figure 2 and Table S3).
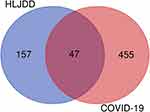 |
Figure 2 Venn diagram presents the overlapping genes between HLJDD and COVID-19-related targets. |
Construction of PPI Network
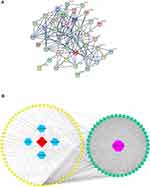
As shown in Figure 3A, we observed that the HLJDD-related targets network contained 175 nodes and 747 edges (average node degree=7.32, average local clustering coefficient=0.448, and PPI enrichment P-value<1.0e-16). Based on the HLJDD-related targets, the herb-compound- target network was also established. This network revealed the relationships among 4 Chinese herbs, 67 compounds and 204 compound-related targets (Figure 3B).
In addition, the potential drug target network was built. As shown in Figure 4A, the drug target network composed of 41 nodes and 146 edges (average node degree=6.21, average local clustering coefficient=0.5, and PPI enrichment P-value<1.0e-16). Furthermore, the herb-compound-drug target network was established based on the potential drug targets of anti-COVID-19. This network included 4 Chinese herbs, 67 compounds, and 47 drug targets (Figure 4B).
Normally, genes with a high intensity of association with other genes usually play a crucial role of hub genes. The greater the degree value of a node is, the more nodes are connected to it and the more important it is in the network. In this analysis, the degree value of all nodes in the drug targets network for COVID-19 is shown in Table S4. R software (version 3.6.2) was used to screen out the hub genes by the degree value method according to the HLJDD-COVID-19 target network. The degree value of a node refers to the number of edges between nodes in a network and the top 10 targets are shown in Table 1. As shown in Figure 5, there were 20 targets, namely, RELA, IL6, IL1B, MAPK14, CXCL8, MAPK1, IL4, IL10, IL2, MAPK8, CCL2, FOS, IFNG, CXCL2, STAT1, BCL2L1, CASP3, CXCL10, EGFR, and PRKCB, whose degree value was greater than the average node degree value. These results indicated that these targets might be hub genes effective in the treatment of COVID-19.
 |
Table 1 The Top 10 Degree Genes in HLJDD-COVID-19 Target Network |
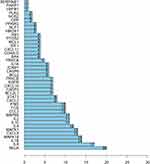 |
Figure 5 Top 20 drug targets with higher degree in the drug target network. |
Functional Enrichment Analysis of Drug Targets
To elucidate the molecular mechanism of the 47 drug targets, GO enrichment and KEGG pathway enrichment analyses were performed via the DAVID database, and the specific enrichment categories are listed in Table S5. The top 10 significant enriched BP, CC, and MF terms are displayed in Figure 6A; meanwhile, the top 30 significant involved KEGG pathways are showed in Figure 6B. In brief, a total of 148 BP terms were identified, such as cellular response to lipopolysaccharide, inflammatory response, immune response, chemotaxis, and regulation of blood pressure; a total of 14 CC terms were identified, including extracellular space, extracellular region, cytosol, Bcl-2 family protein complex, external side of plasma membrane, mitochondrial outer membrane, caveola, perinuclear region of cytoplasm, mitochondrion, cytoplasm, nucleoplasm, intracellular, nucleus, and extracellular exosome; moreover, a total of 17 MF terms were identified, including BH3 domain binding, cytokine activity, identical protein binding, enzyme binding, protein binding, chemokine activity, protein homodimerization activity, transcription factor binding, heme binding, MAP kinase activity, protein heterodimerization activity, tumor necrosis factor receptor binding, prostaglandin-endoperoxide synthase activity, histone kinase activity (H3-T6 specific), ubiquitin protein ligase binding, nitric-oxide synthase activity, and growth factor activity.
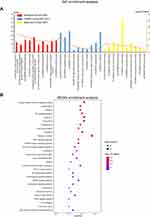 |
Figure 6 Functional enrichment analysis of 47 drug targets against COVID-19. (A) Top 10 terms for GO enrichment analysis. (B) Top 30 pathways for KEGG enrichment analysis. |
In terms of the KEGG enrichment analysis, 77 significant signaling pathways were identified, such as Chagas disease (American trypanosomiasis), Leishmaniasis, Pertussis, TNF signaling pathway, Influenza A, NF-κB signaling pathway, HIF-1 signaling pathway, and MAPK signaling pathway.
Discussion
Starting in December 2019, a novel coronavirus called SARS-CoV-2 began causing a respiratory illness that is sweeping the world. The WHO considers the current outbreak of COVID-19 to be a global pandemic. Currently, COVID-19 has become a serious social problem affecting human health. Since the newly emerged SARS-CoV-2 is a previously unknown pathogen, no specific drug is available. Therefore, there is an urgent need to explore therapeutic drugs and specific vaccines to combat the emerging virus. However, the development of antiviral drugs takes time and resources, so reusing existing drugs to treat emerging viral diseases are effective strategies. In general, these drug options can be roughly divided into virus based and host based treatment.32,33
In China, the combination of TCM treatment and effective life support has been widely used in the clinical treatment of COVID-19. TCM treatment is effective in preventing COVID-19, which can help reduce the severity of COVID-19 and promote the recovery of patients with COVID-19. TCM emphasizes both evil and positive treatment. Based on TCM theory, the main therapeutic principles are invigorating the lungs to prevent dampness, clearing away heat and detoxification, replenishing qi, and nourishing yin.34 However, the molecular mechanism of TCM prescription in treating COVID-19 disease remains not completely clear. In theory, TCMs have a certain therapeutic effect through multiple-molecules, multi-targets and multi-pathways. In the syndrome differentiation and treatment of syndrome type via TCM, the target organ location of COVID-19 is the lung and its pathogenic property is damp and toxin plague, so it should be treated with purging the lung and detoxification method. Notably, HLJDD is the representative prescription for heat clearing and detoxification. In the clinical practice of TCM, HLJDD has been widely used to treat inflammation, hypertension, gastrointestinal diseases and hepatic and cerebral vascular diseases.35 However, the pharmacological mechanisms of HLJDD in the treatment of COVID-19 have not been clearly studied.
Network pharmacology is based on the interaction network relationship of “drug-target-disease”. By analyzing the relationship among genes, diseases, drugs and other network libraries, network pharmacology reveals the mystery of “drug-target”, “target-disease” and “drug-disease” interactions and guides accurate clinical drug use and new drug research and development.36 In this study, we used network pharmacology approach to construct and analyze a network of targets, conduct enrichment analysis on these targets, and reveal their possible mechanism of action to explore the potential targets of HLJDD in the treatment of COVID-19.
In short, 67 active compounds and 204 targets of compounds were screened. We observed that a variety of alkaloids and flavonoids were searched in the HLJDD, such as quercetin, baicalein, wogonin, berberine, and acacetin, which were consistent with existing studies.18,37 Baicalein and wogonin are flavonoids that can inhibit the overactivation of the complement system in vivo and improve acute lung injury induced by influenza A virus.38 In addition, Song et al39 explored the therapeutic effects of baicalein for the treatment of COVID-19 in vivo and in vitro. They found that baicalein could inhibit cell damage induced by SARS-CoV-2; meanwhile, it could also improve the respiratory function of mice with acute lung injury, inhibit pneumonia cell infiltration as well as the replication of the virus, and thus reduce lung tissues damage. Meanwhile, baicalein had an inhibitory effect on SARS-CoV-2 3C-like protease activity, and it also could inhibit the replication of SARS-CoV-2 in Vero cells.40 These studies revealed that baicalein may be a promising drug for the treatment of COVID-19. Berberine has antibacterial and anti-inflammatory pharmacological effects and can inhibit the replication of respiratory syncytial virus and cytomegalovirus to improve the symptoms of viral pneumonia.41,42 Quercetin has anti-inflammatory, antiviral, and immunomodulatory pharmacological effects.43 Another key compound, acacetin, has anti-inflammatory effects by reducing interleukin 6 (IL-6), interleukin 8 (IL-8), and intercellular adhesion molecule-1 (ICAM-1) levels.44,45 Taken together, we speculate that these active compounds in HLJDD may improve lung injury caused by COVID-19 through synergistic anti-inflammatory effects.46
Highly pathogenic coronaviruses, such as SARS coronavirus and MERS coronavirus, can cause fatal pneumonia, which is related mainly to rapid viral replication, a large amount of inflammatory cell infiltration and an enhanced pro-inflammatory cytokine/chemokine response.47 In this analysis, several potential targets of HLJDD against COVID-19 were methodically identified, including RELA, IL6, IL1B, MAPK14, CXCL8, MAPK1, IL4, IL10, IL2, MAPK8, CCL2, FOS, IFNG, CXCL2, STAT1, BCL2L1, CASP3, CXCL10, EGFR, and PRKCB. Reportedly, majority of them have been involved in pulmonary inflammation. Genes such as interleukin 1β (IL-1B) and IL-6 are involved in the recruitment of immune cells to the lungs; meanwhile, IL-1B can activate caspases and other related signaling pathways, induce apoptosis of infected cells, and thus inhibit virus replication. MAPK14, MAPK1, and MAPK8 are members of the MAPK family, which can promote the release of pro-inflammatory cytokines such as IL-1B and IL-6, and it is regarded as a core factor in inflammation regulation.48 Evidence has also indicated that SARS-CoV-2 infection can increase the expression levels of plasma cytokines and chemokines, such as interleukin 2 (IL2), IL6, interleukin 7 (IL7), interleukin 10 (IL10), C-C motif chemokine ligand 2 (CCL2), C-C motif chemokine ligand 3 (CCL3), C-X-C motif chemokine ligand 10 (CXCL10), cytokines interferon γ (IFN-γ), transforming growth factor α (TGF-α), interleukin 1 (IL1), interleukin 12 (IL12), CCL2, C-X-C motif chemokine ligand 9 (CXCL9) and CXCL10, which were associated with pulmonary inflammation and lung damage in COVID-19 patients.49 Furthermore, previous study revealed that the activation of inflammatory signaling pathways and cytokine storm reserved as crucial factors that leaded to ARDS in patients with COVID-19, and the excessive secretion of pro-inflammatory cytokines and chemokines resulted in innate immune system disorders that eventually caused immune damage.50 Therefore, HLJDD may serve the role of anti-inflammatory, anti-immune apoptosis, and alleviation of pneumonia-induced multi-organ damage via regulating the expression of these hub genes, which indirectly inhibit the virus-mediated inflammatory response by regulating the immune function of the body.
In the KEGG analysis of the potential targets against COVID-19, we identified several signaling pathways associated with inflammation and viral infection, including the tumor necrosis factor (TNF) signaling pathway, nuclear factor κB (NF-κB) signaling pathway, Hypoxia-inducible factor 1 (HIF-1) signaling pathway, Influenza A, Hepatitis B, Herpes simplex infection, and Prion diseases. The NF-κB signaling pathway plays a key role in the cellular inflammatory response and immune response, and the regulation of the NF-κB signaling pathway can reduce the injury to the liver, lung and kidney caused by pneumonia.51,52 HIF-1 is a transcription factor and can be used as the main regulator of oxygen homeostasis, and the HIF-1 signaling pathway is related to the release of inflammatory cytokines in viral pneumonia.53 TNF is a proinflammatory cytokine that plays an important role in cell proliferation, differentiation and apoptosis.54 Severe pneumonia with sepsis has been reported to be characterized by an inflammatory response disorder related to endotoxin.55 In the early stage of sepsis, Hypoxia-inducible factor 1α (HIF1-α) is activated under hypoxia and then coactivates signal transducer and activator of transcription 3 (STAT3) to participate in the inflammatory process.55
In this study, network pharmacology was used to explore the feasibility, relevant molecular targets and signaling pathways of HLJDD in the treatment of COVID-19. It is worth noting that network pharmacology has great advantages in studying the way drugs act. Our research has certain limitations. First, the potential targets were only obtained by using the network pharmacology method, and it is a lack of in vivo experimental verification. Second, because of the difficulty of data collection, the database may not include all known or unknown targets and protein-protein interactions. This aspect may improve as additional data become available in the future. Third, HLJDD is a medicine of great bitterness and cold, and it can easily hurt the spleen and stomach if it is taken for a long time or if too much is taken, so it should be applied according to clinical practice based on syndrome differentiation. We hope that our results can provide some theoretical and scientific basis for HLJDD in the treatment of COVID-19 and provide a reference for drug development for SARS-CoV-2 infection.
Conclusions
In this study, 47 potential targets against COVID-19 were identified by network pharmacology method, and the molecular mechanism of HLJDD in the treatment of COVID-19 was further investigated through functional enrichment analysis. Based on the results of enrichment analysis, we revealed the characteristics of multi-target synergistic effects and multi-pathway regulations and found that HLJDD may improve COVID-19 through the TNF signaling pathway, NF-κB signaling pathway and HIF-1 signaling pathway. Inhibiting the overproduction of inflammatory factors is of great significance in the prevention and treatment of COVID-19 and the alleviation of multiorgan injury in COVID-19 patients. As an important adjuvant therapy, the combination of HLJDD and other traditional Chinese medicines with modern medical methods will benefit COVID-19 patients. The effect of HLJDD in the treatment of COVID-19 requires further experiments to verify the specific molecular mechanism.
Data Sharing Statement
The data used to support the findings of this study are included within the article and the supplementary information file(s).
Ethics Approval and Consent to Participate
The Institutional Review Board and Ethics Committee of the Second Affiliated Hospital of Guangdong Medical University deemed that this work is a web-based pharmacologic analysis based on a database of patients who have received ethical approval. Users can download relevant data for free to research and publish related articles. This research is based on open-source data, so review and approval was not required for this research by an institutional review board or ethics committee and there are no ethical issues or other conflicts of interest.
Acknowledgments
We appreciate all the people who are fighting the COVID-19 epidemic around the world. We thank American Journal Experts (AJE) for English language editing. We would like to thank Dr. Weinmin Yao and Dr. Guangfeng He from the Department of Respiration, The Second Affiliated Hospital of Guangdong Medical University, for giving us a lot of suggestions in this revision, including the whole work design, data rearrangement, analysis and interpretation, etc.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
This study is funded by 2021 Zhanjiang Science and Technology Development Special Fund Competitive Allocation Project (High-level Hospital Construction Special Project of the Second Affiliated Hospital of Guangdong Medical University) (No. 2021A05107).
Disclosure
The authors declare that they have no conflicts of interest.
References
1. Zheng J. SARS-CoV-2: an emerging coronavirus that causes a global threat. Int J Biol Sci. 2020;16:1678. doi:10.7150/ijbs.45053
2. Lu R, Zhao X, Li J, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565–574. doi:10.1016/S0140-6736(20)30251-8
3. Isakbaeva ET, Khetsuriani N, Beard RS, et al. SARS-associated coronavirus transmission, United States. Emerg Infect Dis. 2004;10:225–231. doi:10.3201/eid1002.030734
4. Zhai P, Ding Y, Wu X, Long J, Zhong Y, Li Y. The epidemiology, diagnosis and treatment of COVID-19. Int J Antimicrob Agents. 2020;55:105955. doi:10.1016/j.ijantimicag.2020.105955
5. Nkengasong J. China’s response to a novel coronavirus stands in stark contrast to the 2002 SARS outbreak response. Nat Med. 2020;26:310–311.
6. Su S, Wong G, Shi W, et al. Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol. 2016;24:490–502. doi:10.1016/j.tim.2016.03.003
7. Chan JF-W, Kok K-H, Zhu Z, et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microb Infect. 2020;9:221–236. doi:10.1080/22221751.2020.1719902
8. Zhou P, Yang X-L, Wang X-G, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270–273. doi:10.1038/s41586-020-2012-7
9. Richardson P, Griffin I, Tucker C, et al. Baricitinib as potential treatment for 2019-nCoV acute respiratory disease. Lancet. 2020;395:e30. doi:10.1016/S0140-6736(20)30304-4
10. Gralinski LE, Menachery VD. Return of the coronavirus: 2019-nCoV. Viruses. 2020;12:135. doi:10.3390/v12020135
11. Zumla A, Hui DS, Azhar EI, Memish ZA, Maeurer M. Reducing mortality from 2019-nCoV: host-directed therapies should be an option. Lancet. 2020;395:e35–e36. doi:10.1016/S0140-6736(20)30305-6
12. Xu X-W, Wu -X-X, Jiang X-G, et al. Clinical findings in a group of patients infected with the 2019 novel coronavirus (SARS-Cov-2) outside of Wuhan, China: retrospective case series. BMJ. 2020;368:m606.
13. Poston JT, Patel BK, Davis AM. Management of critically ill adults with COVID-19. JAMA. 2020. doi:10.1001/jama.2020.4914
14. Bennardo F, Buffone C, Giudice A. New therapeutic opportunities for COVID-19 patients with Tocilizumab: possible correlation of interleukin-6 receptor inhibitors with osteonecrosis of the jaws. Oral Oncol. 2020:104659. doi:10.1016/j.oraloncology.2020.104659
15. Oshima N, Narukawa Y, Hada N, Kiuchi F. Quantitative analysis of anti-inflammatory activity of orengedokuto: importance of combination of flavonoids in inhibition of PGE 2 production in mouse macrophage-like cell line J774. 1. J Nat Med. 2013;67:281–288. doi:10.1007/s11418-012-0679-2
16. Tanaka K, Nara K, Nishimura T, et al. Fever of unknown origin successfully treated by oren-gedoku-to (huanglian-jie-du-tang). Int J Gen Med. 2013;6:829. doi:10.2147/IJGM.S52488
17. Chen M, Liao Z, Lu B, et al. Huang-Lian-Jie-Du-decoction ameliorates hyperglycemia and insulin resistant in association with gut microbiota modulation. Front Microbiol. 2018;9:2380. doi:10.3389/fmicb.2018.02380
18. Lu J, Wang J-S, Kong L-Y. Anti-inflammatory effects of Huang-Lian-Jie-Du decoction, its two fractions and four typical compounds. J Ethnopharmacol. 2011;134:911–918. doi:10.1016/j.jep.2011.01.049
19. Zhang Q, Fu X, Wang J, Yang M, Kong L. Treatment effects of ischemic stroke by berberine, baicalin, and jasminoidin from Huang-Lian-Jie-Du-Decoction (HLJDD) explored by an integrated metabolomics approach. Oxid Med Cell Longev. 2017;2017:1–20. doi:10.1155/2017/9848594
20. Hopkins AL. Network pharmacology: the next paradigm in drug discovery. Nat Chem Biol. 2008;4:682. doi:10.1038/nchembio.118
21. Ru J, Li P, Wang J, et al. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J Cheminform. 2014;6:13. doi:10.1186/1758-2946-6-13
22. Wang P, Zhou W, Meng J, Huo H, Sui F. Inter-module coupling analysis of Huang-Lian-Jie-Du-Decoction on stroke. Front Pharmacol. 2019;10:1288. doi:10.3389/fphar.2019.01288
23. Chen ML, Shah V, Patnaik R, et al. Bioavailability and bioequivalence: an FDA regulatory overview. Pharm Res. 2001;18:1645–1650. doi:10.1023/a:1013319408893
24. Zuegg J, Cooper MA. Drug-likeness and increased hydrophobicity of commercially available compound libraries for drug screening. Curr Top Med Chem. 2012;12:1500–1513. doi:10.2174/156802612802652466
25. Zhang W, Chen Y, Jiang H, et al. Integrated strategy for accurately screening biomarkers based on metabolomics coupled with network pharmacology. Talanta. 2020;211:120710. doi:10.1016/j.talanta.2020.120710
26. UniProt Consortium. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 2019;47:D506–D515. doi:10.1093/nar/gky1049
27. Stelzer G, Rosen N, Plaschkes I, et al. The GeneCards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics. 2016;54:1–30. doi:10.1002/cpbi.5
28. Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 2005;33:D514–D517. doi:10.1093/nar/gki033
29. Szklarczyk D, Franceschini A, Wyder S, et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015;43:D447–D452. doi:10.1093/nar/gku1003
30. Otasek D, Morris JH, Bouças J, Pico AR, Demchak B. Cytoscape automation: empowering workflow-based network analysis. Genome Biol. 2019;20:1–15. doi:10.1186/s13059-019-1758-4
31. Dennis G, Sherman BT, Hosack DA, et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4:R60. doi:10.1186/gb-2003-4-9-r60
32. Lu H. Drug treatment options for the 2019-new coronavirus (2019-nCoV). Biosci Trends. 2020;14:69–71. doi:10.5582/bst.2020.01020
33. Wang M, Cao R, Zhang L, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020;30:269–271. doi:10.1038/s41422-020-0282-0
34. Zhao Z, Li Y, Zhou L, et al. Prevention and treatment of COVID-19 using Traditional Chinese Medicine: a review. Phytomedicine. 2021;85:153308. doi:10.1016/j.phymed.2020.153308
35. Cao Y, Gao C, Sun J, Wang J, Zhou Q, Liu G. Study on pharmacological effects of Huanglianjiedu Tang extract. J China Pharm Univ. 1996;27:605–608.
36. Tang F, Tang Q, Tian Y, Fan Q, Huang Y, Tan X. Network pharmacology-based prediction of the active ingredients and potential targets of Mahuang Fuzi Xixin decoction for application to allergic rhinitis. J Ethnopharmacol. 2015;176:402–412. doi:10.1016/j.jep.2015.10.040
37. Yang Y, Wang H-J, Yang J, et al. Chemical profiling and quantification of Chinese medicinal formula Huang-Lian-Jie-Du decoction, a systematic quality control strategy using ultra high performance liquid chromatography combined with hybrid quadrupole-orbitrap and triple quadrupole mass spectrometers. J Chromatogr A. 2013;1321:88–99. doi:10.1016/j.chroma.2013.10.072
38. Zhi H, Jin X, Zhu H, et al. Exploring the effective materials of flavonoids-enriched extract from Scutellaria baicalensis roots based on the metabolic activation in influenza A virus induced acute lung injury. J Pharm Biomed Anal. 2020;177:112876. doi:10.1016/j.jpba.2019.112876
39. Song J, Zhang L, Xu Y, et al. The comprehensive study on the therapeutic effects of baicalein for the treatment of COVID-19 in vivo and in vitro. Biochem Pharmacol. 2021;183:114302. doi:10.1016/j.bcp.2020.114302
40. Liu H, Ye F, Sun Q, et al. Scutellaria baicalensis extract and baicalein inhibit replication of SARS-CoV-2 and its 3C-like protease in vitro. J Enzyme Inhib Med Chem. 2021;36:497–503. doi:10.1080/14756366.2021.1873977
41. Luganini A, Mercorelli B, Messa L, Palù G, Gribaudo G, Loregian A. The isoquinoline alkaloid berberine inhibits human cytomegalovirus replication by interfering with the viral Immediate Early-2 (IE2) protein transactivating activity. Antiviral Res. 2019;164:52–60. doi:10.1016/j.antiviral.2019.02.006
42. Lee B-H, Chathuranga K, Uddin MB, et al. Coptidis Rhizoma extract inhibits replication of respiratory syncytial virus in vitro and in vivo by inducing antiviral state. J Microbiol. 2017;55:488–498. doi:10.1007/s12275-017-7088-x
43. Jo S, Kim H, Kim S, Shin DH, Kim MS. Characteristics of flavonoids as potent MERS‐CoV 3C‐like protease inhibitors. Chem Biol Drug Des. 2019;94:2023–2030. doi:10.1111/cbdd.13604
44. Bi C, Dong X, Zhong X, Cai H, Wang D, Wang L. Acacetin protects mice from Staphylococcus aureus bloodstream infection by inhibiting the activity of sortase A. Molecules. 2016;21:1285. doi:10.3390/molecules21101285
45. Huang W-C, Liou C-J. Dietary acacetin reduces airway hyperresponsiveness and eosinophil infiltration by modulating eotaxin-1 and th2 cytokines in a mouse model of asthma. Evid Based Complement Alternat Med. 2012;2012:1–11. doi:10.1155/2012/910520
46. Qi Y, Zhang Q, Zhu H. Huang-Lian Jie-Du decoction: a review on phytochemical, pharmacological and pharmacokinetic investigations. Chin Med. 2019;14:57. doi:10.1186/s13020-019-0277-2
47. Runfeng L, Yunlong H, Jicheng H, et al. Lianhuaqingwen exerts anti-viral and anti-inflammatory activity against novel coronavirus (SARS-CoV-2). Pharmacol Res. 2020;156:104761. doi:10.1016/j.phrs.2020.104761
48. Panteva M, Korkaya H, Jameel S. Hepatitis viruses and the MAPK pathway: is this a survival strategy? Virus Res. 2003;92:131–140.
49. Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497–506. doi:10.1016/S0140-6736(20)30183-5
50. Choudhary S, Sharma K, Silakari O. The interplay between inflammatory pathways and COVID-19: a critical review on pathogenesis and therapeutic options. Microb Pathog. 2021;150:104673. doi:10.1016/j.micpath.2020.104673
51. Du J, Abdel-Razek O, Shi Q, et al. Surfactant protein D attenuates acute lung and kidney injuries in pneumonia-induced sepsis through modulating apoptosis, inflammation and NF-κB signaling. Sci Rep. 2018;8:1–14. doi:10.1038/s41598-018-33828-7
52. Kim Y, Allen E, Baird LA, et al. NF-κB RelA is required for hepatoprotection during pneumonia and sepsis. Infect Immun. 2019;87:e00132–00119. doi:10.1128/IAI.00132-19
53. Guo X, Zhu Z, Zhang W, et al. Nuclear translocation of HIF-1α induced by influenza A (H1N1) infection is critical to the production of proinflammatory cytokines: HIF-1α nuclear translocation induced by H1N1. Emerg Microb Infect. 2017;6:1–8. doi:10.1038/emi.2017.21
54. Rothe J, Gehr G, Loetscher H, Lesslauer W. Tumor necrosis factor receptors-structure and function. Immunol Res. 1992;11:81–90. doi:10.1007/BF02918612
55. Li L-L, Dai B, Sun Y-H, Zhang T-T. Monocytes undergo functional reprogramming to generate immunosuppression through HIF-1α signaling pathway in the late phase of sepsis. Mediators Inflamm. 2020;2020:1–9. doi:10.1155/2020/4235909
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.