Back to Journals » OncoTargets and Therapy » Volume 11
Exosome-mediated delivery of MALAT1 induces cell proliferation in breast cancer
Authors Zhang P, Zhou H, Lu K, Lu Y, Wang Y, Feng T
Received 25 October 2017
Accepted for publication 28 November 2017
Published 9 January 2018 Volume 2018:11 Pages 291—299
DOI https://doi.org/10.2147/OTT.S155134
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Samir Farghaly
Ping Zhang, Hongxing Zhou, Kefeng Lu, Yunou Lu, Yan Wang, Tongbao Feng
Department of Clinical Laboratory, The Affiliated Changzhou No 2 People’s Hospital, Nanjing Medical University, Changzhou, People’s Republic of China
Background: Breast cancer is the most common cancer in women worldwide. Cancer-secreted exosomes have recently been recognized as important mediators of intercellular communication. The aim of this study was to determine the role of exosomal long noncoding RNA metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) in breast cancer progression.
Materials and methods: Breast cancer specimens were obtained with informed consent from patients. Quantitative real-time polymerase chain reaction (qRT-PCR) was used to detect MALAT1 expression, and cellular proliferation was measured using cell counting kit-8 (CCK-8) assay.
Results: MALAT1 was highly expressed in breast cancer tissues and associated with disease progression. Breast cancer exosomes promoted cell proliferation and exosome-mediated MALAT1 to induce cell proliferation.
Conclusion: These findings indicated that exosomal MALAT1 could regulate cancer progression and represent a novel strategy for overcoming breast cancer.
Keywords: breast cancer, exosomes, MALAT1, cell proliferation
Introduction
Breast cancer is the most common cancer affecting women worldwide.1,2 Although diagnostic methods and therapeutic strategies for breast cancer have been improved in last decades, long-term survival of breast cancer patients still remains poor due to high proliferation and/or postsurgical recurrence rate.3–5 In the cancer cell proliferation field, many studies are focused on characteristics of the tumor microenvironment6–8 and on the intercellular communication of malignant and nonmalignant cells of the host.6,9
Exosomes, small vesicles delivered by many cells of the organism, are found in all body fluids. Exosomes are characterized by lipid composition, common and donor cell-specific proteins, mRNA, small noncoding RNA including miRNA, and DNA, which have recently been recognized as important mediators of intercellular communication,10–12 thus allowing the transfer of tumor-associated signaling molecules to surrounding cells.7,13 Importantly, the microRNAs in secreted exosomes can be transferred to a recipient cell where they affect post-transcriptional gene regulation and regulate proliferation, migration, and invasiveness in cancer biology.13,14 Cancer-secreted exosomes and miRNAs can be internalized by other cell types in the primary tumor microenvironment and pre-/metastatic niches.13,15 The cerebrospinal fluid from patients with recurrent glioma-derived exosomal miR-21 levels correlated with tumor spinal/ventricle metastasis and the recurrence with anatomical site preference.14 Human colon cancer patients with more advanced disease show higher levels of circulating exosomal miR-193a, which causes cell cycle G1 arrest and cell proliferation repression through targeting of cytoplasmic activation- and proliferation-associated protein 1 (Caprin1).16 Exosome-miR-210 from hypoxic breast cancer cells is transferred to cells in the tumor microenvironment and that miR-210 is involved in expression of vascular remodeling-related genes, such as Ephrin type-A receptor 3 (ephrin A3) and tyrosine-protein phosphatase nonreceptor type 1 (PTP1B), to promote angiogenesis.17,18 Exosome-1246 from a highly metastatic human oral cancer cell line induces cell growth through the activation of extracellular regulated protein kinases (ERK) and serine/threonine kinase(AKT) as well as promotes cell motility of the poorly metastatic cancer cell.19 In endothelial monolayers, exosome-mediated transfer of cancer-secreted miR-105 efficiently destroys tight junctions and the integrity of these natural barriers against metastasis.20
In this study, we evaluated the effect of breast cancer exosomes on tumor cell proliferation. We found that breast cancer exosomes promoted cell proliferation and exosome-mediated MALAT1 to induce tumor cell proliferation in breast cancer. Therefore, we speculated that the exosome-MALAT1, as a key regulator, might be a promising therapeutic target for the treatment of breast cancer.
Materials and methods
Clinical specimens
Human specimens were obtained with informed consent from patients who had undergone breast cancer surgery at The Affiliated Changzhou No 2 People’s Hospital, Nanjing Medical University, Changzhou, People’s Republic of China. All the samples were shown to be correctly labeled clinically and pathologically and immediately frozen at −80°C until use. The ethical committees of The Affiliated Changzhou No 2 People’s Hospital, Nanjing Medical University, approved this study, and written informed consent was obtained from all the patients.
Cell lines and cell culture
Human breast cancer cell lines (MCF-7, MDA-MB-231, and MDA-MB-435S) were obtained from The Affiliated Changzhou No 2 People’s Hospital, Nanjing Medical University, and was approved by the ethical committees of The Affiliated Changzhou No 2 People’s Hospital, Nanjing Medical University. The cell lines were cultured in Roswell Park Memorial Institute (RPMI) Medium 1640 and Dulbecco’s Modified Eagle’s Medium (DMEM; Thermo Fisher Scientific, Waltham, MA, USA) containing 10% fetal bovine serum (FBS; Thermo Fisher Scientific) and were grown in a humidified 5% CO2 incubator at 37°C.
Exosome purification
For exosomes secreted by cultured cell lines, conditioned media were first prepared by incubating cells grown at subconfluence in growth media containing exosome-depleted FBS (prepared by overnight ultracentrifugation at 100,000× g at 4°C) for 48 hours. For ultracentrifugation isolation, conditioned cell culture media were collected and centrifuged at 300× g for 5 minutes, 2,000× g for 10 minutes, and 10,000× g for 30 minutes at 4°C to remove cells and large debris. The supernatant was filtered using a 0.22 μm pore filter (Merck Millipore, Billerica, MA, USA), and the exosomes were pelleted at 140,000× g for 3 hours at 4°C. The exosome pellet was washed with 10 mL of 1× phosphate-buffered solution (PBS; Thermo Fisher Scientific) and pelleted again by centrifugation at 140,000× g for 2 hours at 4°C. The resulting pellet was either suspended in 1× PBS for whole exosome applications or further processed for RNA or protein extraction. Total exosome RNA was extracted using the TRIzol reagent (Thermo Fisher Scientific) following the manufacturer’s protocol. Recovered exosomes were measured for protein amount using Epoch (BioTek Instruments, Inc. Winooski, VT, USA).
RNA extraction and quantitative real-time polymerase chain reaction (qRT-PCR)
These procedures were performed as described previously.21 Total RNA was extracted using TRIzol reagent, and RNA quality and concentration were determined using a NanoDrop 2000 system (Thermo Fisher Scientific). RNA was converted into cDNA using the HiScript® II Q RT SuperMix (Vazyme, Nanjing, People’s Republic of China). qRT-PCR was performed using the SYBR® Green PCR Master Mix (Vazyme) on ABI VII7 Real-Time RT-PCR system (Bio-Rad Laboratories Inc., Hercules, CA, USA). Primers are listed in Table S1. RNA expression levels were obtained using the comparative cycle threshold 2−ΔΔCt method and analyzed as mean ± SD.
RNA oligoribonucleotides and cell oligoribonucleotides transfection
MALAT1 and all RNA oligoribonucleotides for in vitro studies were purchased from GenePharma (Shanghai, People’s Republic of China). Oligoribonucleotides were performed using Lipofectamine 2000 (Thermo Fisher Scientific). Unless otherwise indicated, 100 nM of RNA duplex or 80 nM of MALAT1-siRNA was used for each transfection, and all the experiments were repeated in triplicate.
Cell proliferation
Cellular proliferation was measured using cell counting kit-8 (CCK-8) assay (Sigma-Aldrich Co., St Louis, MO, USA), according to the manufacturer’s instructions. Briefly, 5×103 cells per well were seeded into a 96-well plate. At particular points in time, 20 μL of CCK-8 solution was added to each well for 4 hours at 37°C. The absorbance was detected by the Benchmark Plus™ Microplate Spectrometer (Bio-Rad Laboratories Inc.).
All experiments were performed in triplicate.
Xenograft mouse model
These procedures were performed as described previously.21 The 5×106 cells stably expressed MALAT1 were subcutaneously injected into flank area of 4-week-old male nude mice. The tumor volume (mm3) was measured every 7 days and was calculated using the following formula: volume = width × width × length/2. Mice were sacrificed after 4 weeks, and tumor tissues were excised. All animal experiments were performed with the approval of the Animal Care and Use Committee of The Affiliated Changzhou No 2 People’s Hospital, Nanjing Medical University, and in accordance with the ethical standards of the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health for Animal Welfare Act.
Statistical analysis
Statistically significant differences between breast cancer tissues and adjacent nontumor tissue samples were determined using the Wilcoxon signed-rank test, and differences between cell lines were determined using the Student’s t-test. Cases were divided into two groups, high and low, using the median value of MALAT1 expression as a cutoff. Associations between MALAT1 expression and clinicopathological characteristics were analyzed using Fisher’s exact test. Survival analyses were performed using the Kaplan–Meier method with the log-rank test. P-value <0.05 was considered statistically significant.
Results
MALAT1 is upregulated in breast cancer and associated with disease progression
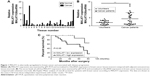
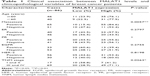
To explore MALAT1 expression levels in breast cancer patients, we used qRT-PCR to measure its expression in breast cancer tissues and the serum of patients with breast cancers. We found that MALAT1 was significantly increased in breast cancer tissues compared with those in adjacent tissues (Figure 1A) and the serum levels of cell-free MALAT1 could significantly differentiate between breast cancer patients and volunteers (Figure 1B). In addition, we analyzed the clinicopathological differences between high and low MALAT1 expression groups. Cases were divided into two groups, high and low, using the median value of MALAT1 expression as a cutoff. As shown in Table 1, MALAT1 expression differed significantly according to metastasis (P<0.01) and TNM stages (P<0.05). No difference was found between MALAT1 expression and other clinical features. Furthermore, we performed Kaplan–Meier curves with log-rank analysis to investigate the association between MALAT1 expression and cancer patient’s overall survival. We found that breast cancer patients with high MALAT1 expression had a significantly shorter survival time compared with low MALAT1 expression (P<0.05, Figure 1C). These data suggested that MALAT1 acted as an oncogene in the development of breast cancer.
MALAT1 induces breast cancer proliferation in vivo and in vitro
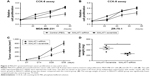
To investigate the role of MALAT1 in breast cancer, we transfected MALAT1-siRNAs into MDA-MB-231 and ZR-75-1 cells. As expected, MALAT1-siRNAs obviously changed the expression of MALAT1 in breast cancer cells (Figure S1). We used MALAT1-siRNA to measure breast cancer cell proliferation. We found that MALAT1-siRNA significantly suppressed cell proliferation in breast cancer (Figure 2A and B). In addition, we subcutaneously injected breast cancer cells with MALAT1-siRNA treated into nude mice. We observed that the tumor growth in MALAT1-siRNA group was slower compared with the control group (Figure 2C). The average weight of tumor in the MALAT1-siRNA group was significantly lower compared with the control group (Figure 2D). These results showed that MALAT1 promoted breast cancer cell proliferation in vivo and in vitro.
MALAT1 is highly expressed and secreted by breast cancer cells
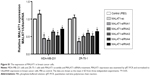
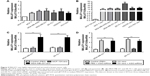
Exosomes have recently been recognized as important mediators of intercellular communication. To identify the exosome-associated RNA inducing proliferation, we used qRT-PCR methods to analyze the expression levels of lncRNAs in tumor cell and exosomes. We found that MALAT1 was significantly highly expressed in breast cancer cells (Figure 3A) and exosomes (Figure 3B).
To evaluate that MDA-MB-231-secreted MALAT1 could be transferred to recipient cells via exosomes, we measured the MALAT1 levels in MDA-MB-231 and ZR-75-1 cells treated with exosomes derived from MCF-10A or MDA-MB-231 cells. The data showed that MALAT1 was highly expressed in recipient MDA-MB-231 and ZR-75-1 cells (Figure 3C). Moreover, to confirm this increase in MALAT1 in recipient cells due to the exosome-mediated lncRNAs transfer, we detected MALAT1 expression in recipient cells with AGO-siRNA (Argonaute protein, core family number protein of RNA-induced silencing complex [RISC]) transfected. We found that MALAT1 was highly expressed in recipient cells in the presence or absence of AGO-siRNA transfection (Figure 3D). These results suggested that MALAT1 was overexpressed in breast cancer exosomes and could be transfected to recipient cells by exosome mediated.
MALAT1 in exosomes derived from cancer cells is involved in cell proliferation
To evaluate the effect of cancer cells secreted exosomes on cell proliferation, we used the MDA-MB-231 metastatic breast cancer cell line, with MALAT1-siRNA treatment and the MCF-10A noncancerous mammary epithelial line as models for studying cancer-secreted exosomes. Exosomes were isolated from culture supernatant using standard differential centrifugations. Breast cancer cells MDA-MB-231 and ZR-75-1 were stimulated with exosomes at the indicated concentrations for 24, 48, and 72 hours. Assessment of proliferation by CCK-8 assay revealed that MDA-MB-231 cancer cell exosomes significantly increased MDA-MB-231 and ZR-75-1 cell proliferations, and MCF-10A exosomes had little effect on tumor cell proliferation (Figure 4A and B).
Furthermore, CCK-8 assay was performed to evaluate the effect of exosome-MALAT1 on regulating cell proliferation. After the inhibition of MALAT1 in cancer cells, the exosomes derived from these cells showed decreased cell proliferation in breast cancer (Figure 4A and B). These data indicated that exosomes derived from cancer cells induced cell proliferation via MALAT1 regulation in breast cancer.
Discussion
Cancer development relies on the interactions between cancer cells and different components of its microenvironment. Recently, exosomes have been implicated as key mediators of intercellular communication within the primary tumor microenvironment, which release the endocytic origin into the extracellular milieu and modify the phenotype of recipient cells.22–24 Previous studies have shown that exosomes contain proteins, miRNAs, mRNAs, DNA fragments, lncRNAs, and various proteins.25–28 The cerebrospinal fluid derived exosomal miR-21 levels correlated with tumor spinal/ventricle metastasis and the recurrence with anatomical site preference, which could be demonstrated as a promising indicator for glioma diagnosis and prognosis, particularly with values to predict tumor recurrence or metastasis.14 Plasma exosomal miR-21 and miR-4257 showed potential as biomarkers of recurrence in NSCLC patients.25 Hypoxic lung cancer cell-derived exosomal miR-23a also inhibited tight junction protein ZO-1, thereby increasing vascular permeability and cancer transendothelial migration.26 Ovarian cancer cell lines secreted the tumor-suppressive miR-940 into the extracellular environment via exosomes to maintain their invasiveness and tumorigenic phenotype.27 Tumor-associated macrophage-derived miR-21 promoted cisplatin resistance in gastric cancer.28 EGFR-containing exosomes derived from gastric cancer cells could favor the development of a liver-like microenvironment promoting liver-specific metastasis.29 Cancer-associated fibroblasts (CAFs)-mediated hepatocellular carcinoma (HCC) progression is partially related to the loss of antitumor miR-320a in the exosomes of CAFs, and that promoting the transfer of stromal cell-derived miR-320a might be a potential treatment option to overcome HCC progression.30
MALAT1 expression levels are dysregulated in a variety of cancers, including HCC,31,32 renal cell carcinoma,33 cholangiocarcinoma,34 pancreatic cancer,35,36 lung cancer,37,38 gastric cancer,39,40 oral squamous cell carcinoma41 and ovarian cancer.42–44 In our previous study, we found that MALAT1 was upregulated in breast cancer tissues and cancer cell lines,45 but MALAT1 interaction with other tumor cells in the tumor microenvironment still needs further study. In the present study, we revealed tumor cells secreted MALAT1 to recipient cells to regulate receptor cell proliferation in tumor microenvironment. We found that MALAT1 was significantly increased in breast cancer tissues compared with adjacent tissues and the serum levels of cell-free MALAT1 could significantly differentiate between breast cancer patients and volunteers. We also found MALAT1 was significantly highly expressed in breast cancer cells and cancer cell exosomes, MDA-MB-231 exosomes significantly increased MDA-MB-231 and ZR-75-1 cell proliferation; however, the exosomes derived from MDA-MB-231 cells treated with MALAT1-siRNA showed decreased cell proliferation in breast cancer.
Conclusion
Our data suggested that exosomes derived from breast cancer could induce cancer cell proliferation. In particular, MALAT1 was secreted by breast cancer cells and could be transfected to recipient cells by cancer cell exosomes. Thus, our findings provided new insights into the mechanism of cell proliferation modulated by exosomal MALAT1 regulation in breast cancer progression and represent a novel strategy for overcoming breast cancer.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (31601156 to TF), the Technology Project of Changzhou Social Development (CE20135044 to TF), the Project of Jiangsu Provincial Medical Youth Talent (QNRC2016267 to TF), and the Projects of Nanjing Medical University Science and Technology Development Fund Key (2016NJMUZD103 to PZ).
Disclosure
The authors report no conflicts of interest in this work.
References
DeSantis CE, Fedewa SA, Goding Sauer A, Kramer JL, Smith RA, Jemal A. Breast cancer statistics, 2015: convergence of incidence rates between black and white women. CA Cancer J Clin. 2016;66(1):31–42. | ||
Donepudi MS, Kondapalli K, Amos SJ, Venkanteshan P. Breast cancer statistics and markers. J Cancer Res Ther. 2014;10(3):506–511. | ||
Raphael J, Gandhi S, Li N, Lu FI, Trudeau M. The role of quantitative estrogen receptor status in predicting tumor response at surgery in breast cancer patients treated with neoadjuvant chemotherapy. Breast Cancer Res Treat. 2017;164(2):285–294. | ||
Jitariu AA, Cimpean AM, Ribatti D, Raica M. Triple negative breast cancer: the kiss of death. Oncotarget. 2017;8(28):46652–46662. | ||
Blackburn HL, Ellsworth DL, Shriver CD, Ellsworth RE. Breast cancer metastasis to the axillary lymph nodes: are changes to the lymph node “Soil” localized or systemic? Breast Cancer (Auckl). 2017;11:1178223417691246. | ||
Forster JC, Harriss-Phillips WM, Douglass MJ, Bezak E. A review of the development of tumor vasculature and its effects on the tumor microenvironment. Hypoxia (Auckl). 2017;5:21–32. | ||
Xie HY, Shao ZM, Li DQ. Tumor microenvironment: driving forces and potential therapeutic targets for breast cancer metastasis. Chin J Cancer. 2017;36(1):36. | ||
Wang M, Zhao J, Zhang L, et al. Role of tumor microenvironment in tumorigenesis. J Cancer. 2017;8(5):761–773. | ||
Rycaj K, Li H, Zhou J, Chen X, Tang DG. Cellular determinants and microenvironmental regulation of prostate cancer metastasis. Semin Cancer Biol. 2017;44:83–97. | ||
Hoshino A, Costa-Silva B, Shen TL, et al. Tumour exosome integrins determine organotropic metastasis. Nature. 2015;527(7578):329–335. | ||
Jia Y, Chen Y, Wang Q, et al. Exosome: emerging biomarker in breast cancer. Oncotarget. 2017;8(25):41717–41733. | ||
Boelens MC, Wu TJ, Nabet BY, et al. Exosome transfer from stromal to breast cancer cells regulates therapy resistance pathways. Cell. 2014;159(3):499–513. | ||
Melo SA, Luecke LB, Kahlert C, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523(7559):177–182. | ||
Shi R, Wang PY, Li XY, et al. Exosomal levels of miRNA-21 from cerebrospinal fluids associated with poor prognosis and tumor recurrence of glioma patients. Oncotarget. 2015;6(29):26971–26981. | ||
Quail DF, Joyce JA. The microenvironmental landscape of brain tumors. Cancer Cell. 2017;31(3):326–341. | ||
Teng Y, Ren Y, Hu X, et al. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat Commun. 2017;8:14448. | ||
King HW, Michael MZ, Gleadle JM. Hypoxic enhancement of exosome release by breast cancer cells. BMC Cancer. 2012;12:421. | ||
Jung KO, Youn H, Lee CH, Kang KW, Chung JK. Visualization of exosome-mediated miR-210 transfer from hypoxic tumor cells. Oncotarget. 2017;8(6):9899–9910. | ||
Sakha S, Muramatsu T, Ueda K, Inazawa J. Exosomal microRNA miR-1246 induces cell motility and invasion through the regulation of DENND2D in oral squamous cell carcinoma. Sci Rep. 2016;6:38750. | ||
Zhou W, Fong MY, Min Y, et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell. 2014;25(4):501–515. | ||
Feng T, Xu D, Tu C, et al. MiR-124 inhibits cell proliferation in breast cancer through downregulation of CDK4. Tumour Biol. 2015;36(8):5987–5997. | ||
Park JE, Tan HS, Datta A, et al. Hypoxic tumor cell modulates its microenvironment to enhance angiogenic and metastatic potential by secretion of proteins and exosomes. Mol Cell Proteomics. 2010;9(6):1085–1099. | ||
Bobrie A, Colombo M, Raposo G, Thery C. Exosome secretion: molecular mechanisms and roles in immune responses. Traffic. 2011;12(12):1659–1668. | ||
Ge R, Tan E, Sharghi-Namini S, Asada HH. Exosomes in cancer microenvironment and beyond: have we overlooked these extracellular messengers? Cancer Microenviron. 2012;5(3):323–332. | ||
Dejima H, Iinuma H, Kanaoka R, Matsutani N, Kawamura M. Exosomal microRNA in plasma as a non-invasive biomarker for the recurrence of non-small cell lung cancer. Oncol Lett. 2017;13(3):1256–1263. | ||
Hsu YL, Hung JY, Chang WA, et al. Hypoxic lung cancer-secreted exosomal miR-23a increased angiogenesis and vascular permeability by targeting prolyl hydroxylase and tight junction protein ZO-1. Oncogene. 2017;36(34):4929–4942. | ||
Rashed MH, Kanlikilicer P, Rodriguez-Aguayo C, et al. Exosomal miR-940 maintains SRC-mediated oncogenic activity in cancer cells: a possible role for exosomal disposal of tumor suppressor miRNAs. Oncotarget. 2017;8(12):20145–20164. | ||
Zheng P, Chen L, Yuan X, et al. Exosomal transfer of tumor-associated macrophage-derived miR-21 confers cisplatin resistance in gastric cancer cells. J Exp Clin Cancer Res. 2017;36(1):53. | ||
Zhang H, Deng T, Liu R, et al. Exosome-delivered EGFR regulates liver microenvironment to promote gastric cancer liver metastasis. Nat Commun. 2017;8:15016. | ||
Zhang Z, Li X, Sun W, et al. Loss of exosomal miR-320a from cancer-associated fibroblasts contributes to HCC proliferation and metastasis. Cancer Lett. 2017;397:33–42. | ||
Hou Z, Xu X, Zhou L, et al. The long non-coding RNA MALAT1 promotes the migration and invasion of hepatocellular carcinoma by sponging miR-204 and releasing SIRT1. Tumour Biol. 2017;39(7):1010428317718135. | ||
Luo F, Sun B, Li H, et al. A MALAT1/HIF-2alpha feedback loop contributes to arsenite carcinogenesis. Oncotarget. 2016;7(5):5769–5787. | ||
Zhang HM, Yang FQ, Chen SJ, Che J, Zheng JH. Upregulation of long non-coding RNA MALAT1 correlates with tumor progression and poor prognosis in clear cell renal cell carcinoma. Tumour Biol. 2015;36(4):2947–2955. | ||
Wang C, Mao ZP, Wang L, et al. Long non-coding RNA MALAT1 promotes cholangiocarcinoma cell proliferation and invasion by activating PI3K/Akt pathway. Neoplasma. 2017;64(5):725–731. | ||
Zhang Y, Tang X, Shi M, Wen C, Shen B. MiR-216a decreases MALAT1 expression, induces G2/M arrest and apoptosis in pancreatic cancer cells. Biochem Biophys Res Commun. 2017;483(2):816–822. | ||
Pang EJ, Yang R, Fu XB, Liu YF. Overexpression of long non-coding RNA MALAT1 is correlated with clinical progression and unfavorable prognosis in pancreatic cancer. Tumour Biol. 2015;36(4):2403–2407. | ||
Jen J, Tang YA, Lu YH, Lin CC, Lai WW, Wang YC. Oct4 transcriptionally regulates the expression of long non-coding RNAs NEAT1 and MALAT1 to promote lung cancer progression. Mol Cancer. 2017;16(1):104. | ||
Liu M, Sun W, Liu Y, Dong X. The role of lncRNA MALAT1 in bone metastasis in patients with non-small cell lung cancer. Oncol Rep. 2016;36(3):1679–1685. | ||
Zhang Y, Chen Z, Li MJ, Guo HY, Jing NC. Long non-coding RNA metastasis-associated lung adenocarcinoma transcript 1 regulates the expression of Gli2 by miR-202 to strengthen gastric cancer progression. Biomed Pharmacother. 2017;85:264–271. | ||
Qi Y, Ooi HS, Wu J, et al. MALAT1 long ncRNA promotes gastric cancer metastasis by suppressing PCDH10. Oncotarget. 2016;7(11):12693–12703. | ||
Zhou X, Liu S, Cai G, et al. Long non coding RNA MALAT1 promotes tumor growth and metastasis by inducing epithelial-mesenchymal transition in oral squamous cell carcinoma. Sci Rep. 2015;5:15972. | ||
Wu L, Wang X, Guo Y. Long non-coding RNA MALAT1 is upregulated and involved in cell proliferation, migration and apoptosis in ovarian cancer. Exp Ther Med. 2017;13(6):3055–3060. | ||
Lei R, Xue M, Zhang L, Lin Z. Long noncoding RNA MALAT1-regulated microRNA 506 modulates ovarian cancer growth by targeting iASPP. Onco Targets Ther. 2017;10:35–46. | ||
Jin Y, Feng SJ, Qiu S, Shao N, Zheng JH. LncRNA MALAT1 promotes proliferation and metastasis in epithelial ovarian cancer via the PI3K-AKT pathway. Eur Rev Med Pharmacol Sci. 2017;21(14):3176–3184. | ||
Feng T, Shao F, Wu Q, et al. miR-124 downregulation leads to breast cancer progression via LncRNA-MALAT1 regulation and CDK4/E2F1 signal activation. Oncotarget. 2016;7(13):16205–16216. |
Supplementary materials
  | Table S1 Primers for qRT-PCR |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.