Back to Journals » International Journal of Nanomedicine » Volume 15
Exosome Derived from Coronary Serum of Patients with Myocardial Infarction Promotes Angiogenesis Through the miRNA-143/IGF-IR Pathway
Authors Geng T, Song ZY, Xing JX, Wang BX, Dai SP, Xu ZS
Received 18 December 2019
Accepted for publication 6 April 2020
Published 21 April 2020 Volume 2020:15 Pages 2647—2658
DOI https://doi.org/10.2147/IJN.S242908
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Mian Wang
Tao Geng, Zhi-Yuan Song, Jing-Xian Xing, Bing-Xun Wang, Shi-Peng Dai, Ze-Sheng Xu
Department of Cardiovascular Disease, Cangzhou Central Hospital of Tianjin Medical University, Cangzhou, Hebei Province, People’s Republic of China
Correspondence: Ze-Sheng Xu
Department of Cardiovascular Disease, Cangzhou Central Hospital of Tianjin Medical University, No. 16, Xinhua West Road, Cangzhou, Hebei Province, People’s Republic of China
Tel +86-15003172868
Email [email protected]
Purpose: Myocardial ischemia-reperfusion injury primarily causes myocardial infarction (MI), which is manifested by cell death. Angiogenesis is essential for repair and regeneration in cardiac tissue after MI. In this study, we aimed to investigate the effect of exosomes derived from the serum of MI patients in angiogenesis and its related mechanism.
Patients and Methods: Exosomes, isolated from serum, were collected from MI (MI-exosome) and control (Con-exosome) patients. After coculturing with human umbilical vein endothelial cells, MI-exosome promoted cell proliferation, migration, and tube formation.
Results: The results revealed that the production and release of MI-exosome were associated with cardiomyocytes. Moreover, microarray assays demonstrated that miRNA-143 was significantly decreased in MI-exosome. Meanwhile, the overexpression and knockdown of miRNA-143 could inhibit and enhance angiogenesis, respectively. Furthermore, the effect of exosomal miRNA-143 on angiogenesis was mediated by its targeting gene, insulin-like growth factor 1 receptor (IGF-IR), and was associated with the production of nitric oxide (NO).
Conclusion: Taken together, exosomes derived from the serum of patients with MI promoted angiogenesis through the IGF-IR/NO signaling pathway. The results provide novel understanding of the function of exosomes in MI.
Keywords: myocardial ischemia, exosome, miRNA-143, insulin-like growth factor 1 receptor, angiogenesis
Introduction
Myocardial ischemia-reperfusion (IR) injury is the leading cause of various devastating cardiovascular diseases all over the world,1 and has emerged as the major cause of mortality and morbidity. As one of the primary outcomes of ischemic injury, myocardial infarction (MI) is manifested by aberrant myocardial cell death due to the imbalance of blood supply-demand.2,3 Also, IR injury leads to the coronary artery narrowing and exacerbating the deficiency of oxygen supply to myocardial cells.4 Currently, percutaneous coronary intervention (PCI) and coronary artery bypass grafting (CABG) have been widely applied to treat symptomatic patients with coronary artery disease and significantly restores the blood flow after IR injury.5,6 However, the pathological changes of injured myocardium have not fully ameliorated since the injured microvascular architecture has not been repaired or rebuilt. Thus, exploring the mechanisms and approaches that trigger microvascular regeneration would greatly contribute to patients with coronary artery disease.
It has been demonstrated that interrupted architecture of microvasculature, including aberrant apoptosis and angiogenesis, is an essential factor for pathogenic progression of multiple IR-initiated cardiovascular diseases,7 thus angiogenesis is regarded as an important process to recreate collateral circulation in ischemic myocardium.8 Moreover, microRNAs (miRNAs) are non-coding, single-stranded RNAs, about 18–24 nucleotides in length.9 In addition, miRNAs are ubiquitously expressed in all cell types and are involved in various cellular processes through suppressing target mRNA translation or enhancing degradation.9 Thus, miRNAs function as key regulators in diverse biological and pathological processes, including angiogenesis.10,11 The pharmacological knockdown of miRNA-92a elevates capillary regeneration and improves heart function in MI mouse models.12 The inhibition of miRNA-24 can promote the recovery of MI mice through downregulating the endothelial transcription factor, GATA-2, and endothelial nitric oxide (NO) synthase.13 Collectively, the therapeutic potential of miRNAs should be further investigated in treating coronary artery disease.
Exosomes are small membrane microvesicles (70–120 nm) of endocytic origin, produced by the majority of cell types,14 and participates in intercellular communication through transferring intracellular cargoes, including enzymes, DNA, and non-coding RNAs.15 In the past decade, exosomes have emerged as therapeutic approaches or biomarkers for cardiovascular diseases.16 Exosomes can mediate the pro-angiogenic role of stem cells to promote angiogenesis through paracrine action.17 Exosomes derived from mesenchymal stromal cells promote angiogenesis in MI mice and repair MI-injured myocardium.18 Also, miRNAs and other functioning molecules can be transported by exosomes to play important roles in cardio-protection, such as the regeneration of blood vessels and inhibition of inflammation.19 However, the role of exosomes derived from IR-injured cardiac cells in angiogenesis is still undetermined.
It has been reported that cardiomyocytes of mice with ischemic injury may generate cell-dependent exosomes to the blood circulation system.20 Thus, we collected human coronary blood from the aortic sinus and isolated exosomes. Then, we aimed to investigate the function of exosomes derived from the coronary serum of patients with MI and determine their effect on angiogenesis and related miRNA-related mechanism. Meanwhile, we reported that cardiomyocytes may be a major origin of exosomes that could be transported to endothelial cells. In addition, exosomes derived from MI patients had lower expression of miRNA-143 and could promote angiogenesis of endothelial cells via IGF-IR/NO signaling pathway.
Patients and Methods
Patients
All patients and control subjects participated in this study were informed and gave written consent. The study protocols were approved by the Ethics Committee of Cangzhou Central Hospital and conformed to the Declaration of Helsinki. Suspected MI patients with electrocardiogram diagonal evidence and chest pain and underwent diagnostic cardiac catheterization in Cangzhou Central Hospital between September 2017 to May 2018 were enrolled in this study. The exclusion and inclusion criteria were based on previously described.21,22 Briefly, the patients with following characteristics were excluded from this study, including: (1) poorly controlled blood pressure; (2) hyperlipidemia (total triglyceride > 2.26 mM); (3) diabetes (fasting glucose > 7.0 mM); (4) infectious diseases; (5) other contraindications, for example, hepatic or nephritic conditions. By the angiography analysis, patients with more than 70% stenosis were classified as experimental group (MI). Patients without stenosis or with less than 50% stenosis were classified as the control group.
Exosome Isolation
Exosomes were isolated from the serum of MI and control patients. 10 mL blood samples were collected from the aortic sinus through cordis 6F catheter (Johnson & Johnson, USA) into the centrifuge tubes. Then blood samples were centrifuged at 2400 g for 10 minutes at 4 °C and then the supernatants were collected and centrifuged at 800 g for 10 minutes at 4 °C. Afterward, the purified serum samples were used to isolate exosomes using the ultracentrifugation method as previously described.23,24 The concentration of exosomes was measured using the Pierce™ BCA Protein Assay Kit (Thermo Fisher Scientific, USA) according to the manufacturer’s instructions.
Transmission Electron Microscopy
Transmission electron microscopy (TEM) was performed to determine the morphology of exosomes as previously described.25,26 Fixed exosomes were photographed by JEOL JEM-1400 transmission electron microscope (JEOL USA, Inc.).
Western Blots
Total protein of exosomes, cells, and cardiac tissues were isolated using the RIPA Lysis Buffer (BosterBio, USA). Western blots were performed as previously described.27 The primary antibodies used in this study were as following: Insulin-like growth factor 1 receptor (IGF-IR) (1:500), CD9 (1:500), CD63 (1: 500), Flotillin (1:1000), Tsg101 (1:1000), Rab11a (1:1000), and β-actin (1:5000) (Santa Cruz, Shanghai, China). The protein band was imaged on a Bio-Rad Chemidoc system (Bio-Rad Laboratories, USA).
Quantitative Real-Time PCR (qRT-PCR)
Total RNAs were isolated by TRIzol reagent (Invitrogen, USA) and first-strand cDNAs were synthesized using the PrimeScript RT reagent kit (TaKaRa, USA) according to the manufacturer’s instructions. MiRNA-143 primers were synthesized from QIAGEN (QIAGEN, Germany). The primers used in this study were as following: IGF-IR: 5′-ACAGAGTACCCTTTCTTT-3′ (F) and 5′-AAGAACACAGGATCTGTCCACGAC-3′ (R); β-actin: 5′-AGGCATCCTCACCCTGAAGTA-3′ (F) and 5′- CACACGCAGCTCATTGTAGA-3′ (R). The PCR reactions were performed using the SYBR Premix Ex Taq II kit (TaKaRa, USA) on a 7500 Real-Time PCR System (Applied Biosystems, USA). The relative expressions were analyzed using the 2−ΔΔCt method.28
Cell Culture
Mouse cardiomyocytes (CMs), cardiac endothelial cells (CEs), cardiac fibroblasts (CFs), and human umbilical vein endothelial cells (HUVECs) were obtained from Sigma-Aldrich. Cell lines and CMs were cultured in Dulbecco’s Modified Eagle Medium (Gibco, USA) and supplemented with 10% Fetal Bovine Serum. The CFs were cultured in Endothelial Cell Growth Medium-2 (Lonza, USA). Cells were incubated in a humidified incubator at 37°C and 5% CO2 (Thermo Scientific, USA). To mimic MI, cells were treated as previously described.29
Confocal Imaging
To visualize the internalization of exosomes in HUVECs, exosomes were stained with PKH26 (Sigma-Aldrich, USA) according to the manufacturer’s protocol. The HUVECs were stained with Phalloidin and DAPI (Sigma-Aldrich, USA) according to the manufacturer’s protocol. Then, exosomes were cocultured with HUVECs for 8 hours and imaged using the Confocal Laser Scanning Microscope (Leica, Germany).
Cell Transfection
The HUVECs were transfected with miRNA-143 mimic, inhibitor, and negative control (Sigma-Aldrich, USA) using the Lipofectamine™ 3000 Transfection Reagent (Invitrogen, Waltham, MA, USA) according to the manufacturer’s instructions.
Luciferase Assay
Luciferase vectors containing wild-type and mutated binding sites of miR-143 in 3ʹUTR of IGF-IR were transfected into HUVECs using the Lipofectamine™ 3000 (Invitrogen, USA) according to the manufacturer’s instructions. After 24 hours’ transfection, relative luciferase activity was measured using the Pierce™ Firefly Luciferase Glow Assay Kit (Thermo Scientific, USA).
Cell Proliferation, Migration, and Tube Formation
Cell proliferation profile of HUVECs was determined using the 96,992 Cell Counting Kit (CCK-8) (Sigma-Aldrich, USA). The OD value was detected using the spectrophotometric microplate reader at 450 nm (BMG Labtech, Germany). The migration ability of HUVECs was evaluated by the Transwell assay (Radius Cell Migration Assays Cell Biolabs, Inc, USA) and scratch wound assay (CytoSelect™ 24-Well Wound Healing Assay, Cell Biolabs, Inc, USA) according to the manufacturer’s instructions. Tube formation was determined using µ-Slide Angiogenesis (ibidi GmbH, Germany) according to the manufacturer’s instructions.
Microarray
Total RNA was isolated from exosomes using TRIzol reagent (Invitrogen, USA). The RNA quality was determined by ND-2000 NanoDrop (Thermo Fisher Scientific, USA). The adapter ligation, cDNA synthesis, PCR, and RNA libraries construction were performed by TruSeq Stranded Total RNA Library Prep Kit (Illumina, USA) according to the manufacturer’s instructions. Microarray assay was performed as Previously described.30,31 Next, the samples were sent and processed in BGI Company (Beijing, China). Briefly, SurePrint Human miRNA Microarrays kit (Agilent, USA) was applied for labeling and hybridization according to the manufacturer’s instructions. The miRNA expression profile was analyzed by using the Agilent Human 8×60 K. The experiments had at least 3 repeats. The differential expression of miRNAs was defined as those with larger or less than 2-fold change meanwhile P < 0.05. Raw data were normalized by the Quantile algorithm in R software. Statistic differences were calculated using the t-test.
Nitric Oxide Level
The NO level of HUVECs was determined using the Nitric Oxide Assay Kit (Abcam plc. USA) according to the manufacturer’s instructions.
Animal Model
All experimental procedures were approved by the Institutional Animal Care and Use Committee of Cangzhou Central Hospital. All animal experiments were performed according to the guidelines of the local regulatory agencies and conformed to the Regulations for the Management of Laboratory Animals published by the Ministry of Science and Technology of the People’s Republic of China. Male nude C57BL/6 mice aged 10–12 weeks (Jackson Laboratory, USA) were housed in a pathogen-free environment and had free access to food and water. The MI mouse model was established as previously described.32 The MI-injured zones were collected 3 days after MI treatment.
Statistical Analyses
Statistical analysis was performed using SPSS v.19.0 software (SPSS, IL, USA). Values were presented as mean ± SD. Each experimental group had at least three replicates. Pearson correlation was used to calculate the correlation between miRNA-143 and IGF-IR. Differences between groups were analyzed with Student’s t-test. P < 0.05 was regarded as statistically significant.
Results
Serum-Derived Exosome Isolation and Identification
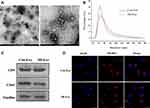
Exosomes were isolated from 14 MI (MI-exosome) and 10 control (Con-exosome) patients. The information on all subjects was summarized in Table 1. The morphology of exosomes was assessed by TEM and exosomes displayed spherical shape (Figure 1A). Nanosight analysis revealed that the diameter distribution of exosomes ranged from 30 to 100 nm (Figure 1B). Next, the exosome markers, CD9, CD63, and Flotillin were detected using Western blots and the results suggested that all three exosome markers were positively expressed in exosomes (Figure 1C). Together, these results demonstrated that the isolated microvesicles from the serum of MI patients were exosomes. Next, to determine whether exosomes could be taken up by HUVECs, exosomes were cocultured with HUVECs for 8 hours. Under the confocal microscope, PKH26-labeled exosomes were found to overlap with DAPI-labeled HUVECs, suggesting exosomes could be successfully internalized by HUVECs (Figure 1D).
 |
Table 1 Characteristics of Patients |
MI-Exosome Promotes Cell Proliferation, Tube Formation, and Migration in HUVECs
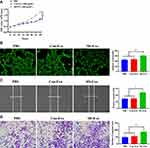
To determine the effect of MI-exosome on HUVECs, CCK-8, tube formation, scratch wound, and Transwell assays were performed in HUVECs. In the CCK-8 assay, MI-exosome significantly increased the proliferative capability of HUVECs compared with Con-exosome (Figure 2A). Next, MI-exosome increased a greater number of vascular branches in HUVECs, relative to Con-exosome, indicating a pro-angiogenetic role of MI-exosome (Figure 2B). In addition, MI-exosome could elevate the migration ability in HUVECs while Con-exosome did not affect HUVECs (Figure 2C and D). Together, exosomes that originated from the coronary serum of MI patients may exert a positive role in angiogenesis of HUVECs.
CMs May Be the Main Origin of MI-Exosome
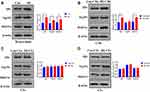
To investigate the origin of MI-exosome, we performed Western blots to assess exosome biogenesis markers, Alix, Tsg101, and Rab11a,33,34 in the myocardium and three types of cardiac cells, including cardiomyocytes, endothelial cells, and fibroblasts. In mouse MI myocardium, all three markers were increased compared with those of the control group (Figure 3A). Also, the expressions of Alix and Rab11a were upregulated in MI-CMs, indicating that CMs may be a primary site to produce exosomes. (Figure 3B). On the other hand, the expressions of three exosome biogenesis markers were not affected by MI in both CEs and CFs (Figure 3C and D). These results indicated that CMs might be the main origin of exosomes in MI.
Exosomal miRNA-143 Is Decreased in MI-Exosome and Promotes Angiogenesis
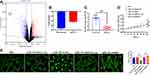
ZTo investigate the potential miRNA that may mediate the role of exosomes in HUVEC angiogenesis, a microarray analysis was performed to determine the miRNA profile in both MI- and Con-exosome. Relatively, miRNA-143 displayed the greatest downregulated level in MI-exosome as shown in the volcano plot (Figure 4A, Table 2), which was also verified by qRT-PCR (Figure 4B), suggesting a potential role of miRNA-143 in the regulation of angiogenesis. In exosomes from all subjects, the results showed that the expression of miRNA-143 was decreased in MI-exosome compared with those of Con-exosome (Figure 4C). To further determine the function of miRNA-143 in HUVECs, the overexpression and knockdown of miRNA-143 was performed via transfecting HUVECs with miRNA-143 mimic and inhibitor, respectively. In CCK-8 and tube formation assays, the overexpression of miRNA-143 significantly inhibited cell proliferation and the ability of tube formation in HUVECs, while the knockdown of miRNA exerted the opposite role (Figure 4D and E). Together, the upregulation of miRNA-143 could suppress proliferation and vascular formation in HUVECs, indicating an anti-angiogenesis role of miRNA-143.
 |
Table 2 Differentially Expressed miRNAs |
MiRNA-143 Directly Targets IGF-IR and Promotes NO Production
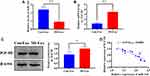
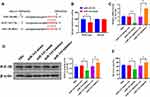
To determine the downstream targeting gene of miRNA-143, we performed bioinformatic analysis to predict the potential target gene of miRNA-143. The results revealed that IGF-IR contained a putative binding site of miRNA-143 in 3ʹUTR (Figure 5A). The luciferase reporter assay revealed that the relative luciferase activity was decreased in HUVECs carrying wild-type 3ʹUTR of IGF-IR compared with those containing mutant 3ʹUTR (Figure 5B), suggesting miRNA-143 could directly target IGF-IR. Furthermore, overexpression and knockdown of miRNA-143 could reduce and elevate the protein and mRNA level of IGF-IR, respectively (Figure 5C and D). Meanwhile, the upregulation of miRNA-143 was associated with reduced NO production (Figure 5E). Therefore, these results suggested that the effect of exosomal miRNA-143 on angiogenesis may be mediated by IGF-IR signaling and NO activity.
Exosomal miRNA-143 Activates IGF-IR Signaling in HUVECs
To determine whether IGF-IR signaling could be activated by exosomal miRNA-143, HUVECs were cocultured with MI-exosome. Compared with Con-exosome, the expression of miRNA-143 was decreased in HUVECs cocultured with MI-exosome (Figure 6A), meanwhile, both mRNA and protein expression of IGF-IR were increased by MI-exosome (Figure 6B and C). Furthermore, a negative correlation between miRNA-143 and IGF-IR was revealed by Pearson correlation analysis (Figure 6D). Together, these results suggested that downregulated miRNA-143 could activate IGF-IR signaling and promote angiogenesis.
Discussion
In the present study, the function of serum-derived exosomal miRNA in angiogenesis was elucidated, providing a novel strategy for the treatment of MI injury. Specifically, exosomes isolated from human coronary blood exerted a pro-angiogenic role in CEs and the origin of these exosomes were primarily from ischemic CMs. Also, the effect of exosomes on CEs was mediated through the downregulation of miRNA-143, which was associated with the IGF-IR/NO pathway.
Currently, previous studies have investigated the functions of exosomes and exosome-carrying intracellular cargoes in various biological and pathological processes,35,36 the most of exosomes, however, are derived from human-cultured cells or animal models, including rats and mice,37 suggesting that there may be limits on the translational value of these studies. Therefore, investigating the role of exosomes obtained from human samples may be an optional solution for this barrier. In this study, we isolated exosomes from coronary blood of both MI and control patients. Relatively, MI-derived exosomes played a significant role in promoting CEs proliferation, abilities of tube formation, as well as migration. Furthermore, our study demonstrated that CMs, but not CEs and CFs, may be an origin of functioning exosomes. Wang et al reported that exosomes secreted from CMs transport miRNA-320 to endothelial cells to inhibit angiogenesis in type 2 diabetic rats.38 Also, exosomal miRNA-30a functions as an important intercellular communication mediator between CMs to regulate autophagy, in response to the hypoxic condition.39 Collectively, CMs are an essential origin of exosomes that play a critical role in the regulation of MI-initiated cellular activities.
To further determine the mechanism underlying the action of MI-associated exosomes in angiogenesis, the microarray assay was performed to investigate functioning miRNAs. Among several candidate miRNAs, miRNA-143 showed the greatest downregulation in MI-exosomes. In the subsequent experiments, the overexpression of miRNA-143 inhibited CEs proliferation and tube formation, whereas the knockdown of miRNA-143 exerted an opposite role in angiogenesis. Moreover, miRNA-143 has been reported to be associated with the angiogenic process in endothelial dysfunction in preeclampsia.40 Also, exosome-transferring miRNA-143 from pulmonary artery smooth muscle cells can be internalized and play a pro-angiogenic role in pulmonary arterial endothelial cells.40 In colorectal cancer, the upregulation of miRNA-143 exerts an inhibitory role in angiogenesis, proliferation, and migration through targeting IGF-IR. Together, these observations revealed the importance of miRNA-143 in angiogenesis in various tissues and cell types.
MiRNAs play important roles in the post-transcriptional regulation of target genes, primarily through binding to the complementary sequences in the 3′ untranslated regions (UTR).41 By combining with bioinformatic analysis and previous studies, IGF-IR was a promising candidate for the targeting gene miRNA-143. By verifying through the luciferase reporter assay, the results showed that miRNA-143 could directly target IGF-IR. Also, the expression of miRNA-143 was negatively correlated with the protein level of IGF-IR. The insulin‐like growth factor (IGF) signaling is essential in diverse physiological processes, including growth, development, and metabolism.42 As one of the main receptors in the IGF system, IGF-IR is activated by autophosphorylation and thereby results in the activation of phosphatidylinositol 3‐kinase (PI3K) and mitogen activated protein kinase (MAPK) signaling.43 Functionally, IGF-IR has been found to play an essential role in angiogenesis. For example, similar to the present study, miRNA-143 can inhibit tumor growth and angiogenesis by targeting IGF-IR in colorectal cancer.44 Also, the overexpression of miR-148a/152 exerts a negative role in cell proliferation and angiogenesis through targeting IGF-IR and insulin receptor substrate 1 (IRS1) in breast cancer.45 Furthermore, the suppression of IGF-IR via NVP-AEW541, a kinase inhibitor, inhibits tumor growth and angiogenesis in pancreatic cancer.46 As such, IGF-IR may function as a signaling hub to mediate angiogenic processes in various tissues, providing a potential biomarker to regulate angiogenesis.
In the present study, the effect of miRNA-143 was associated with the production of NO. Additionally, NO is synthesized by the endothelium and plays a wide range of roles in various biological processes, such as maintaining vascular homeostasis, protection of injured vessels, and regulation of cell growth.47 As a multifunctional factor, NO-related endothelial dysfunction is a notable characteristic of cardiovascular diseases.48 In cardiac endothelial cells, NO has been demonstrated to be involved in promoting cell proliferation and migration, while inhibiting apoptosis.49 In this study, the downregulation of miRNA-143 caused a higher level of NO production, along with increased angiogenesis, indicating NO may exert a cardioprotective effect on CEs in MI, which also extend our understanding for MI.
On the other hand, it should be pointed out that there are still some limitations in this study. For example, due to our relatively small sample size, the findings of this study might not elucidate all functional miRNAs and associated signaling pathways. Also, due to the complexity of clinical characteristics, the exclusion and inclusion criteria used in this study also might not cover all clinical scenarios. Thus, the larger sample size and detailed inclusion criteria should be taken into account in future study. Meanwhile, in this study we only investigated the role of MI-exosome in HUVECs, but not in cardiac cells, such as CM, CF. Therefore, it is worthwhile to further explore the function of MI-exosome in the cardiac tissues and cells. Furthermore, the function of MI-exosome should also be determined in vivo study.
Conclusion
In conclusion, the results suggest that exosomal miRNA-143 derived from the serum of MI patients can promote angiogenesis in CEs through targeting IGF-IR and enhancing NO production. This study may provide a novel understanding of the function of exosomal miRNAs in MI.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Benjamin EJ, Blaha MJ, Chiuve SE, et al. Heart disease and stroke statistics-2017 update: a report from the American Heart Association. Circulation. 2017;135(10):e146–e603.
2. Thygesen K, Alpert JS, White HD. Universal definition of myocardial infarction. J Am Coll Cardiol. 2007;50(22):2173–2195. doi:10.1016/j.jacc.2007.09.011
3. Frangogiannis NG. Pathophysiology of myocardial infarction. Compr Physiol. 2011;5(4):1841–1875.
4. Gao E, Lei YH, Shang X, et al. A novel and efficient model of coronary artery ligation and myocardial infarction in the mouse. Circ Res. 2010;107(12):1445–1453. doi:10.1161/CIRCRESAHA.110.223925
5. Thuijs DJ, Kappetein AP, Serruys PW, et al. Percutaneous coronary intervention versus coronary artery bypass grafting in patients with three-vessel or left main coronary artery disease: 10-year follow-up of the multicentre randomised controlled SYNTAX trial. Lancet. 2019;394(10206):1325–1334. doi:10.1016/S0140-6736(19)31997-X
6. Head SJ, Milojevic M, Daemen J, et al. Mortality after coronary artery bypass grafting versus percutaneous coronary intervention with stenting for coronary artery disease: a pooled analysis of individual patient data. Lancet. 2018;391(10124):939–948. doi:10.1016/S0140-6736(18)30423-9
7. Shimizu T, Okamoto H, Chiba S, et al. VEGF-mediated angiogenesis is impaired by angiotensin type 1 receptor blockade in cardiomyopathic hamster hearts. Cardiovasc Res. 2003;58(1):203–212. doi:10.1016/S0008-6363(02)00843-X
8. Masumura M, Watanabe R, Nagashima A, et al. Anti-salusin-β antibody enhances angiogenesis after myocardial ischemia reperfusion injury. Expert Opin Ther Targets. 2013;17(9):1003–1009. doi:10.1517/14728222.2013.819852
9. Fabbri M, Croce CM, Calin GA. MicroRNAs. Cancer J. 2008;14(1):1–6. doi:10.1097/PPO.0b013e318164145e
10. Ye Y, Perez-Polo JR, Qian J, Birnbaum Y. The role of microRNA in modulating myocardial ischemia-reperfusion injury. Physiol Genomics. 2010;43(10):534–542. doi:10.1152/physiolgenomics.00130.2010
11. Fan Z-X, Yang J. The role of microRNAs in regulating myocardial ischemia reperfusion injury. Saudi Med J. 2015;36(7):787. doi:10.15537/smj.2015.7.11089
12. Bonauer A, Carmona G, Iwasaki M, et al. MicroRNA-92a controls angiogenesis and functional recovery of ischemic tissues in mice. Science. 2009;324(5935):1710–1713. doi:10.1126/science.1174381
13. Meloni M, Marchetti M, Garner K, et al. Local inhibition of microRNA-24 improves reparative angiogenesis and left ventricle remodeling and function in mice with myocardial infarction. Mol Ther. 2013;21(7):1390–1402. doi:10.1038/mt.2013.89
14. Théry C. Exosomes: secreted vesicles and intercellular communications. F1000 Biol Rep. 2011;3:15. doi:10.3410/B3-15
15. Melo SA, Sugimoto H, O’Connell JT, et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 2014;26(5):707–721. doi:10.1016/j.ccell.2014.09.005
16. Yellon DM, Davidson SM. Exosomes: nanoparticles involved in cardioprotection? Circ Res. 2014;114(2):325–332. doi:10.1161/CIRCRESAHA.113.300636
17. Sahoo S, Klychko E, Thorne T, et al. Exosomes from human CD34+ stem cells mediate their proangiogenic paracrine activity. Circ Res. 2011;109(7):724–728. doi:10.1161/CIRCRESAHA.111.253286
18. Lee C, Mitsialis SA, Aslam M, et al. Exosomes mediate the cytoprotective action of mesenchymal stromal cells on hypoxia-induced pulmonary hypertension. Circulation. 2012;126(22):2601–2611. doi:10.1161/CIRCULATIONAHA.112.114173
19. Teng X, Chen L, Chen W, Yang J, Yang Z, Shen Z. Mesenchymal stem cell-derived exosomes improve the microenvironment of infarcted myocardium contributing to angiogenesis and anti-inflammation. Cell Physiol Biochem. 2015;37(6):2415–2424. doi:10.1159/000438594
20. Zhang X, Wang X, Zhu H, et al. Hsp20 functions as a novel cardiokine in promoting angiogenesis via activation of VEGFR2. PLoS One. 2012;7(3):e32765. doi:10.1371/journal.pone.0032765
21. Lunde K, Solheim S, Aakhus S, et al. Intracoronary injection of mononuclear bone marrow cells in acute myocardial infarction. N Engl J Med. 2006;355(12):1199–1209. doi:10.1056/NEJMoa05706
22. Schächinger V, Assmus B, Britten MB, et al. Transplantation of progenitor cells and regeneration enhancement in acute myocardial infarction: final one-year results of the TOPCARE-AMI trial. J Am Coll Cardiol. 2004;44(8):1690–1699. doi:10.1016/j.jacc.2004.08.014
23. Rekker K, Saare M, Roost AM, et al. Comparison of serum exosome isolation methods for microRNA profiling. Clin Biochem. 2014;47(1–2):135–138. doi:10.1016/j.clinbiochem.2013.10.020
24. Wang H, Hou L, Li A, Duan Y, Gao H, Song X. Expression of serum exosomal microRNA-21 in human hepatocellular carcinoma. Biomed Res Int. 2014;2014.
25. Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol. 2008;110(1):13–21. doi:10.1016/j.ygyno.2008.04.033
26. Emanueli C, Shearn AI, Angelini GD, Sahoo S. Exosomes and exosomal miRNAs in cardiovascular protection and repair. Vascul Pharmacol. 2015;71:24–30. doi:10.1016/j.vph.2015.02.008
27. Pulito C, Mori F, Sacconi A, et al. Metformin-induced ablation of microRNA 21-5p releases sestrin-1 and CAB39L antitumoral activities. CELL DISCOV. 2017;3:17022. doi:10.1038/celldisc.2017.22
28. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
29. Yu X, Deng L, Wang D, et al. Mechanism of TNF-α autocrine effects in hypoxic cardiomyocytes: initiated by hypoxia inducible factor 1α, presented by exosomes. J Mol Cell Cardiol. 2012;53(6):848–857. doi:10.1016/j.yjmcc.2012.10.002
30. Izumi H, Tsuda M, Sato Y, et al. Bovine milk exosomes contain microRNA and mRNA and are taken up by human macrophages. J Dairy Sci. 2015;98(5):2920–2933. doi:10.3168/jds.2014-9076
31. Qin X, Yu S, Zhou L, et al. Cisplatin-resistant lung cancer cell–derived exosomes increase cisplatin resistance of recipient cells in exosomal miR-100–5p-dependent manner. Int J Nanomedicine. 2017;12:3721. doi:10.2147/IJN.S131516
32. Tang YL, Zhao Q, Qin X, et al. Paracrine action enhances the effects of autologous mesenchymal stem cell transplantation on vascular regeneration in rat model of myocardial infarction. Ann Thorac Surg. 2005;80(1):229–237. doi:10.1016/j.athoracsur.2005.02.072
33. Kowal J, Tkach M, Théry C. Biogenesis and secretion of exosomes. Curr Opin Cell Biol. 2014;29:116–125. doi:10.1016/j.ceb.2014.05.004
34. Février B, Raposo G. Exosomes: endosomal-derived vesicles shipping extracellular messages. Curr Opin Cell Biol. 2004;16(4):415–421. doi:10.1016/j.ceb.2004.06.003
35. Das CK, Jena BC, Banerjee I, et al. Exosome as a novel shuttle for delivery of therapeutics across biological barriers. Mol Pharm. 2018;16(1):24–40. doi:10.1021/acs.molpharmaceut.8b00901
36. Hood JL, Wickline SA. A systematic approach to exosome-based translational nanomedicine. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2012;4(4):458–467. doi:10.1002/wnan.1174
37. Hu G, Drescher KM, Chen X. Exosomal miRNAs: biological properties and therapeutic potential. Front Genet. 2012;3:56. doi:10.3389/fgene.2012.00056
38. Wang X, Huang W, Liu G, et al. Cardiomyocytes mediate anti-angiogenesis in type 2 diabetic rats through the exosomal transfer of miR-320 into endothelial cells. J Mol Cell Cardiol. 2014;74:139–150. doi:10.1016/j.yjmcc.2014.05.001
39. Yang Y, Li Y, Chen X, Cheng X, Liao Y, Yu X. Exosomal transfer of miR-30a between cardiomyocytes regulates autophagy after hypoxia. J Mol Med. 2016;94(6):711–724. doi:10.1007/s00109-016-1387-2
40. Escudero CA, Herlitz K, Troncoso F, et al. Role of extracellular vesicles and microRNAs on dysfunctional angiogenesis during preeclamptic pregnancies. Front Physiol. 2016;7:98. doi:10.3389/fphys.2016.00098
41. Ebert MS, Sharp PA. Emerging roles for natural microRNA sponges. Curr Biol. 2010;20(19):R858–R861. doi:10.1016/j.cub.2010.08.052
42. Stewart C, Rotwein P. Growth, differentiation, and survival: multiple physiological functions for insulin-like growth factors. Physiol Rev. 1996;76(4):1005–1026. doi:10.1152/physrev.1996.76.4.1005
43. Garrouste F, Remacle-Bonnet M, Fauriat C, Marvaldi J, Luis J, Pommier G. Prevention of cytokine-induced apoptosis by insulin-like growth factor-I is independent of cell adhesion molecules in HT29-D4 colon carcinoma cells–evidence for a NF-κB-dependent survival mechanism. Cell Death Differ. 2002;9(7):768. doi:10.1038/sj.cdd.4401022
44. Qian X, Yu J, Yin Y, et al. MicroRNA-143 inhibits tumor growth and angiogenesis and sensitizes chemosensitivity to oxaliplatin in colorectal cancers. Cell Cycle. 2013;12(9):1385–1394. doi:10.4161/cc.24477
45. Xu Q, Jiang Y, Yin Y, et al. A regulatory circuit of miR-148a/152 and DNMT1 in modulating cell transformation and tumor angiogenesis through IGF-IR and IRS1. J Mol Cell Biol. 2012;5(1):3–13. doi:10.1093/jmcb/mjs049
46. Moser C, Schachtschneider P, Lang SA, et al. Inhibition of insulin-like growth factor-I receptor (IGF-IR) using NVP-AEW541, a small molecule kinase inhibitor, reduces orthotopic pancreatic cancer growth and angiogenesis. Eur J Cancer. 2008;44(11):1577–1586. doi:10.1016/j.ejca.2008.04.003
47. Cannon RO. Role of nitric oxide in cardiovascular disease: focus on the endothelium. Clin Chem. 1998;44(8):1809–1819. doi:10.1093/clinchem/44.8.1809
48. Yetik-Anacak G, Catravas JD. Nitric oxide and the endothelium: history and impact on cardiovascular disease. Vascul Pharmacol. 2006;45(5):268–276. doi:10.1016/j.vph.2006.08.002
49. Lei J, Vodovotz Y, Tzeng E, Billiar TR. Nitric oxide, a protective molecule in the cardiovascular system. Nitric Oxide. 2013;35:175–185. doi:10.1016/j.niox.2013.09.004
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.