Back to Journals » Infection and Drug Resistance » Volume 11
Effects of oral probiotic supplementation on gut Lactobacillus and Bifidobacterium populations and the clinical status of low-birth-weight preterm neonates: a multicenter randomized, double-blind, placebo-controlled trial
Authors Strus M, Helwich E, Lauterbach R, Rzepecka-Węglarz B , Nowicka K, Wilińska M , Szczapa J, Rudnicka M, Sławska H, Szczepański M, Waśko A, Mikołajczyk-Cichońska A, Tomusiak-Plebanek A , Heczko PB
Received 23 February 2018
Accepted for publication 18 May 2018
Published 21 September 2018 Volume 2018:11 Pages 1557—1571
DOI https://doi.org/10.2147/IDR.S166348
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Suresh Antony
Magdalena Strus,1 Ewa Helwich,2 Ryszard Lauterbach,3 Beata Rzepecka-Węglarz,4 Katarzyna Nowicka,2 Maria Wilińska,5 Jerzy Szczapa,6 Małgorzata Rudnicka,7 Helena Sławska,8 Marek Szczepański,9 Aneta Waśko,10 Aleksandra Mikołajczyk-Cichońska,10 Anna Tomusiak-Plebanek,1 Piotr B Heczko1
1Department of Microbiology, Jagiellonian University Medical College, Kraków, Poland; 2Department of Neonatology, Institute of Mother and Child, Warszawa, Poland; 3Clinical Department of Neonatology, University Hospital, Kraków, Poland; 4Department of Neonatal Intensive Care, “UJASTEK” Medical Centre, Kraków, Poland; 5Clinical Department of Neonatology, Independent Public Clinical Hospital CMKP, Warszawa, Poland; 6Department of Neonatology, Gynecology and Obstetrics Clinical Hospital, Poznań, Poland; 7Department of Neonatology, Regional Specialist Hospital, Wrocław, Poland; 8Department of Neonatology, Specialist Hospital No. 2, Bytom, Poland; 9Clinic Department of Neonatology and Neonatal Intensive Care, University Clinical Hospital, Białystok, Poland; 10Medical Department, IBSS BIOMED S.A., Kraków, Poland
Aim: Probiotic bacteria administered directly after birth to preterm neonates may improve gastrointestinal function and may reduce the incidence of late-onset sepsis, which is a frequent complication in this group.
Purpose: The main objective of this study was to evaluate whether a new probiotic bacterial mixture of Lactobacillus rhamnosus KL53A and Bifidobacterium breve PB04 given to preterm, low-birth-weight neonates would influence composition of their gut microbiota and sepsis rates.
Patients and methods: This study was a multicenter, randomized, double-blind, placebo-controlled trial conducted in clinical centers of neonatal care in Poland. A probiotic or placebo preparation was given twice daily to 181 preterm low-birth-weight neonates who were eligible for enteral feeding between July 2012 and July 2013. The probiotic was given to 90 neonates, while placebo was given to 91 neonates. The gut microbiota was monitored by microbiological analysis of stool samples. Sepsis episodes were detected on the basis of clinical and laboratory findings and confirmed by blood cultures.
Results: Tested probiotic administration resulted in continuous increase of the Lactobacillus and Bifidobacterium counts in the gut microbiota. The applied tested strains successfully colonized the neonates gut since they were present in over 90% of stool samples, which was confirmed by molecular analysis. Regardless of the study group (probiotic or placebo), B. breve colonization correlated with lower staphylococcal sepsis incidence, which was irrespective of whether probiotics were given. No sepsis case caused by strains included in study probiotic was recorded.
Conclusion: Appropriately selected and characterized probiotic bacteria may be safely given to preterm neonates to normalize their distorted gut microbiota and may contribute to lower staphylococcal sepsis rates.
Keywords: probiotics, LBW neonates, staphylococcal sepsis, gut microbiota, Lactobacillus, Bifidobacterium
Introduction
Nosocomial infection rates in neonates vary from 6% to 50% depending on their weight and related immune system maturity. Bloodstream infections in very low-birth-weight (VLBW) and low-birth-weight (LBW) neonates are believed to be catheter related because critically ill infants require the delivery of nutrients and drugs over long period of time. A disadvantage of long-term catheter use in neonates is that bloodstream infections (CR-BSIs) caused primarily by coagulase-negative staphylococci (CoNS), such as Staphylococcus epidermidis, are common and responsible for significant global morbidity in VLBW. In contrast, term infants are only rarely infected.1–3
Thus, it is commonly accepted that the skin insertion site and the catheter hub are the most important sources of catheter colonization. From the contaminated hub, the organisms may migrate along the surface of the catheter and enter the bloodstream, which cause late-onset sepsis (LOS). This view is based on in vitro studies demonstrating that CoNS and particularly S. epidermidis are extremely capable of adhering to plastic surfaces of the catheters and biofilm formation. In addition to intrinsically high resistance of bacteria contained in biofilms to antibiotics, methicillin-resistant S. epidermidis clinical isolates (methicillin-resistant S. epidermidis [MRSE]) are widespread and complicates the treatment. Diagnosis of sepsis is confirmed by a positive culture of bacteria obtained from blood. More stringent measures include positive cultures from the catheter hub and a peripheral vessel.4,5
There are, however, reports on LOS were related not to catheter use but to translocation of bacteria from gut lumen to bloodstream due to increased intestinal permeability typical for premature neonates.6–8 Distortions in the establishment of normal gut microbiota with increased populations of commensal microbes that colonize the digestive tract might increase the risk of LOS via disruption of the mucosal barrier with resultant translocation of luminal contents. Mai et al in their study based on matched case–control analysis identified microbiota differences in very premature infants with LOS.9 Consistent with many reports, CoNS was the most common microbe cultured from the blood of infants with LOS. Although they were not able to detect a difference in the abundance of CoNS in the stool samples of LOS and control infants, they detected a higher proportion of Firmicutes phylum (the phylum to which CoNS belongs) in LOS cases 2 weeks before diagnosis of sepsis. Stewart et al noticed earlier that preterm gut microbiota was characterized with the predominance of E. faecalis and CoNS as detected by cultures.10 Both bacterial taxa were found by molecular methods to be associated with LOS and necrotizing enterocolitis (NEC).
Therefore, it is highly probable that at least a part of LOS cases treated in neonatal intensive care units (NICUs) may be related to translocation of commensal bacteria from gut due to impaired tight junctions in the mucosa. Such a mechanism may implicate a positive effect of properly selected probiotics on gut microbiota and subsequently on prevention of LOS. However, this effect remains controversial in the light of the recent reviews. Zhang et al have stated recently that current evidence indicates that probiotic supplementation is safe and may significantly reduce the incidence of LOS in preterm neonates in NICU.11 Conversely, Olsen et al in their meta-analysis on prophylactic probiotics for preterm infants were unable to find significant reduction of sepsis rates, although they noticed a trend toward this effect.12 They found a significant reduction of NEC and a reduction in mortality in the neonates.
Probiotics, that is, lactobacilli and bifidobacteria, have been shown to significantly reduce the risk of NEC, all-cause mortality, LOS, and facilitate feed tolerance in preterm VLBW neonates. The mechanisms of benefits of probiotics include gut barrier enhancement, immune response modulation (eg, TLR4 receptor, nuclear factor-B, inflammatory cytokines), and direct inhibition of gut colonization by pathogens.19
It is postulated that correlation of distortions of the intestinal microbiota is a necessary first step to design novel microbiota-based screening approaches that might lead to early interventions to prevent LOS in high-risk infants.9 Well-characterized and clinically proved probiotics can be used for this purpose.
As it is speculated on perspectives of the progress in managing serious infections in neonatology, clinical research offers the opportunity of adopting preventive strategies such as use of probiotics and lactoferrin to prevent LOS. However, these strategies remain to be commonly accepted and recommended.
Hypothesis
Our assumption is that probiotic bacteria administered directly after birth are beneficial for the development of intestinal microbiota and could prevent or significantly limit gastrointestinal colonization by pathogenic bacteria (PB) and development of abnormal microbiota. Permanent colonization with probiotic bacteria in very early life improves gastrointestinal function in preterm neonates by reducing the onset of or decreasing the severity of the signs and symptoms of feeding intolerance and generalized bacterial infections, including sepsis and NEC.
Objectives
The aim of this study was to evaluate the safety of the tested probiotic preparation and its influence on the development of gut microbiota resembling those of healthy term-born, breast-fed neonates, and on the clinical status of premature LBW neonates. More specific objectives were to determine whether colonization of the neonates gut by the tested probiotic strains would influence populations of the potentially PB and reduce the incidence of sepsis and gastrointestinal disorders in these infants.
Trial design
This was a multicenter, randomized, double-blind, placebo-controlled, and parallel-group study with two randomization arms (1:1). It was conducted in accordance with the original protocol PB-DM/SBK-NEC-01/11.
The amendment of inclusion criteria allowed for a higher birth weight (from up to 1,500 to 1,800 g), higher gestation age (from week 32 up to week 34), and longer study inclusion time (from up to 12 h to 48 h). The protocol was amended to account for differences in standard neonatal care at different study centers, especially in terms of the time of first feeding and insufficient recruitment. The abovementioned modifications were included in order to make the study group more homogeneous and increase recruitment rate. They were introduced following Independent Ethics Committee approval dated September 27, 2012.
Patients and methods
Patients
Eligible were preterm neonates of both sexes, born ≤34 weeks of gestation, and weighing ≥750 g and ≤1,800 g from adult mothers (aged ≥18 years). Children were enrolled by the investigators within 48 h after birth ready for enteral feeding.
Exclusion criteria were major congenital anomalies, including gastrointestinal abnormalities, which might interfere with enteral nourishing, birth weight below 750 g or over 1,800 g, born >34 weeks of gestation, mother’s age <18 years, enteral nutrition contraindicated within the first 48 h of life, birth asphyxia with Apgar score <4, severe clinical condition/disorder that the investigator considers as a contraindication (ie, including the presence of at least three out of four of the following symptoms: necessity of using mechanical ventilation and FiO2 >0.6 elevation to maintain blood oxygen saturation within the range of 88%–93%; metabolic acidosis, pH <7.20 and BE >[-10]; necessity of vasopressor agents to maintain proper arterial blood pressure; signs of failure to at least one vital organ [liver, kidneys, gastrointestinal tract, and adrenal glands]), administration of other probiotic preparations, enrollment in any other clinical study throughout the study period, and absence of parent/legal guardian informed consent.
Ethics-approved written informed consent was obtained from the parents before enrolling a neonate in the trial.
The study took place between April 2012 and December 2013 in eight NICUs in the whole area of Poland. The first participant was enrolled in the trial on July 07, 2012 and the last participant on July 10, 2013. The duration of subject’s participation was ~7 weeks, until they were withdrawn (according to withdrawal criteria) or until discharged from the hospital, whatever occurred earlier. The participant was administered the investigational product over the period of 6 weeks and then observed for a further week.
Data were entered and stored in Microsoft Access and analyzed using JMPÒ software (version 7.0.1, SAS Institute Inc., Cary, NC, USA) and the R 9.2.1 software package (R Foundation for Statistical Computing).
Neonates were monitored in the cooperating centers, and appropriate care was given to them according to standards of medical care for neonates provided by Polish Society of Neonatology.13 LOS was defined according to criteria given by Gastmeier et al:14
- Presence of at least two of the following: temperature >38°C or <36.5°C or temperature instability, tachycardia or bradycardia, apnea, prolonged capillary refill, metabolic acidosis, hypoglycemia, and other signs of bloodstream infections such as lethargy.
- Recognized pathogen cultured from one or more blood cultures or CoNS isolated from at least one blood culture or intravascular line and one of the following: C-reactive protein >2.0 mg/dL, immature/total neutrophil ratio (I/T ratio) >0.2, leukocytes <5,000/μL, and platelets <10,000/μL.
Interventions
Single dose of investigational product (probiotic or placebo) was administered orally twice daily with food (in the morning and in the evening). The first dose was administered within the first 48 h after birth and continued for 6 weeks or shorter, in case of earlier hospital discharge. Discontinuation of the enteral nutrition was equivalent with termination of the administration of the investigational product. Its intake could be discontinued for up to ≤7 days due to suspension of enteral feeding and no additional (supplementary) doses of the investigational product were administered. If the product intake was discontinued for more than 7 days, the patient was withdrawn from the study.
The tested probiotic was food for special medical purposes ( FFbaby® provided by the study sponsor, IBSS BIOMED S.A., Kraków, Poland) in the form of a powder in glass vials. One dose contained not less than 106 CFU of a bacterial mixture including lyophilized Lactobacillus rhamnosus KL53A and Bifidobacterium breve PB04 and auxiliary substances: maltodextrin and ascorbic acid. Before administration, the tested product or placebo was diluted in 1.5 mL of milk (mother’s milk or infant formula) and administered enterally with a feeding tube or a teat (depending on the individual feeding method). The feed containing the tested product or placebo was prepared individually for each participant and used directly after preparation. The strains (L. rhamnosus KL53A and B. breve PB04) were well characterized in in vitro and in vivo studies using methods described elsewhere.13,14 The tested strains have documented human origin. They were isolated from the feces of a healthy breast-fed child and were selected from a larger strain collection because of their in vitro and in vivo probiotic properties. Their strong anti-inflammatory, antipathogenic activity, and tight-junctions stimulating properties were confirmed in laboratory studies. Safety, probiotic properties, and gut mucosa colonizing ability of the tested strains were checked on gnotobiotic (germ-free) mice and on rat neonates. The strains and their composition are subject to patent application.
The probiotic preparation was a composition of an active ingredient, that is, a mixture of L. rhamnosus KL53A and B. breve PB04 strains on maltodextrin as excipient and ascorbic acid as vehicle. Placebo was a composition of maltodextrin and ascorbic acid vehicle. It was produced under the same conditions as active product with the same appearance, net mass, and composition except for bacterial strains. The probiotic preparation and placebo were in the form of a lyophilized powder for suspension.
Microbiological examination of feces
Stool samples were collected before first administration of investigational product and then at 7-day intervals into two preweighted tubes containing liquid transport MRS medium (Oxoid Thermo Scientific, Basingstoke, UK) for probiotic strains and Schaedler Anaerobic Broth (Difco Labs, Detroit, MI, USA) for the remaining bacteria and delivered deep frozen (at –70°C) to the central laboratory. They were then thawed, serially diluted, and cultured on a series of appropriate differential media to measure bacterial populations of Lactobacillus and Bifidobacterium as well as representative potentially PB: Clostridium difficile, Staphylococcus aureus, Enterococcus spp., Klebsiella spp., and Escherichia coli. Following agar media were used: McConkey Agar (Oxoid, UK) for Enterobacteriaceae, Columbia Blood Agar (Difco, USA) with 5% sheep blood for staphylococci and streptococci, Enterococcosel Agar (BD-BBL, Franklin Lakes, NJ, USA) for enterococci, MRS (De man, Rogosa and Sharpe) agar (Oxoid) for lactobacilli and other lactic acid bacteria (LAB), BL Agar (Oxoid) for bifidobacteria, and Wilkins-Chalgren Agar Base with supplements for Clostridium. For Candida fungi, Sabouraud’s substrate Agar (Biocorp, Warsaw, PL, USA) was used. For detailed identification of species of fully formed colonies, phenotypic methods were used and especially API (bioMérieux) tests: API STREP (for Streptococcus and Enterococcus), API STAPH (for Staphylococcus), API 20E (for Enterobacteriaceae), API 20A (for anaerobic bacteria), and API 50CH (for Lactobacillus). The results were analyzed with API LAB software for classification of test bacteria.
Further identification of the isolated strains Lactobacillus and Bifidobacterium genera was based on genotyping (polymerase chain reaction [PCR]) methods.21,22 The species-specific PCR for L. rhamnosus was done using the following primers: PrI 5′ CAG ACT GAA AGT CTG ACG G 3′ and RhaII 5′ GCG ATG CGA ATT TCT ATT ATT 3′ which amplifies a 190 bp fragment. The PCR conditions were as follows: one cycle at 92°C for 2 min; 30 cycles at 95°C for 30 s, at 55°C for 30 s, at 72°C for 30 s; and one cycle at 72°C for 1 min. The following primers IDB31F 5′ TAG GGA GCA AGG CAC TTT GTG T 3′ and IDBC1R 5′ ATC CGA ACT GAG ACC GGT T 3′ were used to identify B. breve. The size of the PCR product was 827 bp and the protocol was as follows: one cycle at 94°C for 5 min; 35 cycles at 94°C for 30 s, at 64°C for 40 s, at 72°C for 30 s; and one cycle at 72°C for 5 min.
Molecular typing of isolates as belonging to strains contained in the product, that is, L. rhamnosus KL53A and B. breve PB04, was done using pulse-field gel electrophoresis. The examined isolates of L. rhamnosus were prepared according to the procedure of Tynkkynen et al.15 Strains belonging to this species were digested with restriction enzyme SgsI (25 U/block) (Thermo Fisher Scientific, Waltham, MA, USA). Preparation of B. breve genomic DNA was performed according to Roy et al.16 In order to digest the chromosomal DNA of this species, we used SpeI restriction enzyme (25 U/block) (Thermo Fisher Scientific).
Electrophoresis was done in CHEF-DR II apparatus (BioRad, Hercules, CA, USA) at the following separation conditions for L. rhamnosus: starting pulse 1 s, final pulse 15 s, voltage 5 V/cm, the temperature was 14°C, and the running time was 22 h and for B. breve: starting pulse 1 s, final pulse 20 s, voltage 6 V/cm, temperature 14°C, and running time 18 h.
Microbiological testing of blood
Diagnosis of sepsis was confirmed by positive blood culture performed according to routine procedures used by local microbiology laboratories in the cooperating clinical centers according to the standards.13 Blood was taken aseptically from all suspected sepsis cases and 1 mL samples were injected into an aerobic blood culture bottle (Bactec Plus 26 Aerobic; BD Microbiology Systems, Franklin Lakes, USA) and cultured on MacConkey agar, horse blood agar (at 37°C, each for 24 h), and Sabourand agar (at 37°C for 38 h). The isolates were characterized using biochemical tests, bioMerieux identification kit API system (bioMerieux, Marcy l’Etoile, France).
Only in cases of positive blood cultures caused by gram-positive rods, blood was plated additionally on MRS (Oxoid, UK) and TPY (Biocorp, Poland) medium for bacteria of the genera Lactobacillus and Bifidobacterium. After inoculation, plates were incubated for 48 h at 37°C under aerobic and anaerobic conditions. All strains of bacteria cultured on MRS and TPY agar were collected in the cooperating centers and were transferred to the central laboratory for molecular identification and typing to compare them with the strains included in tested probiotic.
Outcomes
Primary outcome measures
- Gastrointestinal tract colonization assessed weekly (day 1, 7, 14, 21, 28, 35, 42, and 49) within 8 weeks. Bacteriological tests involved quantitative and qualitative assessments of neonatal gut microbiota during clinical follow-up. Density of the bacterial populations of Lactobacillus and Bifidobacterium as well as following potentially PB and fungi were assessed: E. coli, Klebsiella sp., Clostridium sp., S. aureus including MRSA, S. epidermidis including MRSE, Staphylococcus haemolyticus, Enterococcus faecalis, Streptococcus agalactiae, S. pyogenes, and Candida albicans. Stool samples were collected at 7-day intervals (±2 days) in the period of investigational product intake and on day 7 (±2 days) after the last dose of the investigational product. The first sample was collected before administration of the first dose of investigational product.
- Feeding intolerance episodes, including gastric residuals, vomiting, regurgitation of food, abdominal distension, abdominal rigidity, and gut motility disorders were assessed on a daily basis (from day 1 to 42 and on day 49). Clinical parameters were evaluated throughout the period of care according to standard neonatal care procedures.
- Incidence and type of adverse events and serious adverse events (SAE) with special regard to sepsis caused by bacteria included in investigational product, assessed on a daily basis. Safety was evaluated by determining whether investigational product increases the incidence and influences the type of adverse events and SAE compared to placebo.
Main secondary outcome measures
- LOS caused by gram-positive or gram-negative bacteria assessed on a daily basis (from day 1 to 42 and on day 49). Sepsis was confirmed by positive blood cultures. Clinical parameters were evaluated throughout the period of care according to standard neonatal care procedures.
- NEC incidence, intensity, and severity assessed on a daily basis. Severity of neonatal NEC was determined according to Bell’s staging criteria.17 Clinical parameters were evaluated throughout the period of care according to standard neonatal care procedures.
- Mortality rate, with a focus on deaths attributed to NEC and sepsis assessed on a daily basis. Clinical parameters were evaluated throughout the period of care according to standard neonatal care procedures.
Sample size
As indicated in literature18 and information obtained from the study Medical Expert, the likelihood of the studied symptoms development in the control group is estimated at pplacebo=0.3. On the assumption that the probiotic efficacy is 30%, the likelihood of symptoms development in the probiotic group is estimated at pintervention=0.21. Thus, the least significant sample size was calculated as 138 participants (69 participants per each group).
This sample size refers to participants who completed the entire clinical study cycle; it was therefore necessary to increase the number of participants to prevent random discontinuations from reducing the analyzed sample size below the determined level. In consideration of the likelihood of early patient withdrawal, the baseline sample size was increased by ca. 40%, that is, maximum of 194 participants were planned to be enrolled.
Randomization, allocation, concealment, implementation, and blinding
Participants were enrolled and randomly assigned to one of two study arms (probiotic or placebo) using equal group ratios, according to a computer-generated sequence (R software package). The randomization list was produced by a statistician with no clinical input in the trial and held securely by the study sponsor. Each study center received randomization datasets (sets of randomization numbers to be assigned to study participants) and a set of packages containing the investigational products (probiotic or placebo), labeled with numerical codes assigned to one of the two treatment groups and a letter (A, B, C, or D). Each participant was assigned an ID number composed of a two-digit study center number and a three-digit screening number assigned in chronological order as participants were enrolled in a given center. Upon randomization, the participant’s ID was supplemented with a three-digit randomization number and a letter (A–D) identifying the randomization data set.
The participants, investigators, clinic and central laboratory staff, study monitors, persons entering data to a database, and study statisticians were blinded to group assignment.
Statistical methods
Participants were described by means of morphometric and developmental parameters, such as birth weight, body weight gain, and gestational age (weeks of gestation). Experimental data were described using descriptive statistical methods. For continuous variables, parameters describing the central tendencies (mean with the standard error, median, and modal value) and dispersion parameters: SD, minimum, maximum, and quartile limits were presented. For categorical variables, fractions of cases were determined when the variables have specific values. Inductive statistical tools were used to compare study groups, depending on the type and distribution of the analyzed variables. Quantitative variables for the two groups were compared with the Student’s t-test. For variable distribution significantly deviating from the Gaussian curve, the Student’s t-test was substituted with nonparametric tests (Mann–Whitney, Kolmogorov–Smirnov, or Wilcoxon’s tests) due to method limitations. For comparisons of more than two groups, analysis of variance (ANOVA) and post hoc (a posteriori) Tukey’s and Dunnett’s tests were applied. For variables noncompliant with the assumptions of the ANOVA (such as normal distribution and uniformity of variance groups), it was substituted with a nonparametric equivalent of the Kruskal–Wallis test supplemented with the Steel-Dwass test, a nonparametric equivalent of the post hoc Tukey test. The analysis of categorized variables was based on comparisons between the fractions of values of such variables. These analyses were carried out with frequency tests χ2 (chi-square) or G2 (likelihood ratio). The significance level for statistical analysis was set at p<0.05. The following statistical tools were used: SAS JMP 7.0.1 and R package 9.2.1.
Ethics and trial approval
This trial received ethical approval from the Independent Ethics Committee of Jagiellonian University, Krakow, Poland (no. KBET/127/L/2012 on April 26, 2012) and was conducted according to good clinical practice requirements and the Declaration of Helsinki.
This clinical study has been entered into the clinical trials register retrospectively due to the fact that clinical trials regarding food products are not subject to regulatory approval and there is no legal obligation to include such trials in the register. But, in order to fulfill journal policies, we have obtained needed registration in ClinicalTrials.gov no. NCT02073214.
Results
Altogether 182 participants were enrolled and randomized; one child did not receive investigational product. Four out of 181 were excluded from the analysis due to formal reasons such as incorrect qualification.
For statistical analyses, the following populations were distinguished:
- Baseline characteristics for all study participants enrolled, who provided written informed consent (n=177).
- Intent-to-treat (ITT) group: participants who were randomized and used at least one capsule of the investigated product or placebo (n=177). Safety analysis was performed for this population.
- Per-protocol (PP) group: participants who completed the study according to the protocol and have not fulfilled any of the exclusion criteria throughout the study (n=153). Efficacy analysis, including additional analysis, was carried out for this population. Figure 1 presents a diagram of participant flow.
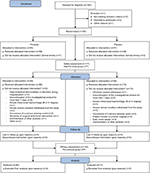  | Figure 1 Participant flow chart. Notes: *According to the protocol, participant could have been withdrawn from the trail due to more than one reason. |
Maternal and neonatal baseline characteristics are presented in Table 1. In general, no significant differences between groups were found. The only significant differences regarding mother’s age and baby head circumference were found when characteristics of probiotic group (AB) and placebo group (CD) neonates were compared. Mean values of the age of mother and head circumference, however, seem to have no influence on the interpretation of obtained results and the statistical significance found results from large data set. It is thus worth stressing that due to randomized allocation of participants to study and control group, it was possible to achieve great similarity of the analyzed groups of participants, which validates further statistical comparisons.
  | Table 1 Maternal and neonatal baseline characteristics Notes: aChi-square test; bStudent’s t-test; cMann–Whitney U-test. Abbreviations: SD, standard deviation; Av, arithmetical mean. |
Efficacy evaluation
Gastrointestinal tract colonization (primary outcome)
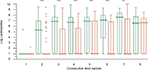
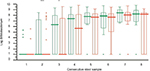
There were no significant differences between probiotic (AB) and placebo (CD) groups regarding numbers of Lactobacillus and Bifidobacterium found in the first collected sample, that is, sample collected before intervention. There were, however, significant differences between groups in favor to AB group in Lactobacillus counts, in the samples collected from weeks 2 to 7 of the study (Figure 2). The differences related to Bifidobacterium counts appeared significant in weeks 2 and 3 (Figure 3).
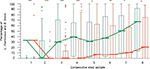
The stool samples analysis revealed that probiotic supplementation was associated with significantly higher populations expressed in percentages of L. rhamnosus and B. breve versus other cultivable bacteria, and especially potentially pathogenic ones in the gut microbiota comparing to the placebo group, predominantly in the first weeks of life (Figure 4). Effectiveness of the colonization was also measured by the number of participants in each group colonized by L. rhamnosus and B. breve. We have also analyzed the last available sample taking into account that some participants were discharged earlier from the hospital (Table 2).
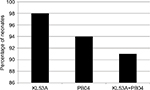
Molecular typing of all isolates belonging to species L. rhamnosus and/or B. breve obtained from feces revealed that nearly all (98%) neonates from study group were colonized with L. rhamnosus KL53A and nearly the same proportion (94%) with B. breve PB04. A very high proportion of the neonates (91%) were colonized with both strains throughout the study period (Figure 5).
There were no significant differences between groups in the abundance of potentially PB (Clostridium spp., E. coli, C. albicans, Klebsiella spp., Enterobacter spp., S. aureus including MRSA, S. epidermidis) present in consecutive samplings.
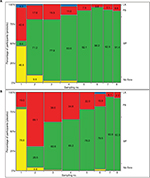
Additional analysis was performed based on percentage of participants, for whom the following parameters were found in subsequent samplings of their feces: i) presence of the Lactobacillus or Bifidobacterium (LAB) genera regarded as representatives of the normal and favorable members of the gut microbiota without coincidental presence of the above listed potentially PB; ii) presence of only potentially PB, without Lactobacillus and Bifidobacterium; iii) presence of both groups of microbes, that is, Lactobacillus, Bifidobacterium, and potentially PB in feces (MF); and iv) absence of the cultivable microorganisms. We found that the percentage of the infants with only PB decreased and the percentage of those with MF increased in subsequent samplings. In the first sampling (1), the percentage of the neonates with only PB only was greater in the probiotic group than in the placebo group. In the next sampling (2), an increase of the percentage of the neonates with MF was observed in the probiotic group and the percentage of infants with PB increased over three times in the placebo group. In subsequent samplings, it was found that the percentage of the patients with PB was decreased, whereas the percentage of those with MF increased. These changes were more pronounced in the probiotic group than in the placebo group. However, no statistical significance between was found between the compared groups as tested by Cochran–Mantel–Haenszel test. The results are shown in Figure 6.
Feeding intolerance episodes (primary outcome)
There was no difference in the number of feeding intolerance episodes, such as gastric residuals, vomiting, regurgitation of food, abdominal distension, abdominal rigidity, and gut motility disorders in the groups.
LOS (secondary outcome)
Overall sepsis analysis in terms of safety was performed on intent to treat group and has been presented in Safety evaluation section while more detailed analysis was performed on a PP group. In this population, there were 18 episodes of late-onset gram-positive sepsis in 18 participants including 11 in the probiotic group (13.8%) and 7 in the placebo group (9.6%). Staphylococci were dominant etiological agents of the late-onset gram-positive sepsis (16 episodes), whereas the remaining sepsis episodes were associated with Klebsiella pneumoniae (1 episode) and yet not identified bacteria (1 episode).
We did not show significant differences between groups in sepsis incidence. Also, analysis of late-onset gram-positive sepsis occurrence in probiotic and placebo groups that involved the difference in the participation of the infants belonging to these groups on individual days of the study, the number of episodes over weeks of the study, and incidence of sepsis diagnosis regardless of the day of stay did not reveal statistically significant differences between the groups. Similarly, probiotic-to-placebo group comparisons did not yield statistically significant increase of risk both in the case of analysis of the whole period of the study and for risk over subsequent weeks when subsequent confounders were involved. Risk analysis (HR) of the first episode of late-onset gram-positive sepsis in the probiotic-to-placebo group comparison did not yield statistically significant results as well. There were also no differences between groups in the occurrence of late-onset gram-negative sepsis. It was diagnosed in total in one baby in the probiotic group and none in the placebo group.
Additional analysis has revealed that regardless of the assignment to either of the study groups (probiotic or placebo), B. breve colonization correlated with the lower staphylococcal sepsis incidence (Figure 7). The probability of sepsis occurrence in a given week and the next week was significantly lower in participants colonized by B. breve than in not colonized participants. Participants without sepsis had significantly higher count of bacteria of Lactobacillus and Bifidobacterium genera in the week in which sepsis was diagnosed (H=8.4150; df=3; p=0.0382). The same dependence was observed for all Bifidobacterium (H=18.6466; df=3; p=0.0003) and B. breve (H=9.5991; df=3; p=0.0223). However, it was not found for Lactobacillus (H=1.0521; df=3; p=0.8787) and L. rhamnosus (H=1.3038; df=3; p=0.7282). When L. rhamnosus and B. breve were considered together, the dependence was close to significance threshold (H=6.7444, df=3, p=0.0805). Moreover, we have also found that the number of days with sepsis symptoms was significantly smaller in participants who were colonized by L. rhamnosus and B. breve prior to the next stool collection than in noncolonized participants.
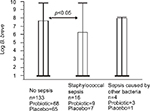  | Figure 7 Correlation between B. breve gut numbers and staphylococcal sepsis (Observed in both groups). Abbreviations: B. breve, Bifidobacterium breve. |
Necrotizing enterocolitis (secondary outcome)
There were no statistically significant differences found between groups neither when the total incidence of NEC over the whole period of the was analyzed nor when numbers of episodes in both groups were analyzed. Logistic regression models were calculated to estimate the risk (OR) of NEC in the period of the study. Results were not statistically significant in the univariate model and after standardization to account for the type of milk the baby was fed, mother’s age and baby head circumference, and additionally body weight at birth. No statistically significant results were found when individual subperiods of the study were analyzed.
Altogether there were six episodes of NEC, found in five participants: five in the probiotic group and one in the placebo group. In the probiotic group, two participants developed NEC of I degree, one participant developed twice NEC of II degree and one participant developed NEC of III degree. In the placebo group, NEC of II degree was diagnosed.
Mortality (secondary outcome)
Mortality rate, in particular deaths attributed to NEC and sepsis, did not reveal differences between the groups. More in-depth analysis of mortality over individual weeks of the study did not show statistically significant differences either. During the 49-day period of the study, six deaths in total were noted, which results in 3.39% mortality rate in the study group. There were two deaths in the probiotic group (mortality rate for this group was 2.25%) and four in the placebo group (mortality rate 4.55%). In the probiotic group, the causes of deaths were intracerebral hemorrhage of III degree, bilateral and intracerebral hemorrhage in cerebellum in one participant and NEC in one participant. In the placebo group, the causes of deaths were intraventricular hemorrhage of III and IV degrees in two participants, malformation syndrome in one participant and sepsis with septic shock and disseminated intravascular coagulation in one participant. Sepsis caused one death in total, it was a LOS in the placebo group.
Safety evaluation (primary outcome)
Safety evaluation was based on the assessment of the number and types of adverse events and SAE in all enrolled participants who were administered at least one dose of the investigational product (ITT group, n=177). It should be also noted that adverse events related to feeding intolerance were analyzed as primary end point and so were excluded from safety analysis. The results are provided in the Efficacy section. The most often observed adverse event was apnea, observed in 43% participants in total. Subsequent adverse events were intraventricular hemorrhage (33%), anemia (19%), respiratory disorders (17.5%), pneumonia (13%), desaturation (12%), conjunctivitis (9%), infection (8.5%), retinopathy (8.5%), patent ductus arteriosus (8%), jaundice (6%), neonatal pneumonia (5.65%), hyponatremia (5%), early onset sepsis (5%), tachycardia (4.5%), skin injury (4.5%), hyperglycemia (4%), edema (4%), respiratory disorders (4%), bronchopulmonary dysplasia (3.4%), and others (with lesser incidence). No statistically significant differences in incidence of adverse events between groups AB (probiotic) and CD (placebo) were found with the exception of hypocalcemia which occurred statistically more often in the CD group (0.0% vs 3.4%; p=0.0394). Hypocalcemia was deemed by the investigator as not related to administration of the investigational product in all reported episodes. Numbers of adverse events of specific types in individual study groups were also compared. The analysis did not reveal any statistically significant differences except conjunctivitis, of which a greater share of babies in the AB group had two episodes diagnosed compared with the CD group (three in seven babies in the AB group with conjunctivitis, zero in nine babies in the CD group with conjunctivitis, p=0.0153). All reported episodes of conjunctivitis were classified by the investigator as not related to administration of the investigational product.
Altogether 565 adverse events were reported during the study in total (excluding feeding intolerance episodes and SAE); 278 and 287 in groups AB and CD, respectively, the investigators qualified 520 of them as not to be related to the investigational product (255 and 265 in groups AB and CD, respectively) and 45 was considered unlikely (23 and 22 in groups AB and CD, respectively). There were no adverse events assessed as probably or certainly related to the investigational product.
SAE occurred in 40 participants (22.6%). The most often was LOS, there were altogether 27 episodes that occurred in 24 participants (13.6%) (ITT group). Differences between groups were not statistically significant. Gram-positive sepsis was diagnosed in 20 cases (20 participants) and gram-negative sepsis in 6 cases (4 participants). LOS was the cause of one death which occurred in placebo group. Other SAE were: intraventricular hemorrhage (3.4% of participants), death (3.4%), NEC (2.8%), pneumothorax (1.7%), early onset sepsis (1.7%), leukomalacia (1.1%), pneumonia (1.1%), bronchopulmonary dysplasia (0.6%), circulatory deficiency (0.6%), respiratory deficiency (0.6%), multiorgan dysfunction (0.6%), and intestinal perforation (0.6%). There were no statistically significant differences between the groups. From among all 61 reported SAE, suspected relation to administration of the investigational product was assessed for 3 events in the probiotic group (two events – NEC, one event – death neonatal) (Table 3). No relation was suspected for 58 events (33 in probiotic group and 25 in the placebo). It is worth to stress that no sepsis was caused by Lactobacillus or Bifidobacterium.
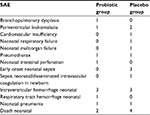  | Table 3 Number of SAE except for sepsis and NEC in probiotic and placebo groups – ITT group Abbreviations: SAE, serious adverse events; NEC, necrotizing enterocolitis; ITT, Intent-to-treat. |
Overall safety analysis confirmed that probiotic including L. rhamnosus KL53A and B. breve PB04 may be safely used in preterm, LBW neonates.
Discussion
This study showing inverse relationship between colonization of the preterm neonates gut by bifidobacteria and staphylococcal sepsis rates indicates that probiotics can influence generalized infections such as sepsis.18,19 This relationship was demonstrated for both study and control groups taken together and obtained results seem to be an important observation indicating that probiotic use in the group of LBW neonates may decrease staphylococcal sepsis rates. Since the clinical effects of probiotics are strain dependent, the probiotic strains applied by us were selected from a larger strain collection of documented human origin because of their in vitro and in vivo characteristics indicating their strong anti-inflammatory activity, ability to exclude PB from their adherent sites on human enterocytes, and tight-junctions stimulating properties for these cells (unpublished). Safety, probiotic properties and colonizing ability of the tested strains were checked on gnotobiotic (germ-free) mice and on rat neonatal models.
Efficacy of probiotics in preventing sepsis in LBW neonates remains controversial. The meta-analysis published by Zhang et al has indicated that probiotics may decrease neonatal sepsis rates at NICU.11 However, another recent meta-analysis performed by Olsen et al on prophylactic usage of probiotics for LBW neonates, based on 12 studies, states that probiotic supplementation is related to significant decrease of NEC but not to sepsis rates.12 It is of interest why these two species were not used in the studies analyzed by Olsen et al. As mentioned before, it is known that each probiotic strain and a mixture of different probiotics is individual and thus results from other even very similar studies cannot be extrapolated.20 Another explanation which can be offered here is that sepsis occurrence was monitored in only 7 of all clinical trials analyzed by Olsen et al, which might result in making analysis of a population of the neonates too small to reach significance of the differences between study versus control groups. The authors mentioned in discussion of their meta-analysis that they observed a trend toward a benefit in the reduction of sepsis. Most probably, this trend could be converted into significance, if the authors of the trials, in which B. breve was used, should break their data according to the pathogens causing sepsis and concentrate over staphylococci as most common cause of the neonatal sepsis. Our data indicate that a high abundance of B. breve in the gut microbiota is related to lower rates of sepsis in LBW neonates. The only possibility to increase numbers of bifidobacteria in the neonates is to supply them orally in a form of a probiotic preparation. It seems that the crucial factor enabling successful gut colonization by these bacteria, not mentioned in protocols of the studies analyzed by Olsen et al, is a time of starting digestive tract colonization by probiotics. Gut of the infants treated in NICU is very rapidly colonized with the hospital pathogens; thus, probiotics should be given shortly after exposition of the infants to hospital environment. Their chances of probiotic bacteria to overcome commensal bacteria when they have already colonized the gut are very limited.
Colonization with probiotic strains given to neonates appeared to be highly efficacious as nearly all of them showed a presence of both probiotic strains in their feces. This effect may also explain general improvement of the gut microbiota composition with predominance of lactobacilli and bifidobacteria over other microbes.
Here, we cannot offer explanation why only sepsis caused by CoNS but not Proteobacteria, that is, gram-negative rods has been influenced by the given probiotic mixture, although gram-negative bacteria appeared to be more susceptible to inhibitory activities of the used probiotics in vitro. However, this is just reflection of the higher numbers of the neonates with sepsis caused by staphylococci. In our study, numbers of the cases of gram-negative sepsis were too low to achieve statistical significance.
Loss of gut barrier function is considered as a primary mechanism of sepsis caused by gram-negative bacteria in VLBW and LBW neonates treated at NICU. It has been demonstrated that the bacteria translocate through gut wall, reach bloodstream either directly by invading submucosal capillary vessels or indirectly by Peyer’s patches and lymphatics to blood and cause bacteremia and then sepsis. There are only a few publications showing that also gram-positive cocci have been able to translocate through inadequately tight GI tract mucosa in critically ill adult patients treated at ICU. No direct studies on translocation of staphylococci from gut into bloodstream in premature neonates were yet published. However, a regulation of intestinal permeability functions by probiotics as a preventive measure for both chronic and acute inflammations of the digestive tract are recently postulated.7 Both L. rhamnosus KL53A and B. breve PB04 used in this study have been shown as potent stimulators of the tight junction proteins by in vitro cultured enterocytes (unpublished). It is therefore possible that at least one of the mechanisms by which probiotics interfere with postulated translocation of staphylococci from gut is based on restoration of the gut barrier functions.
Conclusion
Orally administered, new probiotic preparation containing a mixture of L. rhamnosus KL53A and B. breve PB04 to preterm, LBW neonates is safe and effectively colonizes their distorted gut microbiota and it may also contribute to lower staphylococcal sepsis rates.
Acknowledgments
The authors would like to give special thanks to the participants and their parents. We also wish to thank all investigators, nurses, and cooperating staff. For monitoring, statistics, data management, and analysis, we would like to thank independent external Contract Research Organization – KCRI Sp. z o. o. (Kraków, Poland).
Author contributions
All authors contributed toward data analysis, drafting and critically revising the paper, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
Jean-Baptiste N, Benjamin DK, Cohen-Wolkowiez M, et al. Coagulase-negative staphylococcal infections in the neonatal intensive care unit. Infect Control Hosp Epidemiol. 2011;32(7):679–686. | ||
Rønnestad A, Abrahamsen TG, Medbø S, et al. Sepicemia in the first week of life in a Norwegian national cohort of extremely premature infants. Pediatrics. 2005;115(3):e262–e268. | ||
Wójkowska-Mach J, Gulczyńska E, Nowiczewski M, et al. Late-onset bloodstream infections of very-low-birth-weight infants: data from the Polish neonatology surveillance network in 2009-2011. BMC Infect Dis. 2014;14:339. | ||
Cherifi S, Byl B, Deplano A, et al. Genetic characteristics and antimicrobial resistance of Staphylococcus epidermidis isolates from patients with catheter-related bloodstream infections and from colonized healthcare workers in a Belgian hospital. Ann Clin Microbiol Antimicrob. 2014;13:20. | ||
Maki DG, Weise CE, Sarafin HW. A semiquantitative culture method for identifying intravenous-catheter-related infection. N Engl J Med. 1977;296(23):1305–1309. | ||
Berg RD. Bacterial translocation from the gastrointestinal tract. Adv Exp Med Biol. 1999;473:11–30. | ||
Bischoff SC, Barbara G, Buurman W, et al. Intestinal permeability: a new target for disease prevention and therapy. BMC Gastroenterol. 2014;14:189. | ||
Deitch EA. Gut-origin sepsis: evolution of a concept. Surgeon. 2012;10(6):350–356. | ||
Mai V, Torrazza RM, Ukhanova M, et al. Distortions in development of intestinal microbiota associated with late onset sepsis in preterm infants. PLoS One. 2013;8(1):e52876. | ||
Stewart CJ, Marrs EC, Magorrian S, et al. The preterm gut microbiota: changes associated with necrotizing enterocolitis and infection. Acta Paediatr. 2012;101(11):1121–1127. | ||
Zhang GQ, Hu HJ, Liu CY, Shakya S, Li ZY. Probiotics for preventing late-onset sepsis in preterm neonates: a PRISMA-compliant systematic review and meta-analysis of randomized controlled trials. Medicine (Baltimore). 2016;95(8):e2581. | ||
Olsen R, Greisen G, Schrøder M, Brok J. Prophylactic probiotics for preterm infants: a systematic review and meta-analysis of observational studies. Neonatology. 2016;109(2):105–112. | ||
Polish Society of Neonatology. Standards of the Neonatal Medical Care in Poland (in Polish). Media-Press; 2015. | ||
Gastmeier P, Geffers C, Schwab F, Fitzner J, Obladen M, Rüden H. Development of a surveillance system for nosocomial infections: the component for neonatal intensive care units in Germany. J Hosp Infect. 2004;57(2):126–131. | ||
Tynkkynen S, Satokari R, Saarela M, Mattila-Sandholm T, Saxelin M. Comparison of ribotyping, randomly amplified polymorphic DNA analysis, and pulsed-field gel electrophoresis in typing of Lactobacillus rhamnosus and L. casei strains. Appl Environ Microbiol. 1999;65(9):3908–3914. | ||
Roy D, Ward P, Champagne G. Differentiation of bifidobacteria by use of pulsed-field gel electrophoresis and polymerase chain reaction. Int J Food Microbiol. 1996;29(1):11–29. | ||
Bell MJ, Ternberg JL, Feigin RD, et al. Neonatal necrotizing enterocolitis: therapeutic decisions based upon clinical staging. Ann Surg. 1978;187(1):1–7. | ||
Alfaleh K, Anabrees J, Bassler D, Al-Kharfi T. Probiotics for prevention of necrotizing enterocolitis in preterm infants. Cochrane Database Syst Rev. 2011;16(3):CD005496. | ||
Deshpande G, Rao S, Patole S, Bulsara M. Updated meta-analysis of probiotics for preventing necrotizing enterocolitis in preterm neonates. Pediatrics. 2010;125(5):921–930. | ||
Szajewska H, Shamir R, Turck D, van Goudoever JB, Mihatsch WA, Fewtrell M. Recommendations on probiotics in allergy prevention should not be based on pooling data from different strains. J Allergy Clin Immunol. 2015;136(5):1423–1425. | ||
Walter J, Tannock GW, Tilsala-Timisjarvi A, et al. Detection and identifi-cation of gastrointestinal Lactobacillus species by using denaturing gradient gel electrophoresis and species-specific PCR primers. Appl Environ Microbiol. 2000;66(1):297–303. | ||
Kwon HS, Yang EH, Lee SH, Yeon SW, Kang BH, Kim TY. Rapid identification of potentially probiotic Bifidobacterium species by multiplex PCR using species-specific primers based on the region extending from 16S rRNA through 23S rRNA. FEMS Microbiol Lett. 2005;250(1):55–62. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.