Back to Journals » International Journal of Nanomedicine » Volume 15
Effect of Starvation in Reversing Cancer Chemoresistance Based on Drug-Resistance Detection by Dextran Nanoparticles
Authors Wang C, Gao X, Wang F, Guan W, Dou H, Xu G
Received 24 September 2020
Accepted for publication 24 October 2020
Published 20 November 2020 Volume 2020:15 Pages 9255—9264
DOI https://doi.org/10.2147/IJN.S283430
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Farooq A. Shiekh
Chenglong Wang,1 Xuzhu Gao,1 Fanchen Wang,1 Wencai Guan,1 Hongjing Dou,2 Guoxiong Xu1
1Research Center for Clinical Medicine, Jinshan Hospital, Fudan University, Shanghai 201508, People’s Republic of China; 2The State Key Laboratory of Metal Matrix Composites, School of Materials Science and Engineering, Shanghai Jiao Tong University, Shanghai 200240, People’s Republic of China
Correspondence: Guoxiong Xu
Research Center for Clinical Medicine, Jinshan Hospital, Fudan University, 1508 Longhang Road, Shanghai 201508, People’s Republic of China
Tel +86-21-34189990
Fax +86-21-57039502
Email [email protected]
Hongjing Dou
The State Key Laboratory of Metal Matrix Composites, School of Materials Science and Engineering, Shanghai Jiao Tong University, 800 Dongchuan Road, Shanghai 200240, People’s Republic of China
Tel +86-21-34202956
Fax +86-21-34202745
Email [email protected]
Introduction: Chemoresistance leads to chemotherapy failure in patients with cancer. Multidrug resistance (MDR) in cancer is mainly caused by the high expression of P-glycoprotein encoded by the MDR1 gene, which is an ATP-dependent protease. Keeping the stronger invasion and migration abilities of chemoresistant cells in cancer also requires more ATP consumption. Herein, we aimed to reverse resistance by reducing the glucose supply in the cellular environment.
Methods: A starvation approach in reversing chemoresistance was applied, which was implemented through preparing fluorescent dextran-based nanoparticles to detect the proportion of chemoresistant cells in the chemoresistant/chemosensitive cell mixture after cells cultured in a low-glucose condition.
Results: Chemoresistant cells had higher glucose consumption with higher ATPase expression and stronger glucose dependence compared to chemosensitive cells. Moreover, cancer cells cultured in a low-glucose condition reduced the proportion of chemoresistant cells.
Conclusion: Starvation therapy can be used as a new method to reverse drug resistance in cancer.
Keywords: cancer drug-resistance, P-glycoprotein, starvation therapy, nanoparticles, resistance reversal
Introduction
Chemoresistance is one of the main causes of cancer treatment failure.1 Multidrug resistance (MDR) in cancer is mainly caused by the high expression of P-glycoprotein (P-gp) encoded by the MDR1 gene. Common treatments for reversal of drug resistance include P-gp inhibitor therapy,2 gene therapy,3,4 and multidrug therapy.5 However, these treatments usually have low effectiveness and side effects.6 P-gp inhibitor therapy, for instance, can cause a lot of damages to the human body because P-gp is also expressed in the cells and tissues such as the blood-brain barrier,7 which can help protect body cells from harmful substances. On this basis, the reversal of resistance in a highly effective way with low side effects is necessary.
P-gp is an ATP-dependent molecule.8 Resistant cells need P-gp to expend a lot of energy to excrete drugs to achieve resistance, theoretically requiring more glucose than non-resistant cells. Besides, keeping the stronger invasion and migration abilities of chemoresistant cells in cancer9 also requires more ATP consumption. Therefore, it is possible to reverse resistance by reducing the glucose supply in the cellular environment. Treatment with the reduction of glucose supply in cancer is called starvation therapy. Starvation therapies, including angiogenesis suppression10 and metabolism intervention,11 are increasingly being used in cancer, which can inhibit tumor growth by reducing the carbohydrate supply or by inhibiting glycolysis.12,13
In our previous work, we have developed a facile graft copolymerization induced self-assembly (GISA) method for nanoparticle preparation,14,15 which has been proved that the fluorescence nanoparticles prepared by the GISA method can effectively distinguish the resistant from sensitive cells.9,16 Therefore, in this study, we synthesized fluorescent dextran nanoparticles (FDNPs) and detected the proportion of chemoresistant and chemosensitive cells in the chemoresistant/chemosensitive cell mixture after starvation to find the significance of the starvation approach in reversing cancer drug resistance. As shown in Scheme 1, there are large numbers of chemoresistant cells in cancer cells after treatment, while these resistant cells proliferate more slowly in glucose-deficient environments, thus decreasing the proportion of drug-resistant cells in the drug-surviving cells, which in turn reduces cancer drug resistance. As FDNPs could effectively identify and determine the proportion of chemoresistant cells, we tracked and evaluated the effectiveness of starvation therapy.
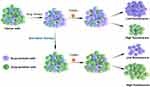 |
Scheme 1 Schematic illustration depicting starvation therapy in reversing cancer drug-resistance detected by fluorescent dextran nanoparticles (FDNPs) assay. |
Results and Discussion
Fluorescent Dextran Nanoparticles Synthesis and Flow Cytometry Assays
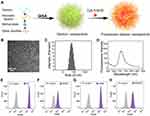
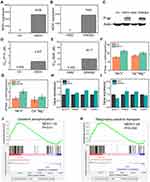
We previously developed a GISA method for the preparation of polysaccharide-based nanocarriers,14,17 and demonstrated that the fluorescent polysaccharide nanoparticles prepared by GISA can effectively distinguish chemoresistant cells from chemosensitive cells.9,16 Here we synthesized FDNPs by the GISA method. The preparation process is shown in Figure 1A. Firstly, dextran was mixed with aminated dextran. Then we carried out graft copolymerization with the polymeric monomer methyl methacrylate. Due to the hydrophobic property of polymethyl methacrylate, the grafted polymer can be spontaneously assembled into nanoparticles in water. Next, the nanoparticles were crosslinked by diallyl disulfide. Finally, fluorescent molecules Cy5.5 were labeled on the amino group of the nanoparticles. The transmission electron microscopy (TEM) image revealed that the FDNPs showed a uniform spherical structure (Figure 1B). Dynamic light scattering (DLS) showed that the FDNPs were uniformly distributed with a particle size of 50–100 nm (Figure 1C). The fluorescence spectrum showed that the FDNPs were successfully labeled by the fluorescent molecule (Figure 1D). All the results indicated that the FDNPs were successfully prepared. We selected two chemoresistant cell lines: one is a human ovarian cancer paclitaxel-resistant (DROV) cell line constructed in this laboratory and another one is a commercialized doxorubicin-resistant human immortalized myelogenous leukemia (DRK562) cell line. Their sensitive counterparts OVCAR-3 (OV) and K562 cells were also used in this study. It was proved that FGNPs can well distinguish chemoresistant cells from chemosensitive cells by flow cytometry (Figure 1E–H). As seen from the flow cytometry data, the chemoresistant cells appeared at the relatively low fluorescence position, while the chemosensitive cells appeared at the relatively high fluorescence position.
Genetic Differences Among the Chemoresistant and Chemosensitive Cancer Cells
The main reason for multi-drug resistance in cancer cells is the high expression of the multi-drug resistance gene MDR1.18,19 Both quantitative real-time polymerase chain reaction (qRT-PCR) and Western blot studies showed the higher expression of MDR1 in DROV and DRK562 cells than their sensitive counterpart OV and K562 cells (Figure 2A–C), which was consistent with the results of drug resistance assays (Figure 2D–E). MDR1 expressed P-gp, which is a drug-efflux transporter and ATP-dependent protein. ATP is an important energy molecule of cells with a high-energy phosphate group.20 The higher ATPase vitality in resistant cells was found by using ATPase kit (Figure 2F–G). The higher ATPase vitality means that drug-resistant cells have higher glucose metabolism.21 qRT-PCR detection also revealed that some genes related to glucose transporter and glycolysis, such as GLUT1,22 HK2,23 PKM2,24 and LDHA,25 were slightly upregulated in chemoresistant cells (Figure 2H–I). Furthermore, mRNA sequencing of OV and DROV cells was performed and analyzed by Gene Set Enrichment Analysis (GSEA) analysis, showing that the oxidative phosphorylation gene signature and respiratory electron transport gene signature were significantly up-regulated in chemoresistant cells (Figure 2J–K), which proved that chemoresistant cells have higher glucose metabolism than chemosensitive cells, suggesting that glucose metabolism in chemoresistant cells was oxidative phosphorylation.
Detection of High Glucose Consumption in Chemoresistant Cancer Cells
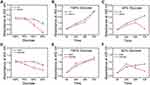
Chemoresistant cells showed higher ATPase vitality and oxidative phosphorylation, indicating that chemoresistant cells have higher energy requirements, which means that chemoresistant cells are more dependent on glucose. In order to know if the concentration of glucose in the cell culture medium had a significant effect on the cell viability of chemoresistant cells, the chemoresistant/chemosensitive cells were cultured in media containing different concentrations of glucose (11.11 mM of the glucose concentration in the RPMI-1640 medium was taken as 100%). After 48 h culture, it was found that the cell viability of chemoresistant cells was significantly decreased with the decrease of glucose concentration compared with that of chemosensitive cells (Figure 3A and D). We then cultured chemoresistant/chemosensitive cells in 100% glucose medium and 50% glucose medium for different times. It was found that the growth rate of chemoresistant/chemosensitive cells in 100% culture medium was almost the same (Figure 3B and E). By comparing to chemosensitive cells, chemoresistant cells in 50% glucose medium showed a process of growth and then rapid decrease (Figure 3C and F), indicating that chemoresistant cells exhausted the glucose in the medium and died. All the results indicated that the chemoresistant cells had higher glucose dependence than the chemosensitive cells.
Reducing Glucose in Reversing Cancer Chemoresistance
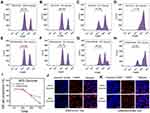
Due to tumor heterogeneity,26,27 there are different types of tumor cells, include resistant and non-resistant cells.28 In order to demonstrate whether the decrease of the concentration of glucose in the cell culture environment, known as starvation therapy, could reduce the proportion of chemoresistant cells in cancer, thus achieving a reversal of drug resistance, we mixed chemoresistant cells with chemosensitive cells in the presence of FGNPs to detect the proportion of chemoresistant cells in mixed cells cultured at different glucose concentrations and to determine whether the starvation therapy is effective in the reversal of drug resistance. As shown in Figure 4, the mixed cells were cultured at different glucose concentrations for 48 h and subsequently incubated with FGNPs for another 4 h. Flow cytometry detection showed that the proportion of chemoresistant cells was decreased with reducing glucose concentration (Figure 4A–H). After the mixed cells were cultured in a medium with a glucose concentration at 50%, the proportion of chemoresistant cells decreased with the increase of culture time (Figure 4I). Laser confocal images also showed that the proportion of chemoresistant cells was decreased with reducing glucose concentration (Figure 4J–K). All the results indicated that lowering the glucose concentration effectively decreased the proportion of chemoresistant cells in the mixed cell population, demonstrating that starvation therapy can reverse cancer resistance.
Application of E. coli in Reversing Cancer Drug-Resistance
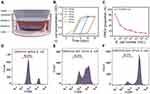
Because of the tumor hypoxic environment,29 anaerobic bacteria can accumulate at tumor areas30 and ways to treat tumors using Escherichia coli (E. coli) bacteria have been developed.31 Therefore, we wanted to know if the bacterial therapy could reduce the proportion of resistant cells by consuming glucose in the tumor microenvironment, and thus, reverse the drug resistance of the tumor. We designed a way for cancer cells to co-culture with bacteria. As shown in Figure 5A, the bacteria were cultured in a transwell chamber with a pore diameter of 0.4 μm and the cancer cells were cultured in the lower chamber. The E. coli with an OD value of 1.77 (number ~ 109/mL) was diluted at different concentrations from 102/mL to 107/mL, and cultured in RPMI-1640 medium for a different period. We found that the exponential growth time of E. coli with different initial concentrations was mainly within 24 h (Figure 5B). There was no significant difference in bacterial concentration after the exponential growth time. After cancer cells were co-cultured with the E. coli for 24 h, FGNPs were added and cultured for 4 h. After dead cells that were suspended were removed, the left cells were digested and collected for flow cytometry detection. It was found that the proportion of resistant cells decreased with the increase of bacterial concentration (Figure 5C–F). These results indicated that bacterial treatment can effectively reduce the proportion of chemoresistant cells and may reverse the drug resistance of the tumor.
Experimental Section
Cell Lines and Cell Cultures
Human ovarian cancer cell line OVCAR-3 and human immortalized myelogenous leukemia cell line K562 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). Human ovarian cancer PTX-resistant cell line OV3R-PTX referred to as DROV in the text was generated in this laboratory at Jinshan Hospital32 and obtained a Chinese Invention Patent (# ZL201410708515.7). Human immortalized myelogenous leukemia doxorubicin (adriamycin, ADR)-resistant cell line DRK562 was purchased from Keygen Biotech (Nanjing, China). The study was approved by the Ethics Committee of Jinshan Hosptial, Fudan University (# JYLLKY-2019-01-01). Sample abbreviations are shown in Table S1. All cells were cultured in RPMI-1640 medium (Gibco) supplemented with 10% fetal bovine serum (FBS, Invitrogen, Carlsbad, CA, USA).
Fluorescent Dextran Nanoparticles (FDNPs) Synthesis and Characterization
The dextran (2.25 g) was mixed with aminated dextran (0.25 g) and dissolved in 50 mL pure water. The mixture was stirred at 30 °C under nitrogen protection for 30 min. Then, 1.16 g of cerium ammonium nitrate was dissolved in 1.26 mL of 0.1 N diluted nitric acid and added to the mixture. The polymerization monomer methyl acrylate (1.02 mL) was added after 5 minutes, and 30 min later the cross-linking agent diallyl disulfide (190 µL dissolved in 5 mL of dimethyl sulfoxide) was added. After the reaction lasted for 4 h, the dextran nanoparticles were obtained by dialysis in pure water for 3 days. After 8 mL of dextran nanoparticles (6.3 mg/mL in pure water) was mixed with 0.024 mL of Cy5.5-NHS (10 mg/mL in dimethyl sulfoxide), the mixture was stirred for 12 h. The fluorescent dextran nanoparticles were obtained after dialyzed in pure water for 3 days. Hydrodynamic diameters of the fluorescent dextran nanoparticles were measured using a Zetasizer Nano ZS90 instrument (Malvern, UK) and transmission electron microscopy image was recorded using a transmission electron microscope (FEI Tecnai G2 Spirit BioTwin, 120 kV, FEI, USA).
RNA Extraction and Quantitative Real-Time PCR
Total RNA was extracted using Trizol according to the manufacturer’s instructions. PCR primer sequences are shown in Table S2. PCR amplification was performed using an SYBR Green Master kit (Roche). The expression levels of genes were normalized to an endogenous control GAPDH. The threshold cycle (Ct) was determined using the sequence detection software V1.4 (7300 real-time PCR system).
Protein Extraction and Western Blot Analysis
The cells were lysed in sodium dodecyl sulfate lysate buffer with 1% benzyl sulfonyl fluoride and 1% phosphatase inhibitor. After SDS-polyacrylamide gel electrophoresis was performed, the primary antibodies of rabbit anti-MDR1 (1:50,000 dilution) and mouse anti-β-actin (1:50,000 dilution) and the secondary antibodies of goat anti-rabbit and mouse IgG (1:10,000 dilution) labeled with horseradish peroxidase were used. The signal was detected by a chemiluminescence imaging system (Tanon Science & Technology, China).
ATPase Vitality Assays
ATPase vitality was detected according to the instructions of the ATPase kit (Keygen Biotech, China). In brief, Na+K+ and Ca++Mg++ ATPase decompose ATP to produce ADP and inorganic phosphorus. Then, the vitality of ATPase was determined by measuring the amount of inorganic phosphorus. ATPase vitality was calculated according to the equation: ATPase vitality

Detection of Cell Vitality and Half-Maximal Inhibitory Concentration (IC50)
Cells were seeded with a density of 1×104 cells/well in 96-well plates and cultured in 100 µL of RPMI-1640 medium (containing various concentrations of glucose between 2.78 and 11.11 mM). At least four parallel wells were performed for each sample. Cell viability was determined using the CCK-8 kit. The optical density (OD) of each well was measured by a microplate reader (BioTek Epoch, USA) at 450 nm. Cells viabilities were calculated according to the equation: Cell viability (%) = ODexperment/ODcontrol×100%. Cells with a density of 1×104 cells/well were cultured in 96-well plates for 24 h. Then the supernatant was removed and 100 μL RPMI-1640 solution (containing PTX with the concentrations ranged from 0.001 to 10 μM or ADR with the concentrations ranged from 0.01 to 100 μM) was added into each well after 48 h. The cell vitality was determined using the CCK-8 assay and the IC50 was calculated using the GraphPad prism software.
RNA Sequencing Analysis
The cells were divided into the drug-resistant cell group and the drug-sensitive cell group. The drug-resistant cell group included DROV monoclonal cells selected from 2 μM of paclitaxel intervention, DROV polyclonal cells selected from 2 μM of paclitaxel intervention, and DROV monoclonal cells selected from 30 μM of paclitaxel intervention. The drug-sensitive cells group included two independently cultured OV cells. mRNA was isolated by Trizol. mRNA sequencing was performed by Shanghai Biochip Co. Ltd. (Shanghai, China) and analyses were performed using Stringtie and HISAT2. Broad GSEA was used to analyze the gene signature enrichment based on the Molecular Signature Database (MSigDB).
Flow Cytometry Analysis
The cells were treated with 10 µg/mL of FDNPs for 4 h and then washed twice with cold PBS (pH 7.4). After cells were collected and suspended by PBS, the cells were immediately detected by flow cytometry (Gallios, Beckman, CA, USA). Flow cytometry data were analyzed using FlowJo (V10).
Laser Confocal Microscopy Assays
The cells were inoculated in a confocal petri dish (diameter 35mm) at 0.5 mL/dish. Cells were then incubated in RPMI-1640 medium with different glucose concentrations for 48 h. Cells were then treated with FDNPs at a concentration of 10 µg/mL for 4 h, washed with PBS for 3 times, stained by DAPI, and imaged with a laser confocal microscope (SP8 STED 3X, Leica, Germany).
Co-Culture of Cells and Bacteria
Co-culture of cells with bacteria was carried out using transwell chambers (Corning, VA, USA) DROV/OV mixed cells (1×106) in 600 μL of RPMI-1640 medium were placed in the lower compartment of the chambers. After incubation for 24 h, 100 µL of RPMI-1640 medium containing various concentrations of E. coli were added to the upper compartment of the chambers. After 24 h of co-culturing, cells were treated with 10 µg/mL of FDNPs for 4 h and then collected and detected by flow cytometry.
Statistical Analyses
Results are presented as the mean ± standard error (SE) for qRT-PCR and IC50 values and the mean ± standard deviation (SD) for cell viability, ATPase assays, and cell counting.
Conclusion
The current study evaluates the effectiveness of starvation therapy. A fluorescent dextran nanoparticle has been prepared to detect the proportion of chemoresistant cells in chemoresistant/chemosensitive cell mixture in real-time. The proportion of chemoresistant cells in the mixed cell population was significantly decreased after reducing glucose concentration in the culture medium, demonstrating that the reversal of drug resistance may be achieved by the starvation approach. The co-culture of cancer cells with E. coli can reduce the proportion of resistant cells. Thus, starvation therapy can be used as a new method to reverse drug resistance in cancer.
Data Sharing Statement
The authors declare that the main data supporting the findings of this study are available within the article files. Extra data are available from the authors on reasonable request.
Acknowledgments
The authors wish to gratefully acknowledge the financial support provided by the National Natural Science Foundation of China (No. 81872121 and 21871180) and the Science and Technology Commission of Shanghai Municipality (No. 17ZR1404100 and 18520710300).
Disclosure
The authors report no conflicts of interest for this work.
References
1. Robey RW, Pluchino KM, Hall MD, Fojo AT, Bates SE. Gottesman, structure of p-glycoprotein reveals a molecular basis for poly-specific drug binding. Nat Rev Cancer. 2018;18(7):452–464. doi:10.1038/s41568-018-0005-8
2. Leopoldo M, Nardulli P, Contino M, Leonetti F, Luurtsema G, Colabufo NA. An updated patent review on P-glycoprotein inhibitors (2011–2018). Expert Opin Ther Pat. 2019;29(6):455–461. doi:10.1080/13543776.2019.1618273
3. Naghizadeh S, Mansoori B, Mohammadi A, Sakhinia E, Baradaran B. Gene silencing strategies in cancer therapy: an update for drug resistance. Curr Med Chem. 2019;26(34):6282–6303.
4. Takahashi S, Ito Y, Hatake K, Sugimoto Y. Review of clinical gene therapy trials for breast cancer and MDR1 gene therapy trial in cancer institute hospital. Breast Cancer. 2006;13(1):8–15.
5. Saneja A, Dubey RD, Alam N, Khare V, Gupta PN. Co-formulation of P-glycoprotein substrate and inhibitor in nanocarriers: an emerging strategy for cancer chemotherapy. Curr Cancer Drug Targets. 2014;14(5):419–433. doi:10.2174/1568009614666140407112034
6. Nikolaou M, Pavlopoulou A, Georgakilas AG, Kyrodimos E. The challenge of drug resistance in cancer treatment: a current overview. Clin Exp Metastasis. 2018;35(4):309–318. doi:10.1007/s10585-018-9903-0
7. Miller DS, Regulation of P-glycoprotein and other ABC drug transporters at the blood-brain barrier. Trends Pharmacol Sci. 2010;31(6):246–254. doi:10.1016/j.tips.2010.03.003
8. Aller SG, Yu J, Ward A, et al. Structure of p-glycoprotein reveals a molecular basis for poly-specific drug binding. Science. 2009;323(5922):1718–1722. doi:10.1126/science.1168750
9. Wang CL, Wang FC, Zhang JG, Liu LS, Xu GX. Dou, fluorescent polysaccharide nanogels for the detection of tumor heterogeneity in drug-surviving cancer cells. Adv Biosyst. 2020;4(2):1900213. doi:10.1002/adbi.201900213
10. Biziota E, Mavroeidis L, Hatzimichael E, Pappas P. Metronomic chemotherapy: A potent macerator of cancer by inducing angiogenesis suppression and antitumor immune activation. Cancer Lett. 2017;400:243–251. doi:10.1016/j.canlet.2016.12.018
11. Vernieri C, Casola S, Foiani M, Pietrantonio F, de Braud F, Longo V. Targeting cancer metabolism: dietary and pharmacologic interventions. Cancer Discov. 2016;6(12):1315–1333. doi:10.1158/2159-8290.CD-16-0615
12. Lee C, Raffaghello L, Longo VD. Starvation, detoxification, and multidrug resistance in cancer therapy. Drug Resist Update. 2012;15(1–2):114–122. doi:10.1016/j.drup.2012.01.004
13. Yu SJ, Chen ZW, Zeng X, Chen XS, Gu Z. Advances in nanomedicine for cancer starvation therapy. Theranostics. 2019;9(26):8026–8047. doi:10.7150/thno.38261
14. Wang H, Dai TT, Li SL, et al. Scalable and cleavable polysaccharide nanocarriers for the delivery of chemotherapy drugs. Acta Biomater. 2018;72:206–216. doi:10.1016/j.actbio.2018.03.024
15. Wang CL, Guan WC, Peng JL, Chen YT, Xu GX, Dou HJ. Gene/paclitaxel co-delivering nanocarriers prepared by framework-induced self-assembly for the inhibition of highly drug-resistant tumors. Acta Biomater. 2020;103:247–258. doi:10.1016/j.actbio.2019.12.015
16. Wang CL, Guan WC, Chen R, et al. Fluorescent glycan nanoparticle-based FACS assays for the identification of genuine drug-resistant cancer cells with differentiation potential. Nano Res. 2020;13:3110–3122. doi:10.1007/s12274-020-2981-8
17. Wang CL, You JY, Gao MM, Zhang PP, Xu GX, Dou HJ. Bio-inspired gene carriers with low cytotoxicity constructed via the assembly of dextran nanogels and nano-coacervates. Nanomedicine. 2020;15:1285–1296. doi:10.2217/nnm-2020-0065
18. Takara K, Sakaeda T, Okumura K. An update on overcoming MDR1-mediated multidrug resistance in cancer chemotherapy. Curr Pharm Des. 2006;12(3):273–286. doi:10.2174/138161206775201965
19. Ieiri I. Functional significance of genetic polymorphisms in P-glycoprotein (MDR1, ABCB1) and breast cancer resistance protein (BCRP, ABCG2). Drug Metab Pharmacokinet. 2012;27(1):85–105. doi:10.2133/dmpk.DMPK-11-RV-098
20. Sak K, Jarv J, Karelson M. ‘Strain effect’ descriptors for ATP and ADP derivatives with modified phosphate groups. Comput Chem. 2002;26(4):341–346.
21. Kato Y, Maeda T, Suzuki A, Baba Y. Cancer metabolism: new insights into classic characteristics. Jpn Dent Sci Rev. 2018;54(1):8–21. doi:10.1016/j.jdsr.2017.08.003
22. Meng Y, Xu X, Luan HY, et al. The progress and development of GLUT1 inhibitors targeting cancer energy metabolism. Future Med Chem. 2019;11(17):2333–2352. doi:10.4155/fmc-2019-0052
23. Garcia SN, Guedes RC, Marques MM. Unlocking the potential of HK2 in cancer metabolism and therapeutics. Curr Med Chem. 2019;26(41):7285–7322.
24. Li YH, Li XF, Liu JT, et al. PKM2, a potential target for regulating cancer. Gene. 2018;668:48–53. doi:10.1016/j.gene.2018.05.038
25. Thongon N, Zucal C, D’Agostino VG, et al. Cancer cell metabolic plasticity allows resistance to NAMPT inhibition but invariably induces dependence on LDHA. Cancer Metab. 2018;6:1. doi:10.1186/s40170-018-0174-7
26. Bedard PL, Hansen AR, Ratain MJ, Siu LL. Tumour heterogeneity in the clinic. Nature. 2013;501(7467):355–364. doi:10.1038/nature12627
27. Maishi N, Annan DA, Kikuchi H, Hida Y, Hida K. Tumor endothelial heterogeneity in cancer progression. Cancers. 2019;11(10):1511. doi:10.3390/cancers11101511
28. Jeon JH, Kim DK, Shin Y, et al. Migration and invasion of drug-resistant lung adenocarcinoma cells are dependent on mitochondrial activity. Exp Mol Med. 2016;48:e277. doi:10.1038/emm.2016.129
29. Petrova V, Annicchiarico-Petruzzelli M, Melino G, Amelio I. The hypoxic tumour microenvironment. Oncogenesis. 2018;7:10. doi:10.1038/s41389-017-0011-9
30. Tirandaz H, Mohammadi E. Efficient tumor targeting by anaerobic butyrate-producing bacteria. Med Hypotheses. 2013;80(5):675–678. doi:10.1016/j.mehy.2013.01.024
31. Zheng DW, Chen Y, Li ZH, et al. Optically-controlled bacterial metabolite for cancer therapy. Nat Commun. 2018;9:1680. doi:10.1038/s41467-018-03233-9
32. Wang FC, Zhang LY, Liu J, Zhang JG, Xu GX. Highly expressed STAT1 contributes to the suppression of sternness properties in human paclitaxel-resistant ovarian cancer cells. Aging-US. 2020;12(11):11042–11060. doi:10.18632/aging.103317
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.