Back to Journals » Cancer Management and Research » Volume 10
Distinct prognosis of mRNA expression of the five RecQ DNA-helicase family members – RECQL, BLM, WRN, RECQL4, and RECQL5 – in patients with breast cancer
Authors Zhu X , Chen H, Yang Y , Xu C , Zhou J, Zhou J, Chen Y
Received 29 August 2018
Accepted for publication 2 November 2018
Published 5 December 2018 Volume 2018:10 Pages 6649—6668
DOI https://doi.org/10.2147/CMAR.S185769
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Chien-Feng Li
Xuan Zhu,1,2,* Huihui Chen,1,* Yi Yang,1,3 Chunjing Xu,1,2 Jun Zhou,4 Jiaojiao Zhou,1,2 Yiding Chen1,2
1Department of Surgical Oncology, Second Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, Zhejiang, People’s Republic of China; 2The Key Laboratory of Cancer Prevention and Intervention of China National Ministry of Education, The Key Laboratory of Molecular Biology in Medical Sciences of Zhejiang Province, Cancer Institute, Zhejiang University School of Medicine, Hangzhou, Zhejiang, People’s Republic of China; 3Department of Breast Surgery, Jiaxing Maternity and Child Health Care Hospital, Jiaxing, Zhejiang, People’s Republic of China; 4Department of Breast Surgery, Women’s Hospital, School of Medicine, Zhejiang University, Hangzhou, Zhejiang, People’s Republic of China
*These authors contributed equally to this work
Background: Five RecQ helicase family members have a role in maintaining genome stability. However, their prognostic roles in breast cancer remain unknown. We aimed to investigate the prognostic values of the RecQ family and clinical outcomes in breast cancer.
Methods: We used the Kaplan–Meier Plotter database (http://kmplot.com/analysis) to analyze prognostic values of RecQ-family mRNA expression in all breast cancers and in different intrinsic subtypes and clinicopathological characteristics. Protein-expression levels of WRN and RECQL4 were confirmed by immunohistochemistry (IHC) in breast cancer tissues.
Results: Increased expression of RECQL mRNA was significantly associated with reduced relapse-free survival (RFS) and postprogression survival (PPS) in all breast cancers, and improved overall survival (OS) in patients with basal-like breast cancer and in mutant-p53-type breast cancer patients. Increased expression of BLM mRNA was correlated with reduced distant metastasis-free survival (DMFS) in all patients. Increased expression of WRN mRNA was associated with improved OS and RFS in breast cancer patients. Increased expression of RECQL4 mRNA was associated with reduced OS, DMFS, and RFS in all breast cancers, and with reduced OS in patients with luminal A, HER2-positive, ER-positive, and PR-positive breast cancer. Increased expression of RECQL5 mRNA was associated with improved RFS in all patients, and with improved OS in patients with lymph-node-negative breast cancer, but with reduced OS in patients with HER2-positive breast cancer. IHC staining confirmed that high expression of WRN was correlated with increased OS and high expression of RECQL4 associated with reduced OS at protein levels.
Conclusion: mRNA-expression levels of RecQ members were significantly correlated with prognosis in breast cancer patients. These preliminary findings require further study to determine whether RecQ-targeting reagents might be developed for clinical application in breast cancer.
Keywords: breast cancer, prognosis, RecQ helicase family, DNA helicase, overall survival, KM plotter
Introduction
Worldwide, breast cancer accounts for 15% of all cancer deaths in women.1 Although early detection through mammography, improved diagnostic technology, and improved treatments have evolved,2 the prevalence of breast cancer continues to rise in many parts of the world, including South America, Africa, and Asia,3 and mortality from breast cancer remains a global issue.4
The RecQ family of DNA helicases is one of the most highly conserved groups of helicases throughout evolution and is necessary for the stability of the whole genome.5 In humans, five RecQ-helicase proteins have been recognized: RECQL, BLM, WRN, RECQL4, and RECQL5. There are three conserved regions in human RecQ proteins: the helicase domain, the RecQ carboxy-terminal (RQC) domain, and the helicase and RNase DC-terminal (HRDC) domain. Most of the DNA-helicase superfamily utilizes energy derived from ATP hydrolysis to unwind dsDNA, including secondary DNA structures, and some helicases also influence protein–DNA complexes.6 RecQ helicases bind to and translocate along one DNA strand in a 3′→5′ direction, and ssDNA stimulates their dsDNA-unwinding activity. RecQ helicases separate double-stranded nucleic acids, and are essential for every aspect of DNA metabolism, including recombination, replication, and DNA repair.7
Mutations in BLM, WRN, or RECQL4, located on chromosomes 15q26.1, 8p12 and 8q24.3 respectively, were found to be associated with some diseases in humans.8 Less clear is whether dysregulation of RecQ-family expression is related to an increased risk of cancer. RECQL is overexpressed in multiple myeloma and associated with poor prognosis,9 and the expression of RECQL is correlated with tumorigenesis in breast cancer.10 Recent studies also showed that abnormal expression of the BLM protein might lead to the development of malignant tumors,11–13 while overexpression of BLM stimulates DNA injury and induces sensitivity to cisplatin chemotherapy in triple-negative breast cancer.14 WRN is considered a tumor suppressor, and it has been shown to inhibit tumor growth in nude mouse xenograft models of breast cancer.15 RECQL4 is overexpressed in breast cancer cell lines, and depletion of RECQL4 facilitates apoptosis.16 Arvind et al assessed RECQL5 expression in breast cancers, and found that expression levels of RECQL5 were associated with aggressive phenotypes and poor survival outcomes.17 The prognostic roles of each RecQ member, especially at the mRNA level, remain poorly understood in breast cancer.
Therefore, the aims of this study were to use an online Kaplan–Meier plotter database to investigate the prognostic role of mRNA expression of the five RecQ DNA-helicase family members – RECQL, BLM, WRN, RECQL4, and RECQL5 – and clinical outcomes in women with breast cancer. We also confirmed the protein-expression levels of WRN and RECQL4 by immunohistochemistry (IHC) in breast cancer tissue.
Methods
Use of online databases, including Kaplan–Meier Plotter
The association of RecQ family member mRNA expression with prognostic value was analyzed using the online Kaplan–Meier Plotter breast cancer database (http://kmplot.com/analysis/index.php), which was produced using gene-expression data and survival information downloaded from the Gene Expression Omnibus (http://www.ncbi.hlm.nih.gov/geo).18
Breast cancer patient-outcome data
At the time of the study, the Kaplan–Meier Plotter database was capable of evaluating the effect of mRNA expression levels of 54,675 genes on patient survival using 10,641 cancer samples, which included breast cancer,18 ovarian cancer,19 lung cancer,20 and gastric cancer.21 There were 3,955 breast cancer patients with relapse-free survival (RFS) data, 1,402 with overall survival (OS) data, 1,747 with distant metastasis-free survival (DMFS) data, and 414 with postprogression survival (PPS) data. ER, PR, HER2, and lymph-node status, tumor grade, intrinsic breast-tumor subtype, and TP53 gene-expression data were collected and analyzed.
Tissue microarrays (TMAs) and IHC staining
TMAs were purchased from Outdo Biotech (Shanghai, China), and contained 160 breast cancer samples. Detailed clinicopathological characteristics of breast cancer samples are provided in Table S1. Approval from the Ethics Committee of the Second Affiliated Hospital, Zhejiang University School of Medicine was obtained for the experiments. IHC staining of the TMAs was performed using primary antibodies against WRN (ab200; Abcam, Cambridge, UK) or RECQL4 (25470002; Novus Biologicals, Littleton, CO, US). Briefly, paraffin slides were baked at 60°C overnight, deparaffinized with 100% xylene, and rehydrated in graded ethanol. After antigen retrieval and inactivation of endogenous peroxidases, slides were preincubated in 10% normal goat serum and incubated with WRN (1:200) or RECQL4 (1:400) overnight at 4°C. Subsequently, slides were serially washed and incubated with biotinylated secondary antibody for 1 hour at room temperature. Finally, slides were stained with DAB and counterstained with hematoxylin.
WRN or RECQL4 expression in TMAs was blindly evaluated by two pathologists using the immunoreactivity score (IRS), based on both the percentage of positive cells and the intensity of staining. The percentage of positive cells was graded as follows: 0 (negative), 1 (<10%), 2 (10%–50%), 3 (51%–80%), 4 (>80%). The intensity of staining was graded as follows: 0 (no color reaction), 1 (mild reaction), 2 (moderate reaction), 3 (intense reaction). IRS ranging from 0 to 12 was calculated by multiplication of the two scores. In this study, WRN or RECQL4 expression was defined as low (IRS ≤6) or high (IRS >6).
Data analysis
In this study, we entered five RecQ DNA-helicase family members (RECQL, BLM, WRN, RECQL4, and RECQL5) into the database (http://kmplot.com/analysis/index.php?P=service%20&cancer%20=%20breast) and analyzed them under different clinical parameters. We only selected the JetSet best probe set of five RecQ DNA-helicase family members to obtain Kaplan–Meier survival plots. mRNA expression of related genes above or below the median divides the cases into high and low expression, respectively. Associations between protein-expression levels of WRN or RECQL4 and clinicopathological features were analyzed using Pearson’s χ2 test. In both Kaplan–Meier Plotter and prognostic relevance of WRN and RECQL4 protein levels in breast cancer patients, survival analysis was estimated according to the Kaplan–Meier method and log-rank test. HR and 95% CI were calculated using the Cox proportional-hazard regression model. P<0.05 was considered statistically significant.
Results
Prognosis in patients with breast cancer and mRNA expression of the five RecQ DNA-helicase family members
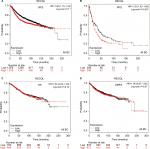
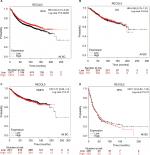
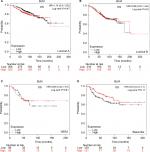
Patient-survival data and mRNA expression of the five RecQ family members in breast cancer patients were obtained from www.kmplot.com. OS, DMFS, RFS, and PPS curves were plotted. The prognostic significance of RECQL mRNA expression was evaluated from the data obtained (Affymetrix ID for RECQL: 212918_at). Increased mRNA-expression levels of RECQL were significantly associated with reduced RFS (HR 1.28, 95% CI 1.15–1.42; P<0.001; Figure 1A), and reduced PPS (HR 1.32, 95% CI 1.03–1.68; P=0.027; Figure 1B). However, mRNA-expression levels of RECQL were not significantly correlated with OS (HR 1.04, 95% CI 0.84–1.28; P=0.74) or DMFS (HR 1.06, 95% CI 0.87–1.28; P=0.57; Figure 1C and D).
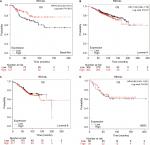
The prognostic significance of BLM mRNA expression was evaluated from the data obtained (Affymetrix ID for BLM: 205733_at). Increased mRNA-expression levels of BLM were significantly associated with reduced DMFS (HR 1.28, 95% CI 1.06–1.56; P=0.011; Figure 2A). However, mRNA-expression levels of BLM were not significantly correlated with OS (HR 1.05, 95% CI 0.85–1.3; P=0.65), RFS (HR 1.02, 95% CI 0.91–1.13; P=0.76), or PPS (HR 1.18, 95% CI 0.92–1.5; P=0.19; Figure 2B–D).
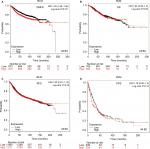
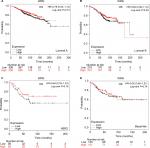
The prognostic significance of WRN mRNA expression was evaluated from the data obtained (Affymetrix ID for WRN: 205667_at). Increased mRNA-expression levels of WRN were associated with improved patient OS (HR 0.76, 95% CI 0.61–0.94; P=0.011; Figure 3A) and increased RFS (HR 0.81, 95% CI 0.73–0.91; P<0.001; Figure 3B). However, WRN mRNA expression was not significantly correlated with DMFS (HR 0.87, 95% CI 0.71–1.05; P=0.15) or PPS (HR 1.21, 95% CI 0.95–1.55; P=0.12; Figure 3C and D).
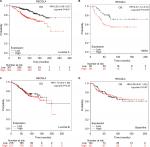
The prognostic significance of RECQL4 mRNA expression was evaluated from the data obtained (Affymetrix ID for RECQL4: 213520_at). Increased mRNA-expression levels of RECQL4 were significantly associated with reduced patient OS (HR 1.74, 95% CI 1.4–2.16; P<0.001; Figure 4A), DMFS (HR 1.53, 95% CI 1.26–1.86; P<0.001; Figure 4B), and RFS (HR 1.65, 95% CI 1.48–1.84; P<0.001; Figure 4C). Also, high expression of RECQL4 mRNA was modestly correlated with poor PPS (HR 1.27, 95% CI 0.99–1.62; P=0.055; Figure 4D), but not significantly.
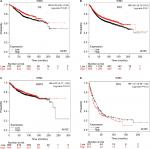
The prognostic significance of RECQL5 mRNA expression was evaluated from the data obtained (Affymetrix ID for RECQL5: 221686_s_at). Increased mRNA-expression levels of RECQL5 were significantly associated with increased RFS (HR 0.83, 95% CI 0.74–0.93; P<0.001; Figure 5A), but not significantly correlated with patient OS (HR 0.98, 95% CI 0.79–1.21; P=0.84), DMFS (HR 1.07, 95% CI 0.88–1.3; P=0.47), or PPS (HR 1.04, 95% CI 0.82–1.33; P=0.72; Figure 5B–D).
Prognosis in patients with mRNA expression of the five RecQ DNA-helicase family members and intrinsic subtypes
To investigate the role of the five RecQ helicase family members further, the prognostic significance of each was evaluated with the different intrinsic breast cancer subtypes identified for patients in the database. The intrinsic subtypes included luminal A, luminal B, HER2-positive, and basal-like breast cancer. Expression levels of mRNA for BLM and WRN and the intrinsic breast cancer subtypes showed no significant prognostic associations (Figures S1 and S2). Increased expression of RECQL mRNA was significantly associated with increased patient OS in patients with basal-like breast cancer (HR 0.54; 95% CI 0.32–0.91; P=0.019; Figure 6A). However, there was no significant difference in patient OS between patients with luminal A, luminal B, and HER2-positive type breast cancer (Figure 6B–D).
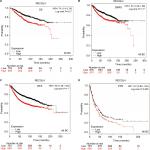
Increased expression levels of RECQL4 mRNA were significantly associated with reduced OS in patients with intrinsic luminal A-type breast cancer (HR 2.26, 95% CI 1.56–3.27; P<0.001; Figure 7A) and HER2-positive breast cancer (HR 2.2, 95% CI 1.12–4.31; P=0.019; Figure 7B). However, mRNA-expression levels of RECQL4 were not significantly correlated with prognosis in patients with intrinsic luminal B or basal-like breast cancer subtypes (Figure 7C, D).
As shown in Figure 8A, increased expression levels of RECQL5 mRNA were significantly associated with reduced patient OS (HR 2.09, 95% CI 1.05–4.17; P=0.032) in patients with HER2-positive breast cancer. However, expression levels of RECQL5 mRNA were not significantly correlated with OS in patients with the other three intrinsic subtypes of breast cancer (Figure 8B–D).
Prognosis in patients with mRNA expression of the five RecQ DNA-helicase family members and patient clinicopathological characteristics
Patient clinicopathological characteristics included tumor ER status, PR status, positive lymph-node status, grade, and TP53-mutation status. Increased RECQL4 mRNA expression was significantly correlated with reduced patient OS in ER-positive breast cancer (HR 1.9, 95% CI 1.33–2.74; P<0.001), and increased WRN mRNA expression was moderately correlated with increased patient OS in ER-negative breast cancer (HR 0.64, 95% CI 0.4–1.02; P=0.056; Table 1).
In patients with PR-positive breast cancer, only RECQL4 mRNA expression was significantly associated with reduced patient prognosis (HR 4.87, 95% CI 1.01–23.52; P=0.029; Table 2). As shown in Table 3, high levels of RECQL5 mRNA expression were associated with improved patient prognosis for patients with breast cancer who were lymph-node-negative (HR 0.68, 95% CI 0.47–0.99; P=0.043).
As shown in Table 4, there was no significant correlation between mRNA-expression levels of the five RecQ DNA helicase family members and histopathological grades of the breast cancers. Table 5 shows that high expression levels of RECQL mRNA were significantly associated with favorable prognosis in mutant-p53-type breast cancer patients (HR 0.33, 95% CI 0.13–0.82; P=0.013).
Prognosis in patients with protein expression of WRN and RECQL4
To analyze the association between mRNA expression of RecQ family members and prognosis in breast cancer patients, we used the Kaplan–Meier Plotter database and found that high mRNA-expression levels of WRN were significantly associated with improved OS and high mRNA-expression levels of RECQL4 significantly associated with reduced OS in patients with breast cancers. mRNA-expression levels of RECQL, BLM, and RECQL5 were not significantly correlated with OS. Then, we investigated if WRN and RECQL4 protein levels were significantly associated with OS in breast cancer patients. We performed IHC in breast cancer TMAs.
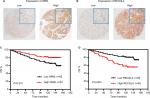
As some cores within the TMAs were missing in the process of IHC staining, there were 152 of 160 tumor cores suitable for analysis of WRN protein expression and 147 of 160 tumor cores suitable for analysis of RECQL4 protein expression. Figure 9A and B show representative images of WRN-stained and RECQL4-stained TMAs. WRN and RECQL4 exhibited high expression rates in breast cancer tissue, with values of 44.1% (67 of 152) and 37.4% (55 of 147), respectively. Moreover, Kaplan–Meier analysis showed that high WRN protein expression was significantly associated with increased OS in breast cancer patients (P=0.031, Figure 9C) and patients with high RECQL4 levels displayed reduced OS (P=0.003, Figure 9D). The results suggested that at both mRNA and protein levels, high WRN expression was correlated with increased OS and high RECQL4 expression with reduced OS.
Discussion
The aims of this study were to use a Kaplan–Meier plotter database to investigate the prognostic value of expression of the five RecQ DNA helicase family members – RECQL, BLM, WRN, RECQL4, and RECQL5 – and clinical outcomes in women with breast cancer. The findings of the study showed that increased mRNA-expression levels of RECQL were significantly associated with reduced RFS and PPS in all breast cancers and improved OS in patients with basal-like breast cancer and mutant-p53-type breast cancer. Increased mRNA-expression levels of BLM were correlated with reduced DMFS in all patients. Increased mRNA-expression levels of WRN were significantly associated with improved OS and RFS in patients with breast cancers. High mRNA-expression levels of RECQL4 were significantly associated with reduced OS, DMFS, and RFS in patients with breast cancers, and reduced OS in patients with luminal A-type breast cancer, HER2-positive breast cancer, ER-positive breast cancer, and PR-positive breast cancer. High mRNA-expression levels of RECQL5 were associated with improved RFS in all breast cancer patients, including those with lymph-node-negative cancer, but with reduced OS in patients with HER2-positive breast cancer. IHC staining confirmed that high WRN expression was correlated with increased OS and high expression of RECQL4 associated with reduced OS, just as the results of mRNA levels showed. As mRNA and protein expression of both members were investigated in two independent breast cancer cohorts, we were unable to correlate directly between mRNA and protein expression in the current study.
mRNA-expression levels of each five RecQ DNA-helicase family members had different prognostic significance in breast cancer, which suggested that each RecQ helicase member has a particular biological function in breast cancer. However, the mechanisms of how each RecQ family member contributes to breast cancer remain unknown, even though each RecQ family member has been shown to have a key role in preserving genome stability by taking effects in DNA replication, recombination, DNA-damage signaling, DNA-repair pathways, and telomere maintenance.6,22,23
RECQL is the smallest, but most abundant member of the human RecQ helicase family, and has been shown to be widely expressed in the cell nucleus.24,25 RECQL has been shown to be highly expressed in solid tumors, including glioblastoma,26 head and neck squamous-cell carcinoma,27 ovarian cancer,28 hepatocellular carcinoma,29 and squamous-cell carcinoma of the tongue.30 RECQL has also been shown to be expressed at high levels in non-small-cell lung cancer cell lines when compared with a normal bronchial epithelial cell line, and it has been proposed to have a role in lung cancer by regulating epithelial–mesenchymal transition.31 RECQL is significantly overexpressed in multiple myeloma cells when compared with normal plasma cells, and has been shown to be associated with poor prognosis.9 It has been hypothesized that tumors may depend on RECQL to tolerate replication-induced DNA damage. Recently, Arora et al showed that high RECQL mRNA expression was correlated with improved breast cancer-specific survival (BCSS) in patients with ER-positive breast cancer.32 However, in this present study, there was no correlation between the mRNA expression of RECQL and prognosis in ER-positive breast cancer, but increased mRNA expression of RECQL was associated with reduced RFS and PPS in all breast cancer patients and improved OS in patients with basal-like breast cancer, and mutant-p53-type breast cancer. The controversial findings could be due to different patient cohorts and limited patient populations.
BLM is focally located to promyelocytic leukemia bodies and telomeres in the cell nucleus.33,34 The expression of BLM varies in the cell cycle, and is increased in the S and G2/M phases.35 BLM is a critical helicase for DNA metabolism, replication, and repair.6 Mutation of BLM is associated with Bloom’s syndrome, which is a rare and autosomal-recessive hereditary clinical disease that is characterized by proportional dwarfism, and which can be associated with the early development of several types of cancer.36 BLM is overexpressed in some tumor cells, and abnormalities in BLM are associated with genome instability, which has been shown in recent studies to lead to tumorigenesis.11–13 Recently, Qian et al showed that BLM was overexpressed in prostate cancer cells and tissue compared with benign prostatic hyperplasia cells and nonprostate cancer tissue.37 In 2015, Arora et al showed that in breast cancer, overexpression of BLM mRNA was significantly correlated with increased histopathological grade, larger tumors, ER-negative tumors, PR-negative tumors, and triple-negative tumor phenotypes, and associated with reduced BCSS.38 BLM has a role in DNA replication and cell proliferation,39 which may be a reason for the presence of high BLM mRNA in invasive breast cancer. The results of this present study showed that high mRNA expression of BLM was correlated with reduced DMFS, although no significant associations were found between the mRNA expression of BLM and tumor grade, ER-negative, PR-negative, or triple-negative breast cancer status. These discrepant results might be due to different methods in determining cutoffs. Second, the number of patients in our study was relatively small.
WRN is located primarily in the nucleolus in normal cells, but after DNA damage it rapidly relocates to other nuclear regions.40–43 The gene responsible for Werner’s syndrome was recognized by positional cloning,44 and in addition to having 3′→5′ helicase activity, the WRN protein also has 3′→5′ exonuclease activity, unlike other RecQ DNA helicases.6 Loss of function of WRN can result in the clinical condition of Werner’s syndrome, which is characterized by an increased incidence of tumors, including follicular thyroid carcinoma, malignant melanoma, and soft-tissue sarcoma.45–47 The WRN gene plays an important role in DNA repair, replication, recombination, and maintenance of nuclear telomeres, which means that reduced expression of WRN results in genomic instability.48–50 Malignant tumor cells might be able to increase WRN expression to protect the stability of their genome, as increased protein expression of WRN has been demonstrated in some cancer cell lines and knockdown of WRN to induce cell death.51 However, there have been few studies on the role of WRN in breast cancer. In this present study, the findings showed that high levels of WRN mRNA expression were significantly associated with increased patient OS and RFS and modestly associated with increased OS in ER-negative breast cancer patients. At the protein level, we observed high WRN levels in 44.1% of breast cancers, and similarly high WRN expression was correlated with increased OS. Research on WRN has primarily focused on in vitro study, and future clinical studies are needed to validate our findings.
RECQL4 is distributed in the nucleus and cytoplasm in different types of cells and tissue.52 RECQL4 is sometimes located in mitochondria, where it may have a role in preserving mitochondrial DNA integrity.53,54 Mutations in the RECQL4 gene can lead to clinical disorders, including Rothmund–Thomson syndrome, which includes growth retardation and a predisposition to cancer, and the RAPADILINO syndrome.52 RECQL4 has been reported to be highly expressed in osteosarcoma cells55 and prostate cancer cells where tumor growth was shown to be reduced by siRNA-mediated suppression of RECQL4.56 Increased expression of RECQL4 has also been previously reported in breast cancer,16 laryngeal cancer,57 cervical cancer,58 and oral cancer.59 In a previous study, increased expression of RECQL4 mRNA (51%), RecQL4 protein (23%), and RECQL4 gene amplification (30.6%) were correlated with invasive tumor characteristics and poor survival in patients with breast cancer.60 Fang et al reported that RECQL4 might interact with survivin to regulate the survival of breast cancer cells.16 The findings of the present study showed that high mRNA expression of RECQL4 was significantly associated with decreased OS, DMFS, and RFS in patients with breast cancer and decreased OS in patients with luminal A-type breast cancer, HER2-positive, ER-positive, and PR-positive breast cancer. At the protein level, we observed high RECQL4 levels in 37.4% of breast cancers, and similarly high expression of RECQL4 was associated with increased OS. This current study was a retrospective study, and our results require confirmation in prospective studies with larger populations.
RECQL5 maintains the stability and integrity of the genome, but there are no reported hereditary diseases associated with defects in the RECQL5 gene. However, RECQL5 genetic polymorphisms have been reported to be associated with laryngeal carcinoma,61 osteosarcoma,62 and breast cancer.63 Studies have shown that expression of RECQL5 is decreased in colorectal cancer64 and osteosarcoma.65 In 2015, Wu et al reported that RECQL5 might have a role as a tumor suppressor in osteosarcoma and as a potential target for the treatment of osteosarcoma.65 In 2016, Patterson et al showed that RECQL5 was highly expressed in urothelial carcinoma of the bladder compared with normal bladder tissue and that overexpression of RECQL5 could promote the proliferation of normal bladder cells. In bladder cancer, increased expression of RECQL5 was associated with poor patient prognosis.66 RECQL5 maintains genomic stability by participating in many DNA metabolic processes including DNA repair.67 High levels of RECQL5 may promote the survival of cancer cells under DNA damage in established tumors. In a previous study in breast cancer, high mRNA-expression levels of RECQL5 were significantly correlated with poor BCSS in a whole cohort of patients with breast cancer and particularly with an ER-positive subgroup.17 Conversely, the findings in this present study showed that increased mRNA expression of RECQL5 was significantly associated with increased RFS, especially in lymph-node-negative patients, but with reduced OS in patients with HER2-positive breast cancer. The different findings between these two studies might be due to diverse patient populations, different follow-up times, and varied methods of data analysis.
Breast cancer is known to be a heterogeneous and complicated disease with many prognostic factors, and several potential prognostic clinicopathological features and molecular markers have been investigated.68 Currently, for patients who are diagnosed with breast cancer, the most important prognostic factors are axillary-node status, tumor size, and histological tumor grade.69 Because intrinsic breast cancer subtypes are also associated with clinical outcome, these were included in this study, as well as including the ER and PR status of the tumors. Low expression of RECQL mRNA has previously been shown to be associated with poor BCSS in patients with ER-positive breast cancer,32 and high BLM mRNA expression was previously reported to be correlated with poor BCSS in patients with ER-positive breast cancer and improved BCSS in patients with ER-negative breast cancer,38 while high RECQL5 mRNA expression has previously been reported to be associated with poor BCSS in patients with ER-positive breast cancer.17 The mechanisms by which the five RecQ helicase members in human cells (RECQL, BLM, WRN, RECQL4, and RECQL5) are activated or inhibited by different hormones, genes, or environmental or epigenetic factors in breast cancer remain to be further investigated.
Conclusion
The findings of this study showed that mRNA-expression levels of the five RecQ DNA helicase family members – RECQL, BLM, WRN, RECQL4, and RECQL5 – were significantly correlated with prognosis in breast cancer patients. These preliminary findings require further study to determine whether diagnostic, prognostic, or therapeutic biomarker targets might be developed for clinical application in breast cancer.
Ethics approval
The Ethics Committee of the Second Affiliated Hospital, Zhejiang University School of Medicine approved this study.
Data availability
Data on associations of RecQ family-member mRNA expression with prognostic value can be downloaded from the online Kaplan–Meier Plotter breast cancer database (http://kmplot.com/analysis/index.php), and data sets used and analyzed during the current study are available from the corresponding author on reasonable request.
Author contributions
XZ, HC, and YY participated in the design and data acquisition of the study. XZ and HC wrote the manuscript. CX, JZ, and Jiaojiao Z participated in data analysis and interpretation of the study. YC modified the study and led scientific discussion. All authors contributed to data analysis, drafting the article, gave final approval of the version to be published, and agreed to be accountable for all aspects of the work.
Abbreviations
IHC, immunohistochemistry; RFS, relapse-free survival; PPS, postprogression survival; OS, overall survival; DMFS, distant metastasis-free survival; RQC, RecQ carboxy-terminal; HRDC, helicase and RNase DC terminal; dsDNA, double-stranded DNA; GEO, Gene Expression Omnibus; TMAs, tissue microarrays; IRS, immunoreactivity scoring system; BCSS, breast cancer-specific survival
Acknowledgments
We acknowledge Zhigang Chen of the Department of Oncology, Second Affiliated Hospital, Zhejiang University School of Medicine for writing guidance. This study was supported by funding from the Key Program of the Natural Science Foundation of Zhejiang Province (grant LZ16H160002), the Zhejiang Provincial Program for the Cultivation of High-Level Innovative Health Talents, and the National Science Foundation for Young Scientists of China (grant 81502270).
Disclosure
The authors report no conflicts of interest in this work.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108. | ||
Althuis MD, Dozier JM, Anderson WF, Devesa SS, Brinton LA. Global trends in breast cancer incidence and mortality 1973-1997. Int J Epidemiol. 2005;34(2):405–412. | ||
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127(12):2893–2917. | ||
Martinson HA, Lyons TR, Giles ED, Borges VF, Schedin P. Developmental windows of breast cancer risk provide opportunities for targeted chemoprevention. Exp Cell Res. 2013;319(11):1671–1678. | ||
Larsen NB, Hickson ID. RecQ helicases: conserved guardians of genomic integrity. Adv Exp Med Biol. 2013;767:161–184. | ||
Croteau DL, Popuri V, Opresko PL, Bohr VA. Human RecQ helicases in DNA repair, recombination, and replication. Annu Rev Biochem. 2014;83:519–552. | ||
Rezazadeh S. RecQ helicases; at the crossroad of genome replication, repair, and recombination. Mol Biol Rep. 2012;39(4):4527–4543. | ||
Chu WK, Hickson ID. RecQ helicases: multifunctional genome caretakers. Nat Rev Cancer. 2009;9(9):644–654. | ||
Viziteu E, Klein B, Basbous J, et al. RECQ1 helicase is involved in replication stress survival and drug resistance in multiple myeloma. Leukemia. 2017;31(10):2104–2113. | ||
Li XL, Lu X, Parvathaneni S, et al. Identification of RECQ1-regulated transcriptome uncovers a role of RECQ1 in regulation of cancer cell migration and invasion. Cell Cycle. 2014;13(15):2431–2445. | ||
de Voer RM, Hahn MM, Mensenkamp AR, et al. Deleterious Germline BLM mutations and the risk for early-onset colorectal cancer. Sci Rep. 2015;5:14060. | ||
Thompson ER, Doyle MA, Ryland GL, et al. Exome sequencing identifies rare deleterious mutations in DNA repair genes FANCC and BLM as potential breast cancer susceptibility alleles. PLoS Genet. 2012;8(9):e1002894. | ||
Davari P, Hebert JL, Albertson DG, et al. Loss of Blm enhances basal cell carcinoma and rhabdomyosarcoma tumorigenesis in Ptch1+/- mice. Carcinogenesis. 2010;31(6):968–973. | ||
Birkbak NJ, Li Y, Pathania S, et al. Overexpression of BLM promotes DNA damage and increased sensitivity to platinum salts in triple-negative breast and serous ovarian cancers. Ann Oncol. 2018;29(4):903–909. | ||
Agrelo R, Cheng WH, Setien F, et al. Epigenetic inactivation of the premature aging Werner syndrome gene in human cancer. Proc Natl Acad Sci USA. 2006;103(23):8822–8827. | ||
Fang H, Nie L, Chi Z, et al. RecQL4 helicase amplification is involved in human breast tumorigenesis. PLoS One. 2013;8(7):e69600. | ||
Arora A, Abdel-Fatah TM, Agarwal D, et al. Clinicopathological and prognostic significance of RECQL5 helicase expression in breast cancers. Carcinogenesis. 2016;37(1):63–71. | ||
Györffy B, Lanczky A, Eklund AC, et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010;123(3):725–731. | ||
Győrffy B, Lánczky A, Szállási Z. Implementing an online tool for genome-wide validation of survival-associated biomarkers in ovarian-cancer using microarray data from 1287 patients. Endocr Relat Cancer. 2012;19(2):197–208. | ||
Győrffy B, Surowiak P, Budczies J, Lánczky A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS One. 2013;8(12):e82241. | ||
Wu X, Liu W, Tang D, et al. Prognostic values of four Notch receptor mRNA expression in gastric cancer. Sci Rep. 2016;6:28044. | ||
Bohr VA. Rising from the RecQ-age: the role of human RecQ helicases in genome maintenance. Trends Biochem Sci. 2008;33(12):609–620. | ||
Sharma S, Doherty KM, Brosh RM Jr. Mechanisms of RecQ helicases in pathways of DNA metabolism and maintenance of genomic stability. Biochem J. 2006;398(3):319–337. | ||
Sharma S, Brosh RM Jr. Unique and important consequences of RECQ1 deficiency in mammalian cells. Cell Cycle. 2008;7(8):989–1000. | ||
Puranam KL, Blackshear PJ. Cloning and characterization of RECQL, a potential human homologue of the Escherichia coli DNA helicase RecQ. J Biol Chem. 1994;269(47):29838–29845. | ||
Mendoza-Maldonado R, Faoro V, Bajpai S, et al. The human RECQ1 helicase is highly expressed in glioblastoma and plays an important role in tumor cell proliferation. Mol Cancer. 2011;10:83. | ||
Arai A, Chano T, Futami K, et al. RECQL1 and WRN proteins are potential therapeutic targets in head and neck squamous cell carcinoma. Cancer Res. 2011;71(13):4598–4607. | ||
Matsushita Y, Yokoyama Y, Yoshida H, et al. The level of RECQL1 expression is a prognostic factor for epithelial ovarian cancer. J Ovarian Res. 2014;7:107. | ||
Futami K, Ogasawara S, Goto H, Yano H, Furuichi Y. RecQL1 DNA repair helicase: A potential tumor marker and therapeutic target against hepatocellular carcinoma. Int J Mol Med. 2010;25(4):537–545. | ||
Tao J, Tao S, Han J, et al. RECQL1 plays an important role in the development of tongue squamous cell carcinoma. Cell Physiol Biochem. 2014;33(5):1579–1590. | ||
Li L, Gao M, Song B, Zhang H, Wang Y. Effects of RECQ1 helicase silencing on non-small cell lung cancer cells. Biomed Pharmacother. 2016;83:1227–1232. | ||
Arora A, Parvathaneni S, Aleskandarany MA, et al. Clinicopathological and functional significance of RECQL1 helicase in sporadic breast cancers. Mol Cancer Ther. 2017;16(1):239–250. | ||
Barefield C, Karlseder J. The BLM helicase contributes to telomere maintenance through processing of late-replicating intermediate structures. Nucleic Acids Res. 2012;40(15):7358–7367. | ||
Bischof O, Kim SH, Irving J, Beresten S, Ellis NA, Campisi J. Regulation and localization of the Bloom syndrome protein in response to DNA damage. J Cell Biol. 2001;153(2):367–380. | ||
Kitao S, Ohsugi I, Ichikawa K, Goto M, Furuichi Y, Shimamoto A. Cloning of two new human helicase genes of the RecQ family: biological significance of multiple species in higher eukaryotes. Genomics. 1998;54(3):443–452. | ||
German J. Bloom’s syndrome. I. Genetical and clinical observations in the first twenty-seven patients. Am J Hum Genet. 1969;21(2):196–227. | ||
Qian X, Feng S, Xie D, Feng D, Jiang Y, Zhang X. RecQ helicase BLM regulates prostate cancer cell proliferation and apoptosis. Oncol Lett. 2017;14(4):4206–4212. | ||
Arora A, Abdel-Fatah TM, Agarwal D, et al. Transcriptomic and Protein Expression Analysis Reveals Clinicopathological Significance of Bloom Syndrome Helicase (BLM) in Breast Cancer. Mol Cancer Ther. 2015;14(4):1057–1065. | ||
Kawabe T, Tsuyama N, Kitao S, et al. Differential regulation of human RecQ family helicases in cell transformation and cell cycle. Oncogene. 2000;19(41):4764–4772. | ||
Singh DK, Karmakar P, Aamann M, et al. The involvement of human RECQL4 in DNA double-strand break repair. Aging Cell. 2010;9(3):358–371. | ||
Lan L, Nakajima S, Komatsu K, et al. Accumulation of Werner protein at DNA double-strand breaks in human cells. J Cell Sci. 2005;118(Pt 18):4153–4162. | ||
Baynton K, Otterlei M, Bjørås M, von Kobbe C, Bohr VA, Seeberg E. WRN interacts physically and functionally with the recombination mediator protein RAD52. J Biol Chem. 2003;278(38):36476–36486. | ||
Constantinou A, Tarsounas M, Karow JK, et al. Werner’s syndrome protein (WRN) migrates Holliday junctions and co-localizes with RPA upon replication arrest. EMBO Rep. 2000;1(1):80–84. | ||
Yu CE, Oshima J, Fu YH, et al. Positional cloning of the Werner’s syndrome gene. Science. 1996;272(5259):258–262. | ||
Lauper JM, Krause A, Vaughan TL, Monnat RJ Jr. Spectrum and risk of neoplasia in Werner syndrome: a systematic review. PLoS One. 2013;8(4):e59709. | ||
Brosh RM Jr. DNA helicases involved in DNA repair and their roles in cancer. Nat Rev Cancer. 2013;13(8):542–558. | ||
Brosh RM Jr, Bohr VA. Human premature aging, DNA repair and RecQ helicases. Nucleic Acids Res. 2007;35(22):7527–7544. | ||
Oshima J, Sidorova JM, Monnat RJ Jr. Werner syndrome: Clinical features, pathogenesis and potential therapeutic interventions. Ageing Res Rev. 2017;33:105–114. | ||
Tang W, Robles AI, Beyer RP, et al. The Werner syndrome RECQ helicase targets G4 DNA in human cells to modulate transcription. Hum Mol Genet. 2016;25(10):2060–2069. | ||
Rossi ML, Ghosh AK, Bohr VA. Roles of Werner syndrome protein in protection of genome integrity. DNA Repair (Amst). 2010;9(3):331–344. | ||
Opresko PL, Calvo JP, von Kobbe C. Role for the Werner syndrome protein in the promotion of tumor cell growth. Mech Ageing Dev. 2007;128(7–8):423–436. | ||
Yin J, Kwon YT, Varshavsky A, Wang W. RECQL4, mutated in the Rothmund-Thomson and RAPADILINO syndromes, interacts with ubiquitin ligases UBR1 and UBR2 of the N-end rule pathway. Hum Mol Genet. 2004;13(20):2421–2430. | ||
Ghosh AK, Rossi ML, Singh DK, et al. RECQL4, the protein mutated in Rothmund-Thomson syndrome, functions in telomere maintenance. J Biol Chem. 2012;287(1):196–209. | ||
Croteau DL, Rossi ML, Canugovi C, et al. RECQL4 localizes to mitochondria and preserves mitochondrial DNA integrity. Aging Cell. 2012;11(3):456–466. | ||
Lu L, Jin W, Liu H, Wang LL. RECQ DNA helicases and osteosarcoma. Adv Exp Med Biol. 2014;804:129–145. | ||
Su Y, Meador JA, Calaf GM, et al. Human RecQL4 helicase plays critical roles in prostate carcinogenesis. Cancer Res. 2010;70(22):9207–9217. | ||
Saglam O, Shah V, Worsham MJ. Molecular differentiation of early and late stage laryngeal squamous cell carcinoma: an exploratory analysis. Diagn Mol Pathol. 2007;16(4):218–221. | ||
Choi YW, Bae SM, Kim YW, et al. Gene expression profiles in squamous cell cervical carcinoma using array-based comparative genomic hybridization analysis. Int J Gynecol Cancer. 2007;17(3):687–696. | ||
Yong ZW, Zaini ZM, Kallarakkal TG, et al. Genetic alterations of chromosome 8 genes in oral cancer. Sci Rep. 2015;4:6073. | ||
Arora A, Agarwal D, Abdel-Fatah TM, et al. RECQL4 helicase has oncogenic potential in sporadic breast cancers. J Pathol. 2016;238(4):495–501. | ||
Qi Y, Zhou X. Haplotype analysis of RECQL5 gene and laryngeal cancer. Tumour Biol. 2014;35(3):2669–2673. | ||
Zhi LQ, Ma W, Zhang H, Zeng SX, Chen B. Association of RECQL5 gene polymorphisms and osteosarcoma in a Chinese Han population. Tumour Biol. 2014;35(4):3255–3259. | ||
He YJ, Qiao ZY, Gao B, Zhang XH, Wen YY. Association between RECQL5 genetic polymorphisms and susceptibility to breast cancer. Tumour Biol. 2014;35(12):12201–12204. | ||
Lao VV, Welcsh P, Luo Y, et al. Altered RECQ Helicase Expression in Sporadic Primary Colorectal Cancers. Transl Oncol. 2013;6(4):458–469. | ||
Wu J, Zhi L, Dai X, Cai Q, Ma W. Decreased RECQL5 correlated with disease progression of osteosarcoma. Biochem Biophys Res Commun. 2015;467(4):617–622. | ||
Patterson K, Arya L, Bottomley S, et al. Altered RECQL5 expression in urothelial bladder carcinoma increases cellular proliferation and makes RECQL5 helicase activity a novel target for chemotherapy. Oncotarget. 2016;7(46):76140–76150. | ||
Tadokoro T, Ramamoorthy M, Popuri V, et al. Human RECQL5 participates in the removal of endogenous DNA damage. Mol Biol Cell. 2012;23(21):4273–4285. | ||
Fisher ER, Costantino J, Fisher B, Redmond C. Pathologic findings from the National Surgical Adjuvant Breast Project (Protocol 4). Discriminants for 15-year survival. National Surgical Adjuvant Breast and Bowel Project Investigators. Cancer. 1993;71(6 Suppl):2141–2150. | ||
Chen XS, Ma CD, Wu JY, et al. Molecular subtype approximated by quantitative estrogen receptor, progesterone receptor and Her2 can predict the prognosis of breast cancer. Tumori. 2010;96(1):103–110. |
Supplementary materials
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.