Back to Journals » OncoTargets and Therapy » Volume 10
Deletion and downregulation of MTAP contribute to the motility of esophageal squamous carcinoma cells
Authors Cheng X, Liu Z, Shang L, Cai H, Zhang Y , Cai Y, Xu X, Hao J, Wang M
Received 18 September 2017
Accepted for publication 6 November 2017
Published 11 December 2017 Volume 2017:10 Pages 5855—5862
DOI https://doi.org/10.2147/OTT.S151953
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Geoffrey Pietersz
Xiao-Yu Cheng,1,2 Zou Liu,1,2 Li Shang,1 Hong-Qing Cai,1 Yu Zhang,1,2 Yan Cai,1,2 Xin Xu,1,2 Jia-Jie Hao,1,2 Ming-Rong Wang1,2
1State Key Laboratory of Molecular Oncology, National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, 2Center for Cancer Precision Medicine, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China
Abstract: Esophageal squamous cell carcinoma (ESCC) is among the most common malignancies, with a low 5-year overall survival rate. In previous studies, we and others have found that 9p21.3 was the most frequently deleted region in ESCC. The MTAP gene, which is located close to CDKN2A/B in 9p21.3, encodes methylthioadenosine phosphorylase. This enzyme plays an important role during the process of adenosine transfer. In the present study, we found that MTAP is deleted at the genomic level in 19.1% (64/341) of primary ESCC tumors, and decreased mRNA and protein expression were present in 31.1% (28/90) and 33.3% (6/18) of ESCCs, respectively. Further statistical analysis showed a positive correlation between deletion and decreased mRNA expression of MTAP in the ESCC tissues tested (coefficient: 0.826; P=1.17×10-23). Knockdown of MTAP expression using small interfering RNA-mediated silencing promoted the invasion and migration of ESCC cells. Also, overexpression of MATP using pcDNA3.1-MTAP plasmid decreased the cell invasion and migration. At the molecular level, MTAP knockdown downregulated E-cadherin and p-GSK3β but upregulated Slug expression. Our results indicated that MTAP deletion results in the decreased expression in ESCCs and that it plays a role in promoting the mobility and inducing the epithelial-to-mesenchymal transition of ESCC cells via the GSK3β/Slug/E-cadherin axis. The data suggest that MTAP might function as a tumor suppressor gene in ESCC.
Keywords: ESCC, MTAP, deletion, invasion, migration
Introduction
Esophageal cancer is among the most common malignancies. Eastern Asia, and Eastern and Southern Africa are the highest-risk areas, and esophageal squamous cell carcinoma (ESCC) is the most prevalent type.1,2 The 5-year overall survival rate for ESCC patients is only 15%–25%.3 Significant progress on the study of gene deletions in human cancers has been made in recent years.4–7 Identification and investigation of genetic deletions might not only help reveal the mechanisms that underlie the tumorigenesis and development of ESCC but also provide potential biomarkers for the detection and therapy of the disease.
In previous studies, we and others have used large-scale genomic techniques to show that 9p21.3 was the most frequently deleted region in ESCC. These techniques included array-based comparative genomic hybridization (array-CGH), single-nucleotide polymorphism (SNP) arrays, and whole-exome sequencing.8–11 Moreover, our multiregional intratumor heterogeneity study showed that 9p21.3 homozygous deletion was an early event10 that was also found especially in the precancerous lesions of the esophageal squamous epithelia.12
CDKN2A and CDKN2B have been reported in various studies to be the most commonly deleted genes within 9p21.3.11 The MTAP gene, which is located close to CDKN2A/B in 9p21.3, encodes methylthioadenosine phosphorylase. This enzyme plays an important role during the process of adenosine transfer. MTAP is frequently deleted in human cancers.13–15 Kim et al found homozygous deletions in MTAP in xenografts that were established from the thoracic duct lymph of ESCC patients.16
In the present study, we analyzed the deletion and expression of MTAP in the primary ESCC tumors and cell lines and assessed the relationship between the deletion and expression of MTAP. Furthermore, we investigated the impact of decreased MTAP expression on the malignant phenotypes of esophageal squamous carcinoma cells.
Materials and methods
Cell culture and tissue specimens
The human ESCC cell lines KYSE30, KYSE150, KYSE180, KYSE410, KYSE450, and KYSE510 were generously provided by Dr Y Shimada (Kyoto University, Kyoto, Japan). The ESCC cell line EC109 was purchased from the cell bank of Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences. The cells were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS; Invitrogen, San Diego, CA, USA), penicillin (100 U/mL), and streptomycin (100 mg/mL).
ESCC tissues and adjacent morphologically normal operative margins were procured from surgical resection specimens collected by the Department of Pathology in the Cancer Hospital, Chinese Academy of Medical Sciences, Beijing, China. All samples used in this study were residual specimens collected after sampling for pathological diagnosis. None of the patients received treatment prior to surgery, and all patients signed the informed consent forms of the Cancer Hospital, CAMS/PUMC for sample collection and molecular analysis. This study was approved by the Ethics Committee/Institutional Review Board of the Cancer Institute/Hospital, PUMC/CAMS (No NCC2015G-06).
Copy number alteration analysis
The MTAP copy number alterations were analyzed using our in-house array-CGH data (GSE4645212), and the data derived from other array-CGH and SNP array platforms are available in Gene Expression Omnibus (GEO; GSE54993 and GSE54994,9 GSE47630,17 GSE1795818) and The Cancer Genome Atlas (TCGA, https://tcga-data.nci.nih.gov/docs/publications/esca_201619) databases. A mean log2 ratio of MTAP <−0.75 was classified as a deletion.
mRNA expression analysis
The MTAP mRNA expression of primary ESCC tumors and cell lines was analyzed using the reads per kilobase per million (RPKM) mapped reads value from the TCGA data (https://tcga-data.nci.nih.gov/docs/publications/esca_201619) and GEO data (GSE23964), respectively. The mean RPKM value of the esophageal epithelial cells is 5.198±0.408, which is available in NCBI website (https://www.ncbi.nlm.nih.gov/gene/4507). The ratio of the RPKM value in each tumor versus that in esophageal epithelial cells was transformed to the log2 ratio value. Mean log2 ratios of MTAP <−1.5 were classified as decreased expression.
Small interfering RNA (siRNA), plasmid construction, and transfection
Two duplex MTAP siRNAs, siRNA-1 (5′-TCACTACCATACCTCAGAT-3′) and siRNA-2 (5′-GGTCTTAAAGACCCTGAAA-3′), and a nonsilencing siRNA (5′-TTCTCCGAACGUGUCACGTTT-3′) were designed and chemically synthesized (GeneChem, Shanghai) for transient transfection.
The CDS region of MTAP was amplified by reverse-transcript PCR using the upstream primer 5′-AAAGGATCCATGGCCTCTGGCACC-3′, and downstream primer 5′-CCCGAATTCTTAATGTCTTGGTAATAAAACAGA-3′, and then cloned into pcDNA3.1.
The cells were transfected with siRNAs or plasmids using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. The cells were harvested 48 hours after transfection. The transfection efficiency was determined by Western blot analysis.
Invasion and migration assays
For the migration assay, 8×104 parental, nonsilencing siRNA- and MTAP siRNA-treated cells were seeded on fibronectin-coated polycarbonate membranes inserted in Transwell (Costar, Cambridge, MA, USA). RPMI-1640 that contained 20% FBS was added to the lower chamber. After incubation for 18 hours at 37°C in a CO2 incubator, the insert was washed with the phosphate-buffered saline, and the cells on the top surface of the insert were removed by wiping with a cotton swab. For the invasion assay, the procedure was similar to the migration assay, except that the transwell membrane was coated with 300 ng/μL matrigel (BD Biosciences, San Jose, CA, USA). The cells that had migrated to the bottom surface of the insert were fixed with methanol, stained with 0.5% crystal violet, and subjected to microscopic inspection.
Western blot analysis
Immunoblotting was conducted with primary antibodies against MTAP (Abcam, ab126770; 1:1,000), E-cadherin (Cell Signaling Technology, #3195; 1:1,000), p-GSK3β (Cell Signaling Technology, #9323; 1:1,000), or GSK3β (Cell Signaling Technology, #12456; 1:1,000). β-actin (Sigma, A19781; 1:5,000) was used as a loading control. The signals were visualized using the super-enhanced chemiluminescence detection reagent (Applygen Technologies, Inc., Beijing, China).
Statistical analysis
Statistical analysis was performed with the SPSS software program (version 17.0). Mann–Whitney test or Kruskal–Wallis test was performed for the evaluation of the association between MTAP deletion and clinicopathological parameters. The correlation between MTAP deletion and mRNA downregulation was analyzed using Spearman’s relative analysis. P-values <0.05 were considered to be statistically significant.
Results
MTAP deletions in ESCC and the association with clinicopathological parameters
We performed in-house array-CGH on 59 primary ESCC tumors12 and further analyzed additional two array-CGH data and two SNP array data measuring ESCCs,9,17–19 for a total of 341 ESCC cases. Overall, MTAP was deleted in 19.1% of the primary ESCC tumors (Table 1; Figure 1).
  | Table 1 MTAP deletions in ESCC |
We then analyzed the relationship between MTAP deletion and the clinicopathological parameters (Table 2). MTAP deletion was significantly correlated with age (P=0.001) but not with gender, pathological T staging (pT), lymph node metastasis (LNM), and grade.
Decreased expression of MTAP mRNA and protein in ESCC
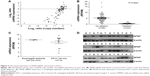
To determine whether the genomic deletion of MTAP results in the downregulation of its expression, we analyzed the relationship between the copy number and mRNA expression of MTAP in 90 ESCC cases using the online data19 in which both copy number alterations and mRNA expression were detected in each case. The mRNA expression was decreased in 31.1% (28/90) of ESCCs. Overall, the MTAP mRNA expression levels were associated with the copy number levels. Reduced MTAP mRNA expression was present in 90.9% (20/22) of the cases with deletions compared with 5.9% (4/68) of the cases without deletions (Figure 2A). Furthermore, the MTAP mRNA levels in MTAP-deleted cases were much lower than those in the cases without the deletions (Figure 2B; mean level: 0.63±0.11 vs 5.34±0.41, P<0.0001). Further statistical analysis showed a positive correlation between deletion and decreased mRNA expression of MTAP in the ESCC tissues tested (coefficient: 0.826; P=1.17×10−23; Table 3). We also analyzed the mRNA levels in 16 cell lines using the GEO data GSE23964 and confirmed that the mean mRNA levels in the ESCC cell lines (6.58±0.69) were lower than those in the esophageal epithelial cell lines (7.64±0.10). Moreover, the mRNA levels in four ESCC cell lines were markedly decreased between 1- and 3.8-fold compared with the levels in the epithelial cell lines (Figure 2C).
  | Table 3 Correlation of deletion and mRNA expression of MTAP in ESCC |
We further examined the expression of MTAP protein in 18 ESCC cases, and the Western blot results showed that the MTAP protein was downregulated in 33.3% (6/18) of the primary ESCC tumors compared with those in operative margin tissues (Figure 2D).
MTAP knockdown increased invasion and migration of ESCC cells
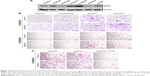
We next investigated whether the decreases in expression affected the malignant phenotype of the ESCC cells. The MTAP protein expression of ESCC cell lines was detected by Western blotting. The levels of MTAP protein are relatively higher in KYSE30, KYSE150, KYSE410, and KYSE450 than those in KYSE180, KYSE510, and EC109 (Figure 3A). We then knocked down MTAP in KYSE150 and KYSE450 using siRNAs, and overexpressed MTAP in KYSE510 using the constructed plasmids pcDNA3.1-MTAP. The transwell assays showed that knockdown of MTAP enhanced the invasion and migration of KYSE150 and KYSE450 cells (Figure 3B), and that overexpression of MTAP decreased the invasion and migration in KYSE510 cells (Figure 3C).
MTAP knockdown regulated the expression of epithelial-to-mesenchymal transition (EMT)-related molecules in ESCC cells
Based on the morphological changes affected by MTAP knockdown, we measured the expression of E-cadherin and Slug, as well as the phosphorylation of GSK3β, which are associated with cell motility and EMT. After MTAP knockdown, both E-cadherin and p-GSK3β were downregulated, whereas Slug was upregulated in the KYSE150 cells (Figure 4).
Discussion
Genomic deletion is one of the major processes that causes tumorigenesis and the development of human cancers. Studies have shown that the most frequent homozygous deletion region in ESCC is 9p21.3,11 which is in an early event during the clonal evolutionary process of ESCC.10 The common deletion peaks are at CDKN2A/B genes,11 inactivation of which are associated with tumorigenesis and cancer development.20–23 The MTAP gene, which is located ~30 kb distal to CDKN2A, is usually co-deleted with CDKN2A in several human cancers.24–27 By analyzing the copy number alterations using high-throughput array-based genomic data from several studies, we found that MTAP deletion occurred in 19.1% of ESCCs. No significant correlation was observed between MTAP deletion and gender, pT, LNM, and grade, except for age. However, a slight higher frequency of MTAP deletion was present in LNM-positive patients than that in LNM-negative ones.
The correlation between MTAP deletion and loss of expression has been found in multiple types of cancers, including gastrointestinal stromal tumors,28 laryngeal squamous cell carcinoma,29 and glioblastoma multiforme.30 In the present study, we established a significantly positive correlation between copy number and mRNA level of MTAP in ESCC. We also found reduced MTAP protein expression in ESCC, which is similar to the observations in lung cancer, liver cancer, lymphoma, and so on.27,31,32
It has been observed that deletion and loss of MTAP expression are associated with poor outcomes for several human cancers,27,33,34 and MTAP inactivation contributes to cell proliferation and invasion of cancer cells.13,14,33–35 However, the role of MTAP in ESCC is currently unknown. In this study, our data indicate that the loss of MTAP expression enhanced the invasion and migration of ESCC cells, and overexpression of MTAP decreased the cell invasion and migration, which suggested that MTAP expression might play a role in the inhibition of cell motility.
EMT is involved in the metastatic process of malignant tumors, and EMT activation promotes the invasion and metastasis of cancer cells.36,37 Our data showed that MTAP knockdown in ESCC cells led to a downregulation of E-cadherin expression, which has been well established as a hallmark of the EMT process in human cancers.38,39 We further found an upregulation of the oncogenic transcriptional repressor Slug in MTAP-knockdown ESCC cells, which indicated that upregulated Slug represses E-cadherin expression through the Slug/E-cadherin axis similar to the process in non-small-cell lung cancers.40,41 It has been reported that Slug expression is stabilized by the inactivation of GSK3β in epithelial cancers.41 In this study, we also detected the decreased phosphorylation of GSK3β at Ser9 after knockdown of MTAP. Collectively, our findings suggest that MTAP expression inhibits cell motility and EMT through GSK3β/Slug/E-cadherin axis. Further investigation should be performed to determine whether deletion of MTAP plays a role in the tumorigenesis or progressions of ESCC.
In summary, our data show that frequent deletion and decreased expression of MTAP occur in primary ESCC tumors and that decreased expression of MTAP enhances the motility and EMT of ESCC cells through the GSK3β/Slug/E-cadherin axis. Together, our current findings suggest that MTAP might act as a tumor suppressor gene in ESCC.
Acknowledgments
This study was supported by the National Natural Science Foundation of China (81520108023), the CAMS Innovation Fund for Medical Sciences (CIFMS) (2016-I2M-001 and 2016-I2M-3-007), Fundamental Research Funds for the Central Public-interest Scientific Institution (2016ZX310178), and the Beijing Nova Program (Z171100001117017).
Disclosure
The authors report no conflicts of interest in this work.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108. | ||
Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136(5):E359–E386. | ||
Pennathur A, Gibson MK, Jobe BA, Luketich JD. Oesophageal carcinoma. Lancet. 2013;381(9864):400–412. | ||
Matsuyama H, Ikemoto K, Eguchi S, et al. Copy number aberrations using multicolour fluorescence in situ hybridization (FISH) for prognostication in non-muscle-invasive bladder cancer (NIMBC). BJU Int. 2014;113(4):662–667. | ||
Hida T, Hamasaki M, Matsumoto S, et al. Immunohistochemical detection of MTAP and BAP1 protein loss for mesothelioma diagnosis: comparison with 9p21 FISH and BAP1 immunohistochemistry. Lung Cancer. 2017;104:98–105. | ||
Perea J, Garcia JL, Perez J, et al. NOMO-1 gene is deleted in early-onset colorectal cancer. Oncotarget. 2017;8(15):24429–24436. | ||
Campos CZ, Losi Guembarovski R, de Oliveira CEC, et al. Glutathione S-transferases deletions may act as prognosis and therapeutic markers in breast cancer. Clin Exp Med. Epub 2017 Apr 28. | ||
Lin DC, Hao JJ, Nagata Y, et al. Genomic and molecular characterization of esophageal squamous cell carcinoma. Nat Genet. 2014;46(5):467–473. | ||
Song Y, Li L, Ou Y, et al. Identification of genomic alterations in oesophageal squamous cell cancer. Nature. 2014;509(7498):91–95. | ||
Hao JJ, Lin DC, Dinh HQ, et al. Spatial intratumoral heterogeneity and temporal clonal evolution in esophageal squamous cell carcinoma. Nat Genet. 2016;48(12):1500–1507. | ||
Lin DC, Wang MR, Koeffler HP. Genomic and epigenomic aberrations in esophageal squamous cell carcinoma and implications for patients. Gastroenterology. Epub 2017 Jul 27. | ||
Shi ZZ, Shang L, Jiang YY, et al. Consistent and differential genetic aberrations between esophageal dysplasia and squamous cell carcinoma detected by array comparative genomic hybridization. Clin Cancer Res. 2013;19(21):5867–5878. | ||
Kryukov GV, Wilson FH, Ruth JR, et al. MTAP deletion confers enhanced dependency on the PRMT5 arginine methyltransferase in cancer cells. Science. 2016;351(6278):1214–1218. | ||
Mavrakis KJ, McDonald ER 3rd, Schlabach MR, et al. Disordered methionine metabolism in MTAP/CDKN2A-deleted cancers leads to dependence on PRMT5. Science. 2016;351(6278):1208–1213. | ||
Zhao M, Zhao Z. Concordance of copy number loss and down-regulation of tumor suppressor genes: a pan-cancer study. BMC Genomics. 2016;17 (Suppl 7):532. | ||
Kim DH, Muto M, Kuwahara Y, et al. Array-based comparative genomic hybridization of circulating esophageal tumor cells. Oncol Rep. 2006;16(5):1053–1059. | ||
Sawada G, Niida A, Uchi R, et al. Genomic landscape of esophageal squamous cell carcinoma in a Japanese population. Gastroenterology. 2016;150(5):1171–1182. | ||
Bass AJ, Watanabe H, Mermel CH, et al. SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet. 2009;41(11):1238–1242. | ||
Cancer Genome Atlas Research Network; Analysis Working Group: Asan University; BC Cancer Agency, et al. Integrated genomic characterization of oesophageal carcinoma. Nature. 2017;541(7636):169–175. | ||
Ortiz B, White JR, Wu WH, Chan TA. Deletion of Ptprd and Cdkn2a cooperate to accelerate tumorigenesis. Oncotarget. 2014;5(16):6976–6982. | ||
McNeal AS, Liu K, Nakhate V, et al. CDKN2B loss promotes progression from benign melanocytic nevus to melanoma. Cancer Discov. 2015;5(10):1072–1085. | ||
Yang H, Kircher DA, Kim KH, et al. Activated MEK cooperates with Cdkn2a and Pten loss to promote the development and maintenance of melanoma. Oncogene. 2017;36(27):3842–3851. | ||
Tu Q, Hao J, Zhou X, et al. CDKN2B deletion is essential for pancreatic cancer development instead of unmeaningful co-deletion due to juxtaposition to CDKN2A. Oncogene. Epub 2017 Sep 11. | ||
Schmid M, Malicki D, Nobori T, et al. Homozygous deletions of methylthioadenosine phosphorylase (MTAP) are more frequent than p16INK4A (CDKN2) homozygous deletions in primary non-small cell lung cancers (NSCLC). Oncogene. 1998;17(20):2669–2675. | ||
Hustinx SR, Hruban RH, Leoni LM, et al. Homozygous deletion of the MTAP gene in invasive adenocarcinoma of the pancreas and in periampullary cancer: a potential new target for therapy. Cancer Biol Ther. 2005;4(1):83–86. | ||
Krasinskas AM, Bartlett DL, Cieply K, Dacic S. CDKN2A and MTAP deletions in peritoneal mesotheliomas are correlated with loss of p16 protein expression and poor survival. Mod Pathol. 2010;23(4):531–538. | ||
Su CY, Chang YC, Chan YC, et al. MTAP is an independent prognosis marker and the concordant loss of MTAP and p16 expression predicts short survival in non-small cell lung cancer patients. Eur J Surg Oncol. 2014;40(9):1143–1150. | ||
Huang HY, Li SH, Yu SC, et al. Homozygous deletion of MTAP gene as a poor prognosticator in gastrointestinal stromal tumors. Clin Cancer Res. 2009;15(22):6963–6972. | ||
Conde L, Vilaseca I, Alos L, et al. Methylthioadenosine phosphorylase inactivation depends on gene deletion in laryngeal squamous cell carcinoma. Histopathology. 2012;61(6):1082–1088. | ||
Crespo I, Tao H, Nieto AB, et al. Amplified and homozygously deleted genes in glioblastoma: impact on gene expression levels. PLoS One. 2012;7(9):e46088. | ||
Czech B, Dettmer K, Valletta D, et al. Expression and function of methylthioadenosine phosphorylase in chronic liver disease. PLoS One. 2013;8(12):e80703. | ||
Marce S, Balague O, Colomo L, et al. Lack of methylthioadenosine phosphorylase expression in mantle cell lymphoma is associated with shorter survival: implications for a potential targeted therapy. Clin Cancer Res. 2006;12(12):3754–3761. | ||
Kim J, Kim MA, Min SY, Jee CD, Lee HE, Kim WH. Downregulation of methylthioadenosin phosphorylase by homozygous deletion in gastric carcinoma. Genes Chromosomes Cancer. 2011;50(6):421–433. | ||
Li CF, Fang FM, Kung HJ, et al. Downregulated MTAP expression in myxofibrosarcoma: a characterization of inactivating mechanisms, tumor suppressive function, and therapeutic relevance. Oncotarget. 2014;5(22):11428–11441. | ||
Marjon K, Cameron MJ, Quang P, et al. MTAP deletions in cancer create vulnerability to targeting of the MAT2A/PRMT5/RIOK1 axis. Cell Rep. 2016;15(3):574–587. | ||
Jolly MK, Ware KE, Gilja S, Somarelli JA, Levine H. EMT and MET: necessary or permissive for metastasis? Mol Oncol. 2017;11(7):755–769. | ||
Santamaria PG, Moreno-Bueno G, Portillo F, Cano A. EMT: present and future in clinical oncology. Mol Oncol. 2017;11(7):718–738. | ||
Thiery JP. Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer. 2002;2(6):442–454. | ||
Bolos V, Peinado H, Perez-Moreno MA, Fraga MF, Esteller M, Cano A. The transcription factor Slug represses E-cadherin expression and induces epithelial to mesenchymal transitions: a comparison with Snail and E47 repressors. J Cell Sci. 2003;116(Pt 3):499–511. | ||
Shih JY, Tsai MF, Chang TH, et al. Transcription repressor slug promotes carcinoma invasion and predicts outcome of patients with lung adenocarcinoma. Clin Cancer Res. 2005;11(22):8070–8078. | ||
Kao SH, Wang WL, Chen CY, et al. GSK3beta controls epithelial-mesenchymal transition and tumor metastasis by CHIP-mediated degradation of Slug. Oncogene. 2014;33(24):3172–3182. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.