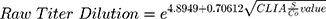Back to Journals » Journal of Blood Medicine » Volume 12
Correlation of Automated Chemiluminescent Method with Enzyme-Linked Immunosorbent Assay (ELISA) Antibody Titers in Convalescent COVID-19 Plasma Samples: Development of Rapid, Cost-Effective Semi-Quantitative Diagnostic Methods
Authors Mendoza R , Silver M, Zuretti AR, Christian M, Das B, Norin AJ, Borgen P, Libien J, Bluth MH
Received 22 December 2020
Accepted for publication 23 February 2021
Published 17 March 2021 Volume 2021:12 Pages 157—164
DOI https://doi.org/10.2147/JBM.S296730
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Michael Shurin
Rachelle Mendoza,1 Michael Silver,2,3 Alejandro R Zuretti,1,4 Manan Christian,4 Ballabh Das,5 Allen J Norin,5 Patrick Borgen,6 Jenny Libien,1,4 Martin H Bluth4
1Department of Pathology, SUNY Downstate Health Sciences University, Brooklyn, NY, USA; 2School of Public Health, SUNY Downstate Health Sciences University, Brooklyn, NY, USA; 3Department of Medicine, Maimonides Medical Center, Brooklyn, NY, USA; 4Department of Pathology, Maimonides Medical Center, Brooklyn, NY, USA; 5Department of Medicine, SUNY Downstate Health Sciences University, Brooklyn, NY, USA; 6Department of Surgery, Maimonides Medical Center, Brooklyn, NY, USA
Correspondence: Martin H Bluth
Blood Transfusion and Donor Services, Clinical Laboratories, Translational Research (Pathology), Maimonides Medical Center, 4802 10 th Avenue, Brooklyn, NY, 11219, USA
Email [email protected]
Background: We investigated the utility of an automated chemiluminescent SARS-CoV-2 IgG antibody assay platform in quantifying the amount of binding antibodies present in donated convalescent plasma.
Methods: A total of 179 convalescent plasma units were analyzed for the presence of SARS-CoV-2 IgG antibodies using the Beckman-Coulter chemiluminescent immunoassay (CLIA) platform. The equipment-derived numerical values (S/Co ratio) were recorded. Aliquots from the same units were subjected to enzyme-linked immunosorbent assay (ELISA) that detects IgG antibodies against the receptor-binding domain (RBD) of the SARS-CoV-2 S1 protein. The relationship between ELISA titers and CLIA S/Co values was analyzed using linear regression and receiver operating characteristics (ROC) curve.
Results: Twenty-one samples (11.7%) had S/Co values of less than 1.0 and were deemed negative for antibodies and convalescent plasma had S/Co values between > 1.0 and 5.0 (70/179, 39.1%). Fifteen units (8.4%) had negative ELISA titer. The majority of the units (95/179. 53.1%) had titers ≥ 1:1024. The sensitivities of ELISA to CLIA were comparable (90.5% vs 88.3%, respectively; p=0.18). There was positive linear correlation between CLIA S/Co values and ELISA IgG titer (Rho = 0.75; Spearman’s rank = 0.82, p-value = < 0.0001). The agreement between the two methods was fair, with a κ index of 0.2741. Using the ROC analysis, we identified a CLIA S/Co cutoff value of 8.2, which gives a sensitivity of 90% and a specificity of 82% in predicting a titer dilution of ≥ 1:1024.
Conclusion: The utility of automated antibody detection systems can be extended from simply a screening method to a semi-quantitative and quantitative functional antibody analysis. CLIA S/Co values can be used to reliably estimate the ELISA antibody titer. Incorporation of chemiluminescent-based methods can provide rapid, cost-effective means of identifying anti-SARS-CoV-2 antibody titers in donated plasma for use in the treatment of COVID-19 infection.
Keywords: COVID-19, ELISA, chemiluminescence, antibody, titers
Introduction
Coronavirus disease 2019 (COVID-19), caused by a novel coronavirus, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is the cause of a pandemic that has infected over 49.7 million people worldwide with a mortality of 2.4% to date.1 Due to the record-breaking economic and social impact of COVID-19, there is a need to optimize all resources to improve outcome and minimize the spread of the virus. A key goal is to increase the availability of treatment options for the severely ill COVID-19 patients, one of which is convalescent plasma.
Initial case reports from China have shown immediate beneficial clinical results in COVID-19 patients transfused with convalescent plasma.2–4 The viral loads were decreased after transfusion, coinciding with increased levels of neutralizing COVID-19 antibody. In a match-controlled study done in the United States,5 patients who received COVID-19 convalescent plasma (CCP) were more likely to have improved oxygen requirement by day 14 post-transfusion as compared to those who did not receive CCP. The recipients also had significantly longer survival, especially non-intubated patients.5 Another study involving approximately 5000 patients observed better survival among recipients who were given CCP units that contain high antibody levels based on chemiluminescent intensity value.6
Recent investigations on recovered COVID-19 patients revealed low antibody levels among those with mild symptoms, especially among the healthy and younger individuals.7,8 Healthy and younger individuals are most likely to be eligible for blood product donation. Another study observed rapid decline in IgG antibodies among recovered COVID-19 patients with mild symptoms from the time they were screened for donation up to plasmapheresis.9 Although the FDA has issued emergency use authorization (EUA) for the use of convalescent plasma, it does not strictly implement antibody testing prior to transfusion of the blood product.10 This has great implication on the quality of convalescent plasma units that are available in blood centers.
Neutralizing antibody assays, considered as the gold standard for detection of anti-COVID function, are assessed using native or pseudotype virus in cellular assay.11 The disadvantages of this analysis include the limited number of facilities that have the capacity and expertise to perform this assay, lengthy turn-around time and high cost.12,13 However, antibody levels can also be estimated using an antibody binding assay such as an enzyme-linked immunoassay (ELISA). A study on COVID-19 patients showed good correlation between ELISA antibody titer and neutralizing antibody levels.14 In particular, the linear correlation with neutralizing antibody levels was seen in ELISA assays that detected immunoglobulins (IgG) antibodies against the receptor-binding domain (RBD) of the viral spike protein S1.14 Although easier to perform than neutralization assays, ELISA assays are limited by long turn-around time and higher costs.
Chemiluminescent immunoassay (CLIA) is considered as a qualitative antibody assay that detects binding antibodies to viral antigens, similar to the principle of an ELISA. The test relies on mixing patient samples with a known viral protein, buffer reagents, and specific enzyme-labeled antibodies that allow a light-based, luminescent read-out.15–17 The amount of light (radiance) emitted from each sample is relative to the number of antibodies present in a patient sample.17 This type of assay can assess the presence of multiple types of antibodies, including IgG, IgM, and IgA, is automated, has a short turn-around time (~45 minutes) and can be implemented with minimal cost in already existing clinical laboratory platforms.18 However, this method has not been optimized to quantify the amount of antibodies. Currently, FDA has not authorized the use of automated chemiluminescent assays for screening of antibodies in potential donors of COVID-19 convalescent plasma.
At Maimonides Medical Center, antibody testing is performed using a Beckman Coulter platform that utilizes CLIA that detects IgG antibodies against the SARS-CoV-2RBD. The intensity value retrieved from the platform, based on the sample-to-cut-off (S/Co) ratio, reflects the amount of anti-SARS-CoV-2 antibodies present in the sample. Since this platform utilizes the RBD of the virus, there is a good chance that the results of this assay may correlate with the number of neutralizing antibodies. However, to date, no study has been published correlating the S/Co value with an ELISA or neutralization titer in the setting of COVID-19. Such a correlation would be crucial to provide a more cost-efficient and rapid antibody testing process that is readily available and can be utilized for convalescent plasma donor screening.
Methods
Plasma Collection
Convalescent plasma was obtained from the Blood Donation Service at Maimonides Medical Center. Convalescent plasma was donated by recovered COVID-19 patients with confirmed diagnosis via clinical laboratory test and were symptom free for at least 28 days. Some CCP donors were initially screened for the presence of antibodies via Dynex AgilityTM instrument through a reference laboratory (Table 1). An aliquot of plasma or serum was collected and stored at −20oC for subsequent antibody testing at the time of plasma donation. This study was approved by the IRB/Research Committee at Maimonides Medical Center. The requirement to obtain informed consent from the subjects has been waived by the IRB in accordance with 45 C.F.R. § 46.116(d) and the guidelines outlined in the Declaration of Helsinki were followed.
 |
Table 1 Characteristics of Volunteer Donors and Their Donated Plasma |
The plasma or serum samples were analyzed for anti-SARS-CoV-2 RBD IgG antibodies using the Beckman Coulter chemiluminescent immunoassay (CLIA) in accordance with manufacturer instructions (https://www.beckmancoulter.com/en/products/immunoassay/access-sars-cov-2-igg-antibody-test). This CLIA test utilizes a recombinant SARS-CoV-2 protein specific for the receptor-binding domain (RBD) of the S1 protein. Results of this assay are based on the sample signal-to-cut-off (S/Co) ratio, with values <1.0 and >/= 1.0 corresponding to negative and positive results, respectively. The S/Co values reflect relative levels of anti-SARS-CoV-2 IgG antibodies.
ELISA IgG
Assay controls and serum samples were diluted to 1:64 followed by 2-fold dilution up to 1:32,768 and added to a 96-well microtiter plate (Thermo Scientific Immulon, Waltham, MA, USA) that was coated with SARS-CoV-2 recombinant RBD (Mount Sinai Medical Center, New York, NY, USA). A secondary anti-human IgG (Fab specific) antibody labeled with horse radish peroxidase (Sigma-Aldrich, St. Louis, MO, USA) was added to each well to form a specific complex of antigen-antibody bound to the plate surface. The binding reaction was then enhanced visually with SIGMAFASTTM OPD (Sigma-Aldrich, St. Louis, MO, USA) substrate generating a yellow color for positive specimens. After application of the stop solution (3M Hydrochloric acid), the color changed from yellow to orange and optical density was measured at 490 nm. When the absorbance value was greater than the cut-off value (OD490 = 0.15) at a minimum dilution of 1:64, the specimen was reported as a positive result and the corresponding titer reported.
Statistical Analysis
The analysis population included COVID-19 convalescent plasma samples collected at the Blood Donation Service, therefore, no sample size calculations were performed. Correlation between CLIA and ELISA titer was analyzed using Spearman’s rank, R square and κ index. The sensitivity and specificity of the two methods were analyzed using a 2x2 table. Linear relationship between the two assays was also assessed using a linear regression model, though due to violation of model assumptions, both CLIA and ELISA values were transformed to maintain the normality of residuals. The square-root of the CLIA assay was used to predict the log-transformed ELISA. The outputs of this model were then transformed back into raw units to show the relationship between the assays. A logistic regression model was created, as well, to predict the probability of a positive ELISA result (titer ≥ 1:1024). A CLIA S/Co cut-off value of 8.2 was determined using the ROC curve to meet 90% sensitivity and 82% specificity.
A two-sided P value of less than 0.05 was considered to indicate statistical significance. Statistical analyses were performed using SAS version 9.4 (Cary, NC) and GraphPad Prism 8 (GraphPad Software, La Jolla, CA).
Results
A total of 179 donated convalescent plasma units were included in this study. The inclusion criteria were mild COVID-19 symptoms and positive RT-qPCR tests on nasopharyngeal swab samples. Females and those younger than 18 years of age were excluded. The characteristics of the donors are summarized in Table 1.
Aliquots of all units were analyzed using the Beckman Coulter CLIA platform and the relative amount of antibodies as measured by the S/Co value was recorded. Twenty-one samples (21/179; 11.7%) had S/Co values of less than 1.0 and were deemed negative for antibodies. Fifteen units that had S/Co values >1.0 (15/179; 8.4%) had negative ELISA titers. The majority of the units had a titer of ≥1:1024 (95/179, 53.1%). The sensitivity of ELISA was comparable to the sensitivity of CLIA (90.5% vs 88.3%, respectively; p-value = 0.18; Table 2). Table 3 shows the distribution of CLIA S/Co values and ELISA IgG antibody titers of all donated convalescent plasma. Samples with S/Co values of less than 10 had ELISA IgG titers ranging from 0 (negative) to 1:4096. Those with S/Co values of 10–20 had titers varying from 1:128 to 1:4096. Samples with CLIA S/Co value higher than 20 had titers of 1:2048 to 1:32,768.
 |
Table 2 Sensitivity of the Antibody Tests Compared to SARS-CoV-2 Viral RNA RT PCR |
 |
Table 3 CLIA S/Co Values and ELISA Titers of Donated Convalescent Plasma |
Using linear regression, there is positive linear correlation between CLIA S/Co values and ELISA IgG titer (Rho = 0.75; Spearman’s rank = 0.82, p-value = <0.0001; Figure 1). The agreement between the two methods was fair, with a κ index of 0.2741. A linear regression model was developed using the square root of the S/Co value and logarithmic transformation of the equivalent ELISA IgG titer (Table 4). Using this linear regression model, an estimate of the equivalent ELISA IgG titer when given a CLIA S/Co value can be computed using the following formula:
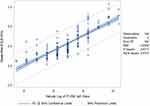 |
Figure 1 Linear regression analysis on the relationship of CLIA S/Co values and ELISA IgG titers. |
 |
Table 4 Linear Regression Model of the Root of Intensity as a Predictor of the Logarithmic Value of ELISA IgG Titer |
Table 5 provides correlation of representative CLIA intensity to its expected titer concentration. Further, a predictive statistical file was created using this formula for more exact conversions from CLIA values to expected titer determinations (Supplementary Table). This file enables the user to enter any CLIA S/Co value and will automatically generate the equivalent ELISA IgG titer with 95% confidence intervals.
 |
Table 5 Representative CLIA Intensity to Expected Titer Concentrations |
A receiver operating characteristic (ROC) curve analysis was utilized to determine the optimal CLIA S/Co value that will reliably correspond to a high ELISA IgG titer (>1:1024). This analysis identified that a CLIA S/Co cutoff value of 8.2 gives a sensitivity of 90% and a specificity of 82% in predicting a titer dilution of more than 1:1024 (Figure 2).
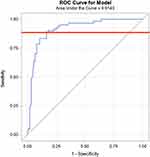 |
Figure 2 Receiver operating characteristic (ROC) curve showing an area under the curve (AUC) of 0.9143. The orange line denotes a CLIA S/Co cutoff value of 8.2. |
Discussion
Our results showed significant linear correlation between the CLIA S/Co values and ELISA IgG titers. This indicates that comparable antibody quantification can be reliably derived from the results of an automated antibody detection system. This observation agrees with the conclusion of a similar study that analyzed the quantification capability of 8 commercially available immunoassays, two of which were CLIA-based.19 They found that the automated systems provided the highest sensitivities (up to 98%), and one of the CLIA platforms had the best overall quantitative correlation to the neutralization titer (Rho=0.729).
Neutralization assays are the gold standard for assessing specific immunity and a benchmark for other antibody assays. These tests are very complex, require incubation times of 5–7 days and a biosafety level 3 laboratory, limiting the routine use of these assays on a large scale.20,21 Previous studies have shown positive correlation between neutralizing titer and ELISA that detects IgG antibodies against the SARS-CoV-2 S1 protein19 or IgG antibodies against the receptor-binding domain of the S1 protein.14 Similar studies correlating CLIA S/CO with ELISA assessment have been reported in other infectious disease spaces such as hepatitis,22 measles23 mycoplasma24 and HIV25 for diagnosis as well as donor infectious disease screening, where applicable. In a similar manner, the results of this study demonstrated correlation between ELISA antibody titer and CLIA S/Co values. Since ELISA titers correlate with the amount of neutralizing antibody, our results extend this association to support the use of automated CLIA-based systems as a fast and convenient method of estimating neutralizing antibody quantities in convalescent plasma.
Only about a third of convalescent plasma units collected at our center had S/Co values of at least 8.2 and antibody titers of more than 1:1024. Preliminary evidence with COVID-19 suggests that patients with mild symptoms may develop very low titer antibodies.14,26,27 One study observed that 18% of convalescent COVID-19 donors had undetectable neutralizing titers in their plasma samples collected an average of 30 days after the onset of symptoms.28 A larger study performed by NYBC showed that more than half of plasma donors had low neutralizing titers and that there was a large variation in antibody titers among donors.29 There is an existing challenge to screen convalescent plasma donors for antibody titers, as evidence shows that the majority of mild COVID-19 patients may not develop an adequate level of anti-SARS- CoV-2 antibodies to provide a therapeutic benefit. Screening of donated convalescent plasma units for antibody is, therefore, an essential step to ensure the efficacy of convalescent plasma as a therapeutic product with viral-neutralizing capacity.
Testing for antibodies should be performed to first confirm the presence of antibodies. Subsequent identification of CCP units that provide higher concentrations of anti-COVID (neutralizing) antibodies may provide a more appropriate logic of selecting optimal CCP units for patient administration. The utility of automated antibody detection systems can be extended from simply a screening method to a semi-quantitative and quantitative functional antibody analysis. CLIA S/Co values can be used to estimate the ELISA antibody titer, and this can immediately provide crucial information on dosing of convalescent plasma. As a semi-quantitative method, a CLIA S/Co cut-off value of 8.2 can be used to reliably detect convalescent plasma donors or units with more than 1:1024 IgG antibody titer (90% sensitivity; 82% specificity); this value is well above the recently published FDA guidance of S/Co) ≥3.3 indicative of high titer CCP further relating the values to functional anti-viral titer-based activity using alternative instrumentation and/or methodologies.30
Application of additional approaches to assess the anti-COVID antibody concentration in CCP is paramount. In addition, methodological differences can introduce differences in results. For example, chemiluminescence methodology is able to determine total Ig (IgM, IgG, IgA) whereas ELISA determines only IgG. This may very well contribute towards the observed differences when a sample may have high percentage of IgM (giving high CLIA number), and low IgG (low titer) and vice versa. Although certain instruments have been promoted and approved to serve this need, many institutions and blood centers lack access to such instruments thereby causing a bottleneck in reference lab identification of appropriate products for transfusion. Expansion to additional CLIA instrumentation can reduce the strain of CCP assessment and offer the ideal plasma product for patients who require such.
Furthermore, the methodology employed by CLIA instrumentation could be used to examine response to COVID-19 vaccination and, depending on target, provide distinction between immune response to COVID-19 infection from the immune response to vaccination.
The rapid turnover and cost benefit of such a CLIA based screening approach provides additional value. The testing can be easily implemented into existing chemiluminescent systems and provides opportunity costs for run time and personal skill set allocation. For example, in certain cases, time to result can be reduced from 180 minutes (ELISA) to 30 minutes (CLIA). In addition, adaptation can be automated in a CLIA setting and may require a lower degree of operator expertise (particularly if automated) than with ELISA.22 This can beneficially impact the operational logic of the clinical laboratory to optimize efficiency of motion, human capital as well as resource allocation.
Acknowledgments
The authors would like to acknowledge Harsha Bajaj, Sergio Valentini and Lilian Castaneda of the transplant laboratory at SUNY Downstate Medical Center and Jean Allen and Lucy Liu Dong of the Blood Transfusion and Donor Services at Maimonides Medical Center for their assistance and support.
Funding
Support for this study has been provided by the Department of Pathology at Maimonides Medical Center and SUNY Downstate Medical Center.
Disclosure
No conflicting relationship exists for any author.
References
1. World Health Organization. Coronavirus disease (COVID-19) Weekly Epidemiological Update and Weekly Operational Update. Available from: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports.
2. Duan K, Liu B, Li C, et al. Effectiveness of convalescent plasma therapy in severe COVID-19 patients. PNAS. 2020;117(17):9490–9496. doi:10.1073/pnas.2004168117
3. Shen C, Wang Z, Zhao F, et al. Treatment of 5 critically Ill patients with COVID-19 with convalescent plasma. JAMA. 2020;323(16):1582–1589. doi:10.1001/jama.2020.4783
4. Chen L, Xiong J, Bao L, Shi Y. Convalescent plasma as a potential therapy for COVID-19. Lancet Infect Dis. 2020;20(4):398–400. doi:10.1016/S1473-3099(20)30141-9
5. Liu STH, Lin HM, Baine I, et al. Convalescent plasma treatment of severe COVID-19: a propensity score–matched control study. Nat Med. 2020;26:1708–1713. doi:10.1038/s41591-020-1088-9
6. Joyner MJ, Senefeld JW, Klasses SA, et al. Effect of convalescent plasma on mortality among hospitalized patients with COVID-19: initial three-month experience. MedRxiv. doi:10.1101/2020.08.12.20169359
7. Wu F, Liu M, Wang A, et al. Evaluating the association of clinical characteristics with neutralizing antibody levels in patients who have recovered from mild COVID-19 in Shanghai, China. JAMA Intern Med. 2020;180(10):1356. doi:10.1001/jamainternmed.2020.4616
8. Choe P, Kang C, Suh H, et al. Antibody responses to SARS-CoV-2 at 8 weeks postinfection in asymptomatic patients. Emerg Infect Dis. 2020;26(10):2484–2487. doi:10.3201/eid2610.202211
9. Ibarrondo FJ, Fulcher JA, Goodman-Meza D, et al. Rapid decay of anti–SARS-CoV-2 antibodies in persons with mild COVID-19. N Engl J Med. 2020;383(11):1085–1087. doi:10.1056/NEJMc2025179
10. FDA. Recommendations for Investigational COVID-19 Convalescent Plasma. Available from: https://www.fda.gov/vaccines-blood-biologics/investigational-new-drug-ind-or-device-exemption-ide-process-cber/recommendations-investigational-COVID-19-convalescent-plasma.
11. Nie J, Li Q, Wu J, et al. Establishment and validation of a pseudovirus neutralization assay for SARS-CoV-2. Emerg. Microbes Infect. 2020;9:680–686. doi:10.1080/22221751.2020.1743767
12. Tang YW, Schmitz JE, Persing DH, Stratton CW. The laboratory diagnosis of COVID-19 infection: current issues and challenges. J Clin Microbiol. 2020;58:e00512–20. doi:10.1128/JCM.00512-20
13. Tan CW, Chia WN, Qin X, et al. A SARS-CoV-2 surrogate virus neutralization test based on antibody-mediated blockage of ACE2–spike protein–protein interaction. Nat Biotechnol. 2020;38(9):1073–1078. doi:10.1038/s41587-020-0631-z
14. Salazar E, Kuchipudi SV, Christensen PA, et al.Relationship between Anti-Spike Protein Antibody Titers and SARS-CoV-2 In Vitro Virus Neutralization in Convalescent Plasma. Preprint. bioRxiv. 2020;. doi:10.1101/2020.06.08.138990.
15. Campbell AK, Patel A. A homogeneous immunoassay for cyclic nucleotides based on chemiluminescence energy transfer. Biochem J. 1983;216(1):185–194. doi:10.1042/bj2160185
16. Vo-Dinh T. Chemiluminescence. Encyclopedia of Applied Physics. 2003. doi:10.1002/3527600434.eap063
17. Wang YFW, Kobayashi M. Antibody Detection: Principles and Applications/Advanced Techniques in Diagnostic Microbiology. Springer US; 2013:53–73.
18. Cinquanta L, Fontana DE, Bizzaro N. Chemiluminescent immunoassay technology: what does it change in autoantibody detection? Auto Immun Highlights. 2017;8:9. doi:10.1007/s13317-017-0097-2
19. Weidner L, Gansdorfer S, Unterweger S, et al. Quantification of SARS-CoV-2 antibodies with eight commercially available immunoassays. J of Clin Vir. 2020;129:104540. doi:10.1016/j.jcv.2020.104540
20. Kohmer N, Westhaus S, Rühl C, Ciesek S, Rabenau HF. Clinical performance of different SARS-CoV-2 IgG antibody tests. J. Med. Virol. 2020;1–5. doi:10.1002/jmv.26145
21. Perera RAPM, Mok CKP, Tsang OTY, et al. Serological assays for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), March 2020. Eurosurveillance. 2020;25(16). doi:10.2807/1560-7917.ES.2020.25.16.2000421
22. Madiyal M, Sagar S, Vishwanath S, Banerjee B, Eshwara VK, Chawla K. Comparing Assay Performance of ELISA and Chemiluminescence Immunoassay in Detecting Antibodies to Hepatitis B Surface Antigen. J Clin Diagn Res. 2016;10:DC22–DC25. doi:10.7860/JCDR/2016/24108.8921
23. de Ory F, Minguito T, Balfagón P, Sanz JC. Comparison of chemiluminescent immunoassay and ELISA for measles IgG and IgM. APMIS. 2015;123(8):648–651. doi:10.1111/apm.12413
24. Chen D, Zhang Y, Xu Y, et al. Comparison of chemiluminescence immunoassay, enzyme-linked immunosorbent assay and passive agglutination for diagnosis of Mycoplasma pneumoniae infection. Ther Clin Risk Manag. 2018;14:1091–1097. doi:10.2147/TCRM.S159227
25. Chang L, Zhao J, Guo F, et al. Comparative evaluation and measure of accuracy of ELISAs, CLIAs, and ECLIAs for the detection of HIV infection among blood donors in China. Can J Infect Dis Med Microbiol. 2020;2020:2164685. doi:10.1155/2020/2164685
26. Klein S, Pekosz A, Park H-S, et al. Sex, age, and hospitalization drive antibody responses in a COVID-19 convalescent plasma donor population. J Clin Invest. 2020;130(11):6141–6150. doi:10.1172/JCI142004
27. Madariaga MLL, Guthmiller JJ, Schrantz S, et al. Clinical predictors of donor antibody titre and correlation with recipient antibody response in a COVID-19 convalescent plasma clinical trial. J Intern Med. 2020. doi:10.1111/joim.13185
28. Robbiani DF, Gaebler C, Muecksch F, et al. Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature. 2020;584(7821):437–442. doi:10.1038/s41586-020-2456-9
29. Luchsinger LL, Ransegnola BP, Jin DK, et al. Serological assays estimate highly variable SARS-CoV_2 neutralizing antibody activity in recovered COVID-19 patients. J. Clin. Microbiol. 2020. doi:10.1128/JCM.02005-20
30. https://www.fda.gov/media/141477/download.
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.