Back to Journals » Infection and Drug Resistance » Volume 11
Comparative analysis of bla KPC-2- and rmtB-carrying IncFII-family pKPC-LK30/pHN7A8 hybrid plasmids from Klebsiella pneumoniae CG258 strains disseminated among multiple Chinese hospitals
Authors Shi L, Feng J, Zhan Z, Zhao Y, Zhou H, Mao H, Gao Y, Zhang Y, Yin Z, Gao B, Tong Y , Luo Y, Zhang D, Zhou D
Received 22 April 2018
Accepted for publication 30 July 2018
Published 11 October 2018 Volume 2018:11 Pages 1783—1793
DOI https://doi.org/10.2147/IDR.S171953
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Lining Shi,1,* Jiao Feng,2,* Zhe Zhan,2 Yuzong Zhao,2,3 Haijian Zhou,4 Haifeng Mao,5 Yingjie Gao,6 Ying Zhang,7 Zhe Yin,2 Bo Gao,2 Yigang Tong,2 Yanping Luo,7 Defu Zhang,2,3 Dongsheng Zhou2
1Institute of Medical Laboratory Sciences, Jinling Hospital, School of Medicine, Nanjing University, Nanjing 210002, China; 2State Key Laboratory of Pathogen & Biosecurity, Beijing Institute of Microbiology & Epidemiology, Beijing 100071, China; 3College of Food Science & Project Engineering, Bohai University, Jinzhou 121013, China; 4National Institute for Communicable Disease Control and Prevention, Chinese Center for Disease Control and Prevention, Beijing 102206, China; 5Department of Laboratory Medicine, The First People’s Hospital of Lianyungang, Lianyungang 222002, China; 6College of Veterinary Medicine, Jilin University, Changchun 130062, China; 7Medical Laboratory Center, Chinese People’s Liberation Army General Hospital, Beijing 100085, China
*These authors contributed equally to this work
Background: We recently reported the complete sequence of a blaKPC-2- and rmtB-carrying IncFII-family plasmid p675920-1 with the pKPC-LK30/pHN7A8 hybrid structure. Comparative genomics of additional sequenced plasmids with similar hybrid structures and their prevalence in blaKPC-carrying Klebsiella pneumoniae strains from China were investigated in this follow-up study.
Methods: A total of 51 blaKPC-carrying K. pneumoniae strains were isolated from 2012 to 2016 from five Chinese hospitals and genotyped by multilocus sequence typing. The blaKPC-carrying plasmids from four representative strains were sequenced and compared with p675920-1 and pCT-KPC. Plasmid transfer, carbapenemase activity determination, and bacterial antimicrobial susceptibility test were performed to characterize resistance phenotypes mediated by these plasmids. The prevalence of pCT-KPC-like plasmids in these blaKPC-carrying K. pneumoniae strains was screened by PCR.
Result: The six KPC-encoding plasmids p1068-KPC, p20049-KPC, p12139-KPC and p64917-KPC (sequenced in this study) and p675920-1 and pCT-KPC slightly differed from one another due to deletion and acquisition of various backbone and accessory regions. Two major accessory resistance regions, which included the blaKPC-2 region harboring blaKPC-2 (carbapenem resistance) and blaSHV-12 (β-lactam resistance), and the MDR region carrying rmtB (aminoglycoside resistance), fosA3 (fosfomycin resistance), blaTEM-1B (β-lactam resistance) and blaCTX-M-65 (β-lactam resistance), were found in each of these six plasmids and exhibited several parallel evolution routes. The pCT-KPC-like plasmids were present in all the 51 K. pneumoniae isolates, all of which belonged to CG258.
Conclusion: There was clonal dissemination of K. pneumoniae CG258 strains, harboring blaKPC-2- and rmtB-carrying IncFII-family pKPC-LK30/pHN7A8 hybrid plasmids, among multiple Chinese hospitals.
Keywords: Klebsiella pneumoniae, multidrug resistance, carbapenemase, plasmids
Introduction
Klebsiella pneumoniae belongs to the family Enterobacteriaceae and is usually found in human gastrointestinal tract and the environment especially in the hospital settings.1 K. pneumoniae has emerged as one of the most important opportunistic bacterial pathogens associated with over 70% of human infections from bloodstream, respiratory tract, and urinary tract.2,3 Carbapenems are used as the first-line therapeutic agents to treat life-threatening infections caused by multidrug-resistant (MDR) K. pneumoniae, but carbapenem-resistant K. pneumoniae strains have spread worldwide and posed a severe threat to public health with the mortality up to 40%–50%.4 The production of carbapenemases, such as K. pneumoniae carbapenemase (KPC), is an important mechanism of carbapenem resistance in K. pneumoniae.5
KPC enzymes belong to Ambler class A and are able to hydrolyze all β-lactam antimicrobial agents including carbapenems.4,6 KPC-2 is the first variant of KPC enzymes found in 1996,5 and, to date, at least 31 different KPC variants have been identified (https://www.ncbi.nlm.nih.gov/pathogens/beta-lactamase-data-resources/). The blaKPC genes are mainly carried on plasmids of different incompatibility (Inc) groups such as IncFII, FIA, I2, A/C, N, X, R, P, U, W, and L/M.5 IncFII plasmids commonly harbor one or more replicons in addition to FII, and they have been frequently isolated from Enterobacteriaceae5 and found to carry various antibiotic resistance genes, including those responsible for resistance to carbapenems (such as blaKPC7 and blaNDM8), extended-spectrum cephalothins (such as blaCTX-M-659 and blaCTX-M-143), aminoglycosides (such as armA10 and rmtB11), and fosfomycins (such as fosA39).
The most predominant isolates of KPC-producing K. pneumoniae belong to the clonal group 258 (CG258), which mainly includes the sequence type 258 (ST258; allelic profile 3–3–1–1–1–1–79 corresponding to the collection of seven housekeeping genes) and its single-locus variation ST11 (allelic profile 3–3–1–1–1–1–4) as defined by multilocus sequence typing (MLST).5 The genome of ST258 is a hybrid of ST11 (80%) and ST442 (20%), and the ST258 strains can be segregated into clades I and II based on the distinct capsule polysaccharide gene (cps) regions; clade I (cps1) has been evolved from clade II (cps2) as a result of horizontal transfer of cps (cps2) region from ST42 to clade II, which is mediated by the mobile element ICEKp258.2 of clade II.5,12 The ST258 and ST11 strains have been identified worldwide – ST258 is mostly prevalent in America and Europe, while ST11 is the highly dominant clone in Asia (especially in China) and Latin America.5
We recently reported the complete nucleotide sequence of a blaKPC-2-carrying IncFII plasmid p675920-1 (accession number, MF133495) from an MDR K. pneumoniae isolate 675920.13 p675920-1 was essentially a hybrid of the rmtB-carrying IncFII plasmid pHN7A8 (accession number, JN232517)9 and the blaKPC-2-carrying IncR plasmid pKPC-LK30 (accession number, KC405622),14 and almost all of the backbone and accessory regions of pKPC-LK30 and pHN7A8 were presented in p675920-1.
This follow-up study presented four fully sequenced blaKPC-2-carrying plasmids, p1068-KPC, p20049-KPC, p12139-KPC, and p64917-KPC, which were genetically closely related to p675920-1. A total of nine available sequenced plasmids (including the abovementioned five plasmids) constituted a novel group of IncFII plasmids with pHN7A8/pKPC-LK30 hybrid structures. This group of plasmids was disseminated among K. pneumoniae CG258 strains from at least nine distinct hospitals in eight different cities of China.
Materials and methods
Identification of bacterial strains
The use of human specimens and all related experimental protocols were approved by the Committee on Human Research of Jinling Hospital and carried out in accordance with the approved guidelines. The selected patients signed the written informed consent. All K. pneumoniae strains, including the 1068, 20049, 12139, and 64917 isolates, were subjected to species identification using 16S rRNA gene sequencing15 and PCR detection of K. pneumoniae-specific gene khe.16 The MLST scheme was derived from the Pasteur Klebsiella MLST database (http://bigsdb.pasteur.fr/klebsiella/klebsiella.html).
Plasmid conjugal transfer
Plasmid conjugal transfer experiments were carried out with the rifampin-resistant Escherichia coli EC600 being used as recipient and each of the 1068, 20049, 12139, and 64917 isolates as donor. Three milliliters of overnight cultures of each of donor and recipient bacteria were mixed together, harvested, and resuspended in 80 µL of brain heart infusion (BHI) broth (BD Biosciences, San Jose, CA, USA). The mixture was spotted on a 1-cm2 hydrophilic nylon membrane filter with a 0.45-µm pore size (EMD Millipore, Billerica, MA, USA) that was placed on BHI agar (BD Biosciences) plate and then incubated for mating at 37°C for 12–18 hours. Bacteria were washed from filter membrane and spotted on Muller–Hinton (MH) agar (BD Biosciences) plates containing 1,000 µg/mL rifampin together with 4 µg/mL meropenem for selecting an E. coli transconjugant-carrying blaKPC.
Plasmid electroporation
To prepare competent cells for plasmid electroporation, 200 mL of overnight culture of E. coli TOP10 in super optimal broth (SOB) at an OD600 of 0.4–0.6 was washed three times with electroporation buffer (0.5 M mannitol and 10% glycerol) and concentrated into a final volume of 2 mL. One microgram of plasmid DNA, which was isolated from each of the 1068, 20049, 12139, and 64917 strains using a Qiagen Plasmid Midi Kit (Qiagen, Hilden, Germany), was mixed with 100 µL of competent cells for electroporation at 25 µF, 200 Ω, and 2.5 kV. The resulting cells were suspended in 500 µL of SOB, and an appropriate aliquot was spotted on SOB agar plates containing 4 µg/mL meropenem for selecting an E. coli electroporant-carrying blaKPC.
DNA sequencing and sequences assemble
Genomic DNA was isolated from the 1068 or 20049 isolate using a Qiagen Blood & Cell Culture DNA Maxi Kit (Qiagen). Plasmid DNA was isolated from the E. coli electroporant, containing p12139-KPC or p64917-KPC, with a Qiagen Large Construct Kit (Qiagen).
For the 1068 isolate, genome sequencing was performed with a sheared DNA library with an average size of 15 kb (ranged from 10 to 20 kb) on a PacBio RSII sequencer (Pacific Biosciences, CA, USA), as well as a paired-end library with an average insert size of 400 bp (ranged from 150 to 600 kb) on a HiSeq sequencer (Illumina, CA, USA). The paired-end short Illumina reads were used to correct the long PacBio reads utilizing proovread,17 and then the corrected PacBio reads were assembled de novo utilizing SMARTdenovo (available from https://github.com/ruanjue/smartdenovo).
Genomic DNA of 20049, or plasmid DNA of p12139-KPC or p64917-KPC, was sequenced from a mate pair library with an average insert size of 5 kb (ranged from 2 to 10 kb) using a MiSeq sequencer (Illumina). DNA contigs that were not matching with the corresponding chromosome sequences were assembled based on their contig coverage values using Newbler 2.6.18 Gaps between contigs were filled using a combination of PCR and Sanger sequencing using an ABI 3730 Sequencer.
Sequence annotation and genome comparison
Open reading frames (ORFs) and pseudogenes were predicted using RAST 2.019 combined with BLASTP/BLASTN searches20 against the UniProtKB/Swiss-Prot21 and RefSeq22 databases. Annotation of resistance genes, mobile elements, and other features was carried out using the online databases including CARD,23 ResFinder,24 ISfinder,25 and INTEGRALL.26 Multiple and pairwise sequence comparisons were performed using MUSCLE 3.8.3127 and BLASTN, respectively. Gene organization diagrams were drawn in Inkscape 0.48.1 (https://inkscape.org/en/).
Phylogenetic analysis
The backbone regions of six plasmids p675920-1, p1068-KPC, p20049-KPC, p12139-KPC, p64917-KPC, and pCT-KPC were compared with that of concatenated pHN7A8+ pKPC-LK30 by MUMmer 3.0.28 Inference of homologous recombination was performed using ClonalFrameML29 to remove recombination-associated single-nucleotide polymorphisms (SNPs). A maximum-likelihood tree was constructed from recombination-free SNPs using MEGA730 with a bootstrap iteration of 1000, using concatenated pHN7A8+ pKPC-LK30 as outgroup.
Detection of carbapenemase activity
The activity of class A/B/D carbapenemases in bacterial cell extracts was determined via a modified CarbaNP test.31 Overnight bacterial cell culture in MH broth was diluted 1:100 into 3 mL of fresh MH broth, and bacteria were allowed to grow at 37°C with shaking at 200 rpm to reach an OD600 of 1.0–1.4. If required, ampicillin was used at 200 µg/mL. Bacterial cells were harvested from 2 mL of the above culture and washed twice with 20 mM Tris-HCl (pH 7.8). Cell pellets were resuspended in 500 µL of 20 mM Tris-HCl (pH 7.8) and lysed by sonication, followed by centrifugation at 10,000× g at 4°C for 5 minutes. Fifty microliters of the supernatant (the enzymatic bacterial suspension) were mixed with 50 µL of substrate I–V, respectively, followed by incubation at 37°C for 2 hours: substrate I: 0.054% phenol red plus 0.1 mM ZnSO4 (pH 7.8), substrate II: 0.054% phenol red plus 0.1 mM ZnSO4 (pH 7.8) and 3 µg/µL imipenem, substrate III: 0.054% phenol red plus 0.1 mM ZnSO4 (pH 7.8), 3 µg/µL imipenem, and 4 µg/µL tazobactam, substrate IV: 0.054% phenol red plus 0.1 mM ZnSO4 (pH 7.8), 3 µg/µL imipenem, and 3 mM EDTA (pH 7.8), substrate V: 0.054% phenol red plus 0.1 mM ZnSO4 (pH 7.8), 3 µg/µL imipenem, 4 µg/µL tazobactam, and 3 mM EDTA (pH 7.8).
Bacterial antimicrobial susceptibility test
Bacterial antimicrobial susceptibility was tested by BioMérieux VITEK 2 and interpreted using Clinical and Laboratory Standards Institute guidelines.32
Nucleotide sequence accession numbers
The complete sequences of plasmids p1068-KPC, p20049-KPC, p12139-KPC, and p64917-KPC were submitted to GenBank under accession numbers MF168402, MF168404, MF168403, and MF168405, respectively.
Results and discussion
Clonal blaKPC-carrying K. pneumoniae isolates
During 2012–2016, 51 blaKPC-positive K. pneumoniae isolates were recovered from five types of specimens (sputum, urine, blood, pus, and peritoneal lavage fluids) of the patients from eight different departments (Intensive Care Units, Department of Respiratory Medicine, Department of Neurosurgery, Department of Neurology, Department of Emergency, Department of Endocrinology, Department of General Surgery, and Department of Hematology) in five distinct public hospitals from Beijing City, Nanjing City, Shijiazhuang City, and Lianyungang City in China (Table S1). These patients were admitted with primary diseases to the hospitals and suffered from different kinds of hospital-acquired infections during hospitalization. They received intravenous treatment of susceptible antimicrobial agents based on the antimicrobial susceptibility test results, and their infection symptoms were progressively improved.
As determined by MLST, these 51 isolates could be assigned into three different STs, namely ST11 (49 strains), ST1711 (one strain), and ST2040 (one strain), all of which belonged to the CG258. The above results indicated the clonal dissemination of blaKPC-carrying K. pneumoniae in these hospitals.
pCT-KPC-like IncFII plasmids in K. pneumoniae isolates
The previously sequenced blaKPC-2-carrying p675920-1 was recovered from K. pneumoniae 675920 from one of the above hospitals.13 In this study, one K. pneumoniae isolate with the widest drug-resistance spectrum was selected from each of the remaining four hospitals to characterize the blaKPC-carrying plasmids in these strains. Repeated attempts through conjugation failed to transfer the blaKPC markers of the four selected strains (1068, 20049, 12139, and 64917) into E. coli EC600. However, these blaKPC markers could be transferred into E. coli TOP10 through electroporation, yielding four corresponding blaKPC-positive E. coli electroporants. All these wild-type and electroporant strains had the Ambler class A carbapenemase activity (data not shown) and were resistant to ampicillin, ampicillin/sulbactam, piperacillin, piperacillin/tazobactam, cefazolin, cefuroxime, cefuroxime axetil, ceftazidime, ceftriaxone, aztreonam, imipenem, and meropenem (Table 1); the production of KPC enzymes accounted for the above phenotypes of these strains.
  | Table 1 Antimicrobial drug susceptibility profiles Abbreviations: I, intermediately resistant; MIC, minimum inhibitory concentration; R, resistant; S, sensitive. |
High-throughput sequencing with indicated wild-type and electroporant strains showed that the 1068, 20049, 12139, and 64917 isolates contained the blaKPC-2-carrying plasmids p1068-KPC, p20049-KPC, p12139-KPC, and p64917-KPC, respectively. These four plasmids have circularly closed sequences, 146,103, 151,653, 169,424, and 169,419 bp in length, with mean G+C contents of 53.7%, 53.5%, 53.6%, and 53.5%, and contained 208, 212, 221, and 219 predicted ORFs in total, respectively (Table 2).
  | Table 2 Major features of plasmids analyzed Note: aThese accessory modules contained resistance genes. Abbreviations: MDR, multidrug resistance; ORF, open reading frame. |
BLAST analysis indicated a total of nine sequenced plasmids (Table 3; including the above five plasmids sequenced in our laboratory) constituted a novel group of plasmids, which had pHN7A8/pKPC-LK30 hybrid structures as characterized previously.13,33 Notably, all of these nine plasmids were recovered from clinical K. pneumoniae isolates from China. The five plasmids p675920-1, p1068-KPC, p20049-KPC, p12139-KPC, and p64917-KPC sequenced in our laboratory together with the first sequenced pCT-KPC were included in a genomic comparison herein. p675920-1 was used as the reference for genomic comparison because its hybrid structure had been characterized in great detail.13
General genomic diversity of pCT-KPC-like plasmids
The molecular structure of each of p675920-1, p1068-KPC, p20049-KPC, p12139-KPC, p64917-KPC, and pCT-KPC was divided into the backbone, and several accessory modules that were defined as acquired DNA regions associated with and bordered with mobile elements and inserted at different sites of the backbone (Figure S1). These six plasmids were highly similar (>82% BLAST query coverage and >99% maximum nucleotide identity) to one another across whole genomes (Table S2).
There were totally 67 SNPs (among them 44 were recombination free) presented within the common (core) backbone regions of these six plasmids. A maximum likelihood phylogenetic tree (Figure S2) was constructed based on the recombination-free SNPs, and accordingly these six plasmids could be assigned into two clades I (p20049-KPC, pCT-KPC, p1068-KPC, and p675920-1) and II (p12139-KPC and p64917-KPC).
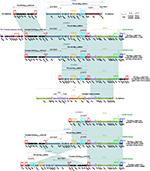
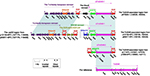
The backbone gene contents of these six plasmids displayed four major modular differences (Figure 1): 1) p20049-KPC contained a unique 4.8-kb maintenance region (from relB to mobB) relative to the other five plasmids; 2) a 15.5-kb plasmid maintenance region (from orf375 to parA) was found in p12139-KPC, p64917-KPC, and p675920-1 but not pCT-KPC, p1068-KPC, and p20049-KPC; 3) 1.9- and 7.3-kb truncations occurred in the conjugal transfer regions of pCT-KPC and p1068-KPC, respectively, which resulted from insertion of the blaKPC-2 region; and 4) the conjugal transfer region of p1068-KPC was interrupted into two separate parts due to insertion of IS26. The recombination of pKPC-LK30 (containing no conjugation transfer genes) with pHN7A8 demolished the conjugation transfer gene organization of pHN7A8, resulting in all these six plasmids to be nonconjugative.
  | Figure 1 Linear comparison of plasmid genome sequences. Notes: Genes are denoted by arrows. Genes, mobile elements, and other features are colored based on the function classification. Shading denotes the regions of homology (>95% nucleotide identity). Accession numbers of pCT-KPC and p675920-113 for reference are KT185451 and MF133495, respectively. |
These six plasmids contained several accessory modules (Figure 1): 1) the blaKPC-2 regions were found in all these plasmids and slightly differed from one another; 2) the MDR regions were observed in all these plasmids and displayed considerable modular diversity; notably, the MDR regions in pCT-KPC, p1068-KPC, and p64917-KPC/p12139-KPC were disrupted into two separated parts (probably due to IS26-mediated homologous recombination), which were located at two different sites of the backbones; and 3) the Tn6346-associated regions were presented in five plasmids except for p20049-KPC, but their gene contents display significant variations. These six plasmids carried eight resistance markers (blaKPC-2, rmtB, fosA3, catA2, blaCTX-M-65, blaTEM-1B, blaSHV-12, and mer) in total, all of which were located in the accessory modules (Table S3).
The blaKPC-2 regions
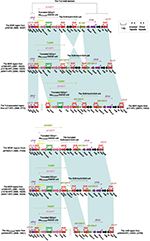
The blaKPC-2 regions of the seven plasmids pKPC-LK30, p1068-KPC, pCT-KPC, p12139-KPC, p64917-KPC, p675920-1, and p20049-KPC exhibited four stages of evolution (Figure 2) and carried three close Tn629634 derivatives (ΔTn6296-1 to ΔTn6296-3) severing as the core genetic environments of blaKPC-2 (Figure 3). First, the blaKPC-2 region of pKPC-LK30 was organized in order of a Tn3-family transposon remnant, a truncated IS26–blaSHV-12–IS26 unit, ΔTn6296-1 generated from Tn6296 after the loss of ΔTn1722-3′ and truncation of Tn6376, ∆Tn21 with insertion of IS5075 into its inverted repeat right (IRR), an IS903B remnant, and ISKpn14, and it evolved into the counterpart of p1068-KPC/pCT-KPC after the loss of the Tn3-family transposon remnant. Second, the deletion of the truncated IS26–blaSHV-12–IS26 unit, the truncation of ΔTn6296-1 into ΔTn6296-2 with deletion of ΔTn1722-5′, and the replacement of IS903B remnant–ISKpn14 by Δint–orf225–parA–orf492–IS26 were observed in p12139-KPC/p64917-KPC relative to p1068-KPC. Third, the evolution of p675920-1 from p1068-KPC involved the deletion of the truncated IS26–blaSHV-12–IS26 unit as well as the truncation of ΔTn6296-1 into ΔTn6296-3 with deletion of ΔTn1722-5′–ΔrepB–klcA–orf6–korC. Fourth, compared to p1068-KPC, p20049-KPC underwent the inversion of a large region composed of ΔTn6296-1 and the truncated IS26–blaSHV-12–IS26 unit.
The MDR regions
Similarly, the MDR regions of the seven plasmids pHN7A8,9 p1068-KPC, p12139-KPC, p64917-KPC, pCT-KPC, p675920-1, and p20049-KPC displayed four stages of evolution (Figures 2 and 4). First, the MDR region of pHN7A8 was generated from insertion of the Tn2–rmtB element at a site within insB of IS1R (making interruption and further truncation of insB), followed by that of a large region (organized sequentially as IS26, ΔTn6377,35 the IS26–fosA3–IS26 unit,36 and IS1294) at a site between tnpA and tnpR of Tn2 (leading to truncation of both tnpA and tnpR and loss of res); it evolved into the counterpart of p1068-KPC/p12139-KPC/p64917-KPC after fragmentation of IS1294 into ΔIS1294-5’ and ΔIS1294-3’ as well as inversion of IS26–fosA3–IS26–ΔIS1294-5’. Second, compared to p1068-KPC, the counterpart of pCT-KPC was split into two separate regions, namely the Tn2-associated region and the MDR region. Third, the evolution from p1068-KPC to p675920-1 involved deletion of IRLIS26–ΔIS1294-5’. Fourth, the split of the MDR region into the two separate parts (ie, the blaCTX-M-65 region and the rmtB region) and the deletion of the IS26–fosA3–IS26 unit were observed in p20019-KPC relative to p1068-KPC.
The Tn6346-associated regions and the catA2 regions
The Tn6346-associated region of p675920-1 could be divided sequentially as a Tn3-family transposon remnant, IS26, a small GIsul2 remnant containing no resistance genes, ΔTn6346-1 and IS26 (Figure 5). The counterparts of p12139-KPC and p64917-KPC split into two separate parts: 1) the catA2 region characteristic of the connection of the above Tn3-family transposon remnant with the IS26–catA2–IS26 unit as observed in p112298-catA (accession number, KY270851)37; and 2) the residue Tn6346-associated region composed in order of ISKpn14, a 288-bp IS903B remnant, ΔTn6346-1 and IS26. The residue Tn6346-associated regions of pCT-KPC and p1068-KPC could be modularly divided into IS26, ΔTn6346-2, and IS26. ΔTn6346-1 and ΔTn6346-213 were the 5′-terminal remnants (containing no resistance genes) of the Tn3-family unit transposon Tn6346.38
Prevalence of pCT-KPC-related plasmids
To illustrate the prevalence of pCT-KPC-like plasmids in the 51 strains, a total of 10 backbone genes or gene loci and two accessory resistance genes across the pCT-KPC genome were selected for PCR detection followed by amplicon sequencing. Ten selected backbone makers could be assigned into three origins: 1) repA1 (IncFII-family replication initiation protein), pemIK (toxin–antitoxin system), stbA (mediator of plasmid stability), and traC and traX (conjugal transfer proteins) from pHN7A8; 2) repB (IncR-type replication initiation protein), parA (partition protein), vagD (toxin of the toxin–antitoxin system), and relEB (toxin/antitoxin system) from pKPC-LK30; and 3) traA (conjugal transfer protein) from a 5.1-kb conjugal transfer region found in pCT-KPC rather than pHN7A8 and pKPC-LK30. The two selected resistance genes blaKPC-2 and rmtB were found in the accessory resistance regions of pKPC-LK30 and pHN7A8, respectively. Of these 51 strains, 49 harbored all 10 backbone markers, while the remaining two contained nine backbone markers except traC. blaKPC-2 and rmtB were detected in all these 51 strains. These results denoted that blaKPC-2- and rmtB-carrying pCT-KPC-like plasmids were present in all these 51 K. pneumoniae strains. In conclusion, the dissemination of blaKPC-2 and rmtB in these hospitals were associated with the spread of genetically closed related pCT-KPC-like plasmids among clonal K. pneumoniae CG258 strains.
Concluding remarks
The six IncFII-family plasmids p675920-1, pCT-KPC, p1068-KPC, p20049-KPC, p12139-KPC, and p64917-KPC displayed similar pHN7A8/pKPC-LK30 hybrid structures but slightly differed from one another due to deletion and acquisition of various backbone and accessory regions. At least seven antibiotic resistance genes, including blaKPC-2, rmtB, fosA3, blaCTX-M-65, blaTEM-1B, and blaSHV-12, were present in each of these six plasmids, mediating MDR of these isolates. pCT-KPC-like plasmids were detected herein in a collection of 51 KPC-positive K. pneumoniae CG258 isolates in five Chinese hospitals. In addition, there were still three available sequenced plasmids, namely pKP1034, pCP018455, and pKPC-CR-HvKP4 (Table 3), which also displayed pHN7A8/pKPC-LK30 hybrid structures and were recovered from K. pneumoniae CG258 isolates from three additional Chinese hospitals.33,39,40 These denoted clonal dissemination of K. pneumoniae CG258 strains, harboring blaKPC-2- and rmtB-carrying IncFII-family pKPC-LK30/pHN7A8 hybrid plasmids, in multiple Chinese hospitals. At least nine copies of IS26 were found in each of these six plasmids and would act as the common regions (ie, adaptors) to mediate homologous recombination, facilitating assembly and mobilization of accessory resistance regions within and among these highly mosaic plasmids.41–43 There were four IS26-flanking mobile elements in total, including Tn6367, the IS26-blaSHV-12-IS26 unit, the IS26-fosA3-IS26 unit, and the IS26-catA2-IS26 unit, and the last three lacked paired short DR sequences (target site duplication signals for transposition) at their both ends and therefore could be annotated as typical IS26-composite transposons.
Acknowledgment
This work was supported by the Natural Science Foundation of Liaoning Province of China (20170540022) and the China Postdoctoral Science Foundation (2015M582803).
Author contributions
All authors contributed toward data analysis, drafting and critically revising the paper and agree to be accountable for all aspects of the work.
Disclosure
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Tzouvelekis LS, Markogiannakis A, Psichogiou M, Tassios PT, Daikos GL. Carbapenemases in Klebsiella pneumoniae and other Enterobacteriaceae: an evolving crisis of global dimensions. Clin Microbiol Rev. 2012;25(4):682–707. | ||
Pitout JD, Nordmann P, Poirel L. Carbapenemase-Producing Klebsiella pneumoniae, a Key Pathogen Set for Global Nosocomial Dominance. Antimicrob Agents Chemother. 2015;59(10):5873–5884. | ||
Zhang D, Yin Z, Zhao Y, et al. p1220-CTXM, a pKP048-related IncFIIK plasmid carrying bla CTX-M-14 and qnrB4. Future Microbiol. 2017;12:1035–1043. | ||
Munoz-Price LS, Poirel L, Bonomo RA, et al. Clinical epidemiology of the global expansion of Klebsiella pneumoniae carbapenemases. Lancet Infect Dis. 2013;13(9):785–796. | ||
Chen L, Mathema B, Chavda KD, DeLeo R, Bonomo RA, Kreiswirth BN. Carbapenemase-producing Klebsiella pneumoniae: molecular and genetic decoding. Trends Microbiol. 2014;22(12):686–696. | ||
Kitchel B, Rasheed JK, Patel JB, et al. Molecular epidemiology of KPC-producing Klebsiella pneumoniae isolates in the United States: clonal expansion of multilocus sequence type 258. Antimicrob Agents Chemother. 2009;53(8):3365–3370. | ||
Chen L, Chavda KD, Melano RG, et al. Complete sequence of a bla(KPC-2)-harboring IncFII(K1) plasmid from a Klebsiella pneumoniae sequence type 258 strain. Antimicrob Agents Chemother. 2013;57(3):1542–1545. | ||
Bonnin RA, Poirel L, Carattoli A, Nordmann P. Characterization of an IncFII plasmid encoding NDM-1 from Escherichia coli ST131. PLoS One. 2012;7(4):e34752. | ||
He L, Partridge SR, Yang X, et al. Complete nucleotide sequence of pHN7A8, an F33:A-:B- type epidemic plasmid carrying blaCTX-M-65, fosA3 and rmtB from China. J Antimicrob Chemother. 2013;68(1):46–50. | ||
Du XD, Li DX, Hu GZ, et al. Tn1548-associated armA is co-located with qnrB2, aac(6’)-Ib-cr and blaCTX-M-3 on an IncFII plasmid in a Salmonella enterica subsp. enterica serovar Paratyphi B strain isolated from chickens in China. J Antimicrob Chemother. 2012;67(1):246–248. | ||
Deng Y, Zeng Z, Chen S, et al. Dissemination of IncFII plasmids carrying rmtB and qepA in Escherichia coli from pigs, farm workers and the environment. Clin Microbiol Infect. 2011;17(11):1740–1745. | ||
Chen L, Mathema B, Pitout JD, Deleo FR, Kreiswirth BN. Epidemic Klebsiella pneumoniae ST258 is a hybrid strain. MBio. 2014;5(3): e01355–14. | ||
Feng J, Yin Z, Zhan Z, et al. Structure genomics of two chimera plasmids p675920-1 and p675920-2 coexisting in a multi-drug resistant Klebsiella pneumoniae isolate. Oncotarget. 2018. Available from: https://doi.org/10.18632/oncotarget.24235. Accessed January 15, 2018. | ||
Chen YT, Lin JC, Fung CP, et al. KPC-2-encoding plasmids from Escherichia coli and Klebsiella pneumoniae in Taiwan. J Antimicrob Chemother. 2014;69(3):628–631. | ||
Frank JA, Reich CI, Sharma S, Weisbaum JS, Wilson BA, Olsen GJ. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl Environ Microbiol. 2008;74(8):2461–2470. | ||
Yin-Ching C, Jer-Horng S, Ching-Nan L, Ming-Chung C. Cloning of a gene encoding a unique haemolysin from Klebsiella pneumoniae and its potential use as a species-specific gene probe. Microb Pathog. 2002;33(1):1–6. | ||
Hackl T, Hedrich R, Schultz J, Förster F. proovread: large-scale high-accuracy PacBio correction through iterative short read consensus. Bioinformatics. 2014;30(21):3004–3011. | ||
Nederbragt AJ. On the middle ground between open source and commercial software - the case of the Newbler program. Genome Biol. 2014;15(4):113. | ||
Brettin T, Davis JJ, Disz T, et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep. 2015;5:8365. | ||
Boratyn GM, Camacho C, Cooper PS, et al. BLAST: a more efficient report with usability improvements. Nucleic Acids Res. 2013;41(Web Server issue):W29–W33. | ||
Boutet E, Lieberherr D, Tognolli M, et al. UniProtKB/Swiss-Prot, the Manually Annotated Section of the UniProt KnowledgeBase: How to Use the Entry View. Methods Mol Biol. 2016;1374:23–54. | ||
O’Leary NA, Wright MW, Brister JR, et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016;44(D1):D733–D745. | ||
Jia B, Raphenya AR, Alcock B, et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017;45(D1):D566–D573. | ||
Zankari E, Hasman H, Cosentino S, et al. Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother. 2012;67(11):2640–2644. | ||
Siguier P, Perochon J, Lestrade L, Mahillon J, Chandler M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006;34(Database issue):D32–D36. | ||
Moura A, Soares M, Pereira C, Leitão N, Henriques I, Correia A. INTEGRALL: a database and search engine for integrons, integrases and gene cassettes. Bioinformatics. 2009;25(8):1096–1098. | ||
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792–1797. | ||
Kurtz S, Phillippy A, Delcher AL, et al. Versatile and open software for comparing large genomes. Genome Biol. 2004;5(2):R12. | ||
Didelot X, Wilson DJ. ClonalFrameML: efficient inference of recombination in whole bacterial genomes. PLoS Comput Biol. 2015;11(2):e1004041. | ||
Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33(7):1870–1874. | ||
Chen Z, Li H, Feng J, et al. NDM-1 encoded by a pNDM-BJ01-like plasmid p3SP-NDM in clinical Enterobacter aerogenes. Front Microbiol. 2015;6:294. | ||
CLSI. Performance Standards for Antimicrobial Susceptibility Testing: Twenty-Fifth Informational Supplement M100-S25. Wayne, PA, USA: CLSI; 2015. | ||
Xiang DR, Li JJ, Sheng ZK, et al. Complete Sequence of a Novel IncR-F33:A-:B- Plasmid, pKP1034, Harboring fosA3, blaKPC-2, blaCTX-M-65, blaSHV-12, and rmtB from an Epidemic Klebsiella pneumoniae Sequence Type 11 Strain in China. Antimicrob Agents Chemother. 2015;60(3):1343–1348. | ||
Jiang Y, Yu D, Wei Z, Shen P, Zhou Z, Yu Y. Complete nucleotide sequence of Klebsiella pneumoniae multidrug resistance plasmid pKP048, carrying blaKPC-2, blaDHA-1, qnrB4, and armA. Antimicrob Agents Chemother. 2010;54(9):3967–3969. | ||
Shen P, Jiang Y, Zhou Z, Zhang J, Yu Y, Li L. Complete nucleotide sequence of pKP96, a 67 850 bp multiresistance plasmid encoding qnrA1, aac(6’)-Ib-cr and blaCTX-M-24 from Klebsiella pneumoniae. J Antimicrob Chemother. 2008;62(6):1252–1256. | ||
Ho PL, Chan J, Lo WU, et al. Dissemination of plasmid-mediated fosfomycin resistance fosA3 among multidrug-resistant Escherichia coli from livestock and other animals. J Appl Microbiol. 2013;114(3):695–702. | ||
Liang Q, Yin Z, Zhao Y, et al. Sequencing and comparative genomics analysis of the IncHI2 plasmids pT5282-mphA and p112298-catA and the IncHI5 plasmid pYNKP001-dfrA. Int J Antimicrob Agents. 2017;49(6):709–718. | ||
Ng SP, Davis B, Palombo EA, Bhave M. A Tn5051-like mer-containing transposon identified in a heavy metal tolerant strain Achromobacter sp. AO22. BMC Res Notes. 2009;2:38. | ||
Zhang L, Li Y, Shen W, Wang SM, Wang G, Zhou Y. Whole-genome sequence of a carbapenem-resistant hypermucoviscous Klebsiella pneumoniae isolate SWU01 with capsular serotype K47 that belongs to ST11 from a patient in China. J Glob Antimicrob Resist. 2017;11:87–89. | ||
Gu D, Dong N, Zheng Z, et al. A fatal outbreak of ST11 carbapenem-resistant hypervirulent Klebsiella pneumoniae in a Chinese hospital: a molecular epidemiological study. Lancet Infect Dis. 2018;18(1):37–46. | ||
Harmer CJ, Moran RA, Hall RM. Movement of IS26-associated antibiotic resistance genes occurs via a translocatable unit that includes a single IS26 and preferentially inserts adjacent to another IS26. MBio. 2014;5(5):e01801–01814. | ||
He S, Hickman AB, Varani AM, et al. Insertion Sequence IS26 Reorganizes Plasmids in Clinically Isolated Multidrug-Resistant Bacteria by Replicative Transposition. MBio. 2015;6(3):e00762. | ||
Harmer CJ, Hall RM. IS26-Mediated Formation of Transposons Carrying Antibiotic Resistance Genes. mSphere. 2016;1(2):e00038–16. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.