Back to Journals » Drug Design, Development and Therapy » Volume 9
Clinicopathological significance and potential drug target of RUNX3 in non-small cell lung cancer: a meta-analysis
Authors Xu L, Lan H, Su Y, Li J, Wan J
Received 24 October 2014
Accepted for publication 19 March 2015
Published 3 June 2015 Volume 2015:9 Pages 2855—2865
DOI https://doi.org/10.2147/DDDT.S76358
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Wei Duan
Lijun Xu,1 Hongwen Lan,1 Yushu Su,1 Jun Li,1 Jingwen Wan2
1Department of Cardiothoracic Surgery, Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, 2Department of Surgery (Operation Room), Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Hubei, People’s Republic of China
Background: Emerging evidence indicates that RUNX3 is a candidate tumor suppressor in several types of human tumors, including non-small cell lung cancer (NSCLC). However, the correlation between RUNX3 hypermethylation and clinicopathological characteristics of NSCLC remains unclear. Here, we conducted a systematic review and meta-analysis to quantitatively evaluate the effects of RUNX3 hypermethylation on the incidence of NSCLC and clinicopathological characteristics.
Methods: A detailed literature search was made using Medline, Embase and Web of Science for related research publications written in English. The methodological quality of the studies was evaluated. The data were extracted and assessed independently by two reviewers. Analysis of pooled data was performed. The odds ratio (OR) and hazard ratio were calculated and summarized.
Results: Final analysis of 911 NSCLC patients from 13 eligible studies was performed. We observed that RUNX3 hypermethylation was significantly higher in NSCLC than in normal lung tissue; the pooled OR from seven studies including 361 NSCLC and 345 normal lung tissue (OR 7.08, confidence interval 4.12–12.17, P<0.00001). RUNX3 hypermethylation may also be associated with pathological types. The pooled OR was obtained from eleven studies including 271 squamous cell carcinoma and 389 adenocarcinoma (OR 0.41, confidence interval 0.19–0.89, P=0.02), which indicated that RUNX3 hypermethylation is significantly higher in adenocarcinoma that in squamous cell carcinoma. We did not find that RUNX3 hypermethylation was correlated with clinical stage or differentiated status. However, NSCLC patients with RUNX3 hypermethylation had a lower survival rate than those without RUNX3 hypermethylation.
Conclusion: The results of this meta-analysis suggest that RUNX3 hypermethylation is associated with an increased risk and worse survival in NSCLC. RUNX3 hypermethylation, which induces inactivation of the RUNX3 gene, plays an important role in lung carcinogenesis and clinical outcome.
Keywords: RUNX3, methylation, lung cancer, tumor suppressor gene, meta-analysis, odds ratio, hazard ratio
Introduction
Non-small cell lung cancer (NSCLC) includes several histological types, including adenocarcinoma, squamous cell carcinoma, large cell carcinoma, and mixed histologies, and accounts for approximately 85% of all lung cancers, making NSCLC the leading cause of cancer-related deaths worldwide.1,2 Despite advances in early detection, radical curative surgery, and multimodal therapeutic modalities, about 80% of NSCLC cases are in an advanced stage at diagnosis; systemic chemotherapy remains the standard care but provides marginal improvement in survival.3 Therefore, further investigation on the mechanism of initiation, progression, and identification of prognostic markers is still needed and will help to select patients with a higher risk of disease recurrence and provide a better prognosis and individualized treatment.
The runt-domain-related (RUNX) family genes, which comprise RUNX1, RUNX2, and RUNX3, have come to prominence recently because of their roles as essential regulators of cell fate in development and their paradoxical effects in cancer, as well as in the regulation of p53-dependent DNA damage response and/or tumorigenesis.4–6 The RUNX3 gene is one of the most important members of the runt-domain family, and plays a critical role in regulation of cell proliferation and cell death by apoptosis, and in angiogenesis, cell adhesion, and invasion.7,8 The RUNX3 gene, localized on chromosome 1p36, a region that exhibits frequent loss of heterozygosity events in breast, colon, gastric, and ovarian cancers, is considered to be a tumor suppressor gene involved in the transforming growth factor-beta signaling pathway.9 RUNX3 and the ubiquitous p53 protein are both principal responders of the p14(ARF)-MDM2 cell surveillance pathway that prevents pathologic consequences of abnormal oncogene activation.10 Its precise function has been intensively studied in several tumors, and it is associated with upregulation of inducing cell cycle arrest, apoptosis, and downregulation of cyclin D1 expression.11–15 Lack of protein expression of RUNX3 by promoter methylation (hypermethylation) has been found to play an important role in regulation of lung alveolar differentiation and epithelial tumorigenesis.16–20 However, its role in NSCLC and its clinical significance have not been thoroughly investigated. In this study, we review and update the published clinical investigations regarding the effect of RUNX3 in patients with NSCLC.
Materials and methods
Search strategy and selection criteria
We searched PubMed, Embase, and ISI Web of Knowledge to identify studies from January 1, 2000 to October, 2014 using the following MeSH terms and keywords: “lung” and “cancer or tumor or neoplasm or carcinoma”, “methylation”, and “RUNX3”. We also manually searched the reference lists of the retrieved articles and reviews for additional articles. Although our search did not have language limits initially, we only included studies published in the English language for full-text reading and final evaluation. Conference abstracts were not selected because of insufficient data reported. After exclusion of non-relevant and/or redundant publications from the databases, the remaining papers were evaluated in the full-text version for inclusion and exclusion and for relevant articles in the reference lists. All searched data were retrieved. Authors’ bibliographies and references of selected studies were also searched for additional relevant studies. The most complete study was chosen to avoid duplication if the same patient population was reported in several publications.
Criteria that an eligible study has to meet were as follows: RUNX3 methylation evaluated in the primary NSCLC tissues; researchers revealed the relationship between RUNX3 methylation, NSCLC clinicopathological parameters, and prognosis; RUNX3 methylation examined by polymerase chain reaction; studies that provide a hazard ratio (HR) and 95% confidence interval (CI); and studies that provided sufficient information to estimate HR for overall survival and 95% CI. Letters, reviews, case reports, conference abstracts, editorials, expert opinion, and all publications regarding in vitro/ex vivo studies, cell lines, and human xenografts were excluded.
Data extraction and methodological assessment
Two authors (LX, HL) independently reviewed and extracted the data from eligible studies. Disagreements were resolved by discussion and consensus. Two authors (YS, JL) reviewed all articles that fitted the inclusion and exclusion criteria. The following information was recorded for each study: the first author’s name, year of publication, sample source, number of cases, clinicopathological parameters, cancer TNM (tumor node metastasis) stage, methylation detection method, methylation rate and/or expression, and follow-up. Data for study characteristics and clinical responses were summarized and the data were converted into table format. Heterogeneity of investigation was evaluated to determine whether the data from the various studies could be subjected to meta-analysis.
For the methodological evaluation of the studies, three investigators (LX, HL, XZ) read through each publication independently, and assessed and scored them according to the REMARK (REporting recommendations for tumor MARKer prognostic studies) guidelines and European Lung Cancer Working Party quality scale.21,22 The three readers provided quality scores and compared them, and then reach a consensus value for each item.
Statistical analysis
The statistical analysis was conducted using Stata version 12.0 (Stata Corporation, College Station, TX, USA) and Review Manager 5.2 (Cochrane Collaboration, Oxford, UK). The pooled frequency of RUNX3 hypermethylation and 95% CI were estimated. The frequency of RUNX3 hypermethylation was compared against different tumor characteristics. Heterogeneity among the studies was evaluated using the Cochran’s Q test 23 and the I2 statistic.24,25 When heterogeneity was not an issue (I2 values <50%), a fixed-effect model was used to calculate parameters. If there was substantial heterogeneity (I2 values ≥50%), a random-effects model was used to pool data and an attempt was made to identify potential sources of heterogeneity based on subgroup analyses. The pooled odds ratio (OR) was estimated for the association between RUNX3 hypermethylation and clinicopathological features. P-values tailed less than 0.05 were considered to be statistically significant.
Publication bias was assessed using a method reported by Egger et al.26 We also explored reasons for statistical heterogeneity using meta-regression, subgroup analysis, and sensitivity analysis. The analysis of meta-regression and publication bias was performed using Stata version 10.0.
Results
Identification of relevant studies
Forty-seven publications were identified by the search method described above. Thirty-four papers were excluded for being laboratory studies, non-original articles (reviews), or studies irrelevant to the current analysis. Eventually, 13 studies were included in the final meta-analysis,27–39 as shown in Figure 1. Of the 13 studies, eleven scored eight points, five scored seven points, and one scored six points. Hence, the studies were of relatively high quality (Table 1).
  | Table 1 Quality assessment of the included studies according to the Newcastle–Ottawa scale |
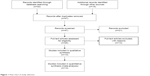  | Figure 1 Flow chart of study selection. |
Study characteristics
Thirteen studies published from 2003 to 2012 were eligible for meta-analysis. A total of 911 NSCLC patients from the People’s Republic of China, Japan, Singapore, South Korea, Spain, and the USA were enrolled. Their basic characteristics are summarized in Table 2.
Correlation of RUNX3 hypermethylation with clinicopathological features
Inactivation of RUNX3 through hypermethylation in NSCLC
The loss of RUNX3 mRNA and/or protein expression is strongly correlated with promoter hypermethylation in several types of cancer, including NSCLC.28,34,40–45 RUNX3 hypermethylation is significantly higher in NSCLC than in normal lung tissue. The pooled OR from seven studies including 361 NSCLC and 345 normal lung tissue is shown in Figure 2 (OR 6.17, CI 3.66–10.41, P<0.00001), indicating that RUNX3 inactivation through hypermethylation plays an important role in the pathogenesis of NSCLC.
Relationship between frequency of RUNX3 hypermethylation and smoking status
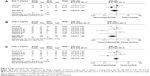
Sato et al reported that the RUNX3 hypermethylation rate in NSCLC patients without a smoking history was significantly higher than that in patients with a smoking history.34 The pooled OR from eight studies including 455 and 202 NSCLC with or without a smoking history is shown in Figure 3A (OR 0.51, CI 0.24–1.08, P=0.08), which indicates that RUNX3 hypermethylation is not strongly associated with smoking status in NSCLC patients.
Relationship between frequency of RUNX3 hypermethylation and pathological types
Previously, Omar et al reported that there was an obvious difference between the activity of RUNX3 in adenocarcinoma and squamous cell carcinoma, indicating that the absence of active RUNX3 in adenocarcinoma and the aberrant downregulation of RUNX3 through hypermethylation may be associated with the development of these tumors.28 The pooled OR from eleven studies including 271 squamous cell carcinomas and 389 adenocarcinomas is shown in Figure 3B (OR 0.41, CI 0.19–0.89, P=0.02), indicating that RUNX3 hypermethylation plays a more important role in the pathogenesis of adenocarcinoma.
Role of RUNX3 hypermethylation in progression of NSCLC
We analyzed 634 NSCLC patients pooled from eight studies to assess whether or not the aberrant RUNX3 hypermethylation in NSCLC was associated with more advanced stage. As shown in Figure 4A, aberrant RUNX3 hypermethylation is not significantly higher in advanced NSCLC (III and IV) than in early stage NSCLC (I and II; OR 1.21, CI 0.61–2.42, P=0.59). In addition, as shown in Figure 4B, aberrant RUNX3 hypermethylation is not significantly higher in poorly differentiated NSCLC than in moderately and highly differentiated NSCLC (OR 0.65, CI 0.14–2.99, P=0.58). These results suggest that epigenetic silencing of RUNX3 gene expression by promoter hypermethylation may not play an important role in the progression and development of NSCLC.
RUNX3 hypermethylation as a prognostic factor for NSCLC
Only three studies evaluated the relationship between RUNX3 hypermethylation and overall survival in NSCLC. The pooled HR for overall survival showed that RUNX3 hypermethylation was associated with worse survival in NSCLC patients, as shown in Figure 4C (HR 3.12, 95% CI 1.47–6.62, P=0.003).
Subgroup analysis and meta-regression
For the group in which heterogeneity is strong, we performed subgroup analysis and meta-regression to analyze the sources of the heterogeneity. In the case of the frequency of RUNX3 hypermethylation and smoking status, the pooled OR for two studies from the People’s Republic of China for patients with or without a smoking history is shown in Figure 5A (I2=47%, OR 0.68, CI 0.30–1.50, P=0.33), indicating that heterogeneity was not strong, suggesting that RUNX3 hypermethylation is not strongly associated with smoking status in NSCLC patients. The pooled OR for six studies from Japan for patients with or without smoking history is shown in Figure 5B (I2=76%, OR 0.46, CI 0.17–1.22, P=0.12), indicating that the heterogeneity is still strong, and that RUNX3 hypermethylation was not significantly associated with smoking status in NSCLC patients. However, the pooled OR for four studies from Japan for patients with or without a smoking history is shown in Figure 5C (I2=50%, OR 0.93, CI 0.52–1.67, P=0.82), indicating that the heterogeneity was not strong, and that RUNX3 hypermethylation is not significantly associated with smoking status in NSCLC patients.
Sensitivity analyses and publication bias
A sensitivity analysis, in which one study was removed at a time, was conducted to assess the stability of the results. The pooled ORs and HRs were not significantly changed, indicating that our analyses were stable. In the case of a relationship between RUNX3 hypermethylation in NSCLC and in normal lung tissue, the overall OR ranged from 4.48 (95% CI 2.04–9.87) to 6.17 (95% CI 3.66–10.41). The funnel plots are largely symmetric (Figure 6), suggesting there was no publication bias in the meta-analysis of RUNX3 methylation/expression and clinicopathological features.
Discussion
RUNX3 belongs to the runt-domain family of transcriptional factors, and its inactivation by promoter hypermethylation plays an important role during normal development and in tumorigenesis in several types of tumors, including NSCLC.11,16–18,46–56 To date, there have been some studies describing the precise expression, prognostic impact, and methylation status of RUNX3 in NSCLC; however, the role of inactivation of RUNX3 in NSCLC and its clinical significance have not been thoroughly investigated. We conducted the present meta-analysis to determine the correlation between RUNX3 hypermethylation and clinicopathological characteristics in NSCLC. Analysis of the pooled data showed that: NSCLC had higher hypermethylation than normal lung tissue (OR 6.17); RUNX3 hypermethylation was not strongly associated with smoking status in NSCLC patients (OR 0.51); RUNX3 hypermethylation played a more important role in the pathogenesis of adenocarcinoma (OR 0.41); aberrant RUNX3 hypermethylation was not significantly higher in advanced NSCLC (III and IV) than that in early stage NSCLC (I and II; OR 1.21); aberrant RUNX3 hypermethylation was not significantly higher in poorly differentiated NSCLC than in moderately and highly differentiated NSCLC (OR 0.65); and NSCLC patients with RUNX3 hypermethylation had a lower survival rate than those without RUNX3 hypermethylation (HR 3.12).
The results from the current study demonstrate that the hypermethylation rate of the RUNX3 gene promoter in NSCLC was significantly higher than in normal lung tissues (OR 7.08), indicating that hypermethylation of the RUNX3 promoter is common in NSCLC. Since changes in hypermethylation of the RUNX3 promoter are reversible, drug treatment through demethylation may be useful to inhibit carcinogenesis, progression, and improve prognosis. This approach brings new direction and hope for cancer treatment through gene-targeted therapy. The results of this meta-analysis also suggest that RUNX3 methylation status could be used as a stratified factor for prediction of NSCLC patient survival. Increasing numbers of studies suggest that determination of the DNA methylation status of tumor suppressor genes has the potential to provide additional information over and above existing cancer markers.
For the group in which the heterogeneity was strong, we performed subgroup analysis and meta-regression to analyze the sources of the heterogeneity. In the case of the relationship between frequency of RUNX3 hypermethylation and smoking status, the heterogeneity was determined in patients from the People’s Republic of China and Japan (Figure 5). The strong heterogeneity appeared to be coming from the studies in Japanese. However, the exact sources of the heterogeneity still need to be determined. Due to the small number of studies and limited information available, we were not able to determine the exact sources of the heterogeneity from the frequency of RUNX3 hypermethylation, pathological types, and clinical stages.
Epigenetics is the study of heritable and age-related modifications of the genome that occur without a change in the primary DNA sequence. Epigenetic alterations, particularly aberrant DNA methylation, one of the best-characterized epigenetic modifications, contribute to tumor initiation and progression.57,58 RUNX3 exerts pleiotropic effects during tumor suppression, and also inhibits the oncogenic Wnt signaling pathway via formation of a complex with the TCF4-β-catenin complex and preventing it from binding to target genes such as c-myc and cyclin D1.46,59 RUNX3 interacts with SMAD3/SMAD4 to activate inhibition of transforming growth factor-beta-dependent proliferation and apoptosis via activation of p21 and Bim. In addition, RUNX3 destabilizes the hypoxia-inducible factor-1 alpha (HIF-1α) protein by promoting hydroxylation of proline in HIF-1α through binding to HIF-1α/PHD2; thus, RUNX3 could be a novel suppressor of HIF-1α and of hypoxia-mediated angiogenesis.60 Therefore, RUNX3 can be considered as a tumor suppressor, and its inactivation could contribute to tumor progression and a poor prognosis. Although only three studies evaluated the relationship between overall survival and hypermethylation of RUNX3 in NSCLC, they yielded very similar results.29,32,33 Based on this meta-analysis, we consider that hypermethylation of RUNX3 in NSCLC tends to indicate a poor prognosis.
Consistent results were shown in the sensitivity analyses, and no evidence of publication bias was found. This study has several potential limitations. First, the possibility of information and selection bias along with unidentified confounders could not be completely excluded because all of the included studies were observational. Second, the searching strategy was restricted to articles published in English. Articles with potentially high-quality data that were published in other languages were not included because of the anticipated difficulties in obtaining accurate medical translations. Hence, caution should be taken when our findings are interpreted among the general populations.
Disclosure
The authors have no financial involvement with any organization or entity with a financial interest in the subject matter or materials discussed in this paper. The authors have no other conflicts of interest to report in regards to this paper.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62:10–29. | ||
Guo P, Huang ZL, Yu P, Li K. Trends in cancer mortality in China: an update. Ann Oncol. 2012;23:2755–2762. | ||
Ramalingam S, Belani C. Systemic chemotherapy for advanced non-small cell lung cancer: recent advances and future directions. Oncologist. 2008;13 Suppl 1:5–13. | ||
Ozaki T, Nakagawara A, Nagase H. RUNX family participates in the regulation of p53-dependent DNA damage response. Int J Genomics. 2013;2013:271347. | ||
Ito Y. RUNX genes in development and cancer: regulation of viral gene expression and the discovery of RUNX family genes. Adv Cancer Res. 2008;99:33–76. | ||
Blyth K, Cameron ER, Neil JC. The RUNX genes: gain or loss of function in cancer. Nat Rev Cancer. 2005;5:376–387. | ||
Subramaniam MM, Chan JY, Yeoh KG, Quek T, Ito K, Salto-Tellez M. Molecular pathology of RUNX3 in human carcinogenesis. Biochim Biophys Acta. 2009;1796:315–331. | ||
Lund AH, van Lohuizen M. RUNX: a trilogy of cancer genes. Cancer Cell. 2002;1:213–215. | ||
Levanon D, Negreanu V, Bernstein Y, Bar-Am I, Avivi L, Groner Y. AML1, AML2, and AML3, the human members of the runt domain gene-family: cDNA structure, expression, and chromosomal localization. Genomics. 1994;23:425–432. | ||
Chi XZ, Kim J, Lee YH, et al. Runt-related transcription factor RUNX3 is a target of MDM2-mediated ubiquitination. Cancer Res. 2009;69:8111–8119. | ||
Shiraha H, Nishina S, Yamamoto K. Loss of runt-related transcription factor 3 causes development and progression of hepatocellular carcinoma. J Cell Biochem. 2011;112:745–749. | ||
Li QL, Ito K, Sakakura C, et al. Causal relationship between the loss of RUNX3 expression and gastric cancer. Cell. 2002;109:113–124. | ||
Chi XZ, Yang JO, Lee KY, et al. RUNX3 suppresses gastric epithelial cell growth by inducing p21(WAF1/Cip1) expression in cooperation with transforming growth factor beta-activated SMAD. Mol Cell Biol. 2005;25:8097–8107. | ||
Wei D, Gong W, Oh SC, et al. Loss of RUNX3 expression significantly affects the clinical outcome of gastric cancer patients and its restoration causes drastic suppression of tumor growth and metastasis. Cancer Res. 2005;65:4809–4816. | ||
Chen LF. Tumor suppressor function of RUNX3 in breast cancer. J Cell Biochem. 2012;113:1470–1477. | ||
Araki K, Osaki M, Nagahama Y, et al. Expression of RUNX3 protein in human lung adenocarcinoma: implications for tumor progression and prognosis. Cancer Sci. 2005;96:227–231. | ||
Lee YS, Lee JW, Jang JW, et al. Runx3 inactivation is a crucial early event in the development of lung adenocarcinoma. Cancer Cell. 2013;24:603–616. | ||
Lee JM, Shin JO, Cho KW, et al. Runx3 is a crucial regulator of alveolar differentiation and lung tumorigenesis in mice. Differentiation. 2011;81:261–258. | ||
Lee KS, Lee YS, Lee JM, et al. Runx3 is required for the differentiation of lung epithelial cells and suppression of lung cancer. Oncogene. 2010;29:3349–3361. | ||
Yanada M, Yaoi T, Shimada J, et al. Frequent hemizygous deletion at 1p36 and hypermethylation downregulate RUNX3 expression in human lung cancer cell lines. Oncol Rep. 2005;14:817–822. | ||
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM. Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst. 2005;97:1180–1184. | ||
Steels E, Paesmans M, Berghmans T, et al. Role of p53 as a prognostic factor for survival in lung cancer: a systematic review of the literature with a meta-analysis. Eur Respir J. 2001;18:705–719. | ||
DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986;7:177–188. | ||
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557–560. | ||
DerSimonian R. Meta-analysis in the design and monitoring of clinical trials. Stat Med. 1996;15:1237–1248. | ||
Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997;315:629–634. | ||
Yu GP, Ji Y, Chen GQ, et al. Application of RUNX3 gene promoter methylation in the diagnosis of non-small cell lung cancer. Oncol Lett. 2012;3:159–162. | ||
Omar MF, Ito K, Nga ME, et al. RUNX3 downregulation in human lung adenocarcinoma is independent of p53, EGFR or KRAS status. Pathol Oncol Res. 2012;18:783–792. | ||
Tang Y, Wu F, Hu C. [RUNX3 promoter hypermethylation and prognosis of early surgically resected non-small cell lung cancers]. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2011;36:650–654. Chinese. | ||
Yanagawa N, Tamura G, Oizumi H, Endoh M, Sadahiro M, Motoyama T. Inverse correlation between EGFR mutation and FHIT, RASSF1A and RUNX3 methylation in lung adenocarcinoma: relation with smoking status. Anticancer Res. 2011;31:1211–1214. | ||
Castro M, Grau L, Puerta P, et al. Multiplexed methylation profiles of tumor suppressor genes and clinical outcome in lung cancer. J Transl Med. 2010;8:86. | ||
Yoshino M, Suzuki M, Tian L, et al. Promoter hypermethylation of the p16 and Wif-1 genes as an independent prognostic marker in stage IA non-small cell lung cancers. Int J Oncol. 2009;35:1201–1209. | ||
Yanagawa N, Tamura G, Oizumi H, et al. Promoter hypermethylation of RASSF1A and RUNX3 genes as an independent prognostic prediction marker in surgically resected non-small cell lung cancers. Lung Cancer. 2007;58:131–138. | ||
Sato K, Tomizawa Y, Iijima H, et al. Epigenetic inactivation of the RUNX3 gene in lung cancer. Oncol Rep. 2006;15:129–135. | ||
Suzuki M, Shigematsu H, Shames DS, et al. DNA methylation-associated inactivation of TGF-beta-related genes DRM/Gremlin, RUNX3, and HPP1 in human cancers. Br J Cancer. 2005;93:1029–1037. | ||
Li QL, Kim HR, Kim WJ, et al. Transcriptional silencing of the RUNX3 gene by CpG hypermethylation is associated with lung cancer. Biochem Biophys Res Commun. 2004;314:223–228. | ||
Yanagawa N, Tamura G, Oizumi H, Takahashi N, Shimazaki Y, Motoyama T. Promoter hypermethylation of tumor suppressor and tumor-related genes in non-small cell lung cancers. Cancer Sci. 2003;94:589–592. | ||
Hawes SE, Stern JE, Feng Q, et al. DNA hypermethylation of tumors from non-small cell lung cancer (NSCLC) patients is associated with gender and histologic type. Lung Cancer. 2010;69:172–179. | ||
Feng Q, Hawes SE, Stern JE, et al. DNA methylation in tumor and matched normal tissues from non-small cell lung cancer patients. Cancer Epidemiol Biomarkers Prev. 2008;17:645–654. | ||
Long C, Yin B, Lu Q, et al. Promoter hypermethylation of the RUNX3 gene in esophageal squamous cell carcinoma. Cancer Invest. 2007;25:685–690. | ||
Tonomoto Y, Tachibana M, Dhar DK, et al. Differential expression of RUNX genes in human esophageal squamous cell carcinoma: downregulation of RUNX3 worsens patient prognosis. Oncology. 2007;73:346–356. | ||
Watanabe T, Kobunai T, Ikeuchi H, et al. RUNX3 copy number predicts the development of UC-associated colorectal cancer. Int J Oncol. 2011;38:201–207. | ||
Goel A, Arnold CN, Tassone P, et al. Epigenetic inactivation of RUNX3 in microsatellite unstable sporadic colon cancers. Int J Cancer. 2004;112:754–759. | ||
Ito K, Lim AC, Salto-Tellez M, et al. RUNX3 attenuates beta-catenin/T cell factors in intestinal tumorigenesis. Cancer Cell. 2008;14:226–237. | ||
Chen W, Gao N, Shen Y, Cen JN. Hypermethylation downregulates Runx3 gene expression and its restoration suppresses gastric epithelial cell growth by inducing p27 and caspase3 in human gastric cancer. J Gastroenterol Hepatol. 2010;25:823–831. | ||
Chuang LS, Ito Y. RUNX3 is multifunctional in carcinogenesis of multiple solid tumors. Oncogene. 2010;29:2605–2615. | ||
Jang BG, Kim WH. Molecular pathology of gastric carcinoma. Pathobiology. 2011;78:302–310. | ||
Kohya N, Koga Y, Kitajima Y, Miyazaki K. Aberrant promoter hypermethylation in biliary tract carcinoma. J Hepatobiliary Pancreat Surg. 2006;13:296–305. | ||
Lee JH, Pyon JK, Kim DW, et al. Expression of RUNX3 in skin cancers. Clin Exp Dermatol. 2011;36:769–774. | ||
Lee YM. Control of RUNX3 by histone methyltransferases. J Cell Biochem. 2011;112:394–400. | ||
Levanon D, Brenner O, Otto F, Groner Y. Runx3 knockouts and stomach cancer. EMBO Rep. 2003;4:560–564. | ||
Shima K, Nosho K, Baba Y, et al. Prognostic significance of CDKN2A (p16) promoter methylation and loss of expression in 902 colorectal cancers: cohort study and literature review. Int J Cancer. 2011;128:1080–1094. | ||
Tamura G. Promoter methylation status of tumor suppressor and tumor-related genes in neoplastic and non-neoplastic gastric epithelia. Histol Histopathol. 2004;19:221–228. | ||
Tamura G. Alterations of tumor suppressor and tumor-related genes in the development and progression of gastric cancer. World J Gastroenterol. 2006;12:192–198. | ||
Zhao C, Bu X. Promoter methylation of tumor-related genes in gastric carcinogenesis. Histol Histopathol. 2012;27:1271–1282. | ||
Suzuki M, Shigematsu H, Shames DS, et al. DNA methylation-associated inactivation of TGF-beta-related genes, DRM/Gremlin, RUNX3, and HPP1 in human cancers. Br J Cancer. 2013;109:31–32. | ||
Delpu Y, Cordelier P, Cho WC, Torrisani J. DNA methylation and cancer diagnosis. Int J Mol Sci. 2013;14:15029–15058. | ||
Ma X, Wang YW, Zhang MQ, Gazdar AF. DNA methylation data analysis and its application to cancer research. Epigenomics. 2013;5:301–316. | ||
Voon DC, Wang H, Koo JK, et al. Runx3 protects gastric epithelial cells against epithelial-mesenchymal transition-induced cellular plasticity and tumorigenicity. Stem Cells. 2012;30:2088–2099. | ||
Lee SH, Bae SC, Kim KW, Lee YM. RUNX3 inhibits hypoxia-inducible factor-1alpha protein stability by interacting with prolyl hydroxylases in gastric cancer cells. Oncogene. 2014;33(11):1458–1467. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.






