Back to Journals » Journal of Inflammation Research » Volume 13
Clinical Value of SARS-CoV2 IgM and IgG Antibodies in Diagnosis of COVID-19 in Suspected Cases
Authors Feng Y
Received 20 October 2020
Accepted for publication 18 November 2020
Published 10 December 2020 Volume 2020:13 Pages 1089—1094
DOI https://doi.org/10.2147/JIR.S287733
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Ning Quan
Yangchun Feng
Clinical Laboratory Center, The Third Hospital Affiliated to Xinjiang Medical University (Affiliated Cancer Hospital), Urumqi, 830011, People’s Republic of China
Correspondence: Yangchun Feng
Third Hospital Affiliated to Xinjiang Medical University (Affiliated Cancer Hospital), Urumqi 830011, People’s Republic of China
Email [email protected]
Objective: To explore the clinical value of SARS-CoV2 IgM and IgG antibodies in the diagnosis of COVID-19 in suspected cases by likelihood ratio.
Methods: By reinterpreting data from a previous study, the positive likelihood ratio of IgM and IgG antibodies in COVID-19 pneumonia diagnosis was calculated, and the posterior probability of IgM and IgG antibodies and their tandem detection was calculated finally.
Results: The positive likelihood ratios of single IgM and IgG antibodies were 18.50 and 12.65, respectively, and the posterior probabilities were 90.18% and 86.26%, respectively. However, the posterior probability of the two antibody-tandem test was 99.15%, which could give clinicians more quantitative confidence in the diagnosis of COVID-19 in suspected cases. According to the results of this study, combining the advantages and disadvantages of nucleic acid testing and antibody detection, a feasible clinical path was found for clinicians to diagnose COVID-19 pneumonia from suspected cases.
Conclusion: For suspected cases, IgM- and IgG-antibody tests should first be done at the same time. If all antibody tests are positive, COVID-19 pneumonia could be confirmed. If not, nucleic acid detection (once or more) should be carried out, and in extreme cases high-throughput viral genome sequencing is required.
Keywords: COVID-19, SARS-CoV2, positive likelihood ratio, posterior probability, antibody test, nucleic acid test, clinical pathway
Introduction
Since the outbreak of SARS-CoV2 last December in China, >51 million cases have been confirmed in >180 countries. As the country with the most severe epidemic in the early period, China has accumulated a lot of experience in the diagnosis and treatment of COVID-19.1,2
Based on increased awareness of the disease, the treatment plan for COVID-19 has been constantly updated by the Chinese government. The latest two versions3,4 are are versions 7 and 8. In version 7, the standard indicators used for confirmed diagnosis of suspected cases were nucleic acid detection, gene sequencing, and antibody testing for SARS-CoV2. Of these, antibody testing was first among the diagnostic criteria. In version 8, antibody testing as a diagnostic indicator of suspected cases was further confirmed. On April 2, 2020, the US Food and Drug Administration approved the first SARS-CoV2 antibody–test kit for COVID-19 detection. The Chinese National Medical Products Administration had also approved 25 antibody-test kits by November 11. Test methods included colloidal gold immunochromatography and chemiluminescence.
Bayesian analysis iss based on the sensitivity and specificity of diagnostic testing, combined with the proportion of various diseases in the population (prior probability) and calculating the probability of suffering from various diseases (posterior probability). Posterior probability can provide a basis for individual diagnosis, which could help clinicians in interpreting test results scientifically and improve diagnosis.5 Posterior probability in antibody testing to diagnose COVID-19 in suspected cases has not been reported, so this study intended to use likelihood ratios (LRs) with antibody testing to calculate posterior probability to clarify the significance of antibody testing in the diagnosis of suspected cases.
In addition, how to use nucleic acid detection and antibody test to diagnose suspected COVID-19 patients more efficiently was a question worth pondering. As such, this study also intended to propose a feasible clinical pathway for clinicians to diagnose suspected cases by nucleic acid and antibody detection.
Methods
Data Sources
The data in this study come from articles on COVID-19 published online by the Chinese Journal of Laboratory Medicine, mainly Ming et al6 and Wanzhou et al.7 The subjects were all suspected and confirmed cases of COVID-19 admitted to the People’s Hospital Affiliated with Wuhan University.
Case Definitions
Suspected Cases: Epidemiological History and Clinical Manifestations
Epidemiological history comprises history of travel to or residence in Wuhan and surrounding areas or other communities where cases have been reported within 14 days prior to the onset of the disease, contact with novel coronavirus–infected people (with positive results on nucleic acid testing) within 14 days prior to the onset of the disease, in contact with patients who have fever or respiratory symptoms from Wuhan and surrounding area or from communities where confirmed cases have been reported within 14 days before the onset of the disease, andlustered cases (two or more cases with fever and/or respiratory symptoms in a small area, such as families, offices, and schools, within 2 weeks.
Clinical manifestations comprise fever and/or respiratory symptoms, the aforementioned imaging characteristics of COVID-19, normal or decreased white blood–cell count, and normal or decreased lymphocyte count in the early stage of onset. A suspect case has any of the epidemiological history plus any two clinical manifestations or all three clinical manifestations if there is no clear epidemiological history.
Confirmed Cases
Suspect cases are those one of the following etiological or serological findings: real-time fluorescent PCR positive for SARS-CoV2 with nucleic acid, Viral gene sequence highly homologous with known SARS-CoV2, SARS-CoV2–specific IgM and IgG detectable in serum, and SARS-CoV2–specific IgG detectable or reaches a titration of at least fourfold the acute phase during convalescence.
Data and Testing
Positive LR (LR+) of IgM and IgG in suspected cases for nucleic acid–positive patients was calculated mainly by reinterpreting the data from Wanzhou et al.7 According to relevant data from Ming et al,6 the posterior probability of IgM- and IgG-antibody detection alone and in series for diagnosis of nucleic acid–positive patients was calculated. SARS-CoV2 IgM and IgG were tested with chemiluminescence immunoassay, and nucleic acid tested using RT-PCR.
Statistical Methods
Data were calculated using a four-grid table.
Results
Diagnostic Performance of IgG and IgM in Diagnosing COVID-19
Wanzhou et al7found that 205 patients had coronavirus infection and 79 patients no COVID-19 pneumonia. Combined with the number of IgG- and IgM-antibody detection mentioned in this article, Tables 1 and 2 were obtained.
 |
Table 1 Diagnostic Performance of IgG Testing to Diagnose COVID-19 |
 |
Table 2 Diagnostic Performance of IgM Testing to Diagnose COVID-19 |
As shown in Table 1, sensitivity, specificity, negative predictive value, and positive predictive value of SARS-CoV2–antibody detection in the diagnosis of pneumonia in COVID-19 were 96.10% (197 of 205), 92.40% (73 of 79), 90.10% (73 of 81) and 97.04% (197 of 203), respectively. The LR– was 0.04 and the LR+ 12.65. Accuracy was 95.07% (270 of 284).
As shown in Table 2, sensitivity, specificity, negative predictive value, and positive predictive value of SARS-CoV2 IgM-antibody detection to diagnose COVID-19 were 70.24% (144 of 205), 96.20% (76 of 79), 56.72% (76 of 137), and 97.76% (144 of 147), respectively. The LR– was 0.31 and the LR+18.50. Accuracy was 77.46% (220 of 284).
Posterior Probability of Single-IgG and -IgM and IgM–IgG Tandem Diagnostic Tests
For posterior probability of a single index:
prior ratio = prior probability/(1 – prior probability)
posterior ratio = prior ratio × LR+
Posterior probability = posterior ratio/(1 + posterior ratio)
For posterior probability formula of tandem diagnostic tests of multiple indices:
prior ratio = prior probability/(1 – prior probability)
posterior ratio = prior ratio × LR+ 1 × LR+ 2 × . . . × LR+ n
Posterior probability = posterior ratio/(1+ posterior ratio)
According to the 8,274 suspected cases of COVID-19 explicitly mentioned in Ming et al,6 the proportion of confirmed cases from suspected cases was 33.17%, which was the prior probability. The posterior probability of single testing of IgG and IgM and IgM–IgG tandem diagnostic testing is shown in Table 3.
 |
Table 3 Posterior Probability of Diagnostic COVID-19 on Single-IgG and -IgM and IgM–IgG Tandem Tests |
For suspected cases, after single detection of IgG and IgM antibodies, the clinician’s confidence in the diagnosis of suspected patients as confirmed COVID-19 cases was 86.26%– 90.18%, but after IgM–IgG tandem testing, the clinician’s confidence in the diagnosis of suspected patients as confirmed COVID-19 cases increased to 99.15%.
Clinical Pathway for Clinicians to Diagnose COVID-19 Based on IgM–IgG Antibody-Tandem Testing
According to the results of this study, combining the advantages and disadvantages of nucleic acid detection and antibody detection, the clinical pathway for clinicians to diagnose COVID-19 is shown in Figure 1. It puts forward a feasible clinical pathway for the diagnosis of pneumonia in COVID-19, starting with IgM–IgG tandem test, then using nucleic acid detection, and finally carrying out high-throughput sequencing if necessary.
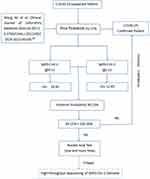 |
Figure 1 Clinical pathway for clinicians to diagnose COVID-19 by SARS-CoV2 antibody. |
Discussion
There is continuous development detection methods for human antibodies after SARS-CoV2 infection.8–10 Nucleic acidand antibody detection are the main laboratory testing methods for COVID-19, But have their own advantages and disadvantages.11,12 First of all, from the aspect of specimen sampling, commonly used specimens for nucleic acid detection are sputum, nasopharynx swab, or various lung-lavage fluids. Sampling poses great infection risk to medical personnel and also potential infection risks for laboratory personnel for nucleic acid detection.13 However, if the type of antibody-test specimen is blood, which is relatively easy to obtain, the risk of infection of medical personnel is relatively low. Secondly, according to the quality of the detection results, nucleic acid detection is restricted by the quality of sample materials and the extraction quality of nucleic acid, and there can be serious undetected phenomena. Different sample types have different detection rates.
Bronchoalveolar lavage–fluid specimens show the highest positive rates (93%), followed by sputum (72%), nasal swabs (63%), fibrobronchoscope brush biopsy (46%), pharyngeal swabs (32%), feces (29%), and blood (1%).14 Repeated nucleic acid detection for for some suspected patients is necessary and can effectively avoid false-negative result. For antibody detection, except for some patients with low immunofunction, antibodies cannot be produced effectively. Antibodies should be present in most infected persons.15 Antibody detection is limited mainly by the sensitivity and related performance of the kit. In addition, there will be some false positives caused by detection interference in tumor patients, patients with autoimmune diseases, and othes with chronic infection. Finally, the convenience of antibody detection has advantages that nucleic acid detection does not have at all, including simplicity and convenience of operation, qualifications of personnel, and requirements of the detection environment. Therefore, it would be of great clinical value to study how to optimize antibody detection and nucleic acid detection in clinical diagnosis.
No clear research on feasible methods of antibody or nucleic acid detection has been found yet. Therefore, this research used published data and adopted the method of LR+ to calculate posterior probability, in order to propose the possibility of optimizing the diagnosis pathway for the first time. Since the original data of the two-antibody tandem test could not be obtained from Wanzhou et al,7 the diagnostic performance index of the tandem test could not be analyzed in this paper. However, this study solved that problem by calculating posterior probability through the LR+, and proved that the posterior probability of the two-antibody tandem test was much higher than each single-antibody detection; however, the posterior probability of single-antibody detection was not different. This showed that the single-antibody test could not significantly increase the confidence of clinicians in a clear diagnosis of confirmed cases from suspected cases. However, the two-antibody tandem test obviously improved this confidence, raising clinicians’ confidence in diagnosis to 99.15%, which was helpful in optimizing the diagnostic process through quantification. The clinical pathway for clinicians to diagnose COVID-19 for suspected cases should be firstly conducting IgM- and IgG-antibody tests. If all antibody tests are positive, COVID-19 can be confirmed. If not, nucleic acid detection (one or more times) can be performed, and in extreme cases high-throughput viral genome sequencing.16
In short, the evaluation of SARS-CoV2 IgM and IgG antibodies serves epidemiological purposes for COVID-19 that are recognized.17 Many conventional diagnostic studies have reported the diagnostic significance of antibody testing,18–20 but in the general population antibody test should not be used as a diagnostic index. It has diagnostic significance only for suspected cases. Compared with the traditional diagnostic performance index, the posterior probability calculated by LR+ can better reflect serial detection of the two antibodies. In practice, it will be more helpful for clinicians in optimizing the diagnostic process. This study was aimed at the secondary excavation of published literature data. In future, the original data should be used and the sample size expanded to further verify the conclusion.
Disclosure
The author declares no conflicts of interest.
References
1. Huang CL, Wang YM, Li XW, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497–506. doi:10.1016/S0140-6736(20)30183-5
2. Wu Z, McGoogan JM. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: summary of a report of 72 314 cases from the Chinese center for disease control and prevention. JAMA. 2020;323(13):1239. doi:10.1001/jama.2020.2648
3. National Health Commission & National Administration of Traditional Chinese Medicine. Diagnosis and treatment protocol for novel coronavirus pneumonia (Trial version 7). Chin Med J. 2020;133. doi:10.3760/cma.j.issn.0366-6999.2020.0027
4. National Health Commission & National Administration of Traditional Chinese Medicine. Diagnosis and treatment protocol for novel coronavirus pneumonia (Trial version 8). Available from: http://www.nhc.gov.cn/yzygj/s7653p/202008/0a7bdf12bd4b46e5bd28ca7f9a7f5e5a.shtml.
5. Chen DT, Schell MJ, Fulp WJ, et al. Application of Bayesian predictive probability for interim futility analysis in single-arm Phase II trial. Transl Cancer Res. 2019;8(Suppl 4):S404–S420. doi:10.21037/tcr.2019.05.17
6. Ming W, Qing W, Wanzhou X, et al. Analysis of novel coronavirus nucleic acid detection and co-infection results of 8274 subjects in Wuhan Area. Chin J Lab Med. 2020;43. doi:10.3760/CMA.J.CN114452-20200222-00106
7. Wanzhou X, Juan L, Xiaoyun H, et al. Diagnostic value of combined detection of serum 2019 novel coronavirus IgM and IgG antibody in novel coronavirus infection. Chin J Lab Med. 2020;43. doi:10.3760/CMA.J.CN114452-20200223-00109
8. Gao HX, Li YN, Xu ZG, et al. Detection of serum immunoglobulin M and immunoglobulin G antibodies in 2019 novel coronavirus infected patients from different stages. Chin Med J (Engl). 2020;133(12):1479–1480.
9. Xiao AT, Gao C, Zhang S. Profile of specific antibodies to SARSCoV-2: the first report. J Infect. 2020;81(1):147–178. doi:10.1016/j.jinf.2020.03.012
10. Pan Y, Li X, Yang G, et al. Serological immunochromatographic approach in diagnosis with SARS-CoV-2 infected COVID-19 patients. J Infect. 2020;81(1):e28–e32. doi:10.1016/j.jinf.2020.03.051
11. Department of Health Science, Technology and Education, National Health Commission of People’s Republic of China. Laboratory biosafety guide for the novel coronavirus. Biosaf Health. 2020. doi:10.3760/cma.j.issn.2096-6962.2020.0001
12. Ping L, Li ZY, Zhao SL, et al. 2019-nCoV serum IgM and IgG antibody used to diagnose the new coronavirus pneumonia preliminary discussion. Chin J Lab Med. 2020;43. doi:10.3760/cmaJcn114452-20200302-00155
13. Li ZT, Yi YX, Luo XM, et al. Development and clinical application of A Rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J Med Virol. 2020. doi:10.1002/jmv.25727
14. Wang W, Xu Y, Gao R, et al. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA. 2020. doi:10.1001/jama.2020.3786
15. Hui-Xia G, Ya-Nan L, Zun-Gui X, et al. Detection of serum immunoglobulin M and immunoglobulin G antibodies in 2019-nCoV infected cases from different stages. Chin Med J. 2020;133. doi:10.1097/CM9.0000000000000820
16. Wen-Da G, Li-Ping C, Feng Y, et al. High-throughput sequencing for confirmation of suspected 2019-nCoV infection identified by fluorescence quantitative polymerase chain reaction. Chin Med J. 2020;133. doi:10.3760/cma.j.issn.0366-6999.2020.0019
17. Gambino CM, Lo Sasso B, Colomba C, et al. Comparison of a rapid immunochromatographic test with a chemiluminescence immunoassay for detection of anti-SARS-CoV-2 IgM and IgG. Biochem Med (Zagreb). 2020;30(3):030901. doi:10.11613/BM.2020.030901
18. Mekonnen D, Mengist HM, Derbie A, et al. Diagnostic accuracy of serological tests and kinetics of severe acute respiratory syndrome coronavirus 2 antibody: a systematic review and meta-analysis. Rev Med Virol. 2020;11:e2181. doi:10.1002/rmv.2181
19. Lisboa Bastos M, Tavaziva G, Abidi SK, et al. Diagnostic accuracy of serological tests for covid-19: systematic review and meta-analysis. BMJ. 2020;370:m2516. doi:10.1136/bmj.m2516
20. Espejo AP, Akgun Y, Al Mana AF, et al. Review of current advances in serologic testing for COVID-19. Am J Clin Pathol. 2020;154(3):293–304. doi:10.1093/ajcp/aqaa112
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
